Abstract
In bacterial chemotaxis, transmembrane chemoreceptors, the CheA histidine kinase, and the CheW coupling protein assemble into signaling complexes that allow bacteria to modulate their swimming behavior in response to environmental stimuli. Among the protein-protein interactions in the ternary complex, CheA-CheW and CheW-receptor interactions were studied previously, whereas CheA-receptor interaction has been less investigated. Here, we characterize the CheA-receptor interaction in Thermotoga maritima by NMR spectroscopy and validate the identified receptor binding site of CheA in Escherichia coli chemotaxis. We find that CheA interacts with chemoreceptor in a manner similar to CheW, and the receptor binding site of CheA's regulatory domain is homologous to that of CheW. Collectively, the receptor binding sites in CheA-CheW complex suggest that conformational changes in CheA are required for assembly of the CheA-CheW-receptor ternary complex and CheA activation.
Keywords: signal transduction, NMR spectroscopy, protein-protein interaction, histidine kinase, MCP
Introduction
Motile bacteria exploit the chemotaxis signal transduction pathway to search for the best living conditions1–4. The components involved in the pathway in Escherichia coli include transmembrane chemoreceptors and six soluble Che proteins. The transmembrane chemoreceptors, also known as methyl-accepting chemotaxis proteins (MCPs), interact with CheA, CheW, CheB, and CheR to allow bacteria to detect, convert, and adjust signals with high sensitivity, dynamic range, and remarkable gain. In complex with CheW and MCPs, the kinase CheA converts the signal transduced by chemoreceptors to a flux of phosphoryl groups in the cytosol at the expense of ATP. Upon negative stimuli (repellent increases or attractant decreases), one stream of the phosphoryl groups is transferred to the response regulator, CheY. The phosphorylated CheY interacts directly with the flagellar motor to promote clockwise flagellar rotation that switches the default swimming behavior of the bacterium to immediate tumbling. The other stream is received as a negative feedback signal by CheB, whose methylesterase activity sets MCPs back to their pre-stimulus state. The methyltransferase, CheR, methylates MCPs at several conserved glutamic acid residues.
In addition to membrane spanning helices, most chemoreceptors consist of periplasmic, HAMP, and cytoplasmic domains. Receptors bind attractants and repellents via the periplasmic domain. The HAMP domain acts as a signal conversion module and transfers the input signal from the periplasmic domain to the cytoplasmic domain where CheA and CheW interact with the receptor. The diversity of the detection by the periplasmic domains of different receptors and the unique signaling pathway in the cytoplasm are well reflected by sequence alignment showing ligand binding domains vary significantly among chemoreceptors, while cytoplasmic signaling domains are highly conserved from species to species and from receptor to receptor.
Dimeric CheA consists of five domains: the histidine-containing phosphortransfer domain (P1), the CheY/CheB binding domain (P2), the dimerization domain (P3), the catalytic/ATP binding domain (P4), and the regulatory domain (P5). CheA utilizes ATP that binds to the P4 domain to phosphorylate a histidine residue in the P1 domain. This trans-autophosphorylation is modulated by the interaction network among chemoreceptors, CheW, and the P5 domain of CheA5–9. CheW serves as an adaptor that is essential for the formation of receptor-CheW-CheA complex and CheA activation7.
The Crane laboratory developed a model for the ternary complex of the receptor-CheW-CheA complex using distances derived from dipolar interactions of nitroxide spin label pairs10. In collaboration with the Jensen laboratory, they have also published a structural model based a combination low-resolution X-ray and electron tomography data11. These models provide a picture of the overall organization of ternary complex showing a planar organization of CheA and CheW molecules organized near the receptor tips. However, these models lack detailed information about the residue-to-residue contacts in the complex.
Here, we examine the direct interaction between CheA and a chemoreceptor, TM001412, from Thermotoga maritima with nuclear magnetic resonance (NMR) techniques. This structural information is further validated in the analogous and well-studied Escherichia coli system. Combined with previous structural studies on the CheA-CheW13 and CheW-receptor14 interactions, these observations suggest that conformational changes in CheA are required for the formation of the signaling complex.
Results
Attempts to study the interaction between chemoreceptor and CheA from E. coli were largely unsuccessful due to solubility properties of E. coli CheA constructs and poor NMR spectral qualities of E. coli chemoreceptor cytoplasmic fragments (data not shown). Instead, we chose to use CheA and a chemoreceptor from the hyperthermophile T. maritima that has a chemotaxis system with components similar to mesophilic bacteria15. These proteins showed higher solubilities and better NMR spectra than their E. coli counterparts. Another advantage of using thermophilic proteins is that we are able to obtain high signal-to-noise ratio in the NMR spectra due to improved NMR relaxation properties at higher temperatures. To further facilitate the high-resolution NMR study of receptor-CheA interaction, we decided to use monomeric CheAΔ354, a construct that contains only P4 and P5 domains of CheA, and TM001490-20614, a truncated form of TM0014 chemoreceptor. CheAΔ354 does not include the dimerization domain (P3), which greatly improved the signal-to-noise ratio of its NMR spectra. TM0014 is a soluble chemoreceptor that lacks the transmembrane region12; a truncated form of this receptor called TM001490-206 was used previously for the receptor-CheW interaction studies and showed structural properties very similar to the full-length counterpart14. Although the use of these constructs has greatly reduced the sizes of the resulting binary and ternary complexes, handling such complexes is still challenging for conventional 15N-based NMR methods. Therefore, we exploited the 13C methyl TROSY methodology that enabled us to study complexes of macromolecules of molecular mass 100 kDa or greater16, 17.
