Abstract
Background
Gene expression studies can be used to help identify disease-associated genes by comparing the levels of expressed transcripts between cases and controls, and to identify functional genetic variants (expression quantitative loci or eQTLs) by comparing expression levels between individuals with different genotypes. While many of these studies are performed in blood or lymphoblastoid cell lines due to tissue accessibility, the relevance of expression differences in tissues that are not the primary site of disease is unclear. Further, many eQTLs are tissue specific. Thus, there is a clear and compelling need to conduct gene expression studies in tissues that are specifically relevant to the disease of interest. One major technical concern about using autopsy-derived tissue is how representative it is of physiologic conditions, given the effect of postmortem interval on tissue degradation.
Results
In this study, we monitored the gene expression of 13 tissue samples harvested from a rapid autopsy heart (non-failed heart) and 7 from a cardiac explant (failed heart) through 24 hours of autolysis. The 24 hour autopsy simulation was designed to reflect a typical autopsy scenario where a body may begin cooling to ambient temperature for ~12 hours, before transportation and storage in a refrigerated room in a morgue. In addition, we also simulated a scenario wherein the body was left at room temperature for up to 24 hours before being found. A small fraction (< 2.5%) of genes showed fluctuations in expression over the 24 hr period and largely belong to immune and signal response and energy metabolism-related processes. Global expression analysis suggests that RNA expression is reproducible over 24 hours of autolysis with 95% genes showing < 1.2 fold change. Comparing the rapid autopsy to the failed heart identified 480 differentially expressed genes, including several types of collagens, lumican (LUM), natriuretic peptide A (NPPA) and connective tissue growth factor (CTGF), which allows for the clear separation between failing and non-failing heart based on gene expression profiles.
Conclusions
Our results demonstrate that RNA from autopsy-derived tissue, even up to 24 hours of autolysis, can be used to identify biologically relevant expression pattern differences, thus serving as a practical source for gene expression experiments.
Background
Gene expression studies monitor the simultaneous transcription levels of genes, and thus bridge the gap between static genomic information and dynamic phenotypes. These studies are used to identify differentially expressed transcripts with respect to measured variables of interest, such as differing environments, treatments, phenotypes, or clinical outcomes, and for the identification of expression quantitative trait loci (eQTL), i.e. genomic regions whose genotype is correlated with the RNA expression in a panel of genetically diverse individuals [1]. Studies of gene expression abnormalities in various disorders often involve the use of postmortem tissue [2]. The factors affecting RNA stability or integrity in postmortem tissues have been most extensively studied in the neuropsychiatric area. Despite the success with identification of sets of differentially expressed genes specific to neuropsychiatric disorders, one can find continued concerns regarding quality and consistency of postmortem RNA analysis [3]. Prior studies in brain have indicated that specific agonal factors, including coma and hypoxia, markedly affect the integrity of RNA, whereas postmortem factors have relatively minor effects on RNA integrity [4]. Of the postmortem factors, pH is noted as a factor of importance for RNA quality in some publications [5].
Recent studies in human skeletal and heart tissues provide examples of variation in protein and enzyme activity, with examples of reduced NOS3 mRNA levels associated with higher postmortem interval (PMI) [6]. These alterations appear to occur more rapidly at higher temperatures, and storing samples at 4°C may delay some of the postmortem changes [7]. There has been a lingering debate over the importance of PMI on RNA degradation. Thus, in addition to variation in expression levels due to technical [8], environmental [9], demographic [10], and genetic factors [11], concerns arise as RNA integrity is affected by various pre- and postmortem events [12], which need to be addressed when quantitative analyses are performed.
These sources of variation raise a major technical concern about the utility of autopsy-derived tissue and how representative it is of physiologic conditions, due to the of the effect of PMI on tissue degradation. In this study we specifically test the utility of devitalized cardiac tissue for gene expression, obtained through both a cardiac explant (failed heart) and a rapid-autopsy program (non-failed heart). We systematically evaluate the variability of RNA message in devitalized tissue in a time dependent manner in an attempt to simulate general autopsy conditions.
Assessing the factors regulating RNA integrity is important to determine the reliability of gene expression patterns in cardiac tissue. Further, the existence of significant biological variability in expression levels of many genes between human hearts is widely recognized. Therefore, in this study we establish the extent of variation as a consequence of autolysis, and secondarily, generate a gene expression profile of a failing human heart.
Methods
Sample Collection
Cardiac tissue was obtained from an explanted failing heart via cardiac transplantation and from a "normal" heart through a metastatic cancer rapid-autopsy program. This study was approved by the IRB of The Johns Hopkins Hospital and all samples provided informed consent. Total RNA was extracted from the tissues using the RNeasy Fibrous Tissue Mini kit (Qiagen) according to the manufacture's protocol.
We obtained left and right ventricle tissue harvested under conditions that simulated general autopsy conditions. The baseline time-point (deemed the 0 hour time-point) for the explanted failing heart was harvested within 1 hour of explantation, while the autopsy heart was harvested 3-4 hours after the patient died. After rapidly collecting an initial 0 hour time-point to establish a baseline measure of gene expression, the hearts were placed in a 37°C incubator that was then turned off and allowed to cool at ~1°C/hr to reach room temperature ~12 hours later. After reaching ambient temperature, 6 heart samples were further cooled to 4°C (cold-24 hour autolysis) and 9 were left at room temperature (warm-24 hour autolysis). These two conditions represent two alternate scenarios of tissue harvesting. The first scenario simulates an autopsy where a body may begin cooling to ambient temperature for ~12 hours, before transportation and storage in a refrigerated room in a morgue which accelerates body cooling. The second we consider a "worst case" scenario where the body is not cooled. At each harvesting time point, tissue was flash frozen in liquid nitrogen or placed in RNAlater from the left and right ventricle free wall. Twenty tissue samples were harvested from each heart at time points 0, 6, 12, 18 (cold and warm) and 24 (cold and warm) hours (Figure 1). The RNA Integrity numbers (RIN) of all the 20 samples were obtained from the Agilent Bioanalyzer 2100 to assess the RNA quality of the samples [13]. Gene expression profiles were determined at time points 0, 6, 12, 18 and 24 hours in 13 tissue samples harvested from the autopsy heart. Profiles were obtained at time points 0, 6, 12, 18 hours in 7 tissue samples from the explant heart.
