Abstract
Alzheimer’s disease is the most common neurodegenerative disorder. Although lipids are major constituents of brain, their role in Alzheimer’s disease pathogenesis is poorly understood. Much attention has been given to cholesterol, but growing evidence suggests that other lipids, such as phospholipids, might play an important role in this disorder. In this review, we will summarize the evidence linking phospholipase D, a phosphatidic acid-synthesizing enzyme, to multiple aspects of normal brain function and to Alzheimer’s disease. The role of phospholipase D in signaling mechanisms downstream of beta-amyloid as well as in the trafficking and processing of amyloid precursor protein will be emphasized.
1. Introduction
Alzheimer’s disease (AD) is a neurodegenerative disease characterized clinically by progressive memory deficits, impaired cognitive function, altered and inappropriate behavior, and a decline in language function [1-3]. AD patients display cortical atrophy, loss of neurons and synapses, as well as extracellular senile plaques and intracellular neurofibrillary tangles (NFTs). While senile plaques are largely composed of aggregated amyloid β-peptide (Aβ) [1, 2], tangles consist of pairs of ~10 nm filaments wound into helices, also called paired helical filaments, and contain hyperphosphorylated forms of the microtubule-associated protein tau (MAPT). Growing evidence indicates that there is crosstalk between Aβ and tau pathogenesis [4]. Genetic studies of families with AD have identified several genes that are important in the pathogenesis of AD. The most widely studied mutations occur in three genes: β-amyloid precursor protein (APP), presenilin 1 (PS1) and presenilin 2 (PS2). APP is the precursor protein of Aβ, and the presenilins have been identified as components of the γ-secretase complex, which, alongside the β-secretase, are responsible for the generation of Aβ [1, 2].
Aβ is derived from the sequential cleavage of type I transmembrane protein APP by membrane-bound proteases, β- and γ-secretase [1, 5, 6]. Beta-site APP cleavage enzyme 1 (BACE1) has been identified as the major β-secretase activity that mediates the first cleavage of APP in the β-amyloidogenic pathway [7, 8]. BACE1-mediated cleavage leads to the release of the APP ectodomain sAPPβ into the extracellular space. The remaining COOH-terminal fragment (CTF), which can be either 99 or 89 amino acids in length (termed “C99” or “C89”), undergoes subsequent cleavage by γ-secretase to release Aβ and the APP intracellular COOH-terminal domain (AICD). The presenilins have been proposed to be the major catalytic component of the γ-secretase complex, whose sequential intramembrane cleavage of APP produces a spectrum of Aβ peptides varying in length by a few amino acids at the COOH-terminus [5, 7]. The majority of Aβ normally ends at amino acid 40 (Aβ40), but the 42-amino acid variant (Aβ42) is more amyloidogenic, and has been hypothesized to nucleate senile plaque formation [1, 5, 7]. In the past few years it has been shown that soluble oligomeric Aβ40/42 in various assembly states better correlate with synaptic malfunction and cognitive impairment than neuritic plaques in vivo, consistent with mounting evidence that oligomeric Aβ is significantly synaptotoxic [1].
As mentioned above, a number of links exist between Aβ and tau pathologies. Importantly, in an AD mouse model associated with a significant Aβ burden, the genetic ablation of tau was neuroprotective for Aβ-induced deficits [9], supporting an intimate crosstalk between Aβ and tau. However, it is not clear whether the precise reasons for the protective role(s) of tau ablation are related directly to tau’s pathogenic potential. Overall, this relationship between the two major pathogenic AD hallmarks remains poorly understood [4]. Nevertheless, development of an improved understanding of tau-related pathology may inform not only our knowledge of Aβ-related dysfunction in AD but the illness as a whole. While no familial genetic studies have shown a direct cause effect for tau mutations in AD, mutations in the MAPT gene were shown to cause a distinct neurodegenerative disorder, frontal temporal dementia, which is not associated with neuritic plaques. There is a single MAPT gene, which, through alternative splicing, can lead to the expression of six different isoforms in the human brain. MAP tau has a well-established known function of stabilizing microtubules. The microtubule-binding capacity of tau is known to be regulated by post-translational modifications. The most well established process is phosphorylation, mediated by a number of kinases including glygogen synthase kinase 3 (GSK3), cyclin-dependent kinase 5 (CDK5) and microtubule-affinity-regulating kinase (MARK). Additionally, oxidative stress has been suggested to play a role in tau modifications. The formation of NFTs is thought to be a sequential process that starts with impaired tau phosphorylation, detachment of tau from microtubules, accumulation of misfolded tau, tau aggregation in pretangles, formation of paired helical filaments and finally NFTs [10].
2. Phospholipid imbalance in Alzheimer’s disease
Lipid-mediated signaling regulates a plethora of physiological processes, including multiple aspects of brain function. Dysregulation of lipid pathways has been involved in a growing number of neurodegenerative disorders. While much attention has been given to the sterol link to AD [11], growing evidence, suggests that other classes of lipids, such as phospholipids, either mediate or modulate key pathological processes associated with AD [12-14]. The first studies to address these questions were based on post-mortem lipid analysis of brain tissue from AD-affected individuals. For instance, it was reported that anterior temporal cortex of brains from patients with AD had significantly lower levels of phosphatidylinositol (PI) and trends for lower levels of PI phosphate (PIP) and PI-4,5-bisphosphate [PI(4,5)P2] [15], a major signaling lipid regulating a variety of biological processes at the plasma membrane and whose normal balance is required for proper synaptic function [16]. Nitsch et al observed that brains of AD patients had decreased levels of phosphatidylcholine (PC) and phosphatidylethanolamine (PE), proposing that AD may be associated with increased membrane phospholipid degradation [17]. Furthermore, using 31P Nuclear Magnetic Resonance to study AD brain preparations, a significant reduction in PE and PI levels was observed, with a trend for a decrease in PC, phosphatidic acid (PA) and cardiolipin and an increase in sphingomyelin and the plasmalogen derivative of PE [18].
The interaction of both tau and Aβ with cellular and artificial membranes has been extensively investigated. Given direct binding of tau to membrane phospholipids, it was hypothesized that this interaction may be relevant for the physiological function of tau as well as AD pathogenesis [19]. Similarly, Aβ can directly alter artificial lipid bilayers by forming pores that are permeable to various ions [20]. Moreover, an in vitro study showed that Aβ disrupts membranes containing acidic phospholipids in a pH-dependent way with more pronounced effects on membranes at acidic pH [21]. More recently, after the discovery of the synaptotoxic properties of soluble Aβ oligomers, studies showed that these oligomers destabilize membranes and trigger Ca2+ influxes through unknown mechanisms [22]. In part reflecting the pathophysiological relevance of oligomer-induced Ca2+ dyshomeostasis, acute and chronic treatments of neurons with soluble oligomers of Aβ were shown to disrupt the metabolism of PI(4,5)P2 in a phenomenon requiring both extracellular Ca2+and PLC activity [13]. Consistent with the view that PI(4,5)P2 destabilization plays a key role in the synapse-impairing actions of Aβ oligomers, hippocampal slices derived from a genetic model lacking one copy of Synj1 (i.e., a gene encoding the PtdIns(4,5)P2 phosphatase synaptojanin) exhibit normal PtdIns(4,5)P2 levels and long-term potentiation (LTP) despite the presence of Aβ42 oligomers [13]. Similarly, a recent study showed that dysregulation of the group IV A phospholipase A2 pathway mediates some aspects of synaptic and neurobehavioral dysfunction in a mouse model of AD [14]. These two studies provide further support to a prior hypothesis maintaining that phospholipases are highly dysregulated in AD [23]. FAD mutations of presenilins were also shown to cause an imbalance in the metabolism of PI(4,5)P2 and that correcting this imbalance decreased amyloidogenesis [12].
Given the body of evidence presented above demonstrating the importance of phospholipids and AD, development of an improved understanding of phospholipid signaling is essential. Increasing evidence has pointed to one enzyme family especially in regulating biosynthesis and metabolism of phospholipids: phospholipase D (PLD). In this review, we will address the role of the PLD pathway in brain regulation as well as discuss its potential implication in AD.
