Abstract
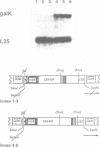
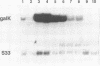
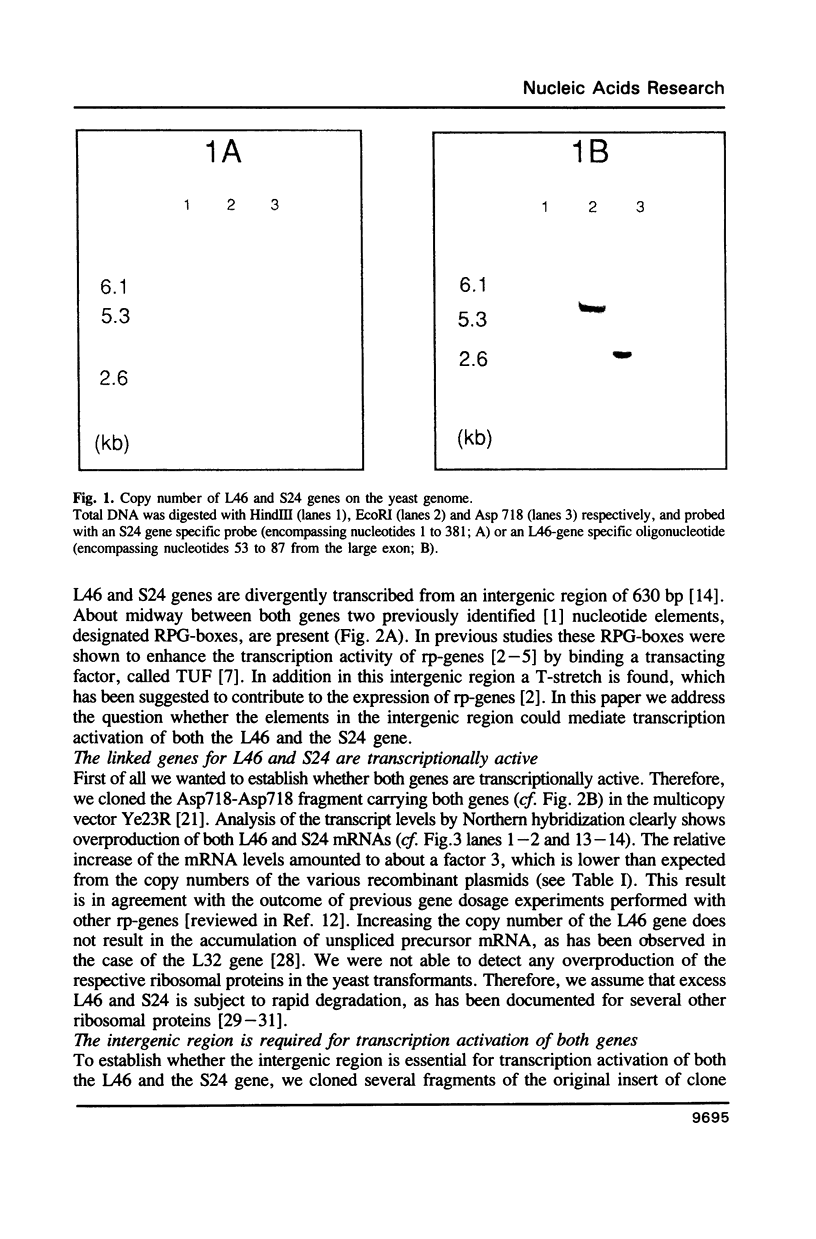
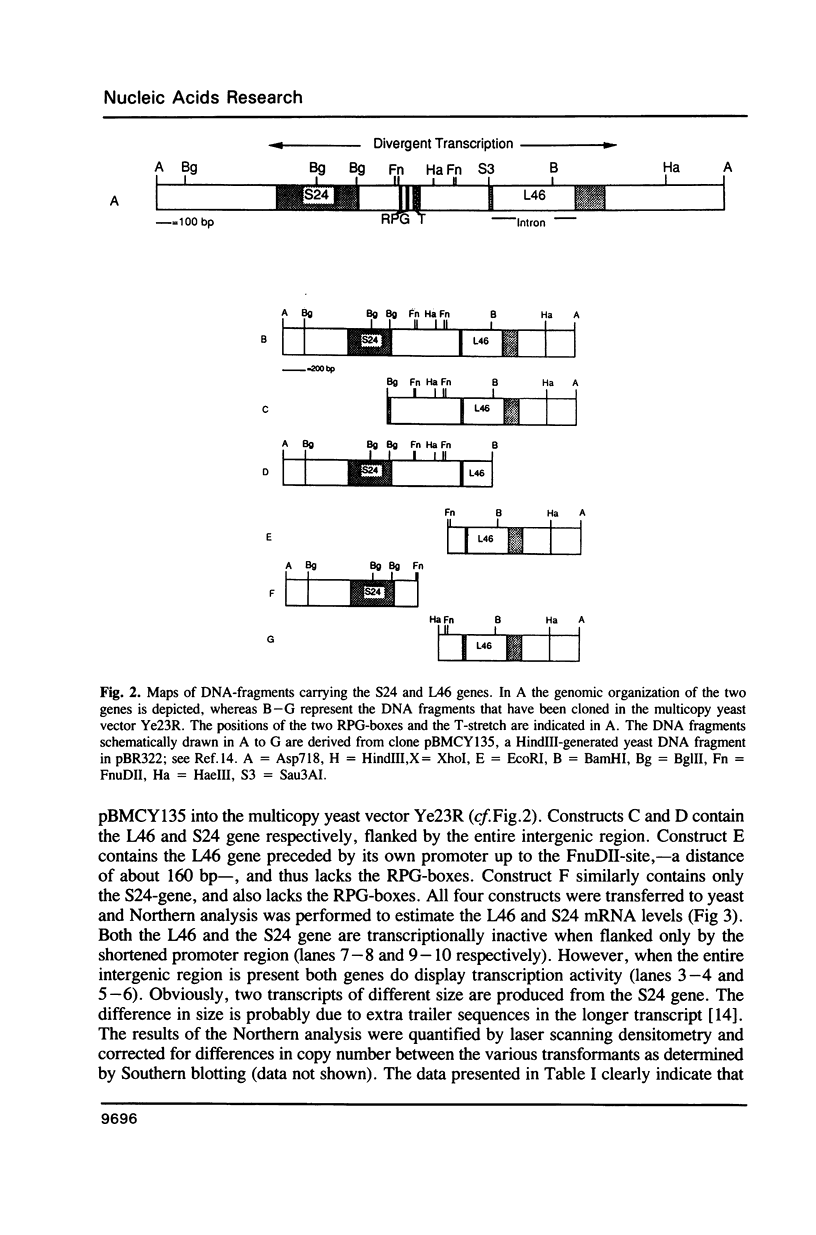
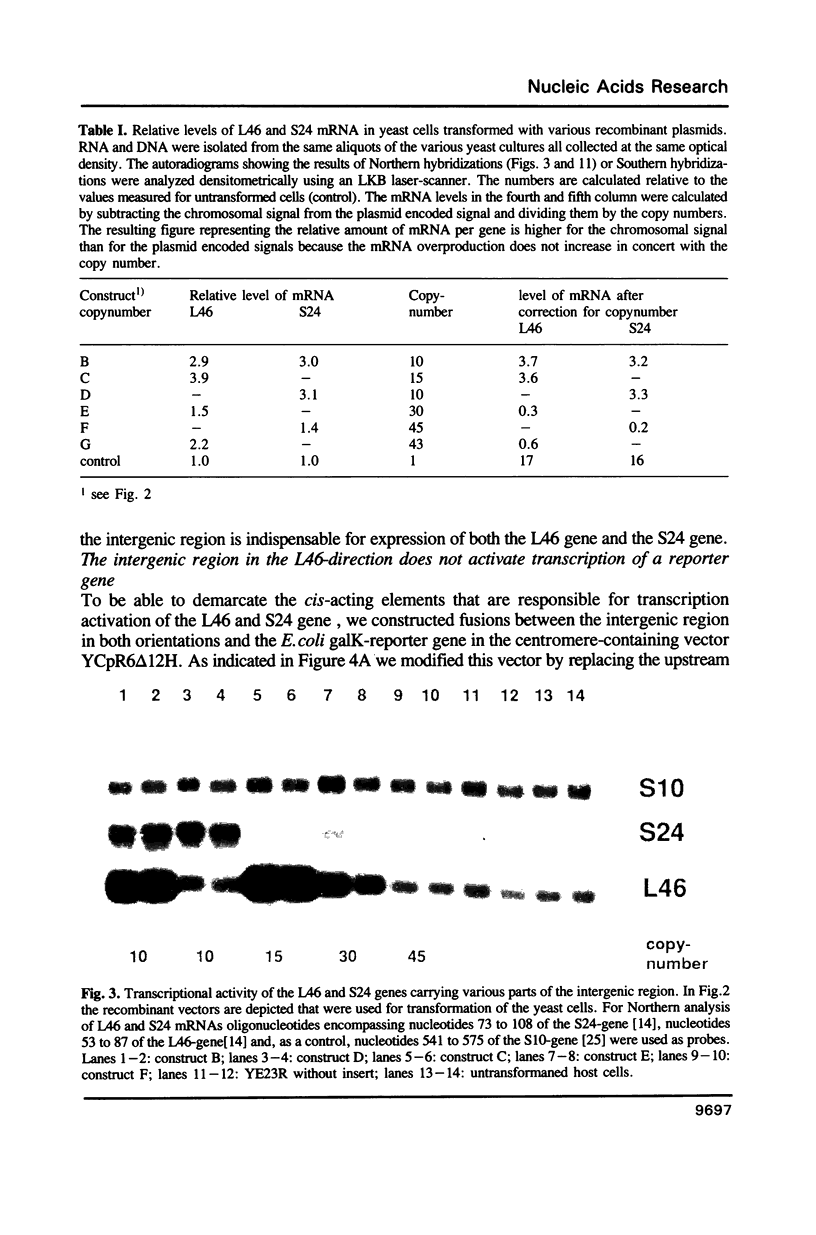
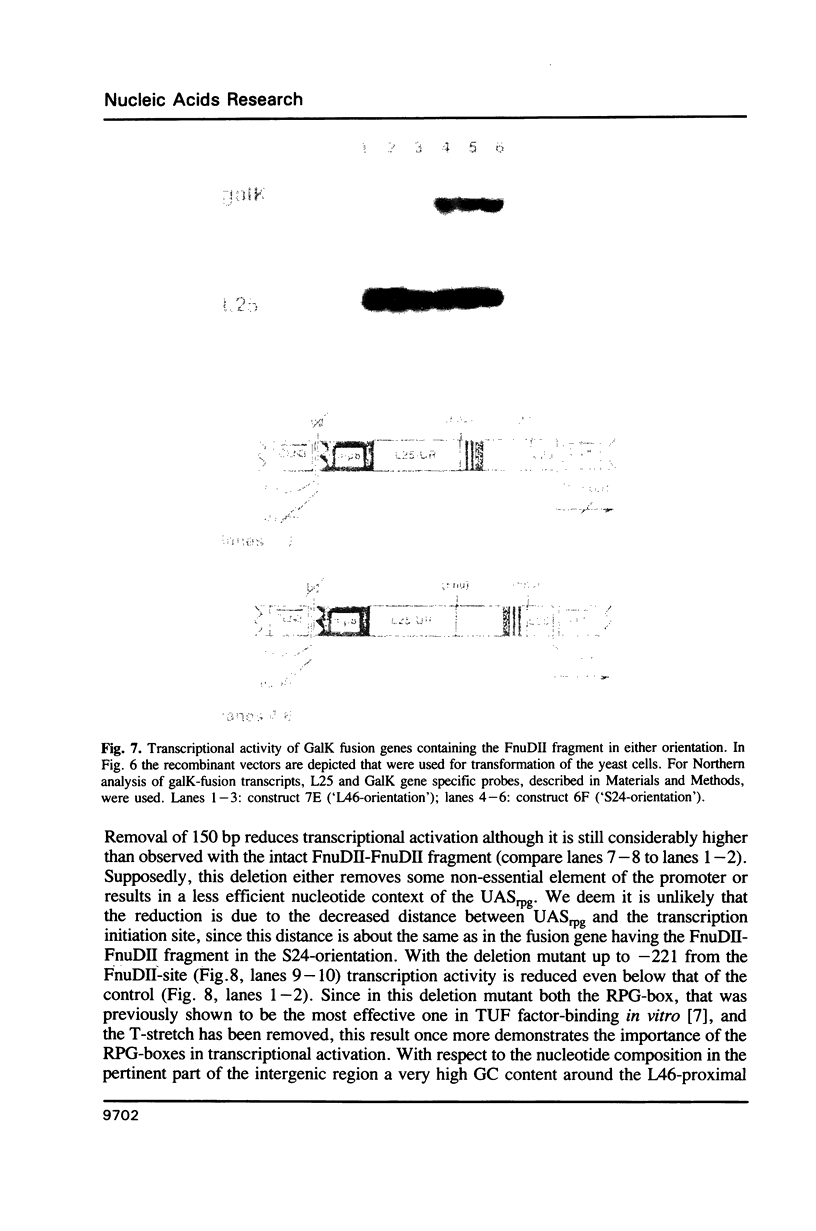
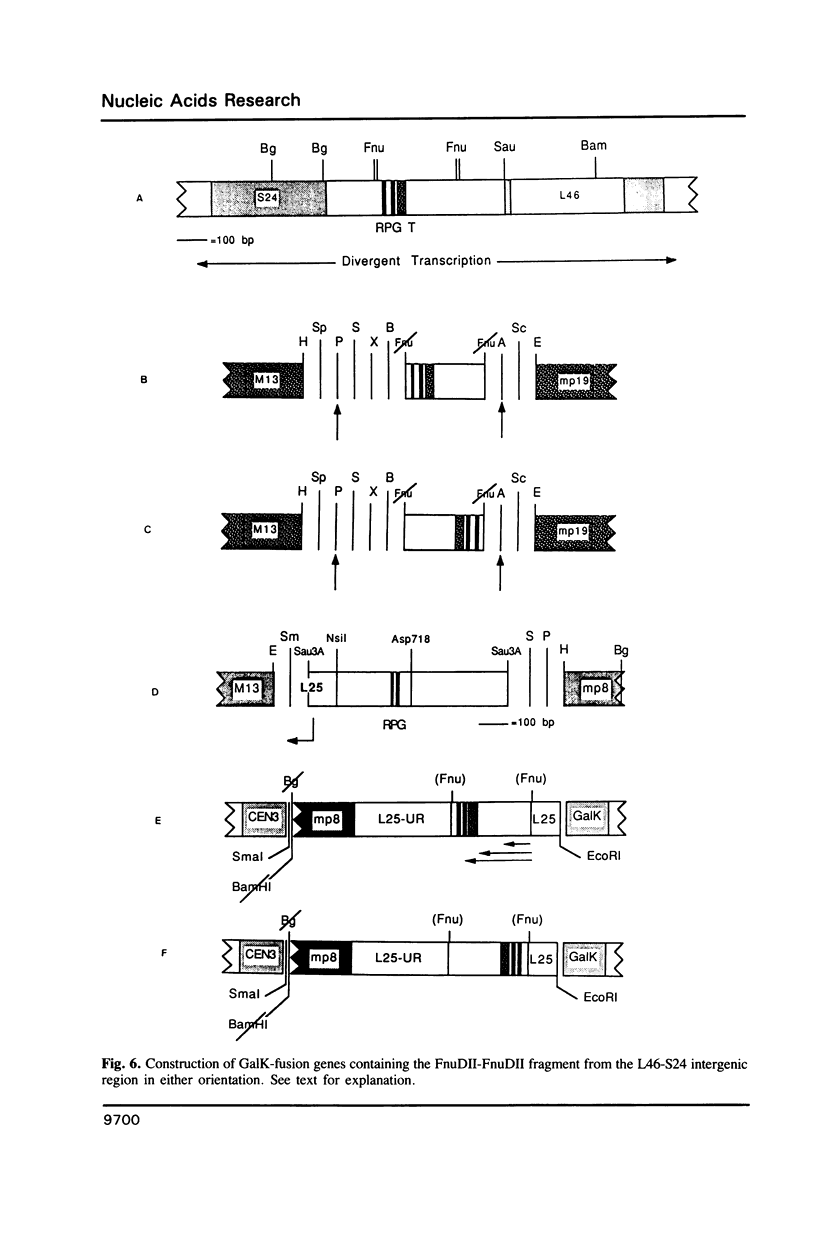
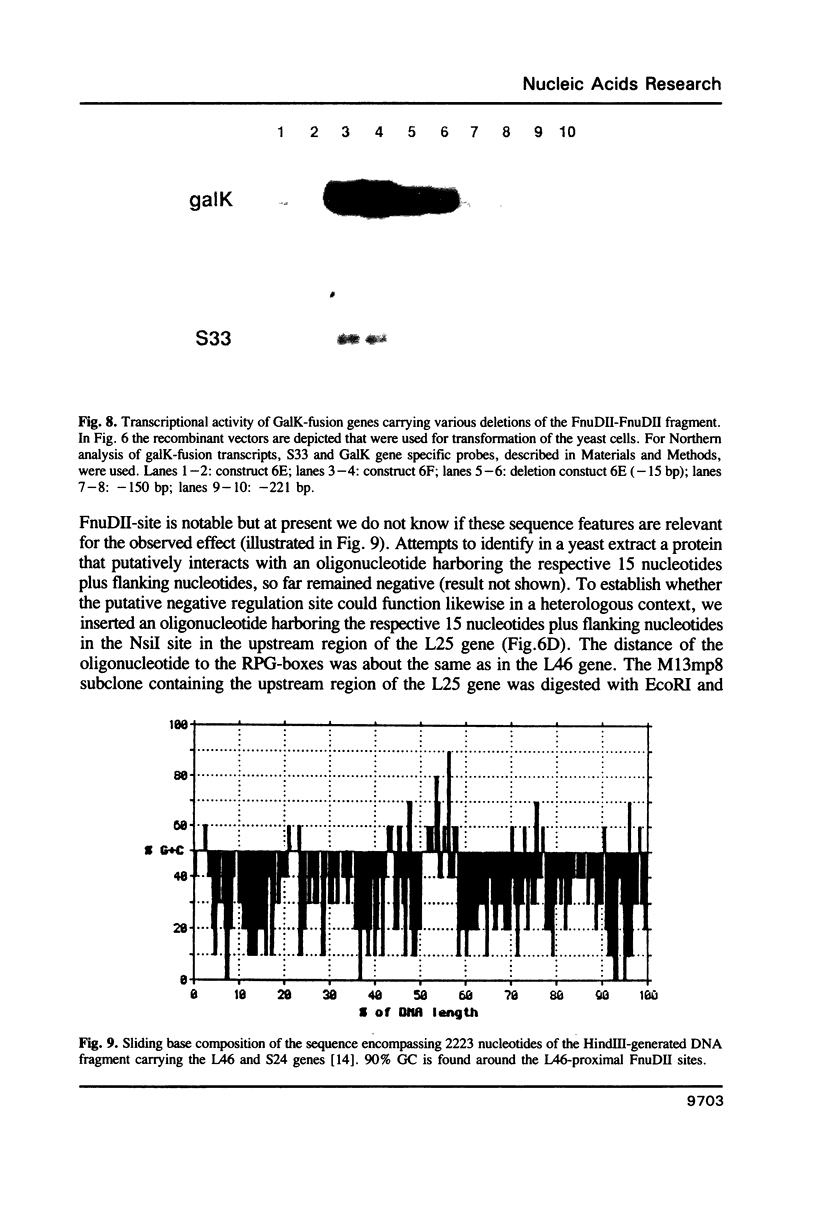
Transcription of the majority of the ribosomal protein (rp) genes in yeast is activated through common cis-acting elements, designated RPG-boxes. These elements have been shown to act as specific binding sites for the protein factor TUF/RAP1/GRF1 in vitro. Two such elements occur in the intergenic region separating the divergently transcribed genes encoding L46 and S24. To investigate whether the two RPG-boxes mediate transcription activation of both the L46 and S24 gene, two experimental strategies were followed: cloning of the respective genes on multicopy vectors and construction of fusion genes. Cloning of the L46 + S24 gene including the intergenic region in a multicopy yeast vector indicated that both genes are transcriptionally active. Using constructs in which only the S24 or the L46 gene is present, with or without the intergenic region, we obtained evidence that the intergenic region is indispensable for transcription activation of either gene. To demarcate the element(s) responsible for this activation, fusions of the intergenic region in either orientation to the galK reporter gene were made. Northern