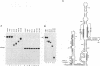Abstract
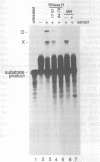
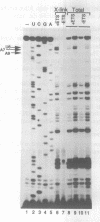
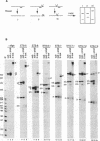
The first cleavage in mammalian pre-rRNA processing occurs within the 5' external transcribed spacer (ETS). We have recently shown that the U3 snRNP is required for this cleavage reaction, binds to the rRNA precursor, and remains complexed with the downstream processing product after the reaction has been completed (1). Using psoralen crosslinking in mouse cell extract we have detected a new interaction between U3 RNA and the mouse ETS processing substrate and its processed product. The crosslinked sites on both U3 and ETS RNAs have been mapped by RNase H cleavage and primer extension analyses. The crosslinked sites in U3 RNA map to C5, U6, and U8. U8 lies within and C5 and U6 are adjacent to an evolutionarily conserved U3 sequence termed box A'. In the ETS the crosslinked sites are U1012 and U1013, 362 nucleotides downstream from the processing site. Although the crosslinked site is dispensable for the primary processing reaction in vitro, a short conserved sequence just 3' to the cleavage site (nucleotides 650-668) is absolutely required for crosslink formation. We conclude that the interaction between U3 RNA and the 5' ETS detected by psoralen crosslinking may play a role in subsequent step(s) of pre-rRNA processing.
Full text
PDF


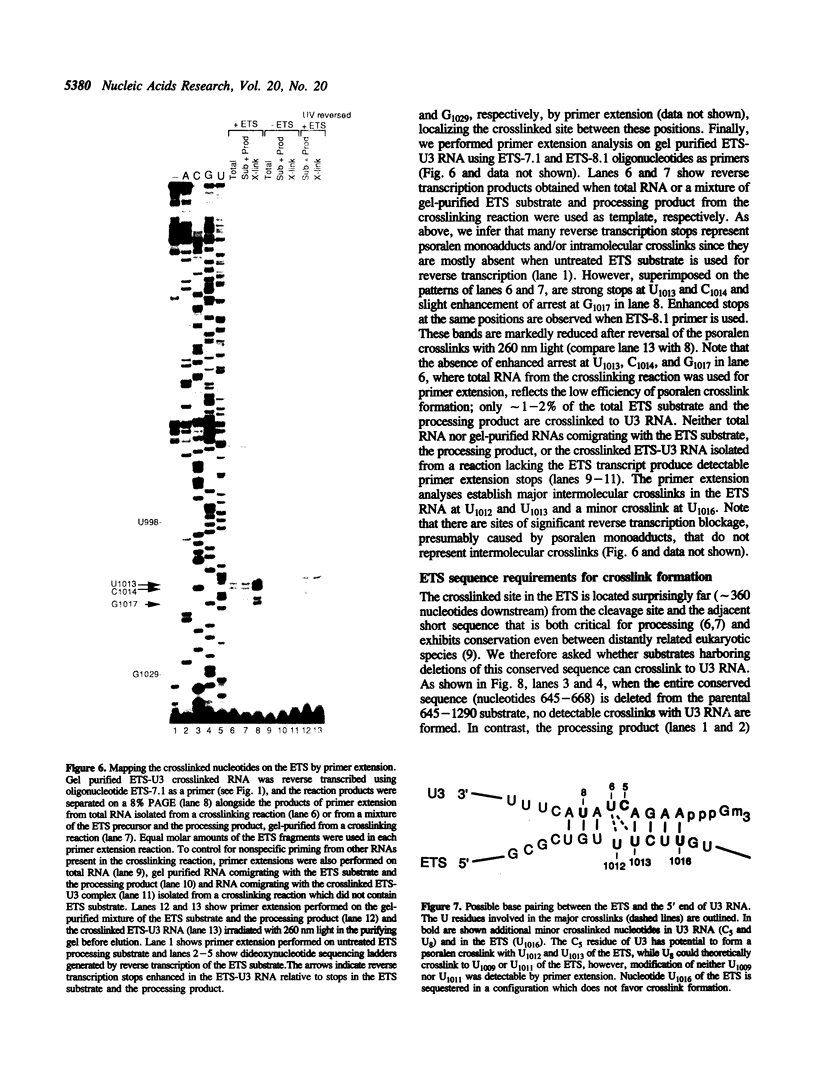
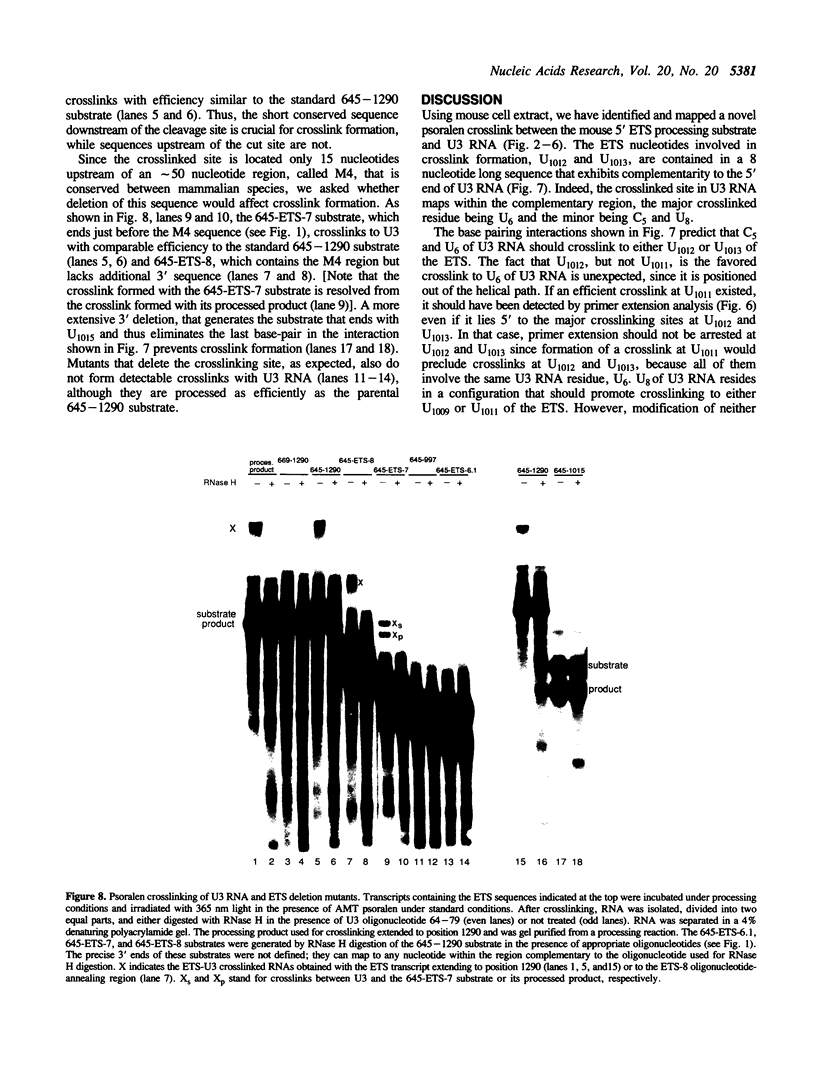

Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Beltrame M., Tollervey D. Identification and functional analysis of two U3 binding sites on yeast pre-ribosomal RNA. EMBO J. 1992 Apr;11(4):1531–1542. doi: 10.1002/j.1460-2075.1992.tb05198.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Calvet J. P., Meyer L. M., Pederson T. Small nuclear RNA U2 is base-paired to heterogeneous nuclear RNA. Science. 1982 Jul 30;217(4558):456–458. doi: 10.1126/science.6178162. [DOI] [PubMed] [Google Scholar]
- Calvet J. P., Pederson T. Base-pairing interactions between small nuclear RNAs and nuclear RNA precursors as revealed by psoralen cross-linking in vivo. Cell. 1981 Nov;26(3 Pt 1):363–370. doi: 10.1016/0092-8674(81)90205-1. [DOI] [PubMed] [Google Scholar]
- Cimino G. D., Gamper H. B., Isaacs S. T., Hearst J. E. Psoralens as photoactive probes of nucleic acid structure and function: organic chemistry, photochemistry, and biochemistry. Annu Rev Biochem. 1985;54:1151–1193. doi: 10.1146/annurev.bi.54.070185.005443. [DOI] [PubMed] [Google Scholar]
