Abstract
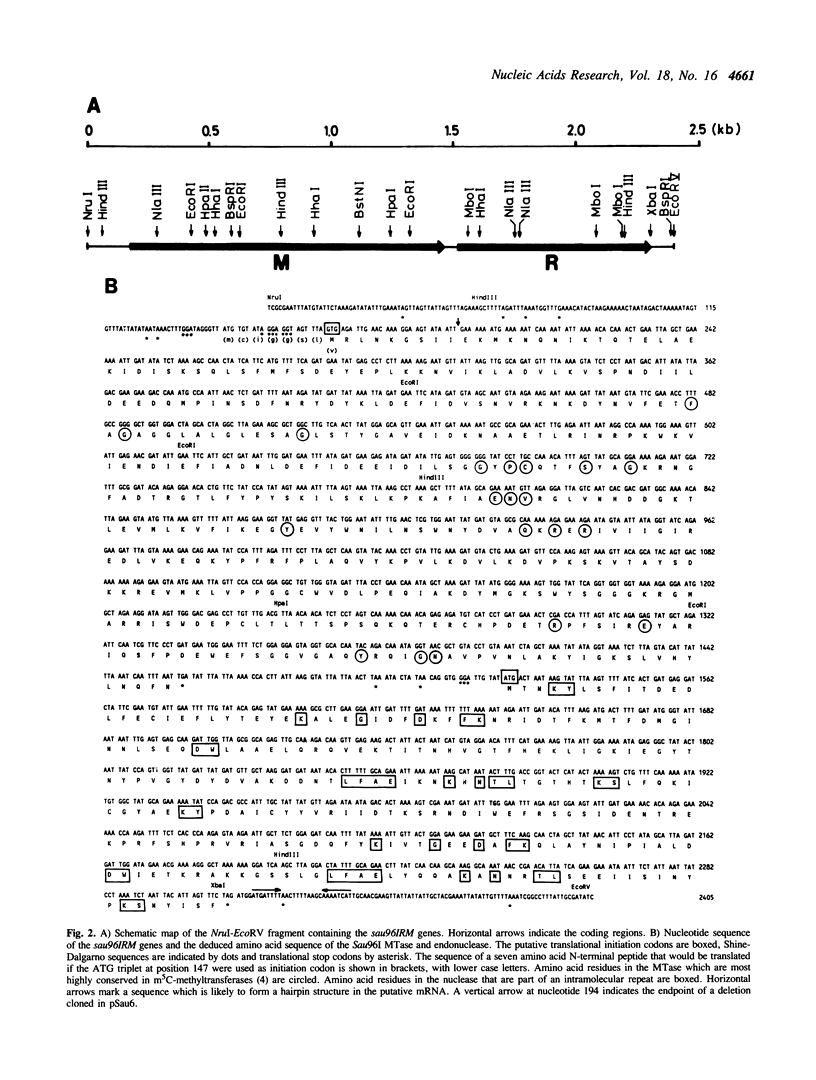
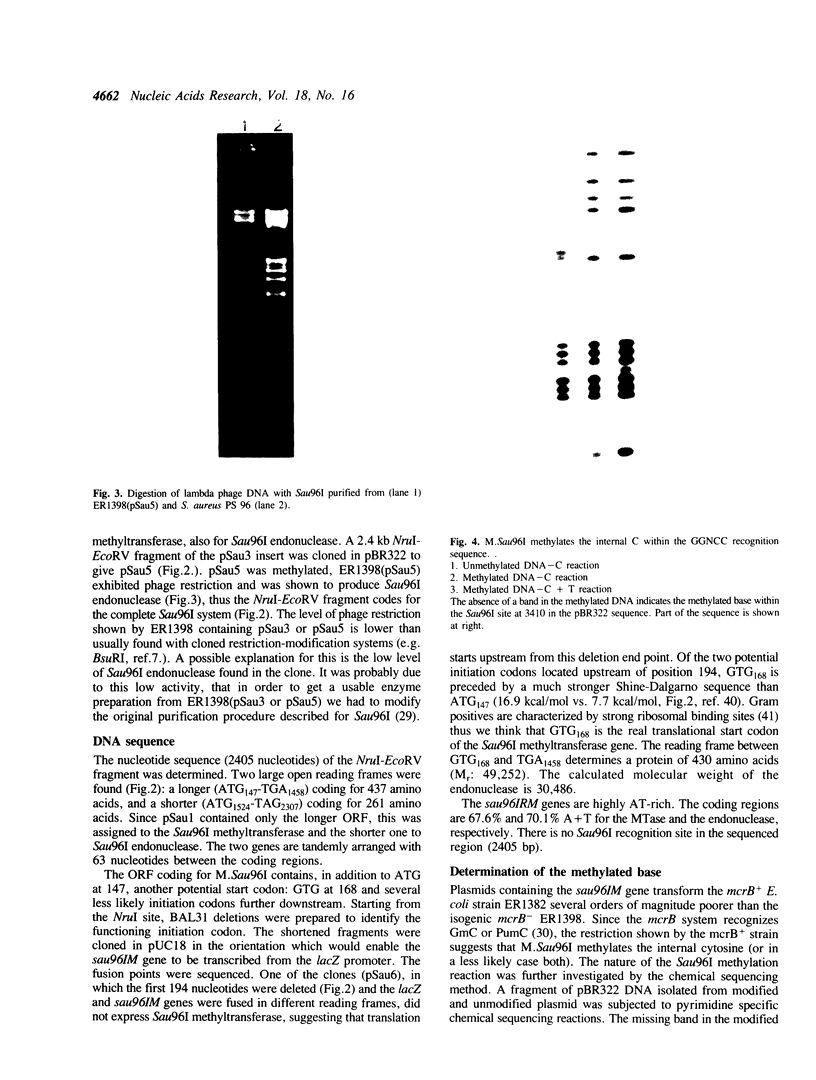
The genes coding for the GGNCC specific Sau96I restriction and modification enzymes were cloned and expressed in E. coli. The DNA sequence predicts a 430 amino acid protein (Mr: 49,252) for the methyltransferase and a 261 amino acid protein (Mr: 30,486) for the endonuclease. No protein sequence similarity was detected between the Sau96I methyltransferase and endonuclease. The methyltransferase contains the sequence elements characteristic for m5C-methyltransferases. In addition to this, M.Sau96I shows similarity, also in the variable region, with one m5C-methyltransferase (M.SinI) which has closely related recognition specificity (GGA/TCC). M.Sau96I methylates the internal cytosine within the GGNCC recognition sequence. The Sau96I endonuclease appears to act as a monomer.