NMR analysis of the binding of CheAΔ354 to TM001490-206
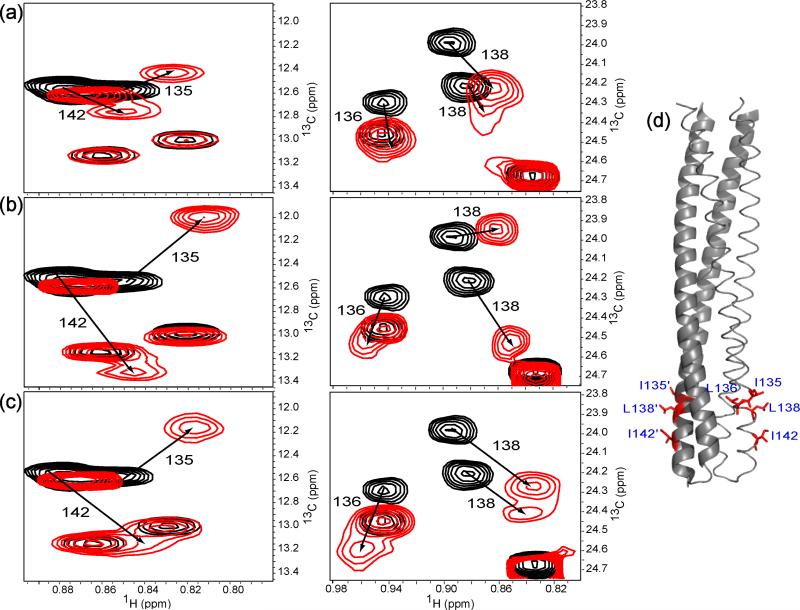
With the published backbone resonance assignment of TM001490-20614, we used the “out-and-back” transfer NMR assignment strategy18 to assign the methyl side chain resonances of isoleucine, leucine, and valine residues on the same construct. All the isoleucine residues were completely assigned; some of the leucine and valine residues were not assigned due to the ambiguity resulting from peak overlap in the spectra. The interaction of TM001490-206 with CheA was observed using a series of methyl-TROSY spectra of 13C methyl-labeled TM001490-206 taken over the course of a titration with the unlabeled CheAΔ354. This interaction was in the fast-exchange regime on the NMR time scale and the resonances of Ile135, Leu136, Leu138, and Ile142 showed significant chemical shift changes (Figure 1a). These residues were mapped onto the crystal structure of TM001412 where they formed a hydrophobic patch at the tip region of the receptor (Figure 1d). Interestingly, the resonances corresponding to these residues were also found to be perturbed as a result of CheW binding to the receptor (Figure 1b).
Fig. 1.
NMR analysis of the bindings of unlabeled CheAΔ354 and CheW to 13C-methyl labeled TM001490-206. (a) Methyl TROSY sections (left, Ile region; right, Leu region) of TM001490-206 (150 μM) in the presence (red, at 1:4 ratio of TM001490-206:CheAΔ354) and absence (black) of CheAΔ354. Residues with significant chemical shift changes are numbered with the arrows indicating the direction of the peak movement. (b) Same Sections of TM001490-206 (150 μM) spectra with (red, at 1:4 ratio of TM001490-206: CheW) and without (black) CheW. (c) Same sections of TM001490-206 (150 μM) spectra before (black) and after (red, at 1:4 ratio of TM001490-206:CheAΔ354-CheW) the addition of CheAΔ354-CheW complex. (d) Residues showing significant chemical shift changes are mapped onto the structure of TM0014 dimer (one monomer shown in cartoon and the other in ribbon format; PDB ID: 3G67).
To further examine the CheA-receptor interaction in the CheA-CheW-receptor ternary complex, 13C methyl-labeled TM001490-206 was titrated with unlabeled CheW-CheAΔ354 binary complex. (CheW and CheAΔ354 form a tight complex with a dissociation constant of ~10 nM13.) We observed that the formation of the CheA-CheW-receptor ternary complex perturbed the chemical shifts of these same residues (Figure 1c). However, the directions and magnitudes of the observed chemical shift changes differed in the ternary complex (Figure 1c) from those in either binary complex (Figure 1 a and b). This suggests that TM001490-206 interacts with both CheAΔ354 and CheW in solution.