3. Overview of Phospholipase D – Structure, Function and Localization
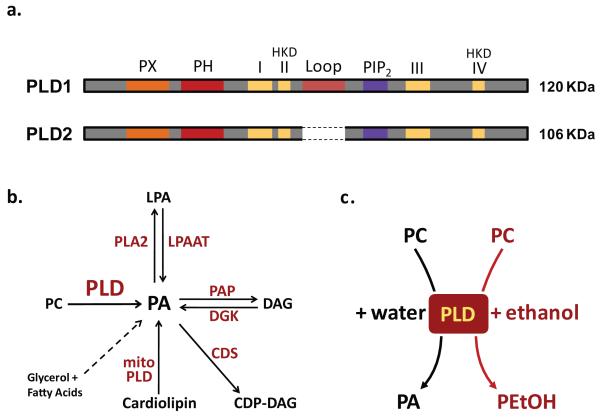
The first evidence for PLD-like activity was reported in plants in 1947 [24], while a mammalian PLD activity was described for the first time in 1973 in rat brain extracts [25]. However, two decades passed until the purification [26] and cloning [27] of PLD from plants. This subsequently led to the identification of SPO14 as a gene that is essential for meiosis and sporulation in yeast and encodes an enzyme harboring PLD activity [28]. The availability of DNA sequences led to the cloning of two PLD isozymes in mammals, PLD1 [29] and PLD2 [30, 31]. In mammals there are three validated isozymes of PLD: PLD1, PLD2 and the recently-identified mitochondrion-associated mitoPLD [32-34]. The majority of studies have so far focused on PLD1 and PLD2 (Fig. 1A), which share: (i) two HxKxxxxD (HKD) motifs that are essential for catalysis; (ii) a phox (PX) consensus sequence and (iii) a pleckstrin homology (PH) domain, which are phosphoinositide-binding modules that are required for proper targeting of PLD; and (iv) a PIP2-binding site, which is fundamental for the enzymatic activity. However, PLD1 differs from PLD2 by the presence of a loop region, which has been proposed to function as a negative regulatory element for catalysis (Fig. 1A) [32]. In the presence of water, both PLD1 and -2 hydrolyze phosphatidylcholine (PC) to generate phosphatidic acid (PA) and free choline. However in the presence of primary alcohols, such as ethanol and 1-butanol, PLD preferentially (~1000-fold) uses these nucleophiles over water for the transphosphatidylation reaction, thus leading to the formation of non-naturally occurring phospholipids, phosphatidylethanol (PtdEtOH) or phosphatidylbutanol (PtdBut), respectively [35] (Fig. 1C). This atypical property of the enzyme has been exploited in a myriad of studies either to block the production of bioactive lipid PA or to measure PLD activity in intact cells or tissues.
Figure 1.
PLD structure, PA metabolism and reactions catalyzed by PLD. (a) Structure of PLD isozymes. Structurally, the two isozymes differ by the presence of a loop domain in the PLD1 isozyme. (b) PA metabolism. Besides the PLD source, PA can be generated from other sources and further metabolized as represented in the figure. The dashed arrow represents the biosynthetic pathway. The enzymes catalyzing the respective reactions are shown in red. (c) PLD activity. In the presence of water, PLD produces PA. In the presence of primary alcohols, such as ethanol, PLD has a 1000-fold higher affinity for primary alcohols as nucleophiles, leading to a preferential generation of phosphatidylethanol (PEtOH). PLD, Phospholipase D; PC, phosphatydilcholine; PA, phosphatidic acid; LPA, lyso-PA; DAG, diacylglycerol; CDP-DAG, cytidine diphosphate-DAG; PLA2, phospholipase A2; LPAAT, LPA acyltransferase; PAP, PA phosphatase; DGK, DAG kinase; CDS, CDP-DAG synthase; mitoPLD, mito-phospholipase D.
PA has unique bioactive properties and can modify both the physical and signalling properties of lipid bilayers. Structurally, it is composed of a three-carbon glycerol backbone, two fatty acid chains and a small phosphate headgroup, thus referred to as a ‘cone shape’ lipid (i.e., a lipid with a small head groups relative to a large hydrophobic domain) [32, 36]. This property not only confers PA a higher affinity for negative curvature within lipid bilayers, but it also reduces the energy barrier for bending membrane, thus acting as a fusogenic lipid. Somewhat reminiscent of phosphoinositides, PA also plays an important role at the membrane-cytosol interface through a direct interaction with effector proteins, such as PIP kinases, mTOR, SNARE proteins and sphingosine kinase [37]. However, unlike a variety of phosphoinositide-binding modules (e.g., pleckstrin homology or FYVE domains), PA-binding (poly)peptides generally do not consist of well defined three dimensional folds, but instead involve basic residues in unstructured parts of effector proteins. Additionally, PA can be metabolized to other lipids with potent bioactivity. For instance, PA can be converted to diacylglycerol (DAG) by PA phosphatases [38, 39] and to lysophosphatidic acid (LPA), which has an inverted cone shape and thus prefers positive curvature. Finally, PA can serve as a precursor for other lipids in the biosynthetic pathway, where it is consumed for the generation of lipids such as PI via the CDP-DAG pathway. It should be noted that PLD is not the only source of PA, as it can be produced by DAG kinases, LPA acid acyltransferase, mitoPLD and other enzymes in the biosynthetic pathway [33, 40] (Fig. 1C). However, the fatty acyl composition of PA pools varies, depending on the specific pathways mediating its production. For example, PLD-derived PA species harbor mainly saturated or monounsaturated fatty acids, rapidly giving rise to a pool of DAG with the same properties (i.e., a cone shape), although the negative charge of PA may confer distinct properties and allow for the binding to a different set of effector proteins compared to DAG. In contrast, the pool of DAG resulting from PI(4,5)P2 cleavage by PLC predominantly harbors polyunsaturated fatty acids, likely achieving different physiological functions [41, 42].
While PLD1 localizes to the Golgi complex, secretory granules and endosomes, PLD2 is concentrated at the plasma membrane, with smaller pools present in the Golgi apparatus, caveolae and in endosomes [43-46]. In part consistent with their subcellular localization, PLD1 has been implicated in the budding and fusion of trans-Golgi-derived secretory vesicles, whereas PLD2 mediates the internalization and recycling of a variety of receptors. Importantly, PLD1 translocates to the plasma membrane upon various stimuli and follows the endosomal internalization route [32, 34, 47]. Our unpublished studies show that PLD1 relocalizes to autophagosomes and modulates autophagy during nutrient deprivation (Claudia Dall’Armi and GDP, unpublished observations).
4. Role of the PLD pathway in brain function
Following the first description of mammalian brain-associated PLD activity in 1973 [25], the same team reported the formation of ethanol metabolite, PEtOH in rat brain synaptosomes through a PLD-mediated transphosphatidylation reaction [48]. This method was further used to measure PLD activity in homogenates of different brain regions, where the hippocampus, hypothalamus and cortex were associated with the highest PLD activity [49]. Both PLD1 and PLD2 are expressed throughout the brain during development and postnatal life, although levels of expression vary from one cell type to another, at least at the mRNA level [30] (see also reference [50]). In agreement with these expression patterns, both PLD1 and PLD2 have been implicated in neurite outgrowth [51-53].
PLD is likely to play a central role in neurophysiology based on its abundance in the brain and on its implication in multiple aspects of cell physiology including membrane trafficking, cytoskeleton regulation and signal transduction [53-55]. Although no genetic or in vivo studies support this idea at this time, several reports involving cell culture models suggest this may be the case, particularly for PLD2. For instance, a study utilizing both RNAi knockdown approach and primary alcohol application has shown that PLD2, likely in concert with small GTPase Ral, regulates the constitutive internalization of metabotropic glutamate receptors, mGluR1a and mGluR5a, as well as colocalizing with these receptors in the endocytic pathway [56]. While this study points to a role of PLD2 in the modulation of excitatory neurotransmission, growing evidence suggests broader involvement in internalization processes. Based on previous findings showing that opioid peptides stimulate PLD activity in chicken embryo cultures [57], more recent studies have shown that PLD regulates the trafficking of opioid receptors, such as the μ- and δ-opioid receptors, with potentially important implications for drug addiction [58, 59]. In this case, however, PLD2 appears to be required for ligand-induced (rather than constitutive) endocytosis and to cooperate with the small GTPase Arf6 which plays an important role in modulating vesicle trafficking to and from the plasma membrane [60, 61]. Interestingly, signalling processes downstream of PLD2 are beginning to be unravelled, as PLD2-activating opioids have been shown to lead to the production of reactive oxygen species (ROS) [62]. Collectively, these studies support a role for PLD2 in signalling processes downstream of the opioid receptors as well as in the regulation of their cell surface levels.
