Abstract
AGA and AGG codons for arginine are the least used codons in Escherichia coli, which are encoded by a rare tRNA, the product of the dnaY gene. We examined the positions of arginine residues encoded by AGA/AGG codons in 678 E. coli proteins. It was found that AGA/AGG codons appear much more frequently within the first 25 codons. This tendency becomes more significant in those proteins containing only one AGA or AGG codon. Other minor codons such as CUA, UCA, AGU, ACA, GGA, CCC and AUA are also found to be preferentially used within the first 25 codons. The effects of the AGG codon on gene expression were examined by inserting one to five AGG codons after the 10th codon from the initiation codon of the lacZ gene. The production of beta-galactosidase decreased as more AGG codons were inserted. With five AGG codons, the production of beta-galactosidase (Gal-AGG5) completely ceased after a mid-log phase of cell growth. After 22 hr induction of the lacZ gene, the overall production of Gal-AGG5 was 11% of the control production (no insertion of arginine codons). When five CGU codons, the major arginine codon were inserted instead of AGG, the production of beta-galactosidase (Gal-CGU5) continued even after stationary phase and the overall production was 66% of the control. The negative effect of the AGG codons on the Gal-AGG5 production was found to be dependent upon the distance between the site of the AGG codons and the initiation codon. As the distance was increased by inserting extra sequences between the two codons, the production of Gal-AGG5 increased almost linearly up to 8 fold. From these results, we propose that the position of the minor codons in an mRNA plays an important role in the regulation of gene expression possibly by modulating the stability of the initiation complex for protein synthesis.
Full text
PDF


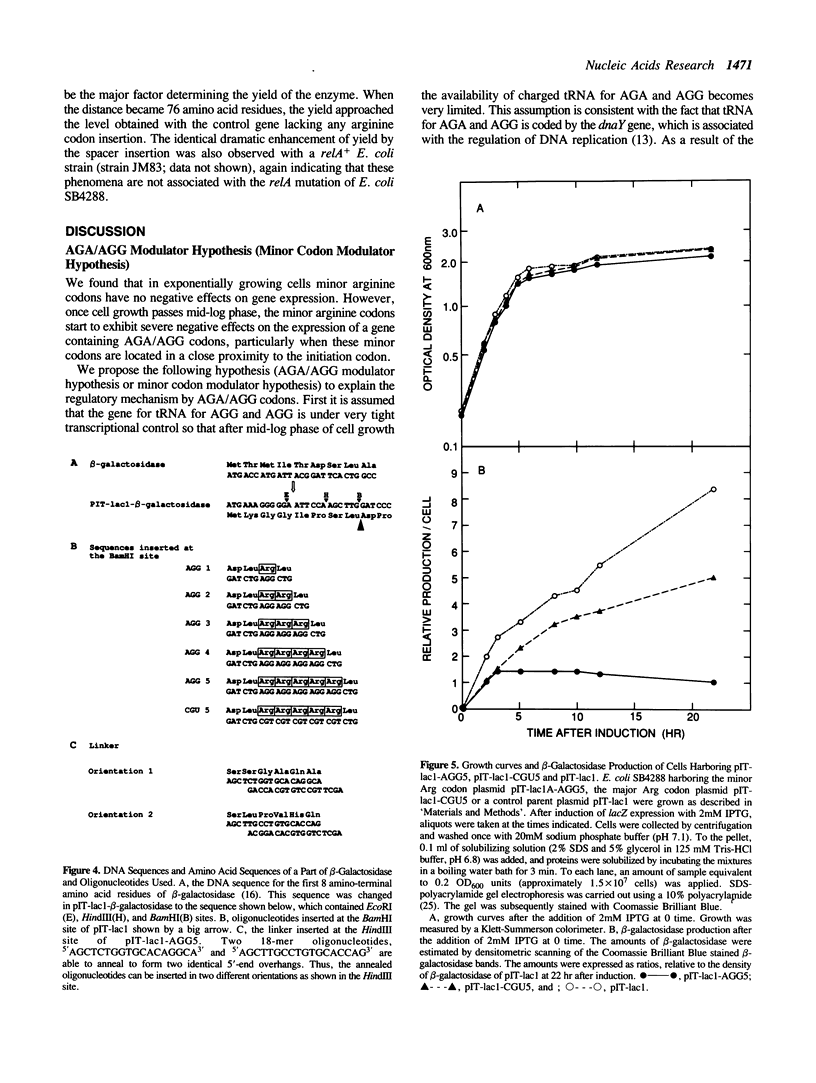
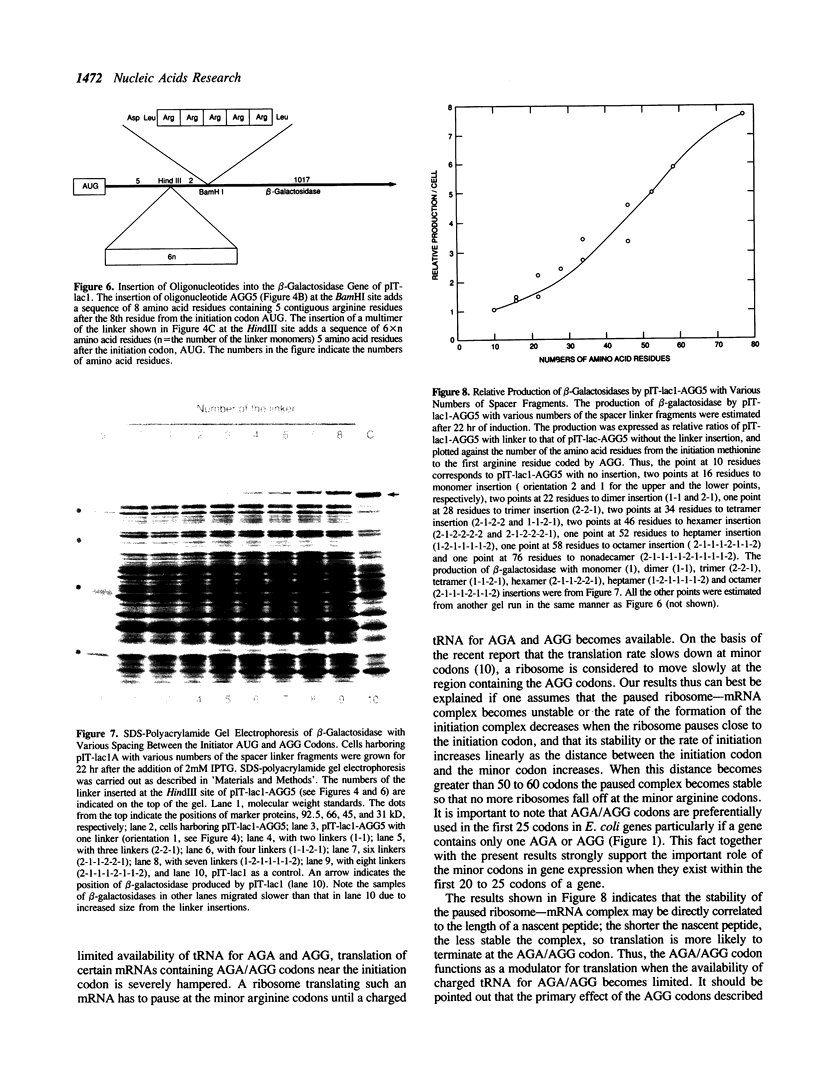

Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Anderson C. W., Baum P. R., Gesteland R. F. Processing of adenovirus 2-induced proteins. J Virol. 1973 Aug;12(2):241–252. doi: 10.1128/jvi.12.2.241-252.1973. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Garcia G. M., Mar P. K., Mullin D. A., Walker J. R., Prather N. E. The E. coli dnaY gene encodes an arginine transfer RNA. Cell. 1986 May 9;45(3):453–459. doi: 10.1016/0092-8674(86)90331-4. [DOI] [PubMed] [Google Scholar]