- Craig N., Kass S., Sollner-Webb B. Nucleotide sequence determining the first cleavage site in the processing of mouse precursor rRNA. Proc Natl Acad Sci U S A. 1987 Feb;84(3):629–633. doi: 10.1073/pnas.84.3.629. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Craig N., Kass S., Sollner-Webb B. Sequence organization and RNA structural motifs directing the mouse primary rRNA-processing event. Mol Cell Biol. 1991 Jan;11(1):458–467. doi: 10.1128/mcb.11.1.458. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hausner T. P., Giglio L. M., Weiner A. M. Evidence for base-pairing between mammalian U2 and U6 small nuclear ribonucleoprotein particles. Genes Dev. 1990 Dec;4(12A):2146–2156. doi: 10.1101/gad.4.12a.2146. [DOI] [PubMed] [Google Scholar]
- Hughes J. M., Ares M., Jr Depletion of U3 small nucleolar RNA inhibits cleavage in the 5' external transcribed spacer of yeast pre-ribosomal RNA and impairs formation of 18S ribosomal RNA. EMBO J. 1991 Dec;10(13):4231–4239. doi: 10.1002/j.1460-2075.1991.tb05001.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kass S., Craig N., Sollner-Webb B. Primary processing of mammalian rRNA involves two adjacent cleavages and is not species specific. Mol Cell Biol. 1987 Aug;7(8):2891–2898. doi: 10.1128/mcb.7.8.2891. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kass S., Sollner-Webb B. The first pre-rRNA-processing event occurs in a large complex: analysis by gel retardation, sedimentation, and UV cross-linking. Mol Cell Biol. 1990 Sep;10(9):4920–4931. doi: 10.1128/mcb.10.9.4920. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kass S., Tyc K., Steitz J. A., Sollner-Webb B. The U3 small nucleolar ribonucleoprotein functions in the first step of preribosomal RNA processing. Cell. 1990 Mar 23;60(6):897–908. doi: 10.1016/0092-8674(90)90338-f. [DOI] [PubMed] [Google Scholar]
- Maser R. L., Calvet J. P. U3 small nuclear RNA can be psoralen-cross-linked in vivo to the 5' external transcribed spacer of pre-ribosomal-RNA. Proc Natl Acad Sci U S A. 1989 Sep;86(17):6523–6527. doi: 10.1073/pnas.86.17.6523. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mazan S., Bachellerie J. P. Structure and organization of mouse U3B RNA functional genes. J Biol Chem. 1988 Dec 25;263(36):19461–19467. [PubMed] [Google Scholar]
- Michot B., Bachellerie J. P. Secondary structure of the 5' external transcribed spacer of vertebrate pre-rRNA. Presence of phylogenetically conserved features. Eur J Biochem. 1991 Feb 14;195(3):601–609. doi: 10.1111/j.1432-1033.1991.tb15743.x. [DOI] [PubMed] [Google Scholar]