5. Links between PLD and Alzheimer’s disease
The first study linking PLD to AD was published by Kanfer et al [63], where a decrease in PLD activity, as measured by an in vitro choline release assay from radiolabeled PC, was first reported in AD brain homogenates relative to those from control patients. However, when PLD activity was re-investigated in AD brain homogenates relative to those from demented patients without AD, no difference was found [64]. The same group subsequently used PEtOH production as a read-out for PLD activity in ethanol-incubated brain homogenates and found an increase in PLD activity in AD brain extracts relative to those from control subjects [65]. Perhaps related to this study, increased protein levels of PLD1 were found in mitochondrial fractions derived from AD brains, although the pathophysiological significance of this finding is unclear [66]. In the next sections, we will summarize more recent evidence connecting PLD to AD as well as to the function of proteins, such as presenilins, which are involved in this neurodegenerative disorder.
5.1 The role of PLD in the Aβ signaling pathway
Following the aforementioned in vitro studies with AD brain homogenates, cell culture models were used to assess the impact of APP overexpression and extracellular Aβ applications on PLD activity. Overexpression of the neuronal isoform of human wild type APP in P19 mouse embryonic cells caused an increase in PLD activity [67]. Aβ application experiments were used as well with various peptide preparation types. In LA-N-2 cells, a neuroblastoma cell line, an increase in PLD activity was observed after incubating cells with Aβ25-35, a peptide sequence whose pathophysiological significance has been questioned [68]. Additionally, the same group showed that indomethacin (a non-steroid anti-inflammatory drug), nordihydroguaiaretic acid (an anti-oxidant drug) and nicotine inhibited the increase in PLD activity produced by Aβ25-35 applications [69, 70]. Furthermore, alanine substitution for the amino acids on the position 29-34 of Aβ25-35 prevented the peptide from having an effect on PLD activity [71]. Finally, pre-treatment with Aβ25-35 desensitized the cells, which did not exhibit a PLD activity increase in response to a new Aβ25-35 treatment, thus prompting the authors to suggest that Aβ25-35 might mediate its effects on PLD upon receptor binding [70]. Aβ1-40 also produced an increase in PLD activity in rat hippocampal primary cultures, but this effect was seen with high concentrations of Aβ aged at 37°C (likely reflecting a requirement for some aggregated state of Aβ for this phenomenon to occur) [72]. The increase in PLD activity induced by Aβ correlated with increased release of cytosolic protein lactate dehydrogenase, suggesting it is associated with Aβ-induced toxicity [72]. Besides neurons, Aβ1-42 used in the low micromolar range caused an increase in PLD activity in astrocytes and microglia in a process dependent upon formyl-peptide-receptor-like 1 (FPRL1). Specifically, Aβ1-42 was internalized alongside the FPRL1 receptor and stimulated a downstream signaling pathway involving the phosphorylation of extracellular signal-regulated kinase (ERK). Because both Aβ1-42 internalization and FPRL1-mediated signaling were abrogated by a primary alcohol, a role for PLD in this process was proposed [73]. Recently, our lab has addressed the effects of soluble oligomeric preparations of Aβ1-42 (oAβ1-42), which are highly synaptotoxic at submicromolar concentrations, on PLD activity. We found that oAβ42 at 200 nM leads to an increase in PLD activity in primary cortical cultures and that this effect is abolished in mouse neurons lacking PLD2. Strikingly, we also found that the genetic of ablation of Pld2 confers protection against the suppressive effects of oAβ1-42 on LTP and the memory-impairing action of SwAPP overexpression in vivo (TGO and GDP, unpublished observations).
5.2 The role of PLD on APP trafficking and Aβ generation
Because Aβ originates from the sequential cleavage of amyloid precursor protein (APP) by β- and γ-secretases, tremendous effort has been put into attempting to understand the molecular mechanisms regulating the subcellular localization and intracellular sorting of APP and its cleaving enzymes, all of which are transmembrane proteins or protein complexes. Generally, an increasingly popular hypothesis in the field is that perturbation of the trafficking of APP, BACE1 and γ-secretase complex [including its catalytic components, presenilins (PS)] may profoundly affect amyloidogenesis and thus impact AD pathogenesis [74].
As with most type 1 transmembrane proteins, APP traffics through the secretory pathway toward the cell surface, where it can be re-internalized and reach the endosomal system. Additionally, there is evidence that APP traffics back to the trans-Golgi network (TGN) and it has been suggested that reducing this transport step results in increased amyloidogenesis. This may reflect the fact that the APP/BACE1 interaction is predominant in the endosomes and that the acidic pH of this compartment is optimal for BACE1-mediated proteolysis [74]. Despite the complexity in the intracellular trafficking pattern of APP, most studies addressing the role of PLD in this process have focused on the cell surface delivery of APP (and particularly, the Swedish mutant of APP) from the TGN. These studies were prompted by a large body of evidence implicating PLD1 as well as its product PA in the budding of secretory vesicles from the TGN [75-77]. Furthermore, while the bulk of the studies addressing the relationship between the γ-secretase complex and APP focus on the APP processing, some evidence indicates that presenilin can also regulate the trafficking of APP independently of its catalytic activity. Specifically, TGN-derived secretory vesicles produced from PS1-deficient cells contain higher levels of APP, thus resulting in increased cell surface delivery of APP. The converse phenomenon was observed in cells expressing an FAD mutant version of PS1 (i.e., ΔE9), suggesting that these findings may be relevant for the pathogenesis of AD [78]. Collectively, these results converged onto a transport pathway involving APP, PS and PLD1, although evidence for a bona fide crosstalk between these three molecules in the context of TGN-to-plasmalemma traffic emerged in subsequent studies.
Accordingly, PLD1 overexpression was shown to promote the formation of APP-containing secretory vesicles from TGN, thus mimicking the effect of PSEN1 nullizygosity [79]. However, primary alcohols failed to rescue this phenotype in PS1 knockout cells, suggesting that the increased biogenesis of APP-containing vesicles observed in PS1-deficient cells is PLD-independent. While these data suggested that WT PS1 and PLD1 may regulate the traffic of APP through independent pathways, there is a clear functional link between these two proteins in the FAD mutant background (at least, for the PS1ΔE9 mutant). For instance, a subcellular fractionation experiment showed that expression of FAD PS1ΔE9 mutant in cells leads to a enrichment of PLD1 in the Golgi/TGN fraction at the expense of lighter fractions (which may in part represent endosomal structures) and that this redistribution was concomitant with a decrease in total PLD activity [79]. Importantly, overexpression of catalytically-active PLD1 (but not the lipase-dead mutant) was found to rescue the defect in the budding of APP-containing vesicles from the TGN as well in the cell surface delivery of APP induced by the PS1 ΔE9 [79]. Whether PLD1 interacts with APP or not is a matter of debate as conflicting data were obtained in independent studies [66, 80].
While the aforementioned study highlights the effects of PLD1 on APP trafficking, Cai et al. also addressed the role of PLD1 in amyloidogenesis. First, it was shown that PLD1 (but not PLD2) physically interacts with the cytoplasmic loop region of PS1 and that through this binding PS1 mediates the recruitment of PLD1 to the Golgi complex. Furthermore, in N2a cells expressing the PS1 mutant ΔE9, overexpression of PLD1 decreased the levels of Aβ, whereas silencing PLD1 and expression of a lipase-dead PLD1 mutant produced the converse effect. Interestingly, co-precipitation experiments showed that PLD1 regulates the assembly of the γ-secretase complex through a direct effect on PS1 (but not the other components of the complex: Pen-2, nicastrin and APH1), although this phenomenon appears to be independent of the lipase activity of PLD1 [81].
