Abstract
The spread of OXA-24/OXA-40 (OXA-24/40)-producing Acinetobacter spp. in the Iberian Peninsula has been strongly influenced by clonal expansion, but the role of horizontal gene transfer has scarcely been explored. blaOXA-24/40-carrying plasmids and genetic environments were characterized in representative (n = 15) Acinetobacter species clinical isolates (obtained between 2001 and 2007) by Acinetobacter baumannii PCR-based replicon typing, sequencing, hybridization, and restriction fragment length polymorphism. Besides the identification of blaOXA-24/40 within the chromosomes of some isolates, the circulation of common blaOXA-24/40-carrying plasmids (30-kb repA_AB; 10-kb aci2) and genetic backbones among Acinetobacter spp. was demonstrated.
TEXT
Carbapenem resistance in Acinetobacter spp. has been mostly associated with the production of OXA-23, OXA-24/OXA-40 (OXA-24/40), and OXA-58 carbapenem-hydrolyzing class D β-lactamases (1, 3, 5–12). Whereas OXA-23- and OXA-58-producing Acinetobacter baumannii are widely disseminated (3, 6, 12), OXA-24/40 producers have been reported mainly as endemic in the Iberian Peninsula, where the ongoing spread of OXA-24/40 since 1995 has been explained by clonal expansion (6, 10). In addition, there have been reports of sporadic cases in Italy and France and of outbreaks in the United States (5, 7, 8). Recently, some studies demonstrated the mobilization of blaOXA-24/40 by lateral gene transfer events (1, 5, 9, 10). In this study, the characterization of the diversity of OXA-24/40-encoding plasmids and the genetic environment of the blaOXA-24/40 genes was conducted in A. baumannii and Acinetobacter haemolyticus clinical isolates from Portugal.
Fifteen representative OXA-24/40-producing Acinetobacter spp. isolates (13 A. baumannii and 2 A. haemolyticus) were selected from a collection of 157 carbapenem-resistant Acinetobacter species clinical isolates from two geographically distant Portuguese hospitals (obtained between 2001 and 2007), (i) Hospital Geral de Santo António (HGSA) and (ii) Centro Hospitalar da Cova da Beira (CHCB) (Table 1) (6). Isolates were identified with the Vitek 2 system and 16S rRNA gene sequencing. Detection of the blaOXA-24/40 gene was performed as previously described (6, 10). Antibiotic susceptibility testing was done by Etest and the disk diffusion method following CLSI guidelines (4). Isolates were chosen on the basis of their ApaI pulsed-field gel electrophoresis (PFGE) patterns (4 subtypes arbitrarily designated A [n = 4; obtained between 2001 and 2004 from HGSA], A1 [n = 3; obtained between 2006 and 2007 from CHCB], A3 [n = 5; obtained in 2006 from HGSA], and A4 [n = 1; obtained in 2003 from HGSA]) and plasmid content (1 to 3 plasmids ranging from 10 kb to 90 kb). They all belonged to the ST98 lineage (6) according to the multilocus sequence typing (MLST) scheme proposed by Bartual et al. (2) (http://pubmlst.org/abaumannii/). This sequence type (ST) is a double-locus variant of the worldwide-disseminated ST92, which is endemic in Portuguese hospitals and has been identified in other European countries associated with the production of different OXA types (6, 12). Conjugative transfer (solid, 37°C) and transformation by electroporation were conducted in a subset of isolates using A. baumannii ATCC 19606, Escherichia coli K-12 BM21 (rifampin and nalidixic acid resistant, plasmid free), or Acinetobacter baylyi ADP1 as the recipient strain, as described previously (5). Transconjugants and transformants were selected using tryptic soy agar (TSA) plates supplemented with rifampin (100 μg/ml) and/or ampicillin (30 μg/ml) or imipenem (0.5 to 1 μg/ml), as reported. The location (chromosome/plasmid) of blaOXA-24/40 was assessed by hybridization of I-CeuI and S1 gels with specific probes (blaOXA-24/40, 16S rRNA genes) in the wild-type and transformant strains. Plasmid characterization was further accomplished by the recently proposed PCR-based replicon typing scheme for A. baumannii plasmids (AB-PBRT), sequencing and hybridization with replicase (rep) probes (3), and restriction fragment length polymorphism (RFLP) (EcoRI, BamHI, or HindIII).
Table 1.