- Green P. J., Inouye M. Roles of the 5' leader region of the ompA mRNA. J Mol Biol. 1984 Jul 5;176(3):431–442. doi: 10.1016/0022-2836(84)90499-6. [DOI] [PubMed] [Google Scholar]
- Hoekema A., Kastelein R. A., Vasser M., de Boer H. A. Codon replacement in the PGK1 gene of Saccharomyces cerevisiae: experimental approach to study the role of biased codon usage in gene expression. Mol Cell Biol. 1987 Aug;7(8):2914–2924. doi: 10.1128/mcb.7.8.2914. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Holm L. Codon usage and gene expression. Nucleic Acids Res. 1986 Apr 11;14(7):3075–3087. doi: 10.1093/nar/14.7.3075. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ikemura T. Correlation between the abundance of Escherichia coli transfer RNAs and the occurrence of the respective codons in its protein genes. J Mol Biol. 1981 Feb 15;146(1):1–21. doi: 10.1016/0022-2836(81)90363-6. [DOI] [PubMed] [Google Scholar]
- Ikemura T. Correlation between the abundance of Escherichia coli transfer RNAs and the occurrence of the respective codons in its protein genes: a proposal for a synonymous codon choice that is optimal for the E. coli translational system. J Mol Biol. 1981 Sep 25;151(3):389–409. doi: 10.1016/0022-2836(81)90003-6. [DOI] [PubMed] [Google Scholar]
- Inouye S., Inouye M. Up-promoter mutations in the lpp gene of Escherichia coli. Nucleic Acids Res. 1985 May 10;13(9):3101–3110. doi: 10.1093/nar/13.9.3101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kalnins A., Otto K., Rüther U., Müller-Hill B. Sequence of the lacZ gene of Escherichia coli. EMBO J. 1983;2(4):593–597. doi: 10.1002/j.1460-2075.1983.tb01468.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maruyama T., Gojobori T., Aota S., Ikemura T. Codon usage tabulated from the GenBank genetic sequence data. Nucleic Acids Res. 1986;14 (Suppl):r151–r197. doi: 10.1093/nar/14.suppl.r151. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nakamura K., Inouye M. DNA sequence of the gene for the outer membrane lipoprotein of E. coli: an extremely AT-rich promoter. Cell. 1979 Dec;18(4):1109–1117. doi: 10.1016/0092-8674(79)90224-1. [DOI] [PubMed] [Google Scholar]
- Nakamura K., Pirtle R. M., Pirtle I. L., Takeishi K., Inouye M. Messenger ribonucleic acid of the lipoprotein of the Escherichia coli outer membrane. II. The complete nucleotide sequence. J Biol Chem. 1980 Jan 10;255(1):210–216. [PubMed] [Google Scholar]