Since PLD1 regulates the traffic of APP and that APP, in turn, may affect the transport of PS1, it was hypothesized that PLD1 could regulate the trafficking of PS1 indirectly, through APP. However, recently published evidence has suggested that PLD1 positively regulates the delivery of PS1 to the cell surface in an APP-independent fashion [80]. Indeed, analysis of PS1 localization in mouse embryonic fibroblasts (MEFs) lacking both APP and its related family member APP-like-protein 2 (APLP2) (APP dKO) showed increased cell surface delivery of PS1. This phenomenon was mimicked by the overexpression of catalytically-active PLD1, but it also occurred in APPdKO cells, thus suggesting that it is independent of an effect of PLD1 on APP transport [80].
In summary, there appears to be a significant crosstalk between PLD1, APP and PS1 with important implications for amyloidogenesis. However, it is unclear whether this crosstalk also occurs in vivo, which should be best addressed with genetic models. Importantly, PLD1 and PLD2 are likely to play distinct roles in AD pathogenesis, perhaps reflecting their differential subcellular localization, expression levels/profile, and regulation as well as their ability to control different aspects in the biology of the key proteins involved in AD.
6. Other potential links between PLD and neurodegeneration
There are at least four additional lines of research potentially linking PLD to degenerative processes. The first involves PLD and a potential link with α-synuclein, the second implicates PLD isozymes as substrates of caspases; the third points to how a perturbation of PA metabolism can alter membrane dynamics with consequences on cell viability; and the fourth involves the connection between PLD and mTOR in survival pathways.
6.1 PLD and Parkinson Disease
In addition to regulating the function of a variety of receptors, PLD has been implicated in brain-associated pathophysiology through an interaction with a disease-relevant cytoplasmic protein, α-synuclein. This aggregate-prone protein is the product of a gene that is mutated in familial cases of Parkinson’s Disease (PD) and also involved in AD pathology based on its presence in Lewy bodies and neuritic plaques, respectively [82]. The original study suggesting a functional link between α-synuclein and PLD2 demonstrated that α-synuclein inhibits the lipase activity of PLD2 in vitro [83]. Subsequent studies have corroborated this link through analyses of the direct physical interaction between these two proteins and of the effects of various PD-linked mutations on PLD activity both in vitro and in intact cells [84, 85]. However the (patho)physiological relevance of this interaction has been questioned, since conflicting data has been obtained by others with respect to the inhibitory action of α-synuclein on PLD activity [86].
6.2 PLD and programmed cell death
Two recent studies have shown that caspases-8, -3 and -7 can cleave PLD isozymes in vitro [87] and that caspase-3 can also cleave these enzymes in vivo [88]. Both demonstrate that PLD1 is cleaved within its loop, which is a region that structurally distinguishes PLD1 from PLD2 and negatively regulates the lipase activity of PLD1. This caspase cleavage produces two fragments of approximately 60 (NH2-terminus) and 55 kDA (at the COOH-terminus), respectively. In contrast, PLD2 is cleaved in proximity to the NH2-terminus, resulting in a shift in PLD2’s molecular weight from 106 to 102 KDa with a release of a small fragment and a larger one still encompassing the bulk of the PLD2 sequence. However, these studies present conflicting conclusions concerning the impact on the activity of the enzymes after caspase-induced cleavage and the role of the various proteolytic products in the modulation of apoptosis outcome [87, 88]. For instance, the first study shows that caspase-cleaved PLD1 was no longer activated by PKC, but resulted in increased activity induced by small GTPases, and PLD2 activity did not change after caspase-3 induced cleavage [87]. However, the second study shows that apoptosis-induced PLD1 cleavage results in lower PLD1 activity levels and agrees with in vitro studies demonstrating that apoptosis-induced PLD2 cleavage had no effect on its activity [88]. Regardless of this discrepancy, caspase-mediated cleavage of PLD may be relevant for AD, as these proteases have been extensively implicated in this disorder (e.g., downstream of Aβ) [89] and both caspase-3 activity and PLD1 fragments matching the size of those generated by caspase-3 are upregulated in AD brain tissue [88].
6.3 PLD overexpression and cellular degeneration
A recent report by Raghu et al provided further insight into the role of PLD in neurodegeneration [90]. In Drosophila melanogaster photoreceptors, PA levels were altered using distinct genetic manipulations targeting PA metabolism, such as (i) overexpression of the only Pld gene; (ii) partial inactivation of cytidine diphosphate (CDP)-DAG synthase (i.e., cds1), which consumes PA for the de novo production of other phospholipids; and (iii) overexpression of rdgA, a gene encoding a fly DAG kinase). These three genetic manipulations produced major developmental defects in the photoreceptors, specifically with reduction in the size of rhabdomeres, a specialized structure in these photoreceptors where sensory transduction occurs. Remarkably, this phenotype correlated with increased levels of a single molecular species of PA (out of 17 analyzed) with a specific fatty acyl composition – 34:2 (16:0/18:2 or 16:1/18:1) and was rescued by co-expression of a PA phosphatase. This study underscores the physiological importance of regulating metabolic pools of PA, and suggests that certain cells, tissues or organisms may not efficiently cope with increased PLD levels/activity, a situation proposed to occur in AD.
6.4 PLD and the mTOR pathway
In the past few years, several studies have highlighted a role for PLD in cell survival. For instance, increased PLD activity was correlated with enhanced survival rates in various human cancer tissues and neoplasic cell lines and also, numerous growth factors have been shown to lead to increased PLD activity [91]. These findings may be related to the observation that PLD can regulate mTOR (mammalian target of rapamycin), which is involved in cell growth and proliferation and has also been proposed to play a role in AD [91, 92]. mTOR is present in two complexes, mTORC1 (when mTOR is bound to Raptor) and mTORC2 (when bound to Rictor). Rapamycin inhibits the formation of the mTORC1 complex through an interaction with FKBP12 (i.e., a modulator of the activity of various complexes in the cell) (review on mTOR). Interestingly, PA was found to compete with rapamycin for binding to the FRB domain of mTOR [93]. Moreover, PA was shown to be required for the stabilization of mTORC1 and 2 complexes, competing with rapamycin [94]. Additionally, PLD1, but not PLD2, was shown to be required for the activation of mTOR by Rheb [95]. Even though the majority of the data suggests a stronger role for PLD1, there is conflicting data concerning the specific role of how each PLD isozyme affects mTOR regulation [96].
7. Conclusions and future directions
Here we review an increasing number of reports showing the involvement of PLD in AD pathogenesis. Although PLD1 and PLD2 have similar protein structures (with the exception of the regulatory loop) and catalyze the same reaction, their different regulation and localization in the cell, implicate these proteins in different cellular processes. In the case of AD pathogenesis, several studies point to a role of PLD1 in the intracellular trafficking of APP and presenilin with important implications for amyloidogenesis. Although PLD2 has not been extensively studied in the context of AD, our unpublished work suggests it may regulate processes associated with Aβ signaling, perhaps due to its predominant localization at the cell surface, a main site of action of this cytotoxic peptide. Importantly, ablation of PLD2 appears to be protective in the context of a transgenic model of AD. Whether PLD has any role in AD-associated tau pathology remains to be discovered. However, the pleiotropic actions of PLD isozymes in the cell, including their implication in many signal transduction pathways, make this scenario plausible. Indeed, a key factor in tau pathology appears to be hyperphosphorylation of microtubule-associated protein by such kinases as Cdk-5 and GSK3β, thus denoting a profound dysregulation of signaling pathways. Interestingly, a link between PLD and these two kinases has been recently published [97, 98]. Furthermore, the implication of PLD in the mTOR pathway, combined with an increasing number of studies exploiting inhibitors of this pathway (e.g., rapamycin analogs) to treat proteinopathies as a result of their stimulatory actions on the macroautophagy pathway, suggests that a manipulation of PLD is likely to affect the clearance of protein aggregates.
PLD isozymes are ubiquitously expressed lipid enzymes implicated in multiple facets of cell physiology, consistent with the pleiotropic roles of their product, PA. Despite nearly two decades of research on these proteins, their physiological (in vivo) functions are still elusive, in part due to the lack of genetic tools in mammalian systems. While efforts along this line are ongoing in several laboratories (including our own), work in lower species suggests that PLD-encoding genes are largely non-essential, unless organisms are undergoing some stress conditions [99]. In this respect, while the discovery that PLD genes are non-essential in mammals contrasts the many studies implicating PLD1 and PLD2 in fundamental cellular processes, it suggests the exciting possibility that PLD isozymes may be used as potential drug targets for therapeutics in several areas, including AD and cancer. The recently-generated pharmacological blockers of these enzymes may prove extremely helpful in this endeavor, particularly if they have different efficacies toward the two isozymes [100-102].