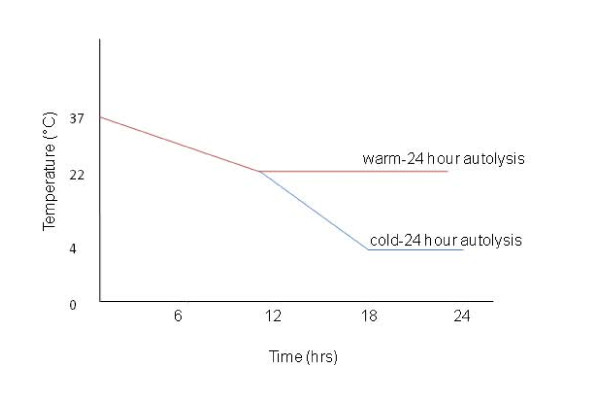
Figure 1.
Simulated autopsy conditions. All heart samples were initially placed in a 37°C incubator that was then turned off and allowed to cool ~1°C/hr to reach room temperature ~12 hours later. Thereafter, to simulate cold and warm-24 hour autolysis tissue harvesting conditions, some hearts were cooled at 2-3°C/hr (to 4°C) (blue line) and others were left at room temperature (red line).
Microarray Expression Analysis
RNA from the 20 samples, along with a HeLa control run in triplicate, were run on the Affymetrix Exon arrays according to the manufacturer's protocol. Array data for this study has been deposited in the National Center for Biotechnology Information's Gene Expression Omnibus (GEO) with accession number GSE32519. Exon arrays have 1.4 million probesets that map to over a million exon clusters. These feature probesets target each transcript throughout, rather than simply at the 3' end, allowing for robust identification of differentially expressed genes [14]. We analyzed expression data from ~200,000 core probesets that are supported by putative mRNAs from the RefSeq database. Signal processing was performed after quantile normalization and median polish summarization of the PM (Perfect Match) probes by using the Affymetrix power tools and the Robust Multi-Array Analysis (RMA) algorithm [15]. Probes with low p values (p < 0.05) have signals that are distinct from background and are considered to be detected above background (DABG). After filtering to retain probes flagged as present (DABG < 0.05) in at least one-fourth of the samples, 162,145 probes remained, which mapped to ~15,000 transcripts. Exon-based and intron-based probesets from putative housekeeping genes serve as positive and negative control sets, respectively. These signals are used to produce an "area under the curve" (AUC) value in a receiver-operating characteristic (ROC) curve as a measure of separation between the positive and negative controls, which is indicative of the quality of hybridization [16]
GNF Expression Atlas 2 [17] allows for the classification of genes depending on their expression in the 79 human organs and tissues covered by the Atlas. Approximately, 8,400 genes expressed in the heart in the GNF Expression Atlas were downloaded via the Biomart tool.
Statistical Analysis
All statistical analysis was performed using R (version 2.9.2). Principal component analysis was performed using the function "prcomp" in the "stats" package of R. Two-class Significance Analysis of Microarrays (SAM) [18] was used to identify genes that were differentially expressed, which is implemented as the "samr" package in R. SAM calculates the test statistic for relative difference in gene expression based on permutation analysis of expression data and calculates a false discovery rate, which gives the proportion of significant genes most likely identified by chance. We used the two class unpaired test statistic in SAM. The two class unpaired test is similar to a between subjects t-tests wherein mean expressions in two groups are determined significant based on the calculated statistic, d, from the original and permuted data for each gene. In this study a false discovery rate (FDR; q < 5%) was used to determine the differential genes between the two classes.
Functional Analysis
Genes were functionally annotated and classified by using the functional annotation tool of database for annotation, visualization, and integrated discovery (DAVID) [19]. The annotations were selected from Gene Ontology's "biological process". The annotations have been selected based on medium stringency (default) setting, with > 2-fold enrichment and Benjamini-Hochberg p-value < 0.05. The fold enrichment for a particular functional category describes the ratio between the number of genes in the gene list belonging to that category and the total number of genes in the gene list, which has at least one functional annotation. This ratio is then compared to the ratio between the total number of genes in that category and the total number of genes in the human genome with at least one functional annotation. It has been observed that it can be difficult to come up with a clear picture of gene function given all the Gene Ontology (GO) terms annotating the genes. Therefore, all the significantly enriched GO categories were assigned their corresponding parent GO Slim annotation. The GO Slims are reduced versions of GO containing high-level, broader parent terms in the whole GO [20] (http://www.geneontology.org/GO.slims.shtml). This is feasible with GO because of the hierarchical parent and child relationships recorded between the terms. Note that we only use the GO Slim terms of biological process.
Results and Discussion
We obtained the expression profiles of 7 cardiac tissue samples from the cardiac transplant operation (explant/failed heart) and 13 from a rapid-autopsy program (autopsy/non-failed heart) harvested under the "cold-" and "warm-24 hour autolysis" conditions (Figure 1). The left and right ventricular tissues showed highly correlated gene expression in both the heart samples (Pearson's coefficient, r > 0.94), and we therefore analyzed the heart regions together. To assess the expression array performance for each sample, we used the ROC AUC summarization score, which is a measure of overall sensitivity and specificity of the expression profile. The AUC scores for the 20 gene expression profiles harvested at increasing autolysis delay all ranged from 0.80 to 0.90, which is within the parameters suggested by Affymetrix [16]. While RIN values are typically used to predict sample performance, we only observed a weak correlation between RIN values and ROC AUC (r2 = 0.27, Figure 2), and note that even samples with RIN values as low as 2.7 displayed robust hybridization. The means of the RIN values in the "warm" and "cold" 24 hour autolysis samples were 5.1 (SD = 1.5) and 6.9 (SD = 1.5), respectively, though this difference was not statistically significant (P = 0.11) (Additional File 1, Figure S1). We further observed that the RIN values were independent of the autolysis time (r2 = 0.002, Figure 3). While this observation is based on a small dataset and is at odds with many, but not all, reports of RIN quality correlating with post mortem interval [21], we note that in our analyses, RIN was not a strong predictor of array performance.