NMR analysis of the binding of TM001490-206 to CheAΔ354
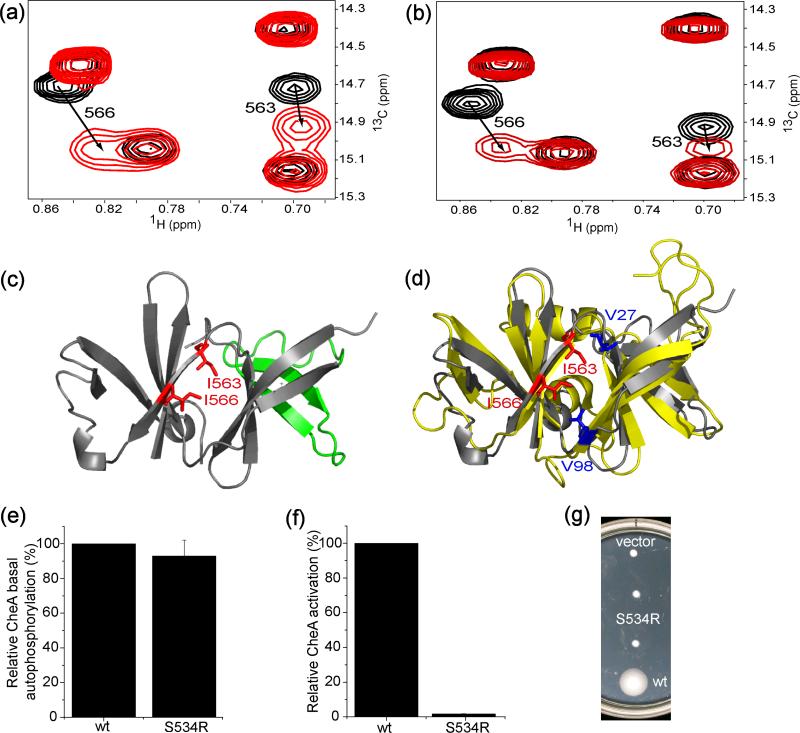
The quality of the backbone amide spectra of the construct of CheAΔ354 (35 kDa) was poor: most amide resonances of the residues in the P5 domain were either very weak or even invisible, and the addition of the P5 domain deteriorated the quality of the resonances of the P4 domain (Supplemental Fig. S1 a and b). However, the δ1 methyl groups of the isoleucine residues in CheAΔ354 showed well-resolved resonances in the methyl TROSY spectrum likely due to the increased sensitivity and faster time scale motions of the methyl side-chains. The 21 isoleucine residues in the 132 amino acid P5 domain provided sufficient probes for NMR interaction studies on this domain. Thus, we performed similar titration experiments with CheAΔ354 13C methyl-labeled at isoleucine residues and unlabeled TM001490-206 to examine the receptor interaction site of CheA. The resonances from the isoleucine residues in the P4 domain, whose assignments were transferred from those made with isolated P4 domain, did not shift at all (Supplemental Fig. S1 c). Two isoleucine methyl resonances from the P5 domain showed significant chemical shift changes (Figure 2a and Supplemental Fig. S1 c), indicating that TM001490-206 made direct contact with the P5 domain. Due to the absence of the amide backbone assignments, we were not able to assign the methyl resonances from the P5 domain using conventional NMR-based assignment strategy. Instead, mutagenesis was used to assign them, especially the two resonances that shifted upon TM001490-206 binding (Supplemental Fig. S1 d and e). Mapping the isoleucine residues (Ile563 and Ile566) that showed chemical shift changes onto the crystal structure of the P5 domain19, we found that they occupied the interface between the two β-barrel subunits of the P5 domain (Figure 2c).
Fig. 2.
NMR and mutational analysis of the receptor binding site of CheA. (a) Methyl TROSY section of 13C-methyl labeled CheAΔ354 (150 μM) in the presence (red, at 1:4 ratio of CheAΔ354:TM001490-206) and absence (black) of TM001490-206. Residues with significant chemical shift changes are numbered with the arrows indicating the direction of the peak movement. (b) Same section of the spectra of methyl-labeled CheAΔ354 (150 μM) in complex with unlabeled CheW before (black) and after (red, 1:4 ratio of CheAΔ354-CheW: TM001490-206) the addition of TM001490-206. (c) Residues (red) showing significant chemical shift changes are mapped onto the structure of the P5 domain of CheA (PDB ID: 1B3Q). The region involved in CheW binding is colored in green13. (d) Structural alignment of CheW (yellow, PDB ID: 1K0S) and the P5 domain of CheA (grey). The two residues (Val27 and Val98) that show the largest shifts in the CheW-receptor interaction14 are highlighted in blue, and Ile563 and Ile566 that shift upon receptor binding are in red. (e) ATPase assays (s.d., n=3) for the basal autophosphorylation activities, (f) in vitro CheA activation assays (s.d., n=3) for the activation abilities, and (g) in vivo swarm assays for the chemotactic abilities of the wild-type E. coli CheA and S534R mutant (duplicate) at the receptor binding site of the P5 domain determined by NMR.
We titrated the binary CheAΔ354-CheW complex with TM001490-206 while observing the methyl resonances of isoleucine labeled CheAΔ354. We found that, upon the addition of the receptor fragment, both Ile563 and Ile566 also underwent chemical shift changes similar to those observed when forming the CheAΔ354-receptor complex (Figure 2b), while other resonances did not shift. We conclude that these CheA residues interact similarly with receptor in both the CheA-receptor binary complex and the CheA-CheW-receptor ternary complex. In a parallel experiment where we titrated the binary CheAΔ354-CheW complex with TM001490-206 while observing the resonances of 13C methyl-labeled CheW, the chemical shifts of the resonances of the CheW residues involved in the CheW- TM001490-206 interaction14 were also perturbed and in fast-exchange regime on the NMR time scale (data not shown). These results show that TM001490-206 rapidly binds to both CheW and the P5 domain in the CheAΔ354-CheW complex.
Interestingly, the P5 domain of CheA shows substantial sequence and folding topology similarities to CheW20. Structural alignment shows that residues Ile563 and Ile566 of CheA are located in a similar region to the residues of CheW that bind receptor (Figure 2d). We conclude that this region of CheW and of the P5 domain of CheA have similar receptor binding function. This conclusion echoes with the previous observation that CheA and CheW bound to the same residues of TM001490-206, providing the structural basis for the competition between CheW and CheA for chemoreceptors21 (see discussion).