ACKNOWLEDGMENTS
We would like to thank Robin Chan, Samuel Frere, Kimberly Robinson, and Zachary Freyberg for critical reading of the manuscript. Work from T.G.O. is supported by the Portuguese Foundation for Science and Technology SFRH/BD/33237/2007 and the Luso-American Development Foundation.
References
- [1].Haass C, Selkoe DJ. Soluble protein oligomers in neurodegeneration: lessons from the Alzheimer’s amyloid beta-peptide. Nat Rev Mol Cell Biol. 2007;8:101–112. doi: 10.1038/nrm2101. [DOI] [PubMed] [Google Scholar]
- [2].Tanzi RE, Bertram L. Twenty years of the Alzheimer’s disease amyloid hypothesis: a genetic perspective. Cell. 2005;120:545–555. doi: 10.1016/j.cell.2005.02.008. [DOI] [PubMed] [Google Scholar]
- [3].Querfurth HW, LaFerla FM. Alzheimer’s disease. N Engl J Med. 2010;362:329–344. doi: 10.1056/NEJMra0909142. [DOI] [PubMed] [Google Scholar]
- [4].Small SA, Duff K. Linking Abeta and tau in late-onset Alzheimer’s disease: a dual pathway hypothesis. Neuron. 2008;60:534–542. doi: 10.1016/j.neuron.2008.11.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [5].Landman N, Kim TW. Got RIP? Presenilin-dependent intramembrane proteolysis in growth factor receptor signaling. Cytokine Growth Factor Rev. 2004;15:337–351. doi: 10.1016/j.cytogfr.2004.04.001. [DOI] [PubMed] [Google Scholar]
- [6].Wilquet V, De Strooper B. Amyloid-beta precursor protein processing in neurodegeneration. Curr Opin Neurobiol. 2004;14:582–588. doi: 10.1016/j.conb.2004.08.001. [DOI] [PubMed] [Google Scholar]
- [7].Haass C. Take five--BACE and the gamma-secretase quartet conduct Alzheimer’s amyloid beta-peptide generation. Embo J. 2004;23:483–488. doi: 10.1038/sj.emboj.7600061. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [8].Sinha S, Lieberburg I. Cellular mechanisms of beta-amyloid production and secretion. Proc Natl Acad Sci U S A. 1999;96:11049–11053. doi: 10.1073/pnas.96.20.11049. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [9].Roberson ED, Scearce-Levie K, Palop JJ, Yan F, Cheng IH, Wu T, Gerstein H, Yu GQ, Mucke L. Reducing endogenous tau ameliorates amyloid beta-induced deficits in an Alzheimer’s disease mouse model. Science. 2007;316:750–754. doi: 10.1126/science.1141736. [DOI] [PubMed] [Google Scholar]
- [10].Ballatore C, Lee VM, Trojanowski JQ. Tau-mediated neurodegeneration in Alzheimer’s disease and related disorders. Nat Rev Neurosci. 2007;8:663–672. doi: 10.1038/nrn2194. [DOI] [PubMed] [Google Scholar]
- [11].Puglielli L, Tanzi RE, Kovacs DM. Alzheimer’s disease: the cholesterol connection. Nat Neurosci. 2003;6:345–351. doi: 10.1038/nn0403-345. [DOI] [PubMed] [Google Scholar]
- [12].Landman N, Jeong SY, Shin SY, Voronov SV, Serban G, Kang MS, Park MK, Di Paolo G, Chung S, Kim TW. Presenilin mutations linked to familial Alzheimer’s disease cause an imbalance in phosphatidylinositol 4,5-bisphosphate metabolism. Proc Natl Acad Sci U S A. 2006;103:19524–19529. doi: 10.1073/pnas.0604954103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [13].Berman DE, Dall’Armi C, Voronov SV, McIntire LB, Zhang H, Moore AZ, Staniszewski A, Arancio O, Kim TW, Di Paolo G. Oligomeric amyloid-beta peptide disrupts phosphatidylinositol-4,5-bisphosphate metabolism. Nat Neurosci. 2008;11:547–554. doi: 10.1038/nn.2100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [14].Sanchez-Mejia RO, Newman JW, Toh S, Yu GQ, Zhou Y, Halabisky B, Cisse M, Scearce-Levie K, Cheng IH, Gan L, Palop JJ, Bonventre JV, Mucke L. Phospholipase A2 reduction ameliorates cognitive deficits in a mouse model of Alzheimer’s disease. Nat Neurosci. 2008;11:1311–1318. doi: 10.1038/nn.2213. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [15].Stokes CE, Hawthorne JN. Reduced phosphoinositide concentrations in anterior temporal cortex of Alzheimer-diseased brains. J Neurochem. 1987;48:1018–1021. doi: 10.1111/j.1471-4159.1987.tb05619.x. [DOI] [PubMed] [Google Scholar]
- [16].Di Paolo G, De Camilli P. Phosphoinositides in cell regulation and membrane dynamics. Nature. 2006;443:651–657. doi: 10.1038/nature05185. [DOI] [PubMed] [Google Scholar]
- [17].Nitsch RM, Blusztajn JK, Pittas AG, Slack BE, Growdon JH, Wurtman RJ. Evidence for a membrane defect in Alzheimer disease brain. Proc Natl Acad Sci U S A. 1992;89:1671–1675. doi: 10.1073/pnas.89.5.1671. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [18].Pettegrew JW, Panchalingam K, Hamilton RL, McClure RJ. Brain membrane phospholipid alterations in Alzheimer’s disease. Neurochem Res. 2001;26:771–782. doi: 10.1023/a:1011603916962. [DOI] [PubMed] [Google Scholar]
- [19].Baudier J, Cole RD. Phosphorylation of tau proteins to a state like that in Alzheimer’s brain is catalyzed by a calcium/calmodulin-dependent kinase and modulated by phospholipids. J Biol Chem. 1987;262:17577–17583. [PubMed] [Google Scholar]
- [20].Arispe N, Rojas E, Pollard HB. Alzheimer disease amyloid beta protein forms calcium channels in bilayer membranes: blockade by tromethamine and aluminum. Proc Natl Acad Sci U S A. 1993;90:567–571. doi: 10.1073/pnas.90.2.567. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [21].McLaurin J, Chakrabartty A. Membrane disruption by Alzheimer beta-amyloid peptides mediated through specific binding to either phospholipids or gangliosides. Implications for neurotoxicity. J Biol Chem. 1996;271:26482–26489. doi: 10.1074/jbc.271.43.26482. [DOI] [PubMed] [Google Scholar]
- [22].Demuro A, Mina E, Kayed R, Milton SC, Parker I, Glabe CG. Calcium dysregulation and membrane disruption as a ubiquitous neurotoxic mechanism of soluble amyloid oligomers. J Biol Chem. 2005;280:17294–17300. doi: 10.1074/jbc.M500997200. [DOI] [PubMed] [Google Scholar]
- [23].Farooqui AA, Rapoport SI, Horrocks LA. Membrane phospholipid alterations in Alzheimer’s disease: deficiency of ethanolamine plasmalogens. Neurochem Res. 1997;22:523–527. doi: 10.1023/a:1027380331807. [DOI] [PubMed] [Google Scholar]
- [24].Hanahan DJ, Chaikoff IL. The Phosphorus-Containing Lipides of the Carrot. Journal of Biological Chemistry. 1947;168:233–240. [PubMed] [Google Scholar]
- [25].Saito M, Kanfer J. Solubilization and Properties of a Membrane-Bound Enzyme from Rat-Brain Catalyzing a Base-Exchange Reaction. Biochemical and Biophysical Research Communications. 1973;53:391–398. doi: 10.1016/0006-291x(73)90674-8. [DOI] [PubMed] [Google Scholar]
- [26].Wang XM, Dyer JH, Zheng L. Purification and Immunological Analysis of Phospholipase-D from Castor Bean Endosperm. Archives of Biochemistry and Biophysics. 1993;306:486–494. doi: 10.1006/abbi.1993.1541. [DOI] [PubMed] [Google Scholar]
- [27].Wang XM, Xu LW, Zheng L. Cloning and Expression of Phosphatidylcholine-Hydrolyzing Phospholipase-D from Ricinus-Communis L. Journal of Biological Chemistry. 1994;269:20312–20317. [PubMed] [Google Scholar]