Figure 2.
RNA quality and hybridization performance. The RIN number, a measure of RNA quality, ranged from 2.7-8.6 in the heart samples. The ROC AUC, a hybridization quality metric, ranged from 0.80 to 0.90, which is within the parameters suggested by Affymetrix. All samples showed robust hybridization irrespective of their RIN values (r2 = 0.27).
Figure 3.
Autolysis interval and RNA quality. The autolysis interval, which ranged from 0 to 24 hours, did not correlate with RNA integrity, measured as RIN numbers (r2 = 0.002).
Autolysis Stability of Gene Expression
As an additional general test of expression array performance with autolysed tissue, we used principal component analysis (PCA) of gene intensities to cluster the 20 autopsy and explant heart samples, along with Hela controls run in triplicate, as well as reference liver and heart data obtained from Affymetrix. Irrespective of the time and temperature of tissue harvest, the expression profiles of the two hearts could clearly be differentiated from non-heart tissues, as observed by the PCA analysis of ~15,000 gene intensities (Figure 4). In Figure 5, we demonstrate that there is a strong correlation between the baseline explant and autopsy hearts compared to their later timepoints (Pearson's correlation ranges from 0.92-0.98). As a comparison, the Pearson's correlation(r), between any two HeLa controls (technical replicates) was 0.98. Additionally, the Pearson's coefficient for HeLa controls and reference liver with respect to a high RIN heart (RIN = 8.6) was 0.65 and 0.66, respectively.
Figure 4.
Principal component analysis (PCA) of heart and non-heart gene expression intensities. A scatterplot of the PC1 and PC2 from the PCA analysis of gene intensities of ~15,000 transcripts from the 20 autopsy and explant heart samples, along with reference triplicate HeLa controls and triplicate heart and liver data (obtained from Affymetrix), clearly separates the samples into heart and non-heart spaces.
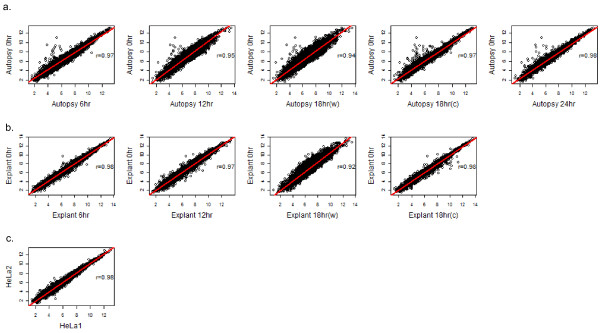
Figure 5.
Correlation between baseline and autolyzed gene expression. a and b. Scatterplots of expression intensity later time-points (x-axis) and the 0 hour time-point (y-axis) for (a) autopsy and (b) explant heart tissues. Red lines represent the linear regression line indicating relationship between expression profiles. Pearson's correlation (r), between the baseline and later timepoints, range from 0.92-0.98. c. Scatterplot of expression intensities between two technical replicates (HeLa Controls) with Pearson's correlation, (r) of 0.98.
The concordance in expression profiles in autopsy tissue was also investigated by pair-wise comparisons of all cardiac expressed genes, expressed as the fold change between time-points, (Figure 6). Specifically, 8,400 genes expressed in the cardiac tissue were retrieved from GNF Expression Atlas 2, which summarizes the expression patterns of human, mouse and rat genes in several selected tissues using whole-genome microarray experiments. To establish a baseline level of expected concordance for technical replicates, we compared expression profile of two HeLa controls, and observed 94.9% of genes with ≤1.25 fold change in expression (red line, Figure 6). The baseline explant and autopsy and their later timepoints, showed similar concordance in expression and we saw no difference from technical replicates (95.9% and 93.4% genes with ≤1.25 fold change, respectively) (purple and brown lines, Figure 6). We observed a difference between the autopsy and explant expression profiles, with only 86.8% genes with ≤1.25 fold change (black line, Figure 6). Comparing, either heart sample to the HeLa controls demonstrated a striking difference in expression, with only 48% with ≤1.25 fold change (green and blue lines, Figure 6). Thus, global gene expression analysis suggests that RNA expression is reproducible over 24 hours of autolysis and we could distinguish between heart and non-heart samples.
Figure 6.
Concordance in expression profiles in heart tissues. Pair-wise comparisons of all cardiac expressed genes (~8,400 based on the GNF anatomical system data: heart), expressed as the fold change between time-points. To establish a baseline level of expected concordance for technical replicates, we compared the expression profile of two HeLa controls, and observed 94.9% of genes with ≤1.25 fold change (red line). Comparing the baseline explant and autopsy to later timepoints, we saw no difference from technical replicates (95.9% and 93.4% ≤1.25 fold change, respectively) (purple and brown lines). However, comparing the autopsy to the explant, we saw a measurable difference (86.8% ≤1.25 fold change) (black line), and comparing either heart sample to the HeLa controls demonstrated a striking difference (48% ≤1.25 fold change) (green and blue lines).
