Abstract
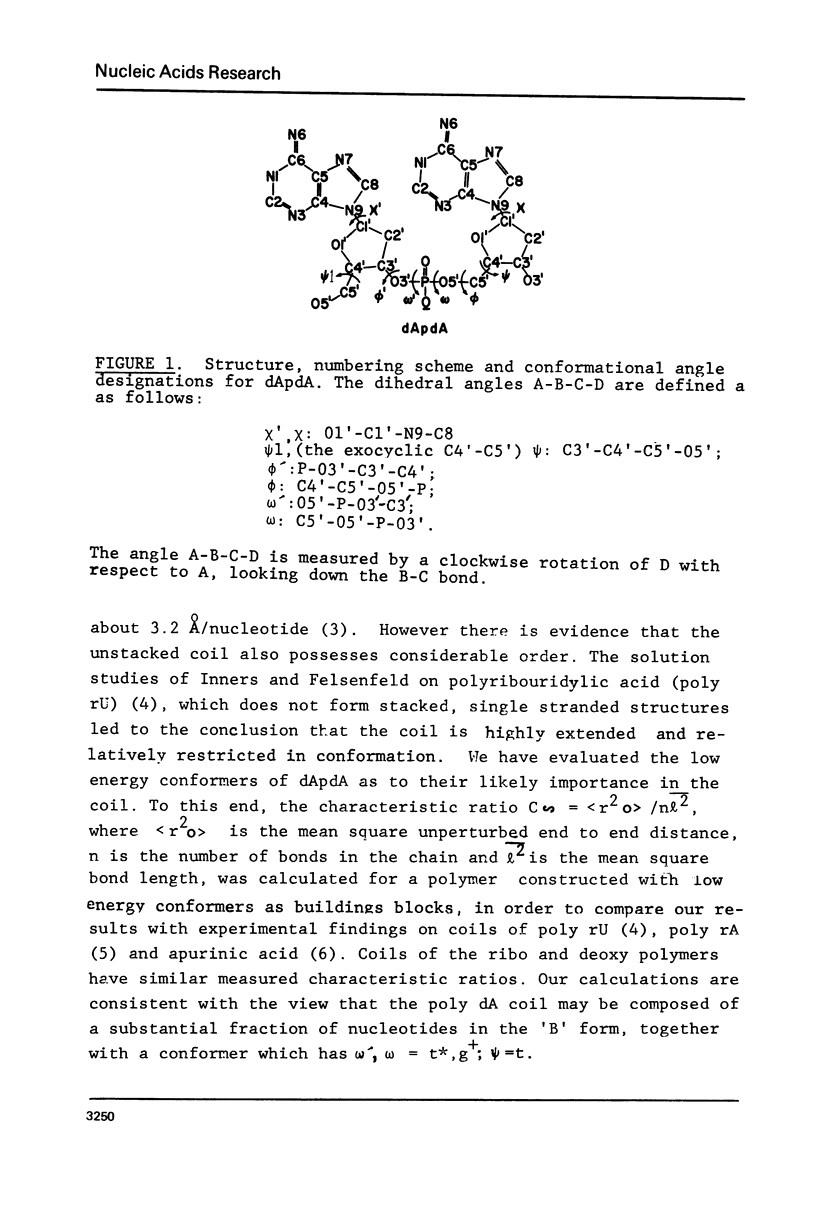
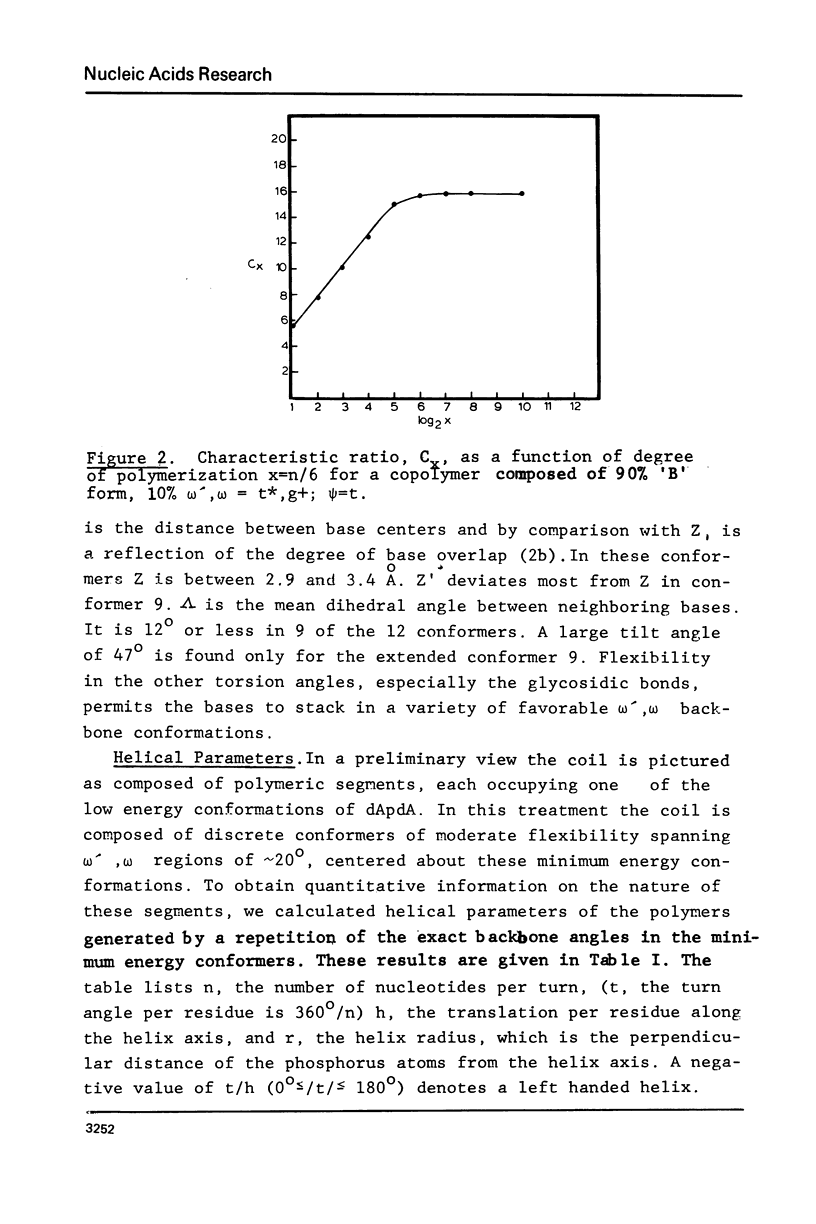
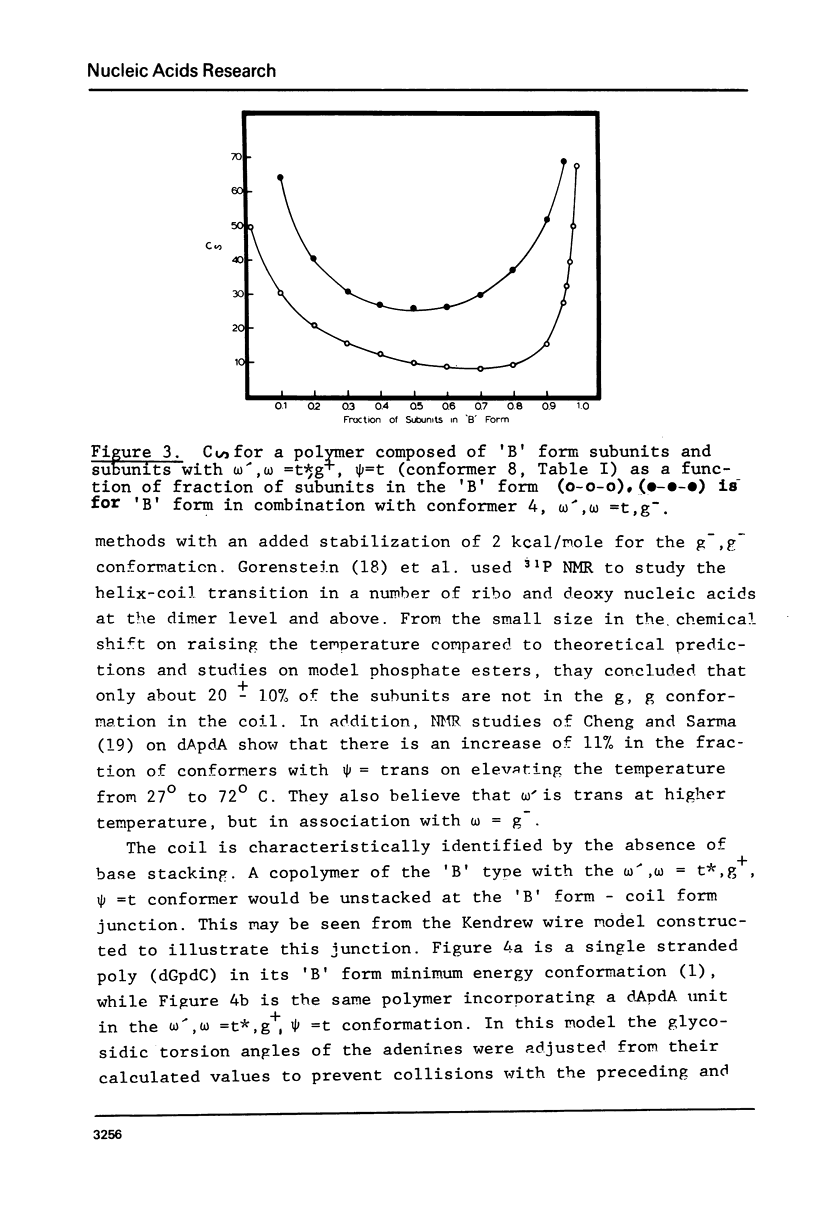
The minimum energy conformations of dApdA have been examined for their suitability as buildings blocks of the single stranded coil form of polynucleotides. Calculations of the characteristic ratio C difference = less than ro greater than 2/n liter2 were made for a polymer generated from all the low energy conformers, as well as for selected combinations. A polymer composed of a conformer with omega', omega = t*,g+,(skewed) psi = t, C-(2)-endo type pucker, in combination with the 'B' form, has a C difference equal to that observed in coils of apurinic