Abstract
We have determined the DNA sequence of intron 1 and flanking exons in the mitochondrial apocytochrome b gene of the Neurospora laboratory strain 74A and the natural isolate North Africa. In contrast to a previous report, we find that this intron contains an open reading frame (ORF) of 951 bases in frame with the upstream exon. The putative intron-encoded protein resembles those of other intron ORFs with respect to length, calculated isoelectric point, and proportion of basic, acidic, polar, and non-polar amino acids; however, no amino acid sequences resembling the "decapeptides" characteristic of maturase-like ORFs were found. Coupled with the previous finding that this intron is capable of self-splicing in vitro in the absence of proteins, the observations discussed here raise the possibility that other introns with long, in-frame ORFs may also be capable of RNA-catalyzed splicing in vitro.
Full text
PDF
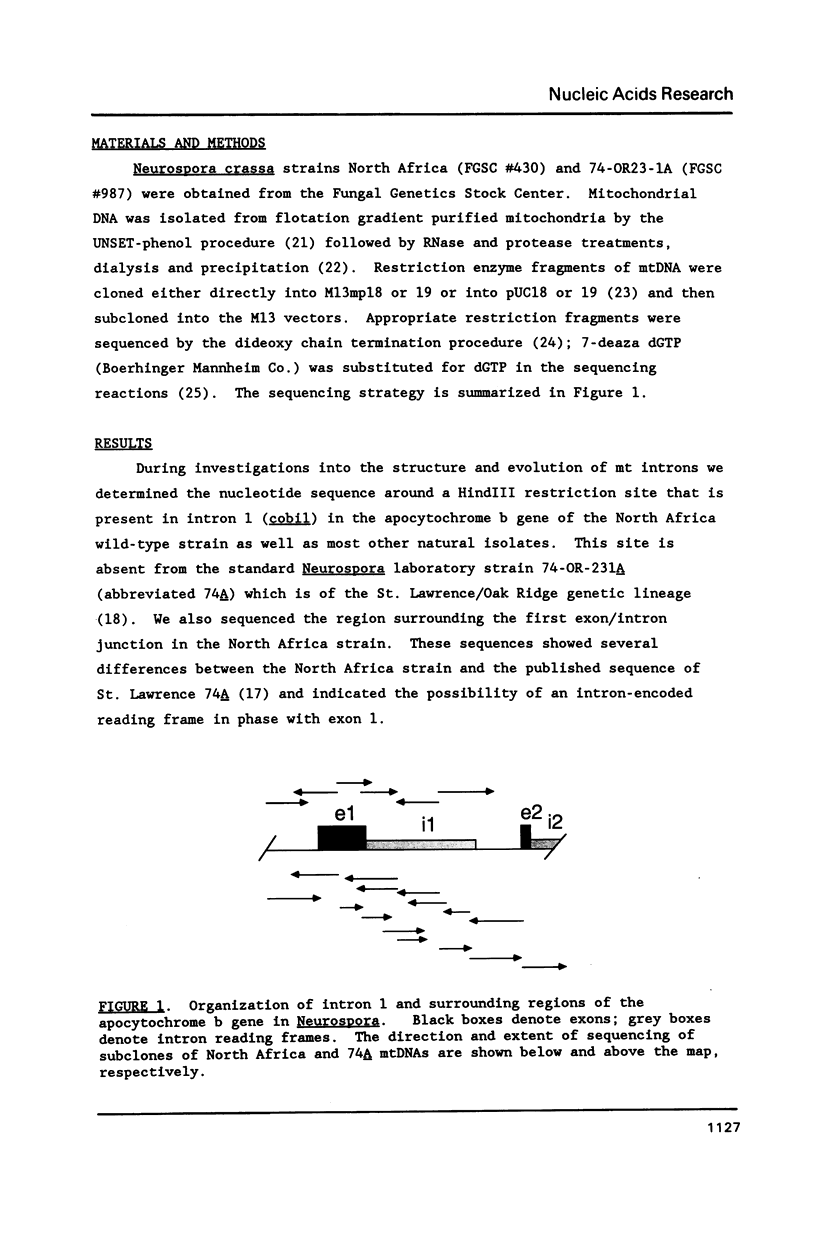








Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Akins R. A., Lambowitz A. M. A protein required for splicing group I introns in Neurospora mitochondria is mitochondrial tyrosyl-tRNA synthetase or a derivative thereof. Cell. 1987 Jul 31;50(3):331–345. doi: 10.1016/0092-8674(87)90488-0. [DOI] [PubMed] [Google Scholar]
- Anziano P. Q., Hanson D. K., Mahler H. R., Perlman P. S. Functional domains in introns: trans-acting and cis-acting regions of intron 4 of the cob gene. Cell. 1982 Oct;30(3):925–932. doi: 10.1016/0092-8674(82)90297-5. [DOI] [PubMed] [Google Scholar]
- Bonitz S. G., Coruzzi G., Thalenfeld B. E., Tzagoloff A., Macino G. Assembly of the mitochondrial membrane system. Structure and nucleotide sequence of the gene coding for subunit 1 of yeast cytochrme oxidase. J Biol Chem. 1980 Dec 25;255(24):11927–11941. [PubMed] [Google Scholar]
- Burger G., Werner S. The mitochondrial URF1 gene in Neurospora crassa has an intron that contains a novel type of URF. J Mol Biol. 1985 Nov 20;186(2):231–242. doi: 10.1016/0022-2836(85)90100-7. [DOI] [PubMed] [Google Scholar]
- Burke J. M., Breitenberger C., Heckman J. E., Dujon B., RajBhandary U. L. Cytochrome b gene of Neurospora crassa mitochondria. Partial sequence and location of introns at sites different from those in Saccharomyces cerevisiae and Aspergillus nidulans. J Biol Chem. 1984 Jan 10;259(1):504–511. [PubMed] [Google Scholar]
- Burke J. M., RajBhandary U. L. Intron within the large rRNA gene of N. crassa mitochondria: a long open reading frame and a consensus sequence possibly important in splicing. Cell. 1982 Dec;31(3 Pt 2):509–520. doi: 10.1016/0092-8674(82)90307-5. [DOI] [PubMed] [Google Scholar]