Full text
PDF





Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Balganesh T. S., Reiners L., Lauster R., Noyer-Weidner M., Wilke K., Trautner T. A. Construction and use of chimeric SPR/phi 3T DNA methyltransferases in the definition of sequence recognizing enzyme regions. EMBO J. 1987 Nov;6(11):3543–3549. doi: 10.1002/j.1460-2075.1987.tb02681.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Behrens B., Noyer-Weidner M., Pawlek B., Lauster R., Balganesh T. S., Trautner T. A. Organization of multispecific DNA methyltransferases encoded by temperate Bacillus subtilis phages. EMBO J. 1987 Apr;6(4):1137–1142. doi: 10.1002/j.1460-2075.1987.tb04869.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bestor T., Laudano A., Mattaliano R., Ingram V. Cloning and sequencing of a cDNA encoding DNA methyltransferase of mouse cells. The carboxyl-terminal domain of the mammalian enzymes is related to bacterial restriction methyltransferases. J Mol Biol. 1988 Oct 20;203(4):971–983. doi: 10.1016/0022-2836(88)90122-2. [DOI] [PubMed] [Google Scholar]
- Blumenthal R. M., Gregory S. A., Cooperider J. S. Cloning of a restriction-modification system from Proteus vulgaris and its use in analyzing a methylase-sensitive phenotype in Escherichia coli. J Bacteriol. 1985 Nov;164(2):501–509. doi: 10.1128/jb.164.2.501-509.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bron S., Hörz W. Purification and properties of the Bsu endonuclease. Methods Enzymol. 1980;65(1):112–132. doi: 10.1016/s0076-6879(80)65017-4. [DOI] [PubMed] [Google Scholar]
- Buhk H. J., Behrens B., Tailor R., Wilke K., Prada J. J., Günthert U., Noyer-Weidner M., Jentsch S., Trautner T. A. Restriction and modification in Bacillus subtilis: nucleotide sequence, functional organization and product of the DNA methyltransferase gene of bacteriophage SPR. Gene. 1984 Jul-Aug;29(1-2):51–61. doi: 10.1016/0378-1119(84)90165-3. [DOI] [PubMed] [Google Scholar]
- Butkus V., Klimasauskas S., Kersulyte D., Vaitkevicius D., Lebionka A., Janulaitis A. Investigation of restriction-modification enzymes from M. varians RFL19 with a new type of specificity toward modification of substrate. Nucleic Acids Res. 1985 Aug 26;13(16):5727–5746. doi: 10.1093/nar/13.16.5727. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Card C. O., Wilson G. G., Weule K., Hasapes J., Kiss A., Roberts R. J. Cloning and characterization of the HpaII methylase gene. Nucleic Acids Res. 1990 Mar 25;18(6):1377–1383. doi: 10.1093/nar/18.6.1377. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Caserta M., Zacharias W., Nwankwo D., Wilson G. G., Wells R. D. Cloning, sequencing, in vivo promoter mapping, and expression in Escherichia coli of the gene for the HhaI methyltransferase. J Biol Chem. 1987 Apr 5;262(10):4770–4777. [PubMed] [Google Scholar]
- Hanck T., Gerwin N., Fritz H. J. Nucleotide sequence of the dcm locus of Escherichia coli K12. Nucleic Acids Res. 1989 Jul 25;17(14):5844–5844. doi: 10.1093/nar/17.14.5844. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Henikoff S. Unidirectional digestion with exonuclease III creates targeted breakpoints for DNA sequencing. Gene. 1984 Jun;28(3):351–359. doi: 10.1016/0378-1119(84)90153-7. [DOI] [PubMed] [Google Scholar]