Validation of the receptor-CheA interaction by mutagenesis
We mutated Ile142 of TM001490-206 and Ile566 of CheAΔ354, the residues that showed the largest chemical shift changes in the CheAΔ354-TM001490-206 titrations, to confirm the binding interfaces of the receptor and CheA determined from our previous NMR experiments. Even when a five fold excess of unlabeled I566R mutant of CheAΔ354 and I142R mutant of TM001490-206 were added to 13C methyl-labeled wild-type TM001490-206 and CheAΔ354, respectively, there were not any significant changes in either the chemical shifts or the intensities of the resonances in the spectra (Supplemental Fig. S1 f and g; S2 a). Moreover, the addition of unlabeled I142R mutant of TM001490-206 did not cause any significant shift or broadening in CheW resonances (Supplemental Fig. S2 b). These results indicated that, Ile566 of CheAΔ354 disrupted receptor-CheA interaction, while Ile142 of TM001490-206 disrupted both receptor-CheA and receptor-CheW interactions.
The ability of receptor fragments to bind CheA and CheW does not necessarily lead to CheA activation. CheA activation appears to need higher oligomers of the complex involving trimers of dimers of receptor. TM0014 does not appear to support the formation of trimers of dimers12 despite the high sequence similarities to their E.coli counterparts and its ability to bind T. maritima CheW and CheA. To investigate whether the interaction observed by NMR using T. maritima proteins was also important for CheA activation in other bacterial chemotaxis systems, we mutated residue 534 (corresponding to Ile566 in T. maritima CheA) at the corresponding receptor binding site in E. coli CheA. The single amino acid replacement did not affect CheA basal autophosphorylation (Figure 2e). However, the kinase activation of the CheA mutant by E. coli chemoreceptor Tsr decreased dramatically (Figure 2f). In vivo chemotaxis assays also showed that the host strain RP9535 (ΔCheA) carrying the CheA mutant was defective in forming chemotactic swarms, in contrast to the complementation by wild-type CheA (Figure 2g). In addition, previous chemical modification studies on S. typhimurium CheA identified that residue 545 (corresponding to Ile560 in T. maritima CheA), which resides in the same region in the structure, was important for in vitro CheA activation22. Therefore, the interaction characterized by NMR in T. maritima was of functional relevance and crucial for CheA receptor-coupled activation in E.coli chemotaxis.
Discussion
Previous studies7, 21 on the interactions among CheA, CheW, and receptor show that CheA and CheW each promote incorporation of the other onto membrane-associating chemoreceptor, and they compete for chemoreceptors at over-stoichiometric ratios. Our results are able to reconcile these seemingly contradictory observations: synergy in binding is the consequence of the multiple interactions between chemoreceptors and the CheW-CheA complex, and the competition between CheA and CheW for chemoreceptors can be explained by the result that CheW and CheA bound to the same site on chemoreceptors due to the structural homology between CheW and the P5 domain of CheA.
Clustering primarily to the poles of E. coli 23, 24, dimeric chemoreceptors interact to form trimers of dimers that are approximately hexagonally arranged in the plane of the inner membrane25-27. Trimers of dimers28 are widely accepted to be the basic structural and functional units that are required for chemotaxis by the cryo-EM images25–27 and by mutagenesis and crosslinking studies on the residues involved in the trimer-of-dimers contact 29–31. Previous mutagenesis and chemical modification studies have shown that the residues on the NMR-determined CheA-binding surface of chemoreceptor are important for CheA activation and chemotaxis30, 32, 33. However, we find that the receptor residues that we identify as the CheW/CheA interaction site are also involved in the Tsr trimer-of-dimers interaction (Supplemental Fig. S3). This suggests that the biochemical role of the residues at the tip region of chemoreceptors is more complex, including interactions with CheA and CheW as well as receptor-receptor interaction. Therefore, assigning one simple biochemical defect to a chemotaxis-defective mutant at these receptor residues seems difficult and requires additional experimentation.
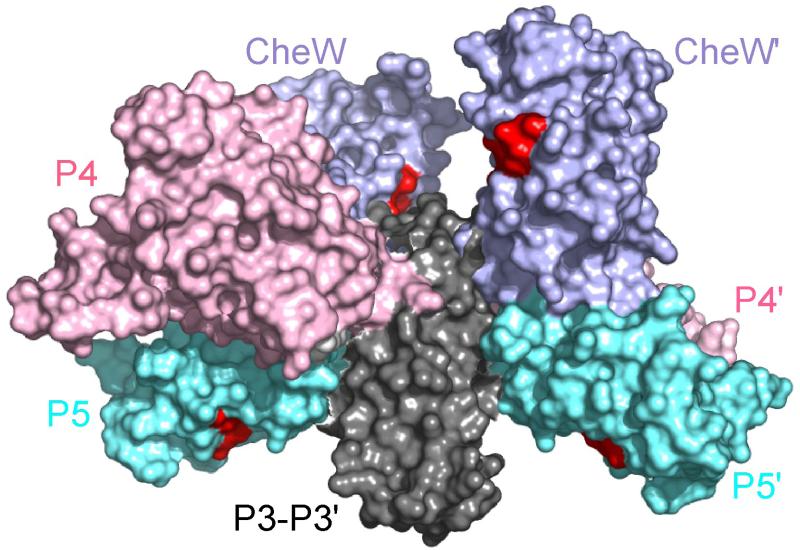
From our chemical shift perturbation results of the receptor-CheA interaction and our previous NMR studies on the interaction between CheW and TM001490-20614, we are able to map the receptor binding sites in the reconstructed structure of dimeric P3P4P5 of CheA complexed with two CheW molecules13, 19 (Figure 3). In this context, the distance between the receptor binding sites on the P5 domain and on CheW is approximately 55 Å. This means that one receptor dimer whose diameter is about 25 Å is not able to bind the two sites simultaneously. This is consistent with the high stoichiometries between chemoreceptors and CheA observed in a series of independent experiments using both the full length and cytoplasmic domains of chemoreceptors34–36.