- Cech T. R. The generality of self-splicing RNA: relationship to nuclear mRNA splicing. Cell. 1986 Jan 31;44(2):207–210. doi: 10.1016/0092-8674(86)90751-8. [DOI] [PubMed] [Google Scholar]
- Collins R. A., Lambowitz A. M. RNA splicing in Neurospora mitochondria. Defective splicing of mitochondrial mRNA precursors in the nuclear mutant cyt18-1. J Mol Biol. 1985 Aug 5;184(3):413–428. doi: 10.1016/0022-2836(85)90291-8. [DOI] [PubMed] [Google Scholar]
- Collins R. A., Lambowitz A. M. Structural variations and optional introns in the mitochondrial DNAs of Neurospora strains isolated from nature. Plasmid. 1983 Jan;9(1):53–70. doi: 10.1016/0147-619x(83)90031-8. [DOI] [PubMed] [Google Scholar]
- Cummings D. J., MacNeil I. A., Domenico J., Matsuura E. T. Excision-amplification of mitochondrial DNA during senescence in Podospora anserina. DNA sequence analysis of three unique "plasmids". J Mol Biol. 1985 Oct 20;185(4):659–680. doi: 10.1016/0022-2836(85)90052-x. [DOI] [PubMed] [Google Scholar]
- Davies R. W., Waring R. B., Ray J. A., Brown T. A., Scazzocchio C. Making ends meet: a model for RNA splicing in fungal mitochondria. Nature. 1982 Dec 23;300(5894):719–724. doi: 10.1038/300719a0. [DOI] [PubMed] [Google Scholar]
- De La Salle H., Jacq C., Slonimski P. P. Critical sequences within mitochondrial introns: pleiotropic mRNA maturase and cis-dominant signals of the box intron controlling reductase and oxidase. Cell. 1982 Apr;28(4):721–732. doi: 10.1016/0092-8674(82)90051-4. [DOI] [PubMed] [Google Scholar]
- Dujon B. Sequence of the intron and flanking exons of the mitochondrial 21S rRNA gene of yeast strains having different alleles at the omega and rib-1 loci. Cell. 1980 May;20(1):185–197. doi: 10.1016/0092-8674(80)90246-9. [DOI] [PubMed] [Google Scholar]
- Garriga G., Bertrand H., Lambowitz A. M. RNA splicing in Neurospora mitochondria: nuclear mutants defective in both splicing and 3' end synthesis of the large rRNA. Cell. 1984 Mar;36(3):623–634. doi: 10.1016/0092-8674(84)90342-8. [DOI] [PubMed] [Google Scholar]
- Garriga G., Lambowitz A. M. RNA splicing in neurospora mitochondria: self-splicing of a mitochondrial intron in vitro. Cell. 1984 Dec;39(3 Pt 2):631–641. doi: 10.1016/0092-8674(84)90470-7. [DOI] [PubMed] [Google Scholar]
- Helmer-Citterich M., Morelli G., Macino G. Nucleotide sequence and intron structure of the apocytochrome b gene of Neurospora crassa mitochondria. EMBO J. 1983;2(8):1235–1242. doi: 10.1002/j.1460-2075.1983.tb01575.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hensgens L. A., Bonen L., de Haan M., van der Horst G., Grivell L. A. Two intron sequences in yeast mitochondrial COX1 gene: homology among URF-containing introns and strain-dependent variation in flanking exons. Cell. 1983 Feb;32(2):379–389. doi: 10.1016/0092-8674(83)90457-9. [DOI] [PubMed] [Google Scholar]