Autolysis Fluctuations of Gene Expression
After, establishing that global expression is largely unaffected by autolysis interval, we looked at the small fraction of genes that were sensitive to autolysis conditions. We selected transcripts with > 2 fold-change (2-FC) between the baseline and later timepoints from each heart to evaluate the change in gene expression during the 24 hours of autolysis in a time dependent manner. For this analysis, we focused on the "cold-24 h autolysis", as this represents a typical autopsy scenario. Only 2.25% (345/15320) of transcripts from the autopsy hearts had a > 2-FC when we compared the transcript levels from the 4 later time-points (6, 12, 18, 24) to the baseline levels. We queried these genes in the DAVID annotation database to find significantly overrepresented GO terms. Table 1, provides the functional categories overrepresented with a > 2-fold enrichment, Benjamini-Hochberg p-value < 0.05 and the broader parent GO Slim term. There was significant overrepresentation of GO terms which associated with two GO Slim categories 1) "cellular metabolism" (76/354) and 2) "response to stimulus" (42/354).
Table 1.
Annotations overrepresented in 2.25% genes fluctuating in the autopsy (non-failed) heart
| Terma | GO Gene Count | GO Slim | GO Slim Gene Count |
|---|---|---|---|
| response to external stimulus | 42 | ||
| response to wounding | 27 | response to stimulus | 42 |
| inflammatory response | 19 | ||
| lipid metabolic process | 41 | ||
| organic acid metabolic process | 33 | ||
| carboxylic acid metabolic process | 33 | ||
| oxoacid metabolic process | 33 | ||
| cellular lipid metabolic process | 30 | ||
| fatty acid metabolic process | 16 | ||
| oxidation reduction | 31 | ||
| alcohol metabolic process | 22 | cellular metabolism | 76 |
| energy derivation by oxidation of organic compounds | 12 | ||
| carbohydrate biosynthetic process | 10 | ||
| generation of precursor metabolites and energy | 18 | ||
| monocarboxylic acid metabolic process | 25 | ||
| cellular ketone metabolic process | 33 | ||
aAll functional annotations are significantly overrepresented with > 2-Fold Enrichment and Benjamini-Hochberg p-value < 0.05
We performed a similar experiment with the explanted heart, to see which categories of genes fluctuate in response to autolysis under the same conditions. One percent (154/15320) of transcripts from the explanted hearts had a > 2-FC, when we compared the fold changes in the transcripts from the 3 later time-points (6, 12, 18) to the baseline. Significantly overrepresented functional categories from the DAVID annotation database and the associated broader GO Slim parent terms are shown in Table 2. In the explant heart the significant enrichment of GO terms associated with the "response to stimulus" category was observed annotating 44/154 genes. Three other significant functional categories were mapped to "circulatory system process" (8/154), "development" (15/154) and "cell differentiation" (27/154). Thus, the broad parent GO Slim term "response to stimulus" was significantly overrepresented in both the autopsy and explant hearts, including 21 genes in common (Additional File 2, Table S1), in this analysis of variation in gene expression during 24 hours of autolysis.
Table 2.
Annotations overrepresented in 1% genes fluctuating in the explant (failed) heart
| Terma | GO Gene Count | GO Slim | GO Slim Gene Count |
|---|---|---|---|
| circulatory system process | 8 | circulatory system process | 8 |
| response to external stimulus | 23 | ||
| regulation of defense response | 9 | ||
| regulation of response to stimulus | 15 | ||
| response to chemical stimulus | 26 | ||
| regulation of inflammatory response | 7 | ||
| response to cytokine stimulus | 7 | ||
| response to corticosteroid stimulus | 7 | ||
| response to organic substance | 18 | ||
| response to extracellular stimulus | 10 | ||
| response to mechanical stimulus | 6 | ||
| response to endogenous stimulus | 13 | ||
| response to stress | 29 | ||
| response to inorganic substance | 9 | response to stimulus | 44 |
| regulation of response to external stimulus | 8 | ||
| response to drug | 9 | ||
| response to glucocorticoid stimulus | 6 | ||
| response to lipopolysaccharide | 6 | ||
| regulation of response to stress | 10 | ||
| response to molecule of bacterial origin | 6 | ||
| response to hormone stimulus | 11 | ||
| response to steroid hormone stimulus | 8 | ||
| response to nutrient levels | 8 | ||
| response to bacterium | 8 | ||
| transforming growth factor beta receptor signaling pathway | 5 | ||
| skeletal system development | 15 | development cell | 15 |
| cell differentiation | 27 | differentiation | 27 |
aAll functional annotations are significantly overrepresented with > 2-Fold Enrichment and Benjamini-Hochberg p-value < 0.05
Gene Expression Profile for a Failing Heart
Having established that the global expression from the individual cardiac tissues is reproducible during 24 hours of autolysis, we next compared the expression profile of the ventricular cardiac tissue from the explant (failing heart) and the autopsy (non-failing heart) by using SAM's two class unpaired test statistic. We focused on the "cold-24 hour autolysis" heart samples, as representative of real-world autopsy conditions. Gene expression profiles from 8 autopsy and 5 explant heart samples were analyzed with the SAM technique, which identified ~480 differentially expressed (DE) transcripts at > 2-FC and q-values < 0.5. The heatmap in Figure 7 demonstrates ~480 transcripts with > 2-FC expression between the two groups with 374 up-regulated and 108 down-regulated transcripts in the explant/failed heart. The list of ~480 genes differentially up- and down-regulated between the explant and autopsy hearts are given in the Additional File 3, Table S2. The results were unchanged after exclusion of the samples harvested at the baseline 0 hour time-point.
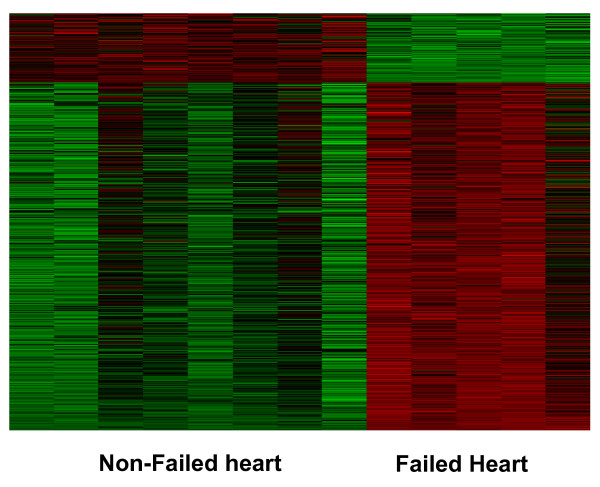
Figure 7.