Epidemiology and plasmid characterization of representative OXA-24/40-producing Acinetobacter species clinical isolates from Portugal
| Species | PFGE subtype (no. of isolates) | Yr(s) of isolation | Hospitala | blaOXA-24/40 plasmid size(s) (kb)b (RFLP profile) (no. of isolates) | rep contentb (plasmid size [kb]) | Antibiotic resistance profiled |
|---|---|---|---|---|---|---|
| A. baumannii | A (4) | 2001–2004 | HGSA | 90, 30 (A) (3) | repA_AB (30), repAci2, repAci6 | CAZ, ATM, IPM, MEM, PIP, FEP, CIP, TOB, TET, MIN |
| 2001–2004 | HGSA | 70, 30 (ND) (1) | repA_AB (30), repAci2, repAci6 | CAZ, ATM, IPM, MEM, PIP, FEP, CIP | ||
| A1 (3) | 2006–2007 | CHCB | 10c (B2) (3) | repA_AB, repAci2 (10), repAci6 | CAZ, ATM, IPM, MEM, PIP, FEP, CIP | |
| A3 (5) | 2006 | HGSA | 90, 10 (B2)c (1) | repA_AB, repAci2 (10), repAci6 | CAZ, ATM, IPM, MEM, PIP, FEP, CIP, (TOB) | |
| 2006 | HGSA | 30 (A1) (4) | repA_AB (30), repAci2, repAci6 | |||
| A4 (1) | 2003 | HGSA | 30 (ND), 10 (B)c (1) | repA_AB, repAci2(10) | CAZ, ATM, IPM, MEM, PIP, FEP, CIP, MIN | |
| A. haemolyticus | (2) | 2002 | HGSA | 90, 30 (A1′)c (2) | repA_AB (30), repAci2 | IPM, MEM, PIP, CIP, TOB, TET |
HGSA, Hospital Geral de Santo António (Northern region); CHCB, Centro Hospitalar da Cova da Beira (Center region).
Plasmid number, size, type, and rep content were determined by RFLP, PCR, and hybridization of S1- and I-CeuI-digested genomic DNA with specific probes (blaOXA-24/40, repAci2, rep_AB, and repAci6). ND, not done. Plasmids harboring blaOXA-24/40 appear in bold. Plasmid transfer by transformation is shown as underlined.
The blaOXA-24/40 probe additionally hybridized in a chromosomal band.
Parentheses indicate variability of a given resistance determinant. Resistance among transformants is represented by underlining. Abbreviations: CAZ, ceftazidime; ATM, aztreonam; IPM, imipenem; MEM, meropenem; PIP, piperacillin; FEP, cefepime; CIP, ciprofloxacin; TOB, tobramycin; TET, tetracycline; MIN, minocycline.
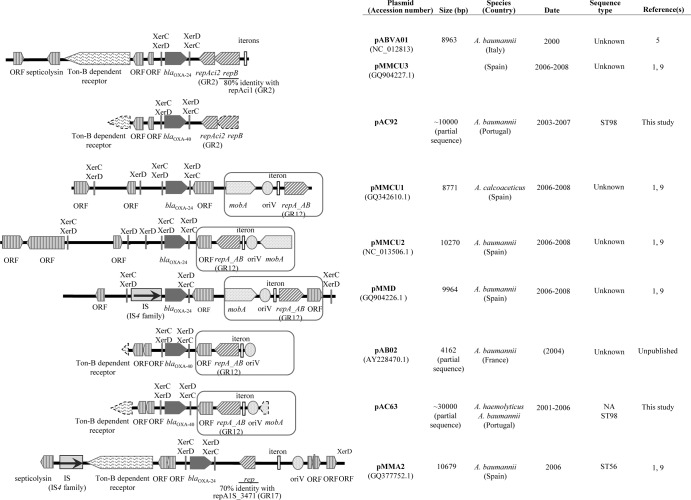
An in silico comparative analysis of six fully sequenced and one partially sequenced Acinetobacter species blaOXA-24/40-carrying plasmids deposited in the GenBank database (pABVA01, pMMCU3, pMMCU1, pMMCU2, pMMD, pAB02, and pMMA2, with the respective accession numbers NC_012813.1, GQ904227.1, GQ342610.1, NC_013506.1, GQ904226.1, AY228470.1, and GQ377752.1) was performed at the NCBI website using the BLAST software and the nucleotide database (http://blast.ncbi.nlm.nih.gov/Blast.cgi). Based on those sequences, the vicinity of the blaOXA-24/40 gene was characterized in representative plasmids from our collection by a PCR mapping strategy and further sequencing (Fig. 1).
Fig 1.
blaOXA-24/40 genetic surroundings observed in clinical Acinetobacter spp. from different countries that were identified in this and previous studies. Parentheses around a year indicate that the year of submission to GenBank is given because the isolation date is unknown. Discontinuous lines represent partial sequences. NA, not applicable; ORF, open reading frame.