- Jacquier A., Dujon B. An intron-encoded protein is active in a gene conversion process that spreads an intron into a mitochondrial gene. Cell. 1985 Jun;41(2):383–394. doi: 10.1016/s0092-8674(85)80011-8. [DOI] [PubMed] [Google Scholar]
- Labouesse M., Herbert C. J., Dujardin G., Slonimski P. P. Three suppressor mutations which cure a mitochondrial RNA maturase deficiency occur at the same codon in the open reading frame of the nuclear NAM2 gene. EMBO J. 1987 Mar;6(3):713–721. doi: 10.1002/j.1460-2075.1987.tb04812.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lang B. F. The mitochondrial genome of the fission yeast Schizosaccharomyces pombe: highly homologous introns are inserted at the same position of the otherwise less conserved cox1 genes in Schizosaccharomyces pombe and Aspergillus nidulans. EMBO J. 1984 Sep;3(9):2129–2136. doi: 10.1002/j.1460-2075.1984.tb02102.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lazowska J., Jacq C., Slonimski P. P. Sequence of introns and flanking exons in wild-type and box3 mutants of cytochrome b reveals an interlaced splicing protein coded by an intron. Cell. 1980 Nov;22(2 Pt 2):333–348. doi: 10.1016/0092-8674(80)90344-x. [DOI] [PubMed] [Google Scholar]
- Macreadie I. G., Scott R. M., Zinn A. R., Butow R. A. Transposition of an intron in yeast mitochondria requires a protein encoded by that intron. Cell. 1985 Jun;41(2):395–402. doi: 10.1016/s0092-8674(85)80012-x. [DOI] [PubMed] [Google Scholar]
- Mannella C. A., Lambowitz A. Interaction of wild-type and poky mitochondrial DNA in heterokaryons of Neurospora. Biochem Biophys Res Commun. 1978 Feb 14;80(3):673–679. doi: 10.1016/0006-291x(78)91621-2. [DOI] [PubMed] [Google Scholar]
- Michel F., Cummings D. J. Analysis of class I introns in a mitochondrial plasmid associated with senescence of Podospora anserina reveals extraordinary resemblance to the Tetrahymena ribosomal intron. Curr Genet. 1985;10(1):69–79. doi: 10.1007/BF00418495. [DOI] [PubMed] [Google Scholar]
- Michel F., Jacquier A., Dujon B. Comparison of fungal mitochondrial introns reveals extensive homologies in RNA secondary structure. Biochimie. 1982 Oct;64(10):867–881. doi: 10.1016/s0300-9084(82)80349-0. [DOI] [PubMed] [Google Scholar]
