Abstract
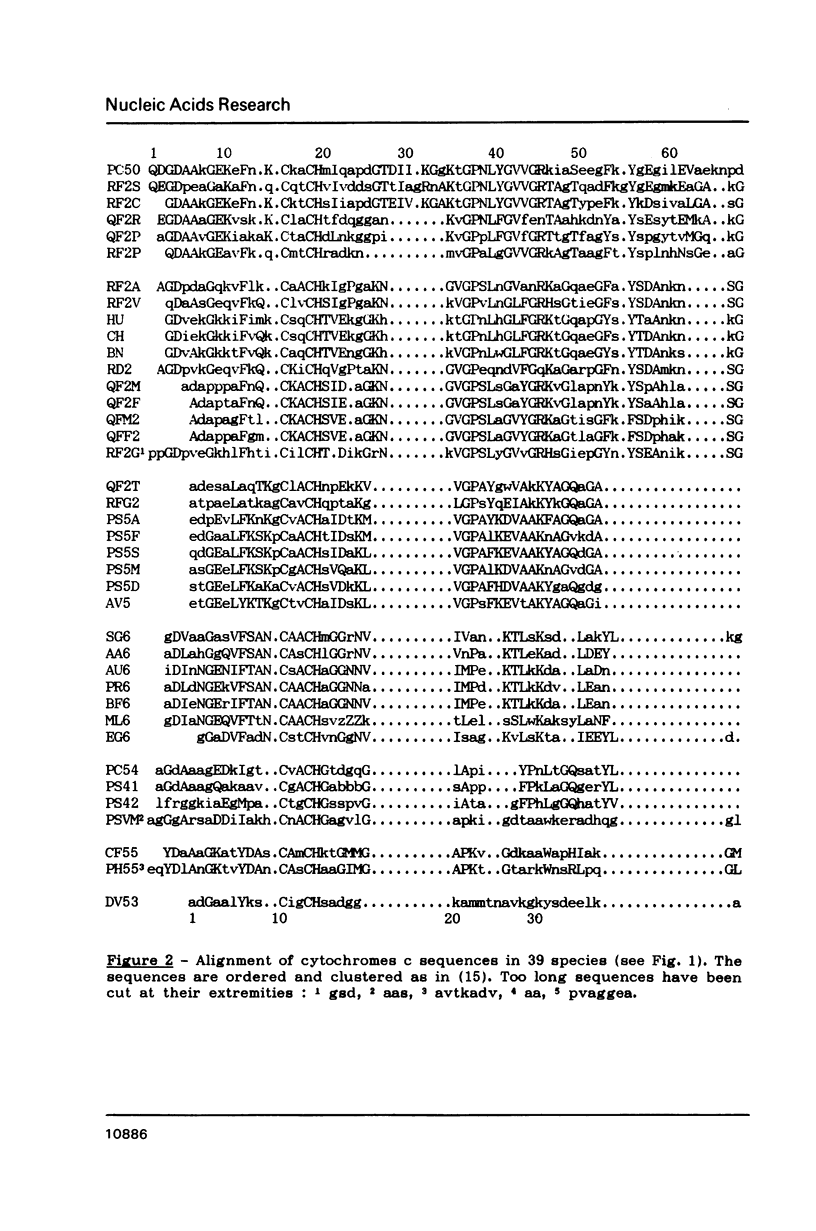
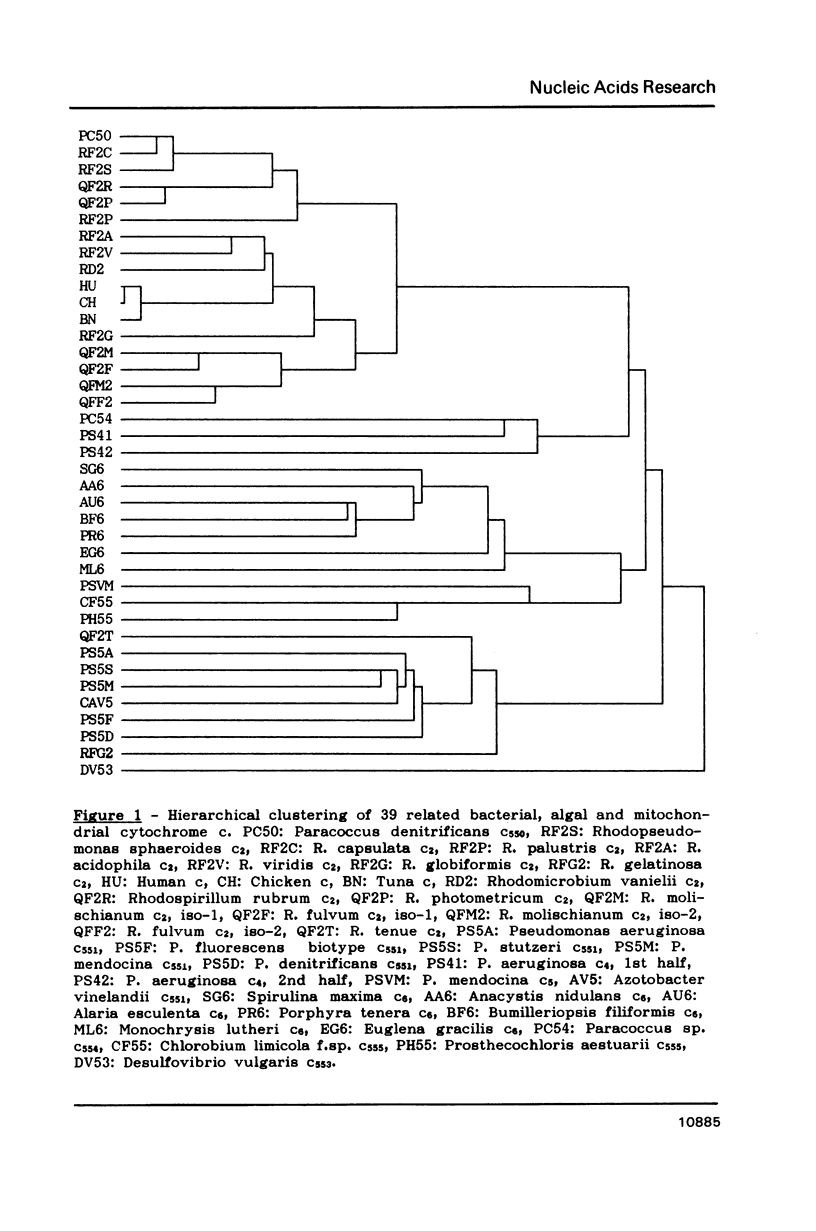
An algorithm is presented for the multiple alignment of sequences, either proteins or nucleic acids, that is both accurate and easy to use on microcomputers. The approach is based on the conventional dynamic-programming method of pairwise alignment. Initially, a hierarchical clustering of the sequences is performed using the matrix of the pairwise alignment scores. The closest sequences are aligned creating groups of aligned sequences. Then close groups are aligned until all sequences are aligned in one group. The pairwise alignments included in the multiple alignment form a new matrix that is used to produce a hierarchical clustering. If it is different from the first one, iteration of the process can be performed. The method is illustrated by an example: a global alignment of 39 sequences of cytochrome c.
Full text
PDF








Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Bacon D. J., Anderson W. F. Multiple sequence alignment. J Mol Biol. 1986 Sep 20;191(2):153–161. doi: 10.1016/0022-2836(86)90252-4. [DOI] [PubMed] [Google Scholar]
- Bains W. MULTAN: a program to align multiple DNA sequences. Nucleic Acids Res. 1986 Jan 10;14(1):159–177. doi: 10.1093/nar/14.1.159. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Barton G. J., Sternberg M. J. A strategy for the rapid multiple alignment of protein sequences. Confidence levels from tertiary structure comparisons. J Mol Biol. 1987 Nov 20;198(2):327–337. doi: 10.1016/0022-2836(87)90316-0. [DOI] [PubMed] [Google Scholar]
- Barton G. J., Sternberg M. J. Evaluation and improvements in the automatic alignment of protein sequences. Protein Eng. 1987 Feb-Mar;1(2):89–94. doi: 10.1093/protein/1.2.89. [DOI] [PubMed] [Google Scholar]
- Feng D. F., Doolittle R. F. Progressive sequence alignment as a prerequisite to correct phylogenetic trees. J Mol Evol. 1987;25(4):351–360. doi: 10.1007/BF02603120. [DOI] [PubMed] [Google Scholar]