Differential gene expression. Heatmap of transcripts with > 2 fold change in expression between the failing and non-failing heart; 374 genes are up-regulated and 108 down-regulated. (red = up-regulated, and green = down-regulated)
Clinical evidence clearly demonstrates the dynamic nature of the components of the extracellular matrix in response to mechanical unloading of the failing heart [22,23] and a quantitative increase in collagen subtypes [24]. We identified ECM proteins collagen I alpha 1 (COL1A1), collagen III alpha 1 (COL3A1) and fibronectin (FN1) to be upregulated > 2-FC in the failing heart. Importantly, we identified natriuretic peptide A (NPPA) which was 11-fold higher in the failing heart. NPPA is part of a conserved adaptive change in molecular phenotype in response to heart failure and serves as both a diagnostic and potential therapeutic marker [25,26]. Recently, higher than normal levels of osteoglycin (OGN, 12-fold increase in the failing heart) were associated with the heart becoming enlarged in rats and mice and in humans [27]. Periostin, (POSTN, 8-fold increase in the failing heart) has also been implicated in cardiac remodeling and therefore heart failure [28]. Additionally, the expression of connective tissue growth factor (CTGF, 4 fold change), transforming growth factor-beta (TGFB2, 3.18 fold change), brain natriuretic peptide precursor (NPPB, 2.29 fold change), Mu-crystallin (CRYM, 2.8 fold change) and clusterin (CLU, 2.5 fold change), were all elevated in the failing heart, consistent with previous reports [29-34]. The expression of Alpha-1-antichymotrypsin (SERPINA3), a known protease inhibitor, responsible for degradation or disassembly of myocardial proteins, was down regulated in the failing heart when compared to the non-failing heart (8 fold decrease), which is consistent with the report by Yang et al [35].
We also queried with the DAVID database to look for enrichment of functions from GO biological processes represented by the DE genes. The significantly over-represented functional annotations (> 2-fold enrichment and Benjamini-Hochberg p-value < 0.05) and the associated GO Slim terms are listed in Tables 3 and 4. Seventy-nine of the 374 up-regulated genes in the failing heart were annotated into overlapping overrepresented GO functional categories, with the associated GO slim terms "cellular metabolism" and "response to stimulus" annotated for 29/374 and 23/374 genes, respectively (Table 3). Fifty-nine of the 108 down-regulated genes belonged to overlapping functionally overrepresented GO categories which related to parent GO Slim terms "response to stimulus" (49/108), "regulation of biological quality" (23/108), "cellular process" (9/108), and "transport" (5/108) (Table 4). Given that we observed an enrichment of genes involved in "response to stimulus" in the analysis of genes that fluctuate during autolysis, we investigated whether the ~480 DE genes between the failing and non-failing heart included genes identified as fluctuating during autolysis. We observed that 13% of the DE genes showed temporal fluctuation in the autopsy hearts, while 9% showed fluctuation in the explants, including important marker genes like NPPA, NPPB and POSTN. We explored how fluctuation of these genes during autolysis influenced the comparison of gene expression between failed and non-failed heart. In all three cases, these genes were clearly able to distinguish failed from non-failed heart at all time points, with the exception of NPPB at the 12 hr timepoint (Figure 8). Thus, despite > 2FC fluctuations in temporal expression in some important genes due to autolysis, it is still possible to clearly identify specific and relevant differentially expressed genes in autolyzed cardiac tissue.
Table 3.
Annotations overrepresented in the up-regulated genes in the explant (failed) heart
| Terma | GO Gene Count | GO Slim | GO Slim Gene Count |
|---|---|---|---|
| cellular ketone metabolic process | 29 | cellular metabolism | 29 |
| extracellular structure organization | 15 | extracellular structure organization and biogenesis | 15 |
| muscle contraction | 12 | muscle system process | 12 |
| regulation of cell adhesion | 13 | regulation of biological process | 13 |
| enzyme linked receptor protein signaling pathway | 23 | response to stimulus | 23 |
aAll functional annotations are significantly overrepresented with > 2-Fold Enrichment and Benjamini-Hochberg p-value < 0.05
Table 4.
Annotations overrepresented in the down-regulated genes in the explant (failed) heart
| Terma | GO Gene Count | GO Slim | GO Slim Gene Count |
|---|---|---|---|
| cellular ion homeostasis | 9 | cellular process | 9 |
| regulation of biological quality | 27 | regulation of biological quality | 27 |
| defense response | 22 | ||
| response to stimulus | 48 | ||
| response to chemical stimulus | 22 | ||
| immune system process | 19 | ||
| response to stress | 34 | ||
| immune response | 14 | ||
| regulation of response to stimulus | 12 | ||
| response to external stimulus | 26 | ||
| response to bacterium | 7 | ||
| response to wounding | 20 | response to stimulus | 49 |
| regulation of immune response | 9 | ||
| regulation of defense response | 7 | ||
| inflammatory response | 16 | ||
| blood coagulation | 6 | ||
| innate immune response | 10 | ||
| positive regulation of defense | 5 | ||
| response | 9 | ||
| humoral immune response | 6 | ||
| complement activation, classical pathway | |||
| transition metal ion transport | 5 | transport | 5 |
aAll functional annotations are significantly overrepresented with > 2-Fold Enrichment and Benjamini-Hochberg p-value < 0.05
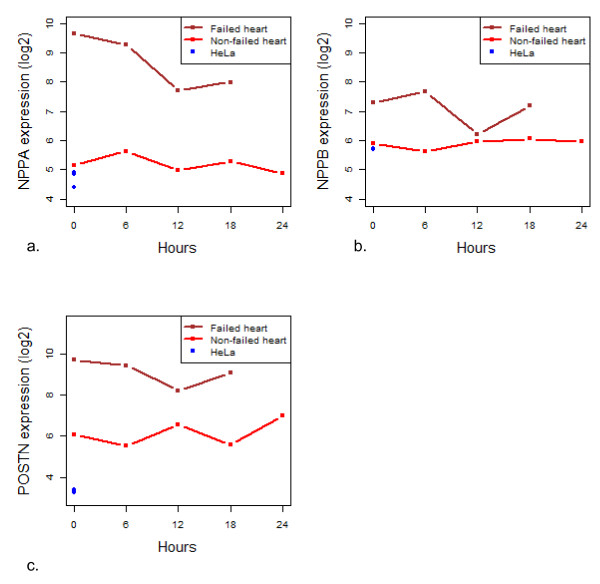
Figure 8.