A. baumannii isolates were resistant to almost all β-lactams, showing variable susceptibility to aztreonam, whereas A. haemolyticus isolates (n = 2) exhibited resistance to all β-lactams except cefepime, ceftazidime, and aztreonam. All isolates were resistant to ciprofloxacin, whereas susceptibility to aminoglycosides and tetracycline was variable, according to CLSI guidelines (Table 1) (4). As consistently observed in other studies, attempts to obtain OXA-24/40-producing transconjugants were unsuccessful (9). Transformants were obtained for A. baumannii isolates belonging to PFGE type A (n = 1) and PFGE type A4 (n = 1) and one A. haemolyticus isolate by using A. baylyi ADP1 as the receptor strain, and they acquired resistance to all β-lactams except ceftazidime, aztreonam, and cefepime. Resistance to other groups of antibiotics was not cotransferred. Variable plasmid size (90 kb, 70 kb, 30 kb and/or 10 kb) and rep content (GR2 group aci2 and/or aci6 and/or GR12 group repA_AB) were observed among the different species and PFGE types. The blaOXA-24/40 gene was identified within distinct plasmids (70 to 90 kb and 30 kb, 90 kb and 10 kb, 30 kb, or 10 kb) and additionally within the chromosomes of some A. baumannii and A. haemolyticus isolates (Table 1).
Highly related 30-kb blaOXA-24/40-carrying plasmids (arbitrarily designated RFLP pattern A) were identified in A. baumannii (PFGE subtypes A and A3) and A. haemolyticus isolates collected within the same hospital (obtained between 2001 and 2006) (Table 1). These plasmids belonged to the GR12 group (3) and showed rep content (100% identity to repA_AB) and sequences surrounding blaOXA-24/40 identical to those of the partially sequenced prototype pAB02 plasmid (GenBank accession number AY228470.1) (Fig. 1). A module consisting of an open reading frame (ORF) of unknown function, the replicase repA_AB, oriV, and mobA (represented within a rectangle in Fig. 1) was also found within that structure. Interestingly, it has been previously identified in other 10-kb repA_AB plasmids and seems to be a recurrent platform in plasmids (10 kb/30 kb) from different A. baumannii clones and Acinetobacter species in several European countries (Portugal, Spain, and France) (Fig. 1) (1, 9). The presence of mob genes probably facilitates plasmid mobilization by self-transmissible plasmids and may ultimately explain their occurrence in non-A. baumannii species (3, 9, 10).
A 10-kb OXA-24/40-encoding plasmid (arbitrarily designated RFLP pattern B) was also frequently identified among A. baumannii isolates (PFGE subtypes A1, A3, and A4) collected from different hospitals (obtained between 2003 and 2007) (Table 1). This plasmid belonged to the GR2 group (3), with a rep sequence identical to that of aci2 and a blaOXA-24/40 genetic environment closely related to those from the pABVA01 and pMMCU3 plasmids (GenBank accession numbers NC_012813 and GQ904227.1, respectively) identified in A. baumannii isolates from Italy and Spain, respectively (1, 5, 9).
The same rep genes (GR2, GR6, and GR12) have been presumptively associated with other OXA-24/40-producing A. baumannii plasmids from Portugal and Spain by AB-PBRT, thus suggesting the circulation of a common plasmid pool (3, 12). Both blaOXA-24/40 genes and larger contiguous modules are flanked by XerC/XerD-like binding sites (Fig. 1), which might be responsible for their mobilization (5, 9) and may explain the identification of blaOXA-24/40 and/or common modules in different plasmid scaffolds or in chromosomes. Furthermore, the variability observed in the number and location of the orf of unknown function, oriV, and iteron sequences suggests frequent recombinatorial events (5, 9). This study not only confirms the horizontal transmission of common resistance modules between Acinetobacter plasmids but also highlights interspecies plasmid transfer. Although conjugative transfer was not detected, the aci6 replicase (GR6; 100% identity to that of plasmid pACICU2 [GenBank accession number NC_010606]) was able to mediate horizontal transfer of antibiotic resistance plasmids, as previously suggested (3, 12). Moreover, a recently described mechanism involving the release of outer membrane vesicles harboring blaOXA-24/40-carrying plasmids might also shed some light on this process (11). It is interesting to highlight that the blaOXA-24/40 genetic environment and plasmid types are different from those observed for blaOXA-23 or blaOXA-58, thus suggesting independent acquisition and subsequent selection events (3, 12).