analysis of the levels of hybrid mRNA demonstrated that the intergenic region can serve as an heterologous promoter when it is in the 'S24-orientation'. Surprisingly, however, when fused in the reverse orientation the intergenic region did hardly confer transcription activity on the fusion gene. Furthermore, a 274 bp FnuDII-FnuDII fragment from the intergenic region that contains the RPG-boxes, could replace the naturally occurring upstream activation site (UASrpg) of the L25 rp-gene only when inserted in the 'S24-orientation'. Removal of 15 bp from the FnuDII fragment appeared to be sufficient to obtain transcription activation in the 'L46 orientation' as well. Analysis of a construct in which the RPG-boxes were selectively deleted from the promoter region of the L46 gene indicated that the RPG-boxes are needed for efficient transcriptional activation of the L46 gene. We conclude that all promoter elements for the S24 gene are located within the intergenic region, where the RPG-boxes are the most likely UAS-elements. However, the intergenic region (including the RPG-boxes) is required but not sufficient to confer transcription activity on the L46 gene.
Full text
PDF









Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Baldari C., Cesareni G. Plasmids pEMBLY: new single-stranded shuttle vectors for the recovery and analysis of yeast DNA sequences. Gene. 1985;35(1-2):27–32. doi: 10.1016/0378-1119(85)90154-4. [DOI] [PubMed] [Google Scholar]
