Abstract
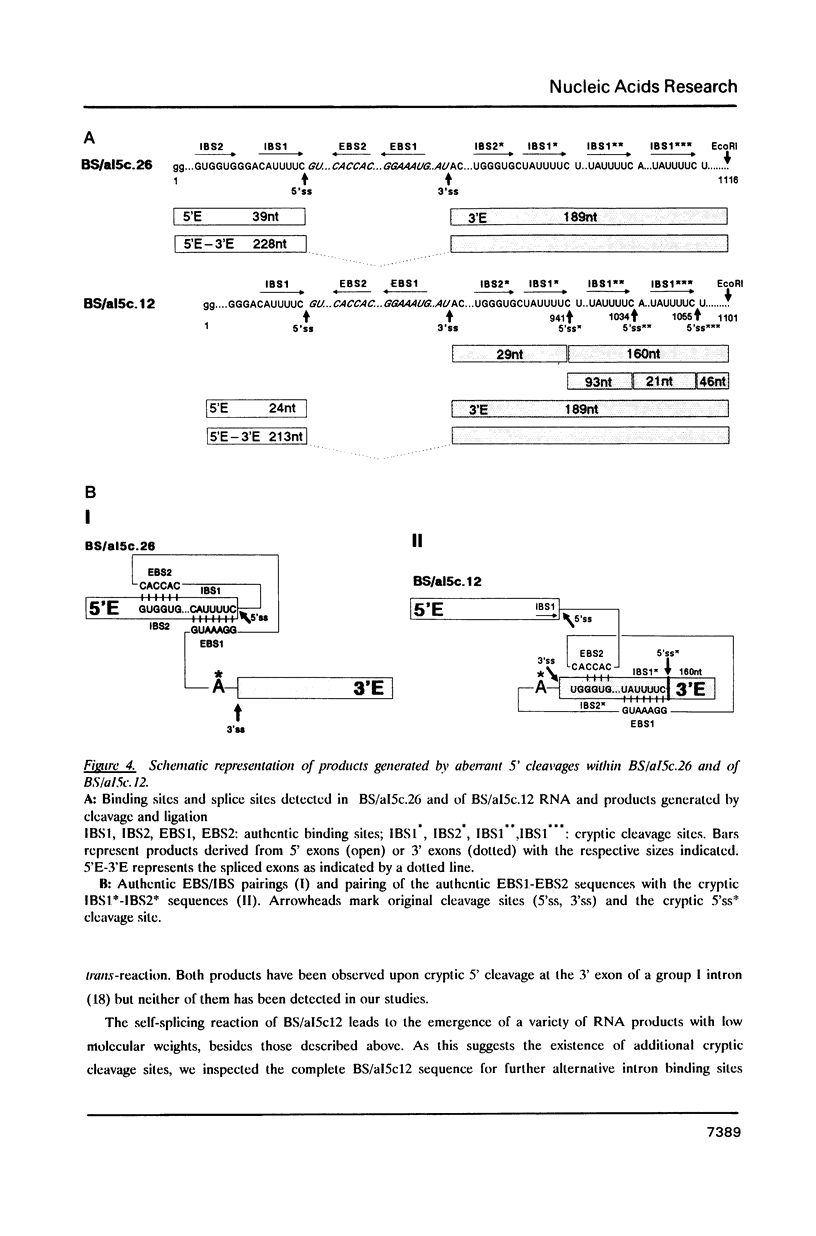
Recognition of 5' splice points by group I and group II self-splicing introns involves the interaction of exon sequences--directly preceding the 5' splice site--with intronic sequence elements. We show here that the exon binding sequences (EBS) of group II intron aI5c can accept various substitutes of the authentic intron binding sites (IBS) provided in cis or in trans. The efficiency of cleavages at these cryptic 5' splice sites was enhanced by deletion of the authentic IBS2 element. All cryptic 5' cleavage sites studied here were preceded by an IBS1 like sequence; indicating that the IBS1/EBS1 pairing alone is sufficient for proper 5' splice site selection by the intronic EBS element. The results are discussed in terms of minimal requirements for 5' cleavages and position effects of IBS sites relative to the intron.