In this study, common blaOXA-24/40-carrying plasmids (30-kb repA_AB and 10-kb aci2) were identified among representative ST98 A. baumannii and A. haemolyticus clinical isolates collected over a large period of time (2001 to 2007). In addition, with a systematic in silico analysis, we were able to identify the same plasmid types and closely related blaOXA-24/40 genetic surroundings in OXA-24/40-producing isolates from different European countries. These findings, together with some previous evidence for horizontal gene transfer of blaOXA-24/40-carrying modules, strongly supports the role of common platforms and plasmids in the dissemination process of OXA-24/40 among Acinetobacter spp.
Nucleotide sequence accession numbers.
The nucleotide sequences obtained in this study that correspond to the representative 30-kb repA_AB (pAC63) and 10-kb aci2 (pAC92) OXA-24/40-encoding plasmids have been submitted to the GenBank database (GenBank accession numbers JN982951 and JN982952, respectively).
ACKNOWLEDGMENTS
The Acinetobacter baylyi ADP1 receptor strain was kindly provided by German Bou.
Filipa Grosso is a Ph.D. student from Fundação para a Ciência e a Tecnologia (SFRH/BD/31647/2006). Ângela Novais was supported by a Marie Curie Intra European Fellowship within the 7th European Community Framework Programme (PIEF-GA-2009-255512). This study was funded by Fundação para a Ciência e a Tecnologia through grant no. PEst-C/EQB/LA0006/2011.
Footnotes
Published ahead of print 23 April 2012
REFERENCES
- 1. Acosta J, et al. 2011. Multidrug-resistant Acinetobacter baumannii harboring OXA-24 carbapenemase, Spain. Emerg. Infect. Dis. 17:1064–1067 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2. Bartual SG, et al. 2005. Development of a multilocus sequence typing scheme for characterization of clinical isolates of Acinetobacter baumannii. J. Clin. Microbiol. 43:4382–4390 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3. Bertini A, et al. 2010. Characterization and PCR-based replicon typing of resistance plasmids in Acinetobacter baumannii. Antimicrob. Agents Chemother. 54:4168–4177 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4. Clinical and Laboratory Standards Institute 2010. Performance standards for antimicrobial susceptibility testing; 20th informational supplement. CLSI M100-S20. Clinical and Laboratory Standards Institute, Wayne, PA [Google Scholar]
- 5. D'Andrea MM, et al. 2009. Characterization of pABVA01, a plasmid encoding the OXA-24 carbapenemase from Italian isolates of Acinetobacter baumannii. Antimicrob. Agents Chemother. 53:3528–3533 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6. Grosso F, Quinteira S, Peixe L. 2011. Understanding the dynamics of imipenem-resistant Acinetobacter baumannii lineages within Portugal. Clin. Microbiol. Infect. 17:1275–1279 [DOI] [PubMed] [Google Scholar]
- 7. Héritier C, Poirel L, Aubert D, Nordmann P. 2003. Genetic and functional analysis of the chromosome-encoded carbapenem-hydrolyzing oxacillinase OXA-40 of Acinetobacter baumannii. Antimicrob. Agents Chemother. 47:268–273 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8. Lolans K, Rice TW, Munoz-Price LS, Quinn JP. 2006. Multicity outbreak of carbapenem-resistant Acinetobacter baumannii isolates producing the carbapenemase OXA-40. Antimicrob. Agents Chemother. 50:2941–2945 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9. Merino M, et al. 2010. OXA-24 carbapenemase gene flanked by XerC/XerD-like recombination sites in different plasmids from different Acinetobacter species isolated during a nosocomial outbreak. Antimicrob. Agents Chemother. 54:2724–2727 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10. Quinteira S, Grosso F, Ramos H, Peixe L. 2007. Molecular epidemiology of imipenem-resistant Acinetobacter haemolyticus and Acinetobacter baumannii isolates carrying plasmid-mediated OXA-40 from a Portuguese hospital. Antimicrob. Agents Chemother. 51:3465–3466 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11. Rumbo C, et al. 2011. Horizontal transfer of the OXA-24 carbapenemase gene via outer membrane vesicles: a new mechanism of dissemination of the carbapenem resistance genes in Acinetobacter baumannii. Antimicrob. Agents Chemother. 55:3084–3090 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12. Towner KJ, et al. 2011. Distribution of intrinsic plasmid replicase genes and their association with carbapenem-hydrolyzing class D β-lactamase genes in European clinical isolates of Acinetobacter baumannii. Antimicrob. Agents Chemother. 55:2154–2159 [DOI] [PMC free article] [PubMed] [Google Scholar]