- Miller K. G., Sollner-Webb B. Transcription of mouse rRNA genes by RNA polymerase I: in vitro and in vivo initiation and processing sites. Cell. 1981 Nov;27(1 Pt 2):165–174. doi: 10.1016/0092-8674(81)90370-6. [DOI] [PubMed] [Google Scholar]
- Musters W., Boon K., van der Sande C. A., van Heerikhuizen H., Planta R. J. Functional analysis of transcribed spacers of yeast ribosomal DNA. EMBO J. 1990 Dec;9(12):3989–3996. doi: 10.1002/j.1460-2075.1990.tb07620.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nakashima K., Darzynkiewicz E., Shatkin A. J. Proximity of mRNA5'-region and 18S rRNA in eukaryotic initiation complexes. Nature. 1980 Jul 17;286(5770):226–230. doi: 10.1038/286226a0. [DOI] [PubMed] [Google Scholar]
- Parker K. A., Steitz J. A. Structural analysis of the human U3 ribonucleoprotein particle reveal a conserved sequence available for base pairing with pre-rRNA. Mol Cell Biol. 1987 Aug;7(8):2899–2913. doi: 10.1128/mcb.7.8.2899. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Piette J. G., Hearst J. E. Termination sites of the in vitro nick-translation reaction on DNA that had photoreacted with psoralen. Proc Natl Acad Sci U S A. 1983 Sep;80(18):5540–5544. doi: 10.1073/pnas.80.18.5540. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Raziuddin, Little R. D., Labella T., Schlessinger D. Transcription and processing of RNA from mouse ribosomal DNA transfected into hamster cells. Mol Cell Biol. 1989 Apr;9(4):1667–1671. doi: 10.1128/mcb.9.4.1667. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Renalier M. H., Mazan S., Joseph N., Michot B., Bachellerie J. P. Structure of the 5'-external transcribed spacer of the human ribosomal RNA gene. FEBS Lett. 1989 Jun 5;249(2):279–284. doi: 10.1016/0014-5793(89)80641-6. [DOI] [PubMed] [Google Scholar]
- Rinke J., Appel B., Digweed M., Lührmann R. Localization of a base-paired interaction between small nuclear RNAs U4 and U6 in intact U4/U6 ribonucleoprotein particles by psoralen cross-linking. J Mol Biol. 1985 Oct 20;185(4):721–731. doi: 10.1016/0022-2836(85)90057-9. [DOI] [PubMed] [Google Scholar]
- Savino R., Gerbi S. A. In vivo disruption of Xenopus U3 snRNA affects ribosomal RNA processing. EMBO J. 1990 Jul;9(7):2299–2308. doi: 10.1002/j.1460-2075.1990.tb07401.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stroke I. L., Weiner A. M. The 5' end of U3 snRNA can be crosslinked in vivo to the external transcribed spacer of rat ribosomal RNA precursors. J Mol Biol. 1989 Dec 5;210(3):497–512. doi: 10.1016/0022-2836(89)90126-5. [DOI] [PubMed] [Google Scholar]