- Bromley S., Hereford L., Rosbash M. Further evidence that the rna2 mutation of Saccharomyces cerevisiae affects mRNA processing. Mol Cell Biol. 1982 Oct;2(10):1205–1211. doi: 10.1128/mcb.2.10.1205. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Buchman A. R., Kimmerly W. J., Rine J., Kornberg R. D. Two DNA-binding factors recognize specific sequences at silencers, upstream activating sequences, autonomously replicating sequences, and telomeres in Saccharomyces cerevisiae. Mol Cell Biol. 1988 Jan;8(1):210–225. doi: 10.1128/mcb.8.1.210. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dabeva M. D., Post-Beittenmiller M. A., Warner J. R. Autogenous regulation of splicing of the transcript of a yeast ribosomal protein gene. Proc Natl Acad Sci U S A. 1986 Aug;83(16):5854–5857. doi: 10.1073/pnas.83.16.5854. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hong S. H., Marmur J. Upstream regulatory regions controlling the expression of the yeast maltase gene. Mol Cell Biol. 1987 Jul;7(7):2477–2483. doi: 10.1128/mcb.7.7.2477. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hu N., Messing J. The making of strand-specific M13 probes. Gene. 1982 Mar;17(3):271–277. doi: 10.1016/0378-1119(82)90143-3. [DOI] [PubMed] [Google Scholar]