Differentially expressed and fluctuating genes. Expression profile of (a) NPPA, (b) NPPB and (c) POSTN genes in the failing and non-failing hearts. The genes are differentially expressed and show fluctuation in expression in the failed heart.
Conclusion
In this study we investigated the gene expression of cardiac tissues in a simulated autopsy experiment. We allowed the heart tissue to autolyze for 24 hours, either at ambient temperature or shifted to 4°C after 12 hours. We chose a limit of 24 hours as we have shown previously that protein expression remains intact for 24 hours by immunohistochemistry, but degradation is noticeable at 48 hours [36]. Thus our findings are applicable to a 24 hour time course and we cannot speculate on late harvesting. We note that, in our experience, 24 hours is a reasonable amount of time to conduct a standard autopsy and is well within the time achievable by organ procurement organizations (OPOs) [37]. The vast majority of genes expressed are stable despite extended autolysis and reduced RNA integrity, indicating that multiple-probe based exon arrays are suited to reliably determine gene expression profiles. We found that most molecular processes are not subject to variation due to tissue procurement time and/or tissue storage temperature. Global expression analysis suggests that RNA expression is reproducible over 24 hours of autolysis with 95% genes showing < 1.2 fold change. As a proof of principle, we used SAM analysis to identify a list of 480 differentially expressed genes, including several types of collagens, lumican (LUM), natriuretic peptide A (NPPA) and connective tissue growth factor (CTGF), which allow for the clear separation between the failing and non-failing heart expression profiles - irrespective of autolysis time.
Our study is consistent with those that suggest that autolysis has relatively minor effects on RNA integrity. Only a small fraction (< 2.5%) of transcripts fluctuate over the 24 hours of autolysis, enriched in functional categories (energy metabolism, immune and signal responses) that are known to be the most adaptable and responsive biological processes and often highly variable between biological specimens [38]. Thus, our results demonstrate that RNA from autopsy-derived tissue, even up to 24 hours of autolysis, can be used to identify biologically relevant expression pattern differences, serving as a practical source for gene expression and eQTL experiments.
Authors' contributions
SG carried out the analysis and drafted the manuscript. MKH obtained samples, contributed to the design of the experiment and helped in the draft of the manuscript; GMH extracted the RNA and conducted the experiments. DEA supervised the project, contributed to the design and coordination and helped draft the manuscript. All authors contributed to manuscript preparation and all approved the final manuscript.
Supplementary Material
Figure S1: RIN range in "warm- and "cold-24 hour autolysis" samples. The range of the RIN values in the warm-24 autolysis samples was 2.7 - 8.6, while the range in the cold- 24 autolysis samples was 3.5 - 8.6. The means of the RIN values in the "warm" and "cold" 24 hour autolysis samples were 5.1 (SD 1.53) and 6.9 (SD 1.45), respectively, and this difference was not statistically significant (P = 0.11).
Table S1: Postmortem fluctuating genes common in autopsy and explant hearts. The gene symbol and description of the 21 genes, which showed variation in gene expression during 24 hours of autolysis in both the autopsy and explant hearts.
Table S2: List of 374 upregulated (Table S2a)and 108 downregulated (Table S2b) genes in the explant/failed heart. List of ~480 genes differentially up- and down-regulated between the explant and autopsy hearts. Among these differentially expressed genes, 374 genes were upregulated and 108 genes were downregulated in the explant/failed heart.
Contributor Information
Simone Gupta, Email: sgupta30@jhmi.edu.
Marc K Halushka, Email: mhalush1@jhmi.edu.
Gina M Hilton, Email: hiltgm0@wfu.edu.
Dan E Arking, Email: arking@jhmi.edu.
Acknowledgements
This work was supported by a grant from the Simons Foundation (SFARI 137603 to DEA). MKH was supported by a Clinician Scientist Award from the Doris Duke Charitable Foundation Grant #2009040.