- [28].Rose K, Rudge SA, Frohman MA, Morris AJ, Engebrecht J. Phospholipase D signaling is essential for meiosis. Proc Natl Acad Sci U S A. 1995;92:12151–12155. doi: 10.1073/pnas.92.26.12151. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [29].Hammond SM, Altshuller YM, Sung TC, Rudge SK, Rose K, Engebrecht J, Morris AJ, Frohman MA. Human Adp-Ribosylation Factor-Activated Phosphatidylcholine-Specific Phospholipase-D Defines a New and Highly Conserved Gene Family. Journal of Biological Chemistry. 1995;270:29640–29643. doi: 10.1074/jbc.270.50.29640. [DOI] [PubMed] [Google Scholar]
- [30].Colley WC, Sung TC, Roll R, Jenco J, Hammond SM, Altshuller Y, Bar-Sagi D, Morris AJ, Frohman MA. Phospholipase D2, a distinct phospholipase D isoform with novel regulatory properties that provokes cytoskeletal reorganization. Curr Biol. 1997;7:191–201. doi: 10.1016/s0960-9822(97)70090-3. [DOI] [PubMed] [Google Scholar]
- [31].Kodaki T, Yamashita S. Cloning, expression, and characterization of a novel phospholipase D complementary DNA from rat brain. Journal of Biological Chemistry. 1997;272:11408–11413. doi: 10.1074/jbc.272.17.11408. [DOI] [PubMed] [Google Scholar]
- [32].Jenkins GM, Frohman MA. Phospholipase D: a lipid centric review. Cell Mol Life Sci. 2005;62:2305–2316. doi: 10.1007/s00018-005-5195-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [33].Choi SY, Huang P, Jenkins GM, Chan DC, Schiller J, Frohman MA. A common lipid links Mfn-mediated mitochondrial fusion and SNARE-regulated exocytosis. Nat Cell Biol. 2006;8:1255–1262. doi: 10.1038/ncb1487. [DOI] [PubMed] [Google Scholar]
- [34].Donaldson JG. Phospholipase D in endocytosis and endosomal recycling pathways. Biochim Biophys Acta. 2009;1791:845–849. doi: 10.1016/j.bbalip.2009.05.011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [35].Gustavsson L. ESBRA 1994 Award Lecture. Phosphatidylethanol formation: specific effects of ethanol mediated via phospholipase D. Alcohol Alcohol. 1995;30:391–406. [PubMed] [Google Scholar]
- [36].Cazzolli R, Shemon AN, Fang MQ, Hughes WE. Phospholipid signalling through phospholipase D and phosphatidic acid. Iubmb Life. 2006;58:457–461. doi: 10.1080/15216540600871142. [DOI] [PubMed] [Google Scholar]
- [37].Stace CL, Ktistakis NT. Phosphatidic acid- and phosphatidylserine-binding proteins. Biochim Biophys Acta. 2006;1761:913–926. doi: 10.1016/j.bbalip.2006.03.006. [DOI] [PubMed] [Google Scholar]
- [38].Sciorra VA, Morris AJ. Roles for lipid phosphate phosphatases in regulation of cellular signaling. Biochimica Et Biophysica Acta-Molecular and Cell Biology of Lipids. 2002;1582:45–51. doi: 10.1016/s1388-1981(02)00136-1. [DOI] [PubMed] [Google Scholar]
- [39].Reue K. The lipin family: mutations and metabolism. Current Opinion in Lipidology. 2009;20:165–170. doi: 10.1097/MOL.0b013e32832adee5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [40].Haucke V, Di Paolo G. Lipids and lipid modifications in the regulation of membrane traffic. Curr Opin Cell Biol. 2007;19:426–435. doi: 10.1016/j.ceb.2007.06.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [41].Pettitt TR, Martin A, Horton T, Liossis C, Lord JM, Wakelam MJ. Diacylglycerol and phosphatidate generated by phospholipases C and D, respectively, have distinct fatty acid compositions and functions. Phospholipase D-derived diacylglycerol does not activate protein kinase C in porcine aortic endothelial cells. J Biol Chem. 1997;272:17354–17359. doi: 10.1074/jbc.272.28.17354. [DOI] [PubMed] [Google Scholar]
- [42].Pettitt TR, McDermott M, Saqib KM, Shimwell N, Wakelam MJ. Phospholipase D1b and D2a generate structurally identical phosphatidic acid species in mammalian cells. Biochem J. 2001;360:707–715. doi: 10.1042/0264-6021:3600707. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [43].Bader MF, Vitale N. Phospholipase D in calcium-regulated exocytosis: lessons from chromaffin cells. Biochim Biophys Acta. 2009;1791:936–941. doi: 10.1016/j.bbalip.2009.02.016. [DOI] [PubMed] [Google Scholar]
- [44].Freyberg Z, Bourgoin S, Shields D. Phospholipase D2 is localized to the rims of the Golgi apparatus in mammalian cells. Mol Biol Cell. 2002;13:3930–3942. doi: 10.1091/mbc.02-04-0059. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [45].Freyberg Z, Siddhanta A, Shields D. “Slip, sliding away”: phospholipase D and the Golgi apparatus. Trends Cell Biol. 2003;13:540–546. doi: 10.1016/j.tcb.2003.08.004. [DOI] [PubMed] [Google Scholar]
- [46].Freyberg Z, Sweeney D, Siddhanta A, Bourgoin S, Frohman M, Shields D. Intracellular localization of phospholipase D1 in mammalian cells. Mol Biol Cell. 2001;12:943–955. doi: 10.1091/mbc.12.4.943. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [47].Roth MG. Molecular mechanisms of PLD function in membrane traffic. Traffic. 2008;9:1233–1239. doi: 10.1111/j.1600-0854.2008.00742.x. [DOI] [PubMed] [Google Scholar]
- [48].Kobayashi M, Kanfer JN. Phosphatidylethanol formation via transphosphatidylation by rat brain synaptosomal phospholipase D. J Neurochem. 1987;48:1597–1603. doi: 10.1111/j.1471-4159.1987.tb05707.x. [DOI] [PubMed] [Google Scholar]
- [49].Kobayashi M, Mccartney DG, Kanfer JN. Developmental-Changes and Regional Distribution of Phospholipase-D and Base-Exchange Enzyme-Activities in Rat-Brain. Neurochemical Research. 1988;13:771–776. doi: 10.1007/BF00971601. [DOI] [PubMed] [Google Scholar]
- [50].Kim M, Moon C, Kim H, Shin MK, Min do S, Shin T. Developmental levels of phospholipase D isozymes in the brain of developing rats. Acta Histochem. 2010;112:81–91. doi: 10.1016/j.acthis.2008.09.004. [DOI] [PubMed] [Google Scholar]
- [51].Zhang Y, Huang P, Du G, Kanaho Y, Frohman MA, Tsirka SE. Increased expression of two phospholipase D isoforms during experimentally induced hippocampal mossy fiber outgrowth. Glia. 2004;46:74–83. doi: 10.1002/glia.10322. [DOI] [PubMed] [Google Scholar]
- [52].Zhang Y, Kanaho Y, Frohman MA, Tsirka SE. Phospholipase D1-promoted release of tissue plasminogen activator facilitates neurite outgrowth. Journal of Neuroscience. 2005;25:1797–1805. doi: 10.1523/JNEUROSCI.4850-04.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [53].Kanaho Y, Funakoshi Y, Hasegawa H. Phospholipase D signalling and its involvement in neurite outgrowth. Biochim Biophys Acta. 2009;1791:898–904. doi: 10.1016/j.bbalip.2009.03.010. [DOI] [PubMed] [Google Scholar]
- [54].Klein J, Chalifa V, Liscovitch M, Loffelholz K. Role of phospholipase D activation in nervous system physiology and pathophysiology. J Neurochem. 1995;65:1445–1455. doi: 10.1046/j.1471-4159.1995.65041445.x. [DOI] [PubMed] [Google Scholar]
- [55].Klein J. Functions and pathophysiological roles of phospholipase D in the brain. J Neurochem. 2005 doi: 10.1111/j.1471-4159.2005.03315.x. [DOI] [PubMed] [Google Scholar]