- Huet J., Cottrelle P., Cool M., Vignais M. L., Thiele D., Marck C., Buhler J. M., Sentenac A., Fromageot P. A general upstream binding factor for genes of the yeast translational apparatus. EMBO J. 1985 Dec 16;4(13A):3539–3547. doi: 10.1002/j.1460-2075.1985.tb04114.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kimmerly W., Buchman A., Kornberg R., Rine J. Roles of two DNA-binding factors in replication, segregation and transcriptional repression mediated by a yeast silencer. EMBO J. 1988 Jul;7(7):2241–2253. doi: 10.1002/j.1460-2075.1988.tb03064.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kimura M., Arndt E., Hatakeyama T., Hatakeyama T., Kimura J. Ribosomal proteins in halobacteria. Can J Microbiol. 1989 Jan;35(1):195–199. doi: 10.1139/m89-030. [DOI] [PubMed] [Google Scholar]
- Larkin J. C., Thompson J. R., Woolford J. L., Jr Structure and expression of the Saccharomyces cerevisiae CRY1 gene: a highly conserved ribosomal protein gene. Mol Cell Biol. 1987 May;7(5):1764–1775. doi: 10.1128/mcb.7.5.1764. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Leer R. J., Van Raamsdonk-Duin M. M., Mager W. H., Planta R. J. Conserved sequences upstream of yeast ribosomal protein genes. Curr Genet. 1985;9(4):273–277. doi: 10.1007/BF00419955. [DOI] [PubMed] [Google Scholar]
- Leer R. J., van Raamsdonk-Duin M. M., Hagendoorn M. J., Mager W. H., Planta R. J. Structural comparison of yeast ribosomal protein genes. Nucleic Acids Res. 1984 Sep 11;12(17):6685–6700. doi: 10.1093/nar/12.17.6685. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Leer R. J., van Raamsdonk-Duin M. M., Kraakman P., Mager W. H., Planta R. J. The genes for yeast ribosomal proteins S24 and L46 are adjacent and divergently transcribed. Nucleic Acids Res. 1985 Feb 11;13(3):701–709. doi: 10.1093/nar/13.3.701. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Leer R. J., van Raamsdonk-Duin M. M., Molenaar C. M., Cohen L. H., Mager W. H., Planta R. J. The structure of the gene coding for the phosphorylated ribosomal protein S10 in yeast. Nucleic Acids Res. 1982 Oct 11;10(19):5869–5878. doi: 10.1093/nar/10.19.5869. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Leer R. J., van Raamsdonk-Duin M. M., Schoppink P. J., Cornelissen M. T., Cohen L. H., Mager W. H., Planta R. J. Yeast ribosomal protein S33 is encoded by an unsplit gene. Nucleic Acids Res. 1983 Nov 25;11(22):7759–7768. doi: 10.1093/nar/11.22.7759. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lin A., McNally J., Wool I. G. The primary structure of rat liver ribosomal protein L39. J Biol Chem. 1984 Jan 10;259(1):487–490. [PubMed] [Google Scholar]
- Maicas E., Pluthero F. G., Friesen J. D. The accumulation of three yeast ribosomal proteins under conditions of excess mRNA is determined primarily by fast protein decay. Mol Cell Biol. 1988 Jan;8(1):169–175. doi: 10.1128/mcb.8.1.169. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McMaster G. K., Carmichael G. G. Analysis of single- and double-stranded nucleic acids on polyacrylamide and agarose gels by using glyoxal and acridine orange. Proc Natl Acad Sci U S A. 1977 Nov;74(11):4835–4838. doi: 10.1073/pnas.74.11.4835. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Molenaar C. M., Woudt L. P., Jansen A. E., Mager W. H., Planta R. J., Donovan D. M., Pearson N. J. Structure and organization of two linked ribosomal protein genes in yeast. Nucleic Acids Res. 1984 Oct 11;12(19):7345–7358. doi: 10.1093/nar/12.19.7345. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nieuwint R. T., Mager W. H., Maurer K. C., Planta R. J. Mutational analysis of the upstream activation site of yeast ribosomal protein genes. Curr Genet. 1989 Apr;15(4):247–251. doi: 10.1007/BF00447039. [DOI] [PubMed] [Google Scholar]
- Osley M. A., Gould J., Kim S., Kane M. Y., Hereford L. Identification of sequences in a yeast histone promoter involved in periodic transcription. Cell. 1986 May 23;45(4):537–544. doi: 10.1016/0092-8674(86)90285-0. [DOI] [PubMed] [Google Scholar]
- Rotenberg M. O., Woolford J. L., Jr Tripartite upstream promoter element essential for expression of Saccharomyces cerevisiae ribosomal protein genes. Mol Cell Biol. 1986 Feb;6(2):674–687. doi: 10.1128/mcb.6.2.674. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rymond B. C., Zitomer R. S., Schümperli D., Rosenberg M. The expression in yeast of the Escherichia coli galK gene on CYC1::galK fusion plasmids. Gene. 1983 Nov;25(2-3):249–262. doi: 10.1016/0378-1119(83)90229-9. [DOI] [PubMed] [Google Scholar]
- Schwindinger W. F., Warner J. R. Transcriptional elements of the yeast ribosomal protein gene CYH2. J Biol Chem. 1987 Apr 25;262(12):5690–5695. [PubMed] [Google Scholar]