- Gribskov M., Homyak M., Edenfield J., Eisenberg D. Profile scanning for three-dimensional structural patterns in protein sequences. Comput Appl Biosci. 1988 Mar;4(1):61–66. doi: 10.1093/bioinformatics/4.1.61. [DOI] [PubMed] [Google Scholar]
- Gribskov M., McLachlan A. D., Eisenberg D. Profile analysis: detection of distantly related proteins. Proc Natl Acad Sci U S A. 1987 Jul;84(13):4355–4358. doi: 10.1073/pnas.84.13.4355. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lipman D. J., Pearson W. R. Rapid and sensitive protein similarity searches. Science. 1985 Mar 22;227(4693):1435–1441. doi: 10.1126/science.2983426. [DOI] [PubMed] [Google Scholar]
- Martinez H. M. A flexible multiple sequence alignment program. Nucleic Acids Res. 1988 Mar 11;16(5):1683–1691. doi: 10.1093/nar/16.5.1683. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Murata M., Richardson J. S., Sussman J. L. Simultaneous comparison of three protein sequences. Proc Natl Acad Sci U S A. 1985 May;82(10):3073–3077. doi: 10.1073/pnas.82.10.3073. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Needleman S. B., Wunsch C. D. A general method applicable to the search for similarities in the amino acid sequence of two proteins. J Mol Biol. 1970 Mar;48(3):443–453. doi: 10.1016/0022-2836(70)90057-4. [DOI] [PubMed] [Google Scholar]
- Sankoff D., Cedergren R. J., Lapalme G. Frequency of insertion-deletion, transversion, and transition in the evolution of 5S ribosomal RNA. J Mol Evol. 1976 Mar 29;7(2):133–149. doi: 10.1007/BF01732471. [DOI] [PubMed] [Google Scholar]
- Santibánez M., Rohde K. A multiple alignment program for protein sequences. Comput Appl Biosci. 1987 Jun;3(2):111–114. doi: 10.1093/bioinformatics/3.2.111. [DOI] [PubMed] [Google Scholar]
- Sobel E., Martinez H. M. A multiple sequence alignment program. Nucleic Acids Res. 1986 Jan 10;14(1):363–374. doi: 10.1093/nar/14.1.363. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Taylor W. R. Multiple sequence alignment by a pairwise algorithm. Comput Appl Biosci. 1987 Jun;3(2):81–87. doi: 10.1093/bioinformatics/3.2.81. [DOI] [PubMed] [Google Scholar]