Fig. 3.
Receptor binding sites of the CheA–CheW complex. The binding sites determined by NMR are shown in red.
The sequence of cytoplasmic domains of chemoreceptors are highly conserved throughout the Bacteria and Archaea37, 38 and contain the determinants for forming complexes with CheA and CheW34, 35, 39, 40. Studies from cryo-EM suggest that the tip regions of chemoreceptors where CheA and CheW bind form a thin layer parallel to the cytoplasmic membrane23, 25-27. However, when we combine the CheW and P5 interaction sites deduced from NMR with the crystal structures of the CheA dimer19 and the CheA-CheW complex13, we see a non-planar distribution of receptor binding sites (Figure 3), apparently inconsistent with the EM images. Studies from bioinformatics as well as cryo-EM27, 37 show that the lengths of the cytoplasmic domains of all the MCPs in a specific species of bacteria are constant, suggesting that the same binding sites shared by CheA and CheW at the tip region of chemoreceptors are planar as well. It also seems unlikely for the cytoplasmic domain of membrane-associating chemoreceptors to undergo a vertical axis displacement in order to bind the non-planar subunits of CheA-CheW complex. Moreover, even if it does, it appears that one of the P4 domains in a CheA dimer would be buried in the receptor stalks or collide with receptor. Since kinase activation by the formation of the signaling complex occurs in CheA, it seems more rational that the CheA-CheW complex undergoes conformational changes driven by binding to the plane defined by the tip region of transmembrane chemoreceptors, which may lead CheA towards activation. Such conformational changes have recently been seen in the low resolution crystal structure of T. maritima P4P5-CheW-Tm14s ternary complex, where CheW and the P5 domain are aligned to the CheA/CheW binding site of the chemoreceptor11. The CheA-receptor interaction sites in the crystal structure are in agreement with the NMR results11. However, in the crystal structure the orientation of the P5-receptor interaction is opposite to that of the CheW-receptor interaction, which conflicts with the trans-membrane nature of chemoreceptors and is likely nonnative. Our NMR-directed mutagenesis studies establish that this binding interface is indeed important for assembly of the CheA-CheW-receptor ternary complex and CheA activation in E. coli system as well.
Finally, this NMR study on the protein-protein interactions in the chemotaxis signaling complex is another example of investigating high molecule weight proteins/complexes with methyl-TROSY41-44. Especially, in this case where the P5 domain is not stable in the absence of the adjoining P4 domain and the backbone amide resonances of the P5 domain could not be obtained in the P4P5 construct, the methyl probes provided high-quality spectra allowing us to investigate the interactions that involve the P5 domain. In subsequent communications, we will continue to use this method to monitor CheA activation by suitable receptor constructs.
Experimental Procedures
Protein preparation
The gene encoding CheAΔ354 (P4-P5 domain, 355-671) was subcloned into the vector pET22b (Novagen), and the expressed protein with a C-terminal His6 tag was purified using nickel-NTA affinity chromatography (Qiagen) and FPLC (Pharmacia). TM001490-206 construct and its purification were previously described in Vu et al 14. Methyl-labeled samples were prepared using the labeling protocol previously created by Kay laboratory45. Purified proteins were dialyzed against the buffer containing 50 mM Na2HPO4 (pH7.4) and 1 mM EDTA and concentrated to a final concentration of 0.2-2 mM. All NMR samples contained 0.02% sodium azide and 10% D2O. E.coli CheA46, CheW47, and CheY48 were expressed in strain RP3098 and purified following published protocols; Tsr-containing membranes were prepared as previously described5. All point mutations were introduced by QuickChange mutagenesis (Stratagene).
Nuclear magnetic resonance (NMR) spectroscopy
NMR data were collected at 40 °C on a Varian 600MHz or a Bruker 800MHz spectrometer each equipped with a 1H/13C/15N/2H cryogenically cooled probe and a Z pulsed-field gradient. The 1H/13C methyl resonances of TM001490-206 were assigned using methyl-TROSY COSY-based out-and-back transfer scheme and the assignment was confirmed by NOE correlation17, 18. All the titration experiments were carried out at 40 °C using 2D methyl-TROSY experiments.