- Karreman C., de Waard A. Agmenellum quadruplicatum M.AquI, a novel modification methylase. J Bacteriol. 1990 Jan;172(1):266–272. doi: 10.1128/jb.172.1.266-272.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Karreman C., de Waard A. Cloning and complete nucleotide sequences of the type II restriction-modification genes of Salmonella infantis. J Bacteriol. 1988 Jun;170(6):2527–2532. doi: 10.1128/jb.170.6.2527-2532.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kiss A., Posfai G., Keller C. C., Venetianer P., Roberts R. J. Nucleotide sequence of the BsuRI restriction-modification system. Nucleic Acids Res. 1985 Sep 25;13(18):6403–6421. doi: 10.1093/nar/13.18.6403. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Klimasauskas S., Timinskas A., Menkevicius S., Butkienè D., Butkus V., Janulaitis A. Sequence motifs characteristic of DNA[cytosine-N4]methyltransferases: similarity to adenine and cytosine-C5 DNA-methylases. Nucleic Acids Res. 1989 Dec 11;17(23):9823–9832. doi: 10.1093/nar/17.23.9823. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kupper D., Zhou J. G., Venetianer P., Kiss A. Cloning and structure of the BepI modification methylase. Nucleic Acids Res. 1989 Feb 11;17(3):1077–1088. doi: 10.1093/nar/17.3.1077. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lauster R. Evolution of type II DNA methyltransferases. A gene duplication model. J Mol Biol. 1989 Mar 20;206(2):313–321. doi: 10.1016/0022-2836(89)90481-6. [DOI] [PubMed] [Google Scholar]
- Lauster R., Trautner T. A., Noyer-Weidner M. Cytosine-specific type II DNA methyltransferases. A conserved enzyme core with variable target-recognizing domains. J Mol Biol. 1989 Mar 20;206(2):305–312. doi: 10.1016/0022-2836(89)90480-4. [DOI] [PubMed] [Google Scholar]
- Lin P. M., Lee C. H., Roberts R. J. Cloning and characterization of the genes encoding the MspI restriction modification system. Nucleic Acids Res. 1989 Apr 25;17(8):3001–3011. doi: 10.1093/nar/17.8.3001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lunnen K. D., Barsomian J. M., Camp R. R., Card C. O., Chen S. Z., Croft R., Looney M. C., Meda M. M., Moran L. S., Nwankwo D. O. Cloning type-II restriction and modification genes. Gene. 1988 Dec 25;74(1):25–32. doi: 10.1016/0378-1119(88)90242-9. [DOI] [PubMed] [Google Scholar]
- Maxam A. M., Gilbert W. Sequencing end-labeled DNA with base-specific chemical cleavages. Methods Enzymol. 1980;65(1):499–560. doi: 10.1016/s0076-6879(80)65059-9. [DOI] [PubMed] [Google Scholar]
- McClarin J. A., Frederick C. A., Wang B. C., Greene P., Boyer H. W., Grable J., Rosenberg J. M. Structure of the DNA-Eco RI endonuclease recognition complex at 3 A resolution. Science. 1986 Dec 19;234(4783):1526–1541. doi: 10.1126/science.3024321. [DOI] [PubMed] [Google Scholar]
- McLaughlin J. R., Murray C. L., Rabinowitz J. C. Unique features in the ribosome binding site sequence of the gram-positive Staphylococcus aureus beta-lactamase gene. J Biol Chem. 1981 Nov 10;256(21):11283–11291. [PubMed] [Google Scholar]
- Norrander J., Kempe T., Messing J. Construction of improved M13 vectors using oligodeoxynucleotide-directed mutagenesis. Gene. 1983 Dec;26(1):101–106. doi: 10.1016/0378-1119(83)90040-9. [DOI] [PubMed] [Google Scholar]
- Ohmori H., Tomizawa J. I., Maxam A. M. Detection of 5-methylcytosine in DNA sequences. Nucleic Acids Res. 1978 May;5(5):1479–1485. doi: 10.1093/nar/5.5.1479. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pósfai G., Baldauf F., Erdei S., Pósfai J., Venetianer P., Kiss A. Structure of the gene coding for the sequence-specific DNA-methyltransferase of the B. subtilis phage SPR. Nucleic Acids Res. 1984 Dec 11;12(23):9039–9049. doi: 10.1093/nar/12.23.9039. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pósfai G., Kiss A., Erdei S., Pósfai J., Venetianer P. Structure of the Bacillus sphaericus R modification methylase gene. J Mol Biol. 1983 Nov 5;170(3):597–610. doi: 10.1016/s0022-2836(83)80123-5. [DOI] [PubMed] [Google Scholar]