Full text
PDF



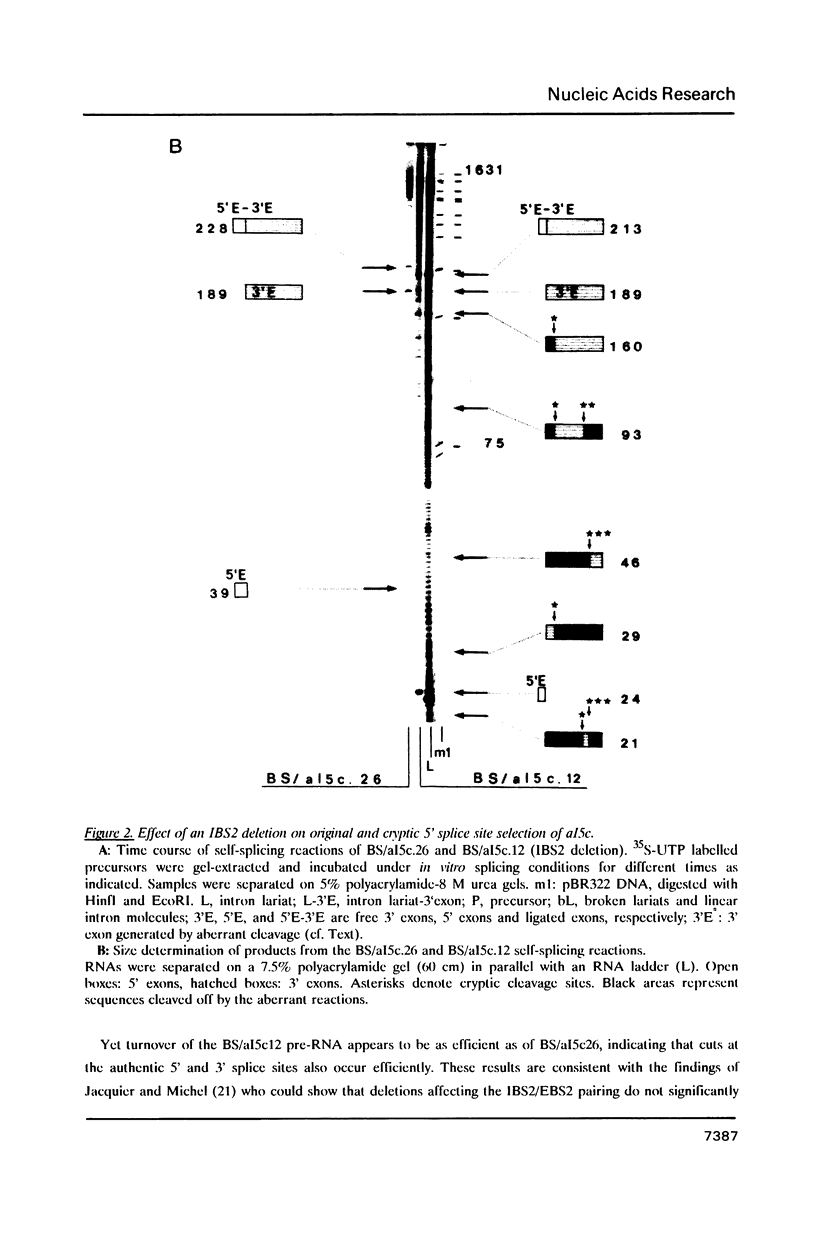






Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Been M. D., Cech T. R. One binding site determines sequence specificity of Tetrahymena pre-rRNA self-splicing, trans-splicing, and RNA enzyme activity. Cell. 1986 Oct 24;47(2):207–216. doi: 10.1016/0092-8674(86)90443-5. [DOI] [PubMed] [Google Scholar]
- Cech T. R., Zaug A. J., Grabowski P. J. In vitro splicing of the ribosomal RNA precursor of Tetrahymena: involvement of a guanosine nucleotide in the excision of the intervening sequence. Cell. 1981 Dec;27(3 Pt 2):487–496. doi: 10.1016/0092-8674(81)90390-1. [DOI] [PubMed] [Google Scholar]
- Chandry P. S., Belfort M. Activation of a cryptic 5' splice site in the upstream exon of the phage T4 td transcript: exon context, missplicing, and mRNA deletion in a fidelity mutant. Genes Dev. 1987 Nov;1(9):1028–1037. doi: 10.1101/gad.1.9.1028. [DOI] [PubMed] [Google Scholar]
- Chu F. K., Maley G. F., West D. K., Belfort M., Maley F. Characterization of the intron in the phage T4 thymidylate synthase gene and evidence for its self-excision from the primary transcript. Cell. 1986 Apr 25;45(2):157–166. doi: 10.1016/0092-8674(86)90379-x. [DOI] [PubMed] [Google Scholar]
- England T. E., Uhlenbeck O. C. 3'-terminal labelling of RNA with T4 RNA ligase. Nature. 1978 Oct 12;275(5680):560–561. doi: 10.1038/275560a0. [DOI] [PubMed] [Google Scholar]
- Frendewey D., Keller W. Stepwise assembly of a pre-mRNA splicing complex requires U-snRNPs and specific intron sequences. Cell. 1985 Aug;42(1):355–367. doi: 10.1016/s0092-8674(85)80131-8. [DOI] [PubMed] [Google Scholar]
- Henikoff S. Unidirectional digestion with exonuclease III creates targeted breakpoints for DNA sequencing. Gene. 1984 Jun;28(3):351–359. doi: 10.1016/0378-1119(84)90153-7. [DOI] [PubMed] [Google Scholar]
- Holl J., Rödel G., Schweyen R. J. Suppressor mutations identify box9 as a central nucleotide sequence in the highly ordered structure of intron RNA in yeast mitochondria. EMBO J. 1985 Aug;4(8):2081–2085. doi: 10.1002/j.1460-2075.1985.tb03895.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jacquier A., Michel F. Multiple exon-binding sites in class II self-splicing introns. Cell. 1987 Jul 3;50(1):17–29. doi: 10.1016/0092-8674(87)90658-1. [DOI] [PubMed] [Google Scholar]
- Jarrell K. A., Peebles C. L., Dietrich R. C., Romiti S. L., Perlman P. S. Group II intron self-splicing. Alternative reaction conditions yield novel products. J Biol Chem. 1988 Mar 5;263(7):3432–3439. [PubMed] [Google Scholar]
- Michel F., Dujon B. Conservation of RNA secondary structures in two intron families including mitochondrial-, chloroplast- and nuclear-encoded members. EMBO J. 1983;2(1):33–38. doi: 10.1002/j.1460-2075.1983.tb01376.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Peebles C. L., Perlman P. S., Mecklenburg K. L., Petrillo M. L., Tabor J. H., Jarrell K. A., Cheng H. L. A self-splicing RNA excises an intron lariat. Cell. 1986 Jan 31;44(2):213–223. doi: 10.1016/0092-8674(86)90755-5. [DOI] [PubMed] [Google Scholar]