- Shore D., Nasmyth K. Purification and cloning of a DNA binding protein from yeast that binds to both silencer and activator elements. Cell. 1987 Dec 4;51(5):721–732. doi: 10.1016/0092-8674(87)90095-x. [DOI] [PubMed] [Google Scholar]
- Siliciano P. G., Tatchell K. Transcription and regulatory signals at the mating type locus in yeast. Cell. 1984 Jul;37(3):969–978. doi: 10.1016/0092-8674(84)90431-8. [DOI] [PubMed] [Google Scholar]
- Sollitti P., Marmur J. Primary structure of the regulatory gene from the MAL6 locus of Saccharomyces carlsbergensis. Mol Gen Genet. 1988 Jul;213(1):56–62. doi: 10.1007/BF00333398. [DOI] [PubMed] [Google Scholar]
- Tsay Y. F., Thompson J. R., Rotenberg M. O., Larkin J. C., Woolford J. L., Jr Ribosomal protein synthesis is not regulated at the translational level in Saccharomyces cerevisiae: balanced accumulation of ribosomal proteins L16 and rp59 is mediated by turnover of excess protein. Genes Dev. 1988 Jun;2(6):664–676. doi: 10.1101/gad.2.6.664. [DOI] [PubMed] [Google Scholar]
- Vignais M. L., Woudt L. P., Wassenaar G. M., Mager W. H., Sentenac A., Planta R. J. Specific binding of TUF factor to upstream activation sites of yeast ribosomal protein genes. EMBO J. 1987 May;6(5):1451–1457. doi: 10.1002/j.1460-2075.1987.tb02386.