ATPase Assay
The auto-phosphorylation rates of CheA variants were determined using the ATPase assay described in Ninfa et al 49. Briefly, the reactions were initiated by the addition of E. coli CheA proteins (2-10 μM, final concentration) to the solution containing 100 mM potassium phosphate (pH 7.5), 5 mM MgCl2, 0.1 mM EDTA, 0.1 mM dithiothreitol, 1.5 mM ATP, 1.5 mM phosphoenolpyruvate, 0.25 mM NADH, 6.0 units of pyruvate kinase, 7.5 units of lactate dehydrogenase, and excess (100 μM) CheY. NADH oxidation was monitored in the Amersham Ultrospec3300 spectrophotometer at room temperature. The ATP hydrolysis rate was reflected by the slope of a linear decay in NADH absorbance at 340 nm and determined using a value of 6,220 M-1cm-1 for the extinction coefficient of NADH.
CheA activation Assay
CheA activation assays were done with Tsr-containing membranes as previously described5. Briefly, the phosphorylation reaction mixtures contained 50mM Tris-HCl (pH 7.5), 100 mM KCl, 10 mM MgCl2, 2.5 μM CheA, 5 μM CheW, 100 μM CheY and 13 μM Tsr in membranes in a total volume of 18 μl. The mixtures were pre-incubated at room temperature for 15 min, and then the reactions were initiated by the addition of 2 μl of 10 mM [γ-32P] ATP (8,000 cpm/pmol). After 15 s, the reactions were quenched by mixing the samples with 20 μl of 2× Laemmli SDS buffer supplemented with 100 mM EDTA. The samples were immediately subjected to SDS-PAGE. Gels were dried, and the amount of phosphorylated CheY was quantitated by phosphor-imaging.
Swarm Assay
The chemotactic abilities of RP9535 (ΔcheA1643) cells carrying mutant pKJ9 plasmids were tested on tryptone soft agar, according to the protocol developed by Parkinson Laboratory50.
Supplementary Material
Highlights.
CheA, CheW, and chemoreceptors form chemotaxis signaling complexes.
CheA interacts with chemoreceptor in a manner similar to CheW.
Receptor binding site of CheA's regulatory domain is homologous to that of CheW.
Receptor binding sites suggest conformational changes in CheA during CheA activation.
Acknowledgements
We thank all the members in the Parkinson laboratory (University of Utah) for providing strains and advice, and Hongjun Zhou (University of California Santa Barbara) for critical reading of the manuscript. This work was supported by National Institutes of Health grant GM59544 to F.W.D.
Abbreviations
- TROSY
transverse relaxation optimized spectroscopy
- MCP
methyl-accepting chemotaxis proteins
- HAMP
histidine kinases, adenyl cyclases, methyl-accepting proteins and phosphatases
- EM
electron microscopy
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
References
- 1.Wadhams GH, Armitage JP. Making sense of it all: bacterial chemotaxis. Nat. Rev. Mol. Cell. Biol. 2004;5:1024–1037. doi: 10.1038/nrm1524. [DOI] [PubMed] [Google Scholar]
- 2.Hazelbauer GL, Falke JJ, Parkinson JS. Bacterial chemoreceptors: high-performance signaling in networked arrays. Trends Biochem. Sci. 2008;33:9–19. doi: 10.1016/j.tibs.2007.09.014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Hazelbauer GL, Lai WC. Bacterial chemoreceptors: providing enhanced features to two-component signaling. Curr. Opin. Microbiol. 2010;13:124–132. doi: 10.1016/j.mib.2009.12.014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Sourjik V, Armitage JP. Spatial organization in bacterial chemotaxis. EMBO J. 2010;29:2724–2733. doi: 10.1038/emboj.2010.178. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Borkovich KA, Kaplan N, Hess JF, Simon MI. Transmembrane signal transduction in bacterial chemotaxis involves ligand-dependent activation of phosphate group transfer. Proc. Natl. Acad. Sci. USA. 1989;86:1208–1212. doi: 10.1073/pnas.86.4.1208. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Borkovich KA, Simon MI. The dynamics of protein phosphorylation in bacterial chemotaxis. Cell. 1990;63:1339–1348. doi: 10.1016/0092-8674(90)90429-i. [DOI] [PubMed] [Google Scholar]
- 7.Gegner JA, Graham DR, Roth AF, Dahlquist FW. Assembly of an MCP receptor, CheW, and kinase CheA complex in the bacterial chemotaxis signal transduction pathway. Cell. 1992;70:975–982. doi: 10.1016/0092-8674(92)90247-a. [DOI] [PubMed] [Google Scholar]
- 8.Bourret RB, Davagnino J, Simon MI. The carboxy-terminal portion of the CheA kinase mediates regulation of autophosphorylation by transducer and CheW. J. Bacteriol. 1993;175:2097–2101. doi: 10.1128/jb.175.7.2097-2101.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Swanson RV, Bourret RB, Simon MI. Intermolecular complementation of the kinase activity of CheA. Mol. Microbiol. 1993;8:435–441. doi: 10.1111/j.1365-2958.1993.tb01588.x. [DOI] [PubMed] [Google Scholar]