- Mizusawa S., Nishimura S., Seela F. Improvement of the dideoxy chain termination method of DNA sequencing by use of deoxy-7-deazaguanosine triphosphate in place of dGTP. Nucleic Acids Res. 1986 Feb 11;14(3):1319–1324. doi: 10.1093/nar/14.3.1319. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Morelli G., Macino G. Two intervening sequences in the ATPase subunit 6 gene of Neurospora crassa. A short intron (93 base-pairs) and a long intron that is stable after excision. J Mol Biol. 1984 Sep 25;178(3):491–507. doi: 10.1016/0022-2836(84)90235-3. [DOI] [PubMed] [Google Scholar]
- Nelson M. A., Macino G. Structure and expression of the overlapping ND4L and ND5 genes of Neurospora crassa mitochondria. Mol Gen Genet. 1987 Feb;206(2):307–317. doi: 10.1007/BF00333589. [DOI] [PubMed] [Google Scholar]
- Nobrega F. G., Tzagoloff A. Assembly of the mitochondrial membrane system. DNA sequence and organization of the cytochrome b gene in Saccharomyces cerevisiae D273-10B. J Biol Chem. 1980 Oct 25;255(20):9828–9837. [PubMed] [Google Scholar]
- Sanger F., Nicklen S., Coulson A. R. DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci U S A. 1977 Dec;74(12):5463–5467. doi: 10.1073/pnas.74.12.5463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schmelzer C., Schweyen R. J. Evidence for ribosomes involved in splicing of yeast mitochondrial transcripts. Nucleic Acids Res. 1982 Jan 22;10(2):513–524. doi: 10.1093/nar/10.2.513. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Taylor J. W., Smolich B. D. Molecular cloning and physical mapping of the Neurospora crassa 74-OR23-1A mitochondrial genome. Curr Genet. 1985;9(7):597–603. doi: 10.1007/BF00381173. [DOI] [PubMed] [Google Scholar]
- Waring R. B., Brown T. A., Ray J. A., Scazzocchio C., Davies R. W. Three variant introns of the same general class in the mitochondrial gene for cytochrome oxidase subunit 1 in Aspergillus nidulans. EMBO J. 1984 Sep;3(9):2121–2128. doi: 10.1002/j.1460-2075.1984.tb02100.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Waring R. B., Davies R. W., Scazzocchio C., Brown T. A. Internal structure of a mitochondrial intron of Aspergillus nidulans. Proc Natl Acad Sci U S A. 1982 Oct;79(20):6332–6336. doi: 10.1073/pnas.79.20.6332. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weiss-Brummer B., Rödel G., Schweyen R. J., Kaudewitz F. Expression of the split gene cob in yeast: evidence for a precursor of a "maturase" protein translated from intron 4 and preceding exons. Cell. 1982 Jun;29(2):527–536. doi: 10.1016/0092-8674(82)90169-6. [DOI] [PubMed] [Google Scholar]
- Yanisch-Perron C., Vieira J., Messing J. Improved M13 phage cloning vectors and host strains: nucleotide sequences of the M13mp18 and pUC19 vectors. Gene. 1985;33(1):103–119. doi: 10.1016/0378-1119(85)90120-9. [DOI] [PubMed] [Google Scholar]