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Warner J. R., Mitra G., Schwindinger W. F., Studeny M., Fried H. M. Saccharomyces cerevisiae coordinates accumulation of yeast ribosomal proteins by modulating mRNA splicing, translational initiation, and protein turnover. Mol Cell Biol. 1985 Jun;5(6):1512–1521. doi: 10.1128/mcb.5.6.1512. [DOI] [PMC free article] [PubMed] [Google Scholar]
- West R. W., Jr, Yocum R. R., Ptashne M. Saccharomyces cerevisiae GAL1-GAL10 divergent promoter region: location and function of the upstream activating sequence UASG. Mol Cell Biol. 1984 Nov;4(11):2467–2478. doi: 10.1128/mcb.4.11.2467. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Woudt L. P., Mager W. H., Nieuwint R. T., Wassenaar G. M., van der Kuyl A. C., Murre J. J., Hoekman M. F., Brockhoff P. G., Planta R. J. Analysis of upstream activation sites of yeast ribosomal protein genes. Nucleic Acids Res. 1987 Aug 11;15(15):6037–6048. doi: 10.1093/nar/15.15.6037. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Woudt L. P., Smit A. B., Mager W. H., Planta R. J. Conserved sequence elements upstream of the gene encoding yeast ribosomal protein L25 are involved in transcription activation. EMBO J. 1986 May;5(5):1037–1040. doi: 10.1002/j.1460-2075.1986.tb04319.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- elBaradi T. T., van der Sande C. A., Mager W. H., Raué H. A., Planta R. J. The cellular level of yeast ribosomal protein L25 is controlled principally by rapid degradation of excess protein. Curr Genet. 1986;10(10):733–739. doi: 10.1007/BF00405095. [DOI] [PubMed] [Google Scholar]