- 10.Bhatnagar J, Borbat PP, Pollard AM, Bilwes AM, Freed JH, Crane BR. Structure of the ternary complex formed by a chemotaxis receptor signaling domain, the CheA histidine kinase, and the coupling protein CheW as determined by pulsed dipolar ESR spectroscopy. Biochemistry. 2011;49:3824–3841. doi: 10.1021/bi100055m. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Briegel A, Li X, Bilwes AM, Hughes KT, Jensen GJ, Crane BR. Bacterial chemoreceptor arrays are hexagonally packed trimers of receptor dimers networked by rings of kinase and coupling proteins. Proc. Natl. Acad. Sci. USA. 2012;109:3766–3771. doi: 10.1073/pnas.1115719109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Pollard AM, Bilwes AM, Crane BR. The structure of a soluble chemoreceptor suggests a mechanism for propagating conformational signals. Biochemistry. 2009;48:1936–1944. doi: 10.1021/bi801727m. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Park SY, Borbat PP, Gonzalez-Bonet G, Bhatnagar J, Pollard AM, Freed JH, Bilwes AM, Crane BR. Reconstruction of the chemotaxis receptor-kinase assembly. Nat. Struct. Mol. Biol. 2006;13:400–407. doi: 10.1038/nsmb1085. [DOI] [PubMed] [Google Scholar]
- 14.Vu A, Wang X, Zhou H, Dahlquist FW. The receptor-CheW binding interface in bacterial chemotaxis. J. Mol. Biol. 2012;415:759–767. doi: 10.1016/j.jmb.2011.11.043. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Swanson RV, Sanna MG, Simon MI. Thermostable chemotaxis proteins from the hyperthermophilic bacterium Thermotoga maritima. J. Bacteriol. 1996;178:484–489. doi: 10.1128/jb.178.2.484-489.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Hamel DJ, Dahlquist FW. The contact interface of a 120 kD CheA-CheW complex by methyl TROSY interaction spectroscopy. J. Am. Chem. Soc. 2005;127:9676–9677. doi: 10.1021/ja052517m. [DOI] [PubMed] [Google Scholar]
- 17.Sprangers R, Kay LE. Quantitative dynamics and binding studies of the 20S proteasome by NMR. Nature. 2007;445:618–622. doi: 10.1038/nature05512. [DOI] [PubMed] [Google Scholar]
- 18.Tugarinov V, Kay LE. Ile, Leu, and Val methyl assignments of the 723-residue malate synthase G using a new labeling strategy and novel NMR methods. J. Am. Chem. Soc. 2003;125:13868–13878. doi: 10.1021/ja030345s. [DOI] [PubMed] [Google Scholar]
- 19.Bilwes AM, Alex LA, Crane BR, Simon MI. Structure of CheA, a signal-transducing histidine kinase. Cell. 1999;96:131–141. doi: 10.1016/s0092-8674(00)80966-6. [DOI] [PubMed] [Google Scholar]
- 20.Griswold IJ, Zhou H, Matison M, Swanson RV, McIntosh LP, Simon MI, Dahlquist FW. The solution structure and interactions of CheW from Thermotoga maritima. Nat. Struct. Biol. 2002;9:121–125. doi: 10.1038/nsb753. [DOI] [PubMed] [Google Scholar]
- 21.Levit MN, Grebe TW, Stock JB. Organization of the receptor-kinase signaling array that regulates Escherichia coli chemotaxis. J. Biol. Chem. 2002;277:36748–36754. doi: 10.1074/jbc.M204317200. [DOI] [PubMed] [Google Scholar]
- 22.Miller AS, Kohout SC, Gilman KA, Falke JJ. CheA Kinase of bacterial chemotaxis: chemical mapping of four essential docking sites. Biochemistry. 2006;45:8699–8711. doi: 10.1021/bi060580y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Zhang P, Khursigara CM, Hartnell LM, Subramaniam S. Direct visualization of Escherichia coli chemotaxis receptor arrays using cryo-electron microscopy. Proc. Natl. Acad. Sci. USA. 2007;104:3777–3781. doi: 10.1073/pnas.0610106104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Maddock JR, Shapiro L. Polar location of the chemoreceptor complex in the Escherichia coli cell. Science. 1993;259:1717–1723. doi: 10.1126/science.8456299. [DOI] [PubMed] [Google Scholar]
- 25.Briegel A, Ding HJ, Li Z, Werner J, Gitai Z, Dias DP, Jensen RB, Jensen GJ. Location and architecture of the Caulobacter crescentus chemoreceptor array. Mol. Microbiol. 2008;69:30–41. doi: 10.1111/j.1365-2958.2008.06219.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Khursigara CM, Wu X, Subramaniam S. Chemoreceptors in Caulobacter crescentus: trimers of receptor dimers in a partially ordered hexagonally packed array. J. Bacteriol. 2008;190:6805–6810. doi: 10.1128/JB.00640-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Briegel A, Ortega DR, Tocheva EI, Wuichet K, Li Z, Chen S, Müller A, Iancu CV, Murphy GE, Dobro MJ, et al. Universal architecture of bacterial chemoreceptor arrays. Proc. Natl. Acad. Sci. USA. 2009;106:17181–17186. doi: 10.1073/pnas.0905181106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Kim KK, Yokota H, Kim SH. Four-helical-bundle structure of the cytoplasmic domain of a serine chemotaxis receptor. Nature. 1999;400:787–792. doi: 10.1038/23512. [DOI] [PubMed] [Google Scholar]