References
- Michaelson JJ, Loguercio S, Beyer A. Detection and interpretation of expression quantitative trait loci (eQTL) Methods. 2009;48:265–276. doi: 10.1016/j.ymeth.2009.03.004. [DOI] [PubMed] [Google Scholar]
- Li JZ, Vawter MP, Walsh DM, Tomita H, Evans SJ, Choudary PV, Lopez JF, Avelar A, Shokoohi V, Chung T, Mesarwi O, Jones EG, Watson SJ, Akil H, Bunney WE Jr, Myers RM. Systematic changes in gene expression in postmortem human brains associated with tissue pH and terminal medical conditions. Hum Mol Genet. 2004;13:609–616. doi: 10.1093/hmg/ddh065. [DOI] [PubMed] [Google Scholar]
- Weinberger DR. Genetic mechanisms of psychosis: in vivo and postmortem genomics. Clin Ther. 2005;27(Suppl A):8–15. doi: 10.1016/j.clinthera.2005.07.016. [DOI] [PubMed] [Google Scholar]
- Tomita H, Vawter MP, Walsh DM, Evans SJ, Choudary PV, Li J, Overman KM, Atz ME, Myers RM, Jones EG, Watson SJ, Akil H, Bunney WE Jr. Effect of agonal and postmortem factors on gene expression profile: quality control in microarray analyses of postmortem human brain. Biol Psychiatry. 2004;55:346–352. doi: 10.1016/j.biopsych.2003.10.013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stan AD, Ghose S, Gao XM, Roberts RC, Lewis-Amezcua K, Hatanpaa KJ, Tamminga CA. Human postmortem tissue: what quality markers matter? Brain Res. 2006;1123:1–11. doi: 10.1016/j.brainres.2006.09.025. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Partemi S, Berne PM, Batlle M, Berruezo A, Mont L, Riuró H, Ortiz JT, Roig E, Pascali VL, Brugada R, Brugada J, Oliva A. Analysis of mRNA from human heart tissue and putative applications in forensic molecular pathology. Forensic Sci Int. 2010;203:99–105. doi: 10.1016/j.forsciint.2010.07.005. [DOI] [PubMed] [Google Scholar]
- Tavichakorntrakool R, Prasongwattana V, Sriboonlue P, Puapairoj A, Pongskul J, Khuntikeo N, Hanpanich W, Yenchitsomanus PT, Wongkham C, Thongboonkerd V. Serial analyses of postmortem changes in human skeletal muscle: A case study of alterations in proteome profile, histology, electrolyte contents, water composition, and enzyme activity. Proteomics Clin Appl. 2008;2:1255–1264. doi: 10.1002/prca.200800051. [DOI] [PubMed] [Google Scholar]
- Kerr MK, Martin M, Churchill GA. Analysis of variance for gene expression microarray data. J Comput Biol. 2000;7:819–837. doi: 10.1089/10665270050514954. [DOI] [PubMed] [Google Scholar]
- Holter NS, Mitra M, Maritan A, Cieplak M, Banavar JR, Fedoroff NV. Fundamental patterns underlying gene expression profiles: simplicity from complexity. Proc Natl Acad Sci USA. 2000;97:8409–8414. doi: 10.1073/pnas.150242097. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rodwell GE, Sonu R, Zahn JM, Lund J, Wilhelmy J, Wang L, Xiao W, Mindrinos M, Crane E, Segal E, Myers BD, Brooks JD, Davis RW, Higgins J, Owen AB, Kim SK. A transcriptional profile of aging in the human kidney. PLoS Biol. 2004;2:2191–2201. doi: 10.1371/journal.pbio.0020427. [DOI] [PMC free article] [PubMed] [Google Scholar]
- DeRisi JL, Iyer VR, Brown PO. Exploring the metabolic and genetic control of gene expression on a genomic scale. Science. 1997;278:680–686. doi: 10.1126/science.278.5338.680. [DOI] [PubMed] [Google Scholar]
- Barton AJ, Pearson RC, Najlerahim A, Harrison PJ. Pre- and postmortem influences on brain RNA. J Neurochem. 1993;61:1–11. doi: 10.1111/j.1471-4159.1993.tb03532.x. [DOI] [PubMed] [Google Scholar]
- Copois V, bibeau F, Bascoul-Mollevi C, Salvetat N, Chabos P, Bareli C, Candeil L, Fraslon C, Conseiller E, Granci V, Maziere P, Kramar A, Ychou M, Pau B, Martineau P, Molina F, Del Rio M. Impact of RNA degradation on gene expression profiles: Assessment of different methods of reliability determines RNA quality. Journal of Biotechnology. 2007;127:549–559. doi: 10.1016/j.jbiotec.2006.07.032. [DOI] [PubMed] [Google Scholar]
- Lockstone HE. Exon array data analysis using Affymetrix power tools and R statistical software. Brief Bioinform. 2011. [DOI] [PMC free article] [PubMed]
- Bolstad BM, Irizarry RA, Astrand M, Speed TP. A comparison of normalization methods for high density oligonucleotide array data based on variance and bias. Bioinformatics. 2003;19:185–193. doi: 10.1093/bioinformatics/19.2.185. [DOI] [PubMed] [Google Scholar]
- Affymetrix. Quality Assessment of Exon and Gene Arrays. Affymetrix Whitepaper 2005. http://media.affymetrix.com/support/technical/whitepapers
- Su A, Wiltshire T, Batalov S, Lapp H, Ching K, Block D, Zhang J, Soden R, Hayakawa M, Kreiman G, Cooke M, Walker J, Hogenesch J. A gene atlas of the mouse and human protein-encoding transcriptomes. Proc Natl Acad Sci USA. 2004;101:6062–6067. doi: 10.1073/pnas.0400782101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tusher VG, Tibshirani R, Chu G. Significance analysis of microarrays applied to the ionizing radiation response. Proc Natl Acad Sci USA. 2001;98:5116–5121. doi: 10.1073/pnas.091062498. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dennis G Jr, Sherman BT, Hosack DA, Yang J, Gao W, Lane HC, Lempicki RA. DAVID: Database for Annotation, Visualization, and Integrated Discovery. Genome Biol. 2003;4:3. doi: 10.1186/gb-2003-4-5-p3. http://david.abcc.ncifcrf.gov/ [DOI] [PubMed] [Google Scholar]
- GO Slim and Subset Guide. http://www.geneontology.org/GO.slims.shtml
- Vennemann M, Kopplekamm A. mRNA profiling in forensic genetics I: Possibilities and limitations. Forensic Sci Int. 2010;15:71–75. doi: 10.1016/j.forsciint.2010.07.006. [DOI] [PubMed] [Google Scholar]