- [56].Bhattacharya M, Babwah AV, Godin C, Anborgh PH, Dale LB, Poulter MO, Ferguson SS. Ral and phospholipase D2-dependent pathway for constitutive metabotropic glutamate receptor endocytosis. J Neurosci. 2004;24:8752–8761. doi: 10.1523/JNEUROSCI.3155-04.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [57].Mangoura D, Dawson G. Opioid peptides activate phospholipase D and protein kinase C-epsilon in chicken embryo neuron cultures. Proc Natl Acad Sci U S A. 1993;90:2915–2919. doi: 10.1073/pnas.90.7.2915. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [58].Koch T, Brandenburg LO, Schulz S, Liang Y, Klein J, Hollt V. ADP-ribosylation factor-dependent phospholipase D2 activation is required for agonist-induced mu-opioid receptor endocytosis. J Biol Chem. 2003;278:9979–9985. doi: 10.1074/jbc.M206709200. [DOI] [PubMed] [Google Scholar]
- [59].Koch T, Brandenburg LO, Liang Y, Schulz S, Beyer A, Schroder H, Hollt V. Phospholipase D2 modulates agonist-induced mu-opioid receptor desensitization and resensitization. J Neurochem. 2004;88:680–688. doi: 10.1046/j.1471-4159.2003.02189.x. [DOI] [PubMed] [Google Scholar]
- [60].Koch T, Wu DF, Yang LQ, Brandenburg LO, Hollt V. Role of phospholipase D2 in the agonist-induced and constitutive endocytosis of G-protein coupled receptors. J Neurochem. 2006;97:365–372. doi: 10.1111/j.1471-4159.2006.03736.x. [DOI] [PubMed] [Google Scholar]
- [61].Rankovic M, Jacob L, Rankovic V, Brandenburg LO, Schroder H, Hollt V, Koch T. ADP-ribosylation factor 6 regulates mu-opioid receptor trafficking and signaling via activation of phospholipase D2. Cellular Signalling. 2009;21:1784–1793. doi: 10.1016/j.cellsig.2009.07.014. [DOI] [PubMed] [Google Scholar]
- [62].Koch T, Seifert A, Wu DF, Rankovic M, Kraus J, Borner C, Brandenburg LO, Schroder H, Hollt V. mu-opioid receptor-stimulated synthesis of reactive oxygen species is mediated via phospholipase D2. J Neurochem. 2009;110:1288–1296. doi: 10.1111/j.1471-4159.2009.06217.x. [DOI] [PubMed] [Google Scholar]
- [63].Kanfer JN, Hattori H, Orihel D. Reduced phospholipase D activity in brain tissue samples from Alzheimer’s disease patients. Ann Neurol. 1986;20:265–267. doi: 10.1002/ana.410200214. [DOI] [PubMed] [Google Scholar]
- [64].Kanfer JN, Pettegrew JW, Moossy J, McCartney DG. Alterations of selected enzymes of phospholipid metabolism in Alzheimer’s disease brain tissue as compared to non-Alzheimer’s demented controls. Neurochem Res. 1993;18:331–334. doi: 10.1007/BF00969091. [DOI] [PubMed] [Google Scholar]
- [65].Kanfer JN, Singh IN, Pettegrew JW, McCartney DG, Sorrentino G. Phospholipid metabolism in Alzheimer’s disease and in a human cholinergic cell. J Lipid Mediat Cell Signal. 1996;14:361–363. doi: 10.1016/0929-7855(96)00545-7. [DOI] [PubMed] [Google Scholar]
- [66].Jin JK, Kim NH, Lee YJ, Kim YS, Choi EK, Kozlowski PB, Park MH, Kim HS, Min DS. Phospholipase D1 is up-regulated in the mitochondrial fraction from the brains of Alzheimer’s disease patients. Neuroscience Letters. 2006;407:263–267. doi: 10.1016/j.neulet.2006.08.062. [DOI] [PubMed] [Google Scholar]
- [67].Lee MJ, Oh JY, Park HT, Uhlinger DJ, Kwak JY. Enhancement of phospholipase D activity by overexpression of amyloid precursor protein in P19 mouse embryonic carcinoma cells. Neurosci Lett. 2001;315:159–163. doi: 10.1016/s0304-3940(01)02339-4. [DOI] [PubMed] [Google Scholar]
- [68].Singh IN, Mccartney DG, Kanfer JN. Amyloid-Beta Protein (25-35) Stimulation of Phospholipases a, C and D Activities of La-N-2 Cells. Febs Letters. 1995;365:125–128. doi: 10.1016/0014-5793(95)00432-9. [DOI] [PubMed] [Google Scholar]
- [69].Singh IN, Sorrentino G, Sitar DS, Kanfer JN. Indomethacin and nordihydroguaiaretic acid inhibition of amyloid beta protein (25-35) activation of phospholipases A(2) and D of LA-N-2 cells. Neuroscience Letters. 1997;222:5–8. doi: 10.1016/s0304-3940(97)13327-4. [DOI] [PubMed] [Google Scholar]
- [70].Singh IN, Sorrentino G, Sitar DS, Kanfer JN. (-)Nicotine inhibits the activations of phospholipases A2 and D by amyloid beta peptide. Brain Research. 1998;800:275–281. doi: 10.1016/s0006-8993(98)00532-0. [DOI] [PubMed] [Google Scholar]
- [71].Singh IN, Sato K, Takashima A, Kanfer JN. Activation of LA-N-2 cell phospholipases by single alanine substitution analogs of amyloid beta peptide (25-35) Febs Letters. 1997;405:65–67. doi: 10.1016/s0014-5793(97)00154-3. [DOI] [PubMed] [Google Scholar]
- [72].Cox DA, Cohen ML. Amyloid beta-induced neurotoxicity is associated with phospholipase D activation in cultured rat hippocampal cells. Neurosci Lett. 1997;229:37–40. doi: 10.1016/s0304-3940(97)00407-2. [DOI] [PubMed] [Google Scholar]
- [73].Brandenburg LO, Konrad M, Wruck C, Koch T, Pufe T, Lucius R. Involvement of formyl-peptide-receptor-like-1 and phospholipase D in the internalization and signal transduction of amyloid beta 1-42 in glial cells. Neuroscience. 2008;156:266–276. doi: 10.1016/j.neuroscience.2008.07.042. [DOI] [PubMed] [Google Scholar]
- [74].Small SA, Gandy S. Sorting through the cell biology of Alzheimer’s disease: intracellular pathways to pathogenesis. Neuron. 2006;52:15–31. doi: 10.1016/j.neuron.2006.09.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [75].Riebeling C, Morris AJ, Shields D. Phospholipase D in the Golgi apparatus. Biochimica Et Biophysica Acta-Molecular and Cell Biology of Lipids. 2009;1791:876–880. doi: 10.1016/j.bbalip.2009.04.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [76].Chen YG, Siddhanta A, Austin CD, Hammond SM, Sung TC, Frohman MA, Morris AJ, Shields D. Phospholipase D stimulates release of nascent secretory vesicles from the trans-Golgi network. J Cell Biol. 1997;138:495–504. doi: 10.1083/jcb.138.3.495. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [77].Ktistakis NT, Brown HA, Sternweis PC, Roth MG. Phospholipase D is present on Golgi-enriched membranes and its activation by ADP ribosylation factor is sensitive to brefeldin A. Proceedings of the National Academy of Sciences of the United States of America. 1995;92:4952–4956. doi: 10.1073/pnas.92.11.4952. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [78].Cai DM, Leem JY, Greenfield JP, Wang P, Kim BS, Wang RS, Lopes KO, Kim SH, Zheng H, Greengard P, Sisodia SS, Thinakaran G, Xu HX. Presenilin-1 regulates intracellular trafficking and cell surface delivery of beta-amyloid precursor protein. Journal of Biological Chemistry. 2003;278:3446–3454. doi: 10.1074/jbc.M209065200. [DOI] [PubMed] [Google Scholar]