- 29.Studdert CA, Parkinson JS. Insights into the organization and dynamics of bacterial chemoreceptor clusters through in vivo crosslinking studies. Proc. Natl. Acad. Sci. USA. 2005;102:15623–15628. doi: 10.1073/pnas.0506040102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Ames P, Studdert CA, Reiser RH, Parkinson JS. Collaborative signaling by mixed chemoreceptor teams in Escherichia coli. Proc. Natl. Acad. Sci. USA. 2002;99:7060–7065. doi: 10.1073/pnas.092071899. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Studdert CA, Parkinson JS. Crosslinking snapshots of bacterial chemoreceptor squads. Proc. Natl. Acad. Sci. USA. 2004;101:2117–2122. doi: 10.1073/pnas.0308622100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Mehan RS, White NC, Falke JJ. Mapping out regions on the surface of the aspartate receptor that are essential for kinase activation. Biochemistry. 2003;42:2952–2959. doi: 10.1021/bi027127g. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Gosink KK, Zhao Y, Parkinson JS. Mutational analysis of N381, a key trimer contact residue in Tsr, the Escherichia coli serine chemoreceptor. J Bacteriol. 2011;193:6452–6460. doi: 10.1128/JB.05887-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Ames P, Parkinson JS. Constitutively signaling fragments of Tsr, the Escherichia coli serine chemoreceptor. J. Bacteriol. 1994;176:6340–6348. doi: 10.1128/jb.176.20.6340-6348.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Liu Y, Levit M, Lurz R, Surette MG, Stock JB. Receptor-mediated protein kinase activation and the mechanism of transmembrane signaling in bacterial chemotaxis. EMBO J. 1997;16:7231–7240. doi: 10.1093/emboj/16.24.7231. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Li M, Hazelbauer GL. Cellular stoichiometry of the components of the chemotaxis signaling complex. J. Bacteriol. 2004;186:3687–3694. doi: 10.1128/JB.186.12.3687-3694.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Le Moual H, Koshland DE., Jr Molecular evolution of the C-terminal cytoplasmic domain of a superfamily of bacterial receptors involved in taxis. J. Mol. Biol. 1996;261:568–585. doi: 10.1006/jmbi.1996.0483. [DOI] [PubMed] [Google Scholar]
- 38.Alexander RP, Zhulin IB. Evolutionary genomics reveals conserved structural determinants of signaling and adaptation in microbial chemoreceptors. Proc. Natl. Acad. Sci. USA. 2007;104:2885–2890. doi: 10.1073/pnas.0609359104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Francis NR, Wolanin PM, Stock JB, Derosier DJ, Thomas DR. Three-dimensional structure and organization of a receptor/signaling complex. Proc. Natl. Acad. Sci. USA. 2004;101:17480–17485. doi: 10.1073/pnas.0407826101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Wolanin PM, Baker MD, Francis NR, Thomas DR, DeRosier DJ, Stock JB. Self-assembly of receptor/signaling complexes in bacterial chemotaxis. Proc. Natl. Acad. Sci. USA. 2006;103:14313–14318. doi: 10.1073/pnas.0606350103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Religa TL, Sprangers R, Kay LE. Dynamic regulation of archaeal proteasome gate opening as studied by TROSY NMR. Science. 2010;328:98–102. doi: 10.1126/science.1184991. [DOI] [PubMed] [Google Scholar]
- 42.Ruschak AM, Religa TL, Breuer S, Witt S, Kay LE. The proteasome antechamber maintains substrates in an unfolded state. Nature. 2010;467:868–871. doi: 10.1038/nature09444. [DOI] [PubMed] [Google Scholar]
- 43.Velyvis A, Yang YR, Schachman HK, Kay LE. A solution NMR study showing that active site ligands and nucleotides directly perturb the allosteric equilibrium in aspartate transcarbamoylase. Proc. Natl. Acad. Sci. USA. 2007;104:8815–8820. doi: 10.1073/pnas.0703347104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Gelis I, Bonvin AM, Keramisanou D, Koukaki M, Gouridis G, Karamanou S, Economou A, Kalodimos CG. Structural basis for signal-sequence recognition by the translocase motor SecA as determined by NMR. Cell. 2007;131:756–769. doi: 10.1016/j.cell.2007.09.039. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Tugarinov V, Kanelis V, Kay LE. Isotope labeling strategies for the study of high-molecular-weight proteins by solution NMR spectroscopy. Nat. Protoc. 2006;1:749–754. doi: 10.1038/nprot.2006.101. [DOI] [PubMed] [Google Scholar]
- 46.Stock A, Chen T, Welsh D, Stock J. CheA protein, a central regulator of bacterial chemotaxis, belongs to a family of proteins that control gene expression in response to changing environmental conditions. Proc. Natl. Acad. Sci. USA. 1988;85:1403–1407. doi: 10.1073/pnas.85.5.1403. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Gegner JA, Dahlquist FW. Signal transduction in bacteria: CheW forms a reversible complex with the protein kinase CheA. Proc. Natl. Acad. Sci. USA. 1991;88:750–754. doi: 10.1073/pnas.88.3.750. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Lowry DF, Roth AF, Rupert PB, Dahlquist FW, Moy FJ, Domaille PJ, Matsumura P. Signal transduction in chemotaxis. A propagating conformation change upon phosphorylation of CheY. J. Biol. Chem. 1994;269:26358–26362. [PubMed] [Google Scholar]
- 49.Ninfa EG, Stock A, Mowbray S, Stock J. Reconstitution of the bacterial chemotaxis signal transduction system from purified components. J. Biol. Chem. 1991;266:9764–9770. [PubMed] [Google Scholar]
- 50.Zhao J, Parkinson JS. Mutational analysis of the chemoreceptor-coupling domain of the Escherichia coli chemotaxis signaling kinase CheA. J. Bacteriol. 2006;188:3299–3307. doi: 10.1128/JB.188.9.3299-3307.2006. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.