- Nomura M., Yates J. L., Dean D., Post L. E. Feedback regulation of ribosomal protein gene expression in Escherichia coli: structural homology of ribosomal RNA and ribosomal protein MRNA. Proc Natl Acad Sci U S A. 1980 Dec;77(12):7084–7088. doi: 10.1073/pnas.77.12.7084. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Post L. E., Strycharz G. D., Nomura M., Lewis H., Dennis P. P. Nucleotide sequence of the ribosomal protein gene cluster adjacent to the gene for RNA polymerase subunit beta in Escherichia coli. Proc Natl Acad Sci U S A. 1979 Apr;76(4):1697–1701. doi: 10.1073/pnas.76.4.1697. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Robinson M., Lilley R., Little S., Emtage J. S., Yarranton G., Stephens P., Millican A., Eaton M., Humphreys G. Codon usage can affect efficiency of translation of genes in Escherichia coli. Nucleic Acids Res. 1984 Sep 11;12(17):6663–6671. doi: 10.1093/nar/12.17.6663. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sanger F., Nicklen S., Coulson A. R. DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci U S A. 1977 Dec;74(12):5463–5467. doi: 10.1073/pnas.74.12.5463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sharp P. M., Li W. H. Codon usage in regulatory genes in Escherichia coli does not reflect selection for 'rare' codons. Nucleic Acids Res. 1986 Oct 10;14(19):7737–7749. doi: 10.1093/nar/14.19.7737. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Spanjaard R. A., van Duin J. Translation of the sequence AGG-AGG yields 50% ribosomal frameshift. Proc Natl Acad Sci U S A. 1988 Nov;85(21):7967–7971. doi: 10.1073/pnas.85.21.7967. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sørensen M. A., Kurland C. G., Pedersen S. Codon usage determines translation rate in Escherichia coli. J Mol Biol. 1989 May 20;207(2):365–377. doi: 10.1016/0022-2836(89)90260-x. [DOI] [PubMed] [Google Scholar]
- Tsung K., Brissette R. E., Inouye M. Identification of the DNA-binding domain of the OmpR protein required for transcriptional activation of the ompF and ompC genes of Escherichia coli by in vivo DNA footprinting. J Biol Chem. 1989 Jun 15;264(17):10104–10109. [PubMed] [Google Scholar]
- Varenne S., Lazdunski C. Effect of distribution of unfavourable codons on the maximum rate of gene expression by an heterologous organism. J Theor Biol. 1986 May 7;120(1):99–110. doi: 10.1016/s0022-5193(86)80020-0. [DOI] [PubMed] [Google Scholar]
- Vieira J., Messing J. The pUC plasmids, an M13mp7-derived system for insertion mutagenesis and sequencing with synthetic universal primers. Gene. 1982 Oct;19(3):259–268. doi: 10.1016/0378-1119(82)90015-4. [DOI] [PubMed] [Google Scholar]