- Li YY, Feng Y, McTiernan CF, Pei W, Moravec CS, Wang P, Rosenblum W, Kormos RL, Feldman AM. Down-regulation of matrix metalloproteinases and reduction in collagen damage in the failing human heart after support with left ventricular assist devices. Circulation. 2001;104:1147–1152. doi: 10.1161/hc3501.095215. [DOI] [PubMed] [Google Scholar]
- Weber KT, Pick R, Janicki JS, Gadodia G, Lakier JB. Inadequate collagen tethers in dilated cardiopathy. Am Heart J. 1988;116:1641–1646. doi: 10.1016/0002-8703(88)90763-6. [DOI] [PubMed] [Google Scholar]
- Schaper J, Speiser B. The extracellular matrix in the failing human heart. Basic Res Cardiol. 1992;87(Suppl 1):303–309. doi: 10.1007/978-3-642-72474-9_26. [DOI] [PubMed] [Google Scholar]
- Schmitt M, Cockcroft JR, Frenneaux MP. Modulation of the natriuretic peptide system in heart failure: from bench to bedside? Clin Sci (Lond) 2003;105:141–160. doi: 10.1042/CS20030044. [DOI] [PubMed] [Google Scholar]
- Chien KR, Knowlton KU, Zhu H, Chien S. Regulation of cardiac gene expression during myocardial growth and hypertrophy: molecular studies of an adaptive physiologic response. FASEB J. 1991;5:3037–3046. doi: 10.1096/fasebj.5.15.1835945. [DOI] [PubMed] [Google Scholar]
- Petretto E, Sarwar R, Grieve I, Lu H, Kumaran MK, Muckett PJ, Mangion J, Schroen B, Benson M, Punjabi PP, Prasad SK, Pennell DJ, Kiesewetter C, Tasheva ES, Corpuz LM, Webb MD, Conrad GW, Kurtz TW, Kren V, Fischer J, Hubner N, Pinto YM, Pravenec M, Aitman TJ, Cook SA. Integrated genomic approaches implicate osteoglycin (Ogn) in the regulation of left ventricular mass. Nat Genet. 2008;40:546–552. doi: 10.1038/ng.134. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stansfield WE, Andersen NM, Tang RH, Selzman CH. Periostin is a novel factor in cardiac remodeling after experimental and clinical unloading of the failing heart. Ann Thorac Surg. 2009;88:1916–1921. doi: 10.1016/j.athoracsur.2009.07.038. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ahmed MS, Øie E, Vinge LE, von Lueder TG, Attramadal T, Attramadal H. Induction of pulmonary connective tissue growth factor in heart failure is associated with pulmonary parenchymal and vascular remodeling. Cardiovasc Res. 2007;74:323–333. doi: 10.1016/j.cardiores.2006.12.010. [DOI] [PubMed] [Google Scholar]
- Krijnen PA, Cillessen SA, Manoe R, Muller A, Visser CA, Meijer CJ, Musters RJ, Hack CE, Aarden LA, Niessen HW. Clusterin: a protective mediator for ischemic cardiomyocytes? Am J Physiol Heart Circ Physiol. 2005;289:H2193–2202. doi: 10.1152/ajpheart.00355.2005. [DOI] [PubMed] [Google Scholar]
- Lim H, Zhu YZ. Role of transforming growth factor-beta in the progression of heart failure. Cell Mol Life Sci. 2006;63:2584–2596. doi: 10.1007/s00018-006-6085-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Asakura M, Kitakaze M. Global gene expression profiling in the failing myocardium. Circ J. 2009;73:1568–1576. doi: 10.1253/circj.CJ-09-0465. [DOI] [PubMed] [Google Scholar]
- Pilbrow AP, Ellmers LJ, Black MA, Moravec CS, Sweet WE, Troughton RW, Richards AM, Frampton CM, Cameron VA. Genomic selection of reference genes for real-time PCR in human myocardium. BMC Med Genomics. 2008;1:64. doi: 10.1186/1755-8794-1-64. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tan FL, Moravec CS, Li J, Apperson-Hansen C, McCarthy PM, Young JB, Bond M. The gene expression fingerprint of human heart failure. Proc Natl Acad Sci USA. 2002;99:11387–11392. doi: 10.1073/pnas.162370099. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yang J, Moravec CS, Sussman MA, DiPaola NR, Fu D, Hawthorn L, Mitchell CA, Young JB, Francis GS, McCarthy PM, Bond M. Decreased SLIM1 expression and increased gelsolin expression in failing human hearts measured by high-density oligonucleotide arrays. Circulation. 2000;102:3046–3052. doi: 10.1161/01.cir.102.25.3046. [DOI] [PubMed] [Google Scholar]
- Maleszewski J, Lu J, Fox-Talbot K, Halushka MK. Robust immunohistochemical staining of several classes of proteins in tissues subjected to autolysis. J Histochem Cytochem. 2007;55:597–606. doi: 10.1369/jhc.6A7152.2007. [DOI] [PubMed] [Google Scholar]
- Halushka MK, Cornish TC, Lu J, Selvin S, Selvin E. Creation, validation, and quantitative analysis of protein expression in vascular tissue microarrays. Cardiovasc Pathol. 2010;19:136–146. doi: 10.1016/j.carpath.2008.12.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pritchard CC, Hsu L, Delrow J, Nelson PS. Project normal: defining normal variance in mouse gene expression. Proc Natl Acad Sci USA. 2001;98:13266–13271. doi: 10.1073/pnas.221465998. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Figure S1: RIN range in "warm- and "cold-24 hour autolysis" samples. The range of the RIN values in the warm-24 autolysis samples was 2.7 - 8.6, while the range in the cold- 24 autolysis samples was 3.5 - 8.6. The means of the RIN values in the "warm" and "cold" 24 hour autolysis samples were 5.1 (SD 1.53) and 6.9 (SD 1.45), respectively, and this difference was not statistically significant (P = 0.11).
Table S1: Postmortem fluctuating genes common in autopsy and explant hearts. The gene symbol and description of the 21 genes, which showed variation in gene expression during 24 hours of autolysis in both the autopsy and explant hearts.
Table S2: List of 374 upregulated (Table S2a)and 108 downregulated (Table S2b) genes in the explant/failed heart. List of ~480 genes differentially up- and down-regulated between the explant and autopsy hearts. Among these differentially expressed genes, 374 genes were upregulated and 108 genes were downregulated in the explant/failed heart.