- Perea J., Jacq C. Role of the 5' hairpin structure in the splicing accuracy of the fourth intron of the yeast cob-box gene. EMBO J. 1985 Dec 1;4(12):3281–3288. doi: 10.1002/j.1460-2075.1985.tb04078.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Price J. V., Engberg J., Cech T. R. 5' exon requirement for self-splicing of the Tetrahymena thermophila pre-ribosomal RNA and identification of a cryptic 5' splice site in the 3' exon. J Mol Biol. 1987 Jul 5;196(1):49–60. doi: 10.1016/0022-2836(87)90510-9. [DOI] [PubMed] [Google Scholar]
- Schmelzer C., Müller M. W. Self-splicing of group II introns in vitro: lariat formation and 3' splice site selection in mutant RNAs. Cell. 1987 Dec 4;51(5):753–762. doi: 10.1016/0092-8674(87)90098-5. [DOI] [PubMed] [Google Scholar]
- Schmelzer C., Schmidt C., May K., Schweyen R. J. Determination of functional domains in intron bI1 of yeast mitochondrial RNA by studies of mitochondrial mutations and a nuclear suppressor. EMBO J. 1983;2(11):2047–2052. doi: 10.1002/j.1460-2075.1983.tb01698.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tabak H. F., Van der Horst G., Osinga K. A., Arnberg A. C. Splicing of large ribosomal precursor RNA and processing of intron RNA in yeast mitochondria. Cell. 1984 Dec;39(3 Pt 2):623–629. doi: 10.1016/0092-8674(84)90469-0. [DOI] [PubMed] [Google Scholar]
- Waring R. B., Ray J. A., Edwards S. W., Scazzocchio C., Davies R. W. The Tetrahymena rRNA intron self-splices in E. coli: in vivo evidence for the importance of key base-paired regions of RNA for RNA enzyme function. Cell. 1985 Feb;40(2):371–380. doi: 10.1016/0092-8674(85)90151-5. [DOI] [PubMed] [Google Scholar]
- Waring R. B., Scazzocchio C., Brown T. A., Davies R. W. Close relationship between certain nuclear and mitochondrial introns. Implications for the mechanism of RNA splicing. J Mol Biol. 1983 Jul 5;167(3):595–605. doi: 10.1016/s0022-2836(83)80100-4. [DOI] [PubMed] [Google Scholar]
- Weiss-Brummer B., Holl J., Schweyen R. J., Rödel G., Kaudewitz F. Processing of yeast mitochondrial RNA: involvement of intramolecular hybrids in splicing of cob intron 4 RNA by mutation and reversion. Cell. 1983 May;33(1):195–202. doi: 10.1016/0092-8674(83)90348-3. [DOI] [PubMed] [Google Scholar]
- van der Horst G., Tabak H. F. Self-splicing of yeast mitochondrial ribosomal and messenger RNA precursors. Cell. 1985 Apr;40(4):759–766. doi: 10.1016/0092-8674(85)90335-6. [DOI] [PubMed] [Google Scholar]
- van der Veen R., Arnberg A. C., Grivell L. A. Self-splicing of a group II intron in yeast mitochondria: dependence on 5' exon sequences. EMBO J. 1987 Apr;6(4):1079–1084. doi: 10.1002/j.1460-2075.1987.tb04861.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- van der Veen R., Arnberg A. C., van der Horst G., Bonen L., Tabak H. F., Grivell L. A. Excised group II introns in yeast mitochondria are lariats and can be formed by self-splicing in vitro. Cell. 1986 Jan 31;44(2):225–234. doi: 10.1016/0092-8674(86)90756-7. [DOI] [PubMed] [Google Scholar]