- [79].Cai D, Zhong M, Wang R, Netzer WJ, Shields D, Zheng H, Sisodia SS, Foster DA, Gorelick FS, Xu H, Greengard P. Phospholipase D1 corrects impaired betaAPP trafficking and neurite outgrowth in familial Alzheimer’s disease-linked presenilin-1 mutant neurons. Proc Natl Acad Sci U S A. 2006;103:1936–1940. doi: 10.1073/pnas.0510710103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [80].Liu Y, Zhang YW, Wang X, Zhang H, You XQ, Liao FF, Xu HX. Intracellular Trafficking of Presenilin 1 Is Regulated by beta-Amyloid Precursor Protein and Phospholipase D1. Journal of Biological Chemistry. 2009;284:12145–12152. doi: 10.1074/jbc.M808497200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [81].Cai D, Netzer WJ, Zhong M, Lin Y, Du G, Frohman M, Foster DA, Sisodia SS, Xu H, Gorelick FS, Greengard P. Presenilin-1 uses phospholipase D1 as a negative regulator of beta-amyloid formation. Proc Natl Acad Sci U S A. 2006;103:1941–1946. doi: 10.1073/pnas.0510708103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [82].Leong SL, Cappai R, Barnham KJ, Pham L. Cle, Modulation of alpha-synuclein aggregation by dopamine: a review. Neurochem Res. 2009;34:1838–1846. doi: 10.1007/s11064-009-9986-8. [DOI] [PubMed] [Google Scholar]
- [83].Jenco JM, Rawlingson A, Daniels B, Morris AJ. Regulation of phospholipase D2: selective inhibition of mammalian phospholipase D isoenzymes by alpha- and beta-synucleins. Biochemistry. 1998;37:4901–4909. doi: 10.1021/bi972776r. [DOI] [PubMed] [Google Scholar]
- [84].Ahn BH, Rhim H, Kim SY, Sung YM, Lee MY, Choi JY, Wolozin B, Chang JS, Lee YH, Kwon TK, Chung KC, Yoon SH, Hahn SJ, Kim MS, Jo YH, Min DS. alpha-Synuclein interacts with phospholipase D isozymes and inhibits pervanadate-induced phospholipase D activation in human embryonic kidney-293 cells. J Biol Chem. 2002;277:12334–12342. doi: 10.1074/jbc.M110414200. [DOI] [PubMed] [Google Scholar]
- [85].Payton JE, Perrin RJ, Woods WS, George JM. Structural determinants of PLD2 inhibition by alpha-synuclein. J Mol Biol. 2004;337:1001–1009. doi: 10.1016/j.jmb.2004.02.014. [DOI] [PubMed] [Google Scholar]
- [86].Rappley I, Gitler AD, Selvy PE, LaVoie MJ, Levy BD, Brown HA, Lindquist S, Selkoe DJ. Evidence that alpha-synuclein does not inhibit phospholipase D. Biochemistry. 2009;48:1077–1083. doi: 10.1021/bi801871h. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [87].Riebeling C, Bourgoin S, Shields D. Caspase cleavage of phospholipase D1 in vitro alters its regulation and reveals a novel property of the “loop” region. Biochimica Et Biophysica Acta-Molecular and Cell Biology of Lipids. 2008;1781:376–382. doi: 10.1016/j.bbalip.2008.05.007. [DOI] [PubMed] [Google Scholar]
- [88].Jang YH, Ahn BH, Namkoong S, Kim YM, Jin JK, Kim YS, Min do S. Differential regulation of apoptosis by caspase-mediated cleavage of phospholipase D isozymes. Cellular Signalling. 2008;20:2198–2207. doi: 10.1016/j.cellsig.2008.07.010. [DOI] [PubMed] [Google Scholar]
- [89].Ribe EM, Serrano-Saiz E, Akpan N, Troy CM. Mechanisms of neuronal death in disease: defining the models and the players. Biochemical Journal. 2008;415:165–182. doi: 10.1042/BJ20081118. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [90].Raghu P, Coessens E, Manifava M, Georgiev P, Pettitt T, Wood E, Garcia-Murillas I, Okkenhaug H, Trivedi D, Zhang QF, Razzaq A, Zaid O, Wakelam M, O’Kane CJ, Ktistakis N. Rhabdomere biogenesis in Drosophila photoreceptors is acutely sensitive to phosphatidic acid levels. Journal of Cell Biology. 2009;185:129–145. doi: 10.1083/jcb.200807027. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [91].Foster DA. Phosphatidic acid signaling to mTOR: Signals for the survival of human cancer cells. Biochimica Et Biophysica Acta-Molecular and Cell Biology of Lipids. 2009;1791:949–955. doi: 10.1016/j.bbalip.2009.02.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [92].Pei JJ, Hugon J. mTOR-dependent signalling in Alzheimer’s disease. Journal of Cellular and Molecular Medicine. 2008;12:2525–2532. doi: 10.1111/j.1582-4934.2008.00509.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [93].Fang YM, Vilella-Bach M, Bachmann R, Flanigan A, Chen J. Phosphatidic acid-mediated mitogenic activation of mTOR signaling. Science. 2001;294:1942–1945. doi: 10.1126/science.1066015. [DOI] [PubMed] [Google Scholar]
- [94].Toschi A, Lee E, Xu LM, Garcia A, Gadir N, Foster DA. Regulation of mTORC1 and mTORC2 Complex Assembly by Phosphatidic Acid: Competition with Rapamycin. Molecular and Cellular Biology. 2009;29:1411–1420. doi: 10.1128/MCB.00782-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [95].Sun Y, Fang Y, Yoon MS, Zhang C, Roccio M, Zwartkruis FJ, Armstrong M, Brown HA, Chen J. Phospholipase D1 is an effector of Rheb in the mTOR pathway. Proceedings of the National Academy of Sciences of the United States of America. 2008;105:8286–8291. doi: 10.1073/pnas.0712268105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [96].Ha SH, Kim DH, Kim IS, Kim JH, Lee MN, Lee HJ, Kim JH, Jang SK, Suh PG, Ryu SH. PLD2 forms a functional complex with mTOR/raptor to transduce mitogenic signals. Cellular Signalling. 2006;18:2283–2291. doi: 10.1016/j.cellsig.2006.05.021. [DOI] [PubMed] [Google Scholar]
- [97].Lee HY, Jung H, Jang IH, Suh PG, Ryu SH. Cdk5 phosphorylates PLD2 to mediate EGF-dependent insulin secretion. Cellular Signalling. 2008;20:1787–1794. doi: 10.1016/j.cellsig.2008.06.009. [DOI] [PubMed] [Google Scholar]
- [98].Rauch C, Loughna PT. C2C12 skeletal muscle cells exposure to phosphatidylcholine triggers IGF-1 like-responses. Cellular Physiology and Biochemistry. 2005;15:211–224. doi: 10.1159/000086408. [DOI] [PubMed] [Google Scholar]
- [99].Raghu P, Manifava M, Coadwell J, Ktistakis NT. Emerging findings from studies of phospholipase D in model organisms (and a short update on phosphatidic acid effectors) Biochimica Et Biophysica Acta-Molecular and Cell Biology of Lipids. 2009;1791:889–897. doi: 10.1016/j.bbalip.2009.03.013. [DOI] [PubMed] [Google Scholar]
- [100].Monovich L, Mugrage B, Quadros E, Toscano K, Tommasi R, LaVoie S, Liu E, Du Z, LaSala D, Boyar W, Steed P. Optimization of halopemide for phospholipase D2 inhibition. Bioorg Med Chem Lett. 2007;17:2310–2311. doi: 10.1016/j.bmcl.2007.01.059. [DOI] [PubMed] [Google Scholar]
- [101].Su W, Yeku O, Olepu S, Genna A, Park JS, Ren HM, Du G, Gelb MH, Morris AJ, Frohman MA. 5-Fluoro-2-indolyl des-chlorohalopemide (FIPI), a Phospholipase D Pharmacological Inhibitor That Alters Cell Spreading and Inhibits Chemotaxis. Molecular Pharmacology. 2009;75:437–446. doi: 10.1124/mol.108.053298. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [102].Scott SA, Selvy PE, Buck JR, Cho HP, Criswell TL, Thomas AL, Armstrong MD, Arteaga CL, Lindsley CW, Brown HA. Design of isoform-selective phospholipase D inhibitors that modulate cancer cell invasiveness. Nature Chemical Biology. 2009;5:108–117. doi: 10.1038/nchembio.140. [DOI] [PMC free article] [PubMed] [Google Scholar]