- Pósfai J., Bhagwat A. S., Pósfai G., Roberts R. J. Predictive motifs derived from cytosine methyltransferases. Nucleic Acids Res. 1989 Apr 11;17(7):2421–2435. doi: 10.1093/nar/17.7.2421. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Raleigh E. A., Wilson G. Escherichia coli K-12 restricts DNA containing 5-methylcytosine. Proc Natl Acad Sci U S A. 1986 Dec;83(23):9070–9074. doi: 10.1073/pnas.83.23.9070. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Renbaum P., Abrahamove D., Fainsod A., Wilson G. G., Rottem S., Razin A. Cloning, characterization, and expression in Escherichia coli of the gene coding for the CpG DNA methylase from Spiroplasma sp. strain MQ1(M.SssI). Nucleic Acids Res. 1990 Mar 11;18(5):1145–1152. doi: 10.1093/nar/18.5.1145. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sanger F., Nicklen S., Coulson A. R. DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci U S A. 1977 Dec;74(12):5463–5467. doi: 10.1073/pnas.74.12.5463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Slatko B. E., Croft R., Moran L. S., Wilson G. G. Cloning and analysis of the HaeIII and HaeII methyltransferase genes. Gene. 1988 Dec 25;74(1):45–50. doi: 10.1016/0378-1119(88)90248-x. [DOI] [PubMed] [Google Scholar]
- Som S., Bhagwat A. S., Friedman S. Nucleotide sequence and expression of the gene encoding the EcoRII modification enzyme. Nucleic Acids Res. 1987 Jan 12;15(1):313–332. doi: 10.1093/nar/15.1.313. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stephenson F. H., Ballard B. T., Boyer H. W., Rosenberg J. M., Greene P. J. Comparison of the nucleotide and amino acid sequences of the RsrI and EcoRI restriction endonucleases. Gene. 1989 Dec 21;85(1):1–13. doi: 10.1016/0378-1119(89)90458-7. [DOI] [PubMed] [Google Scholar]
- Sullivan K. M., Saunders J. R. Sequence analysis of the NgoPII methyltransferase gene from Neisseria gonorrhoeae P9: homologies with other enzymes recognizing the sequence 5'-GGCC-3'. Nucleic Acids Res. 1988 May 25;16(10):4369–4387. doi: 10.1093/nar/16.10.4369. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sussenbach J. S., Steenbergh P. H., Rost J. A., van Leeuwen W. J., van Embden J. D. A second site-specific restriction endonuclease from Staphylococcus aureus. Nucleic Acids Res. 1978 Apr;5(4):1153–1163. doi: 10.1093/nar/5.4.1153. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sznyter L. A., Slatko B., Moran L., O'Donnell K. H., Brooks J. E. Nucleotide sequence of the DdeI restriction-modification system and characterization of the methylase protein. Nucleic Acids Res. 1987 Oct 26;15(20):8249–8266. doi: 10.1093/nar/15.20.8249. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tinoco I., Jr, Borer P. N., Dengler B., Levin M. D., Uhlenbeck O. C., Crothers D. M., Bralla J. Improved estimation of secondary structure in ribonucleic acids. Nat New Biol. 1973 Nov 14;246(150):40–41. doi: 10.1038/newbio246040a0. [DOI] [PubMed] [Google Scholar]
- Tran-Betcke A., Behrens B., Noyer-Weidner M., Trautner T. A. DNA methyltransferase genes of Bacillus subtilis phages: comparison of their nucleotide sequences. Gene. 1986;42(1):89–96. doi: 10.1016/0378-1119(86)90153-8. [DOI] [PubMed] [Google Scholar]
- Walter J., Noyer-Weidner M., Trautner T. A. The amino acid sequence of the CCGG recognizing DNA methyltransferase M.BsuFI: implications for the analysis of sequence recognition by cytosine DNA methyltransferases. EMBO J. 1990 Apr;9(4):1007–1013. doi: 10.1002/j.1460-2075.1990.tb08203.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wu J. C., Santi D. V. Kinetic and catalytic mechanism of HhaI methyltransferase. J Biol Chem. 1987 Apr 5;262(10):4778–4786. [PubMed] [Google Scholar]
- Yanisch-Perron C., Vieira J., Messing J. Improved M13 phage cloning vectors and host strains: nucleotide sequences of the M13mp18 and pUC19 vectors. Gene. 1985;33(1):103–119. doi: 10.1016/0378-1119(85)90120-9. [DOI] [PubMed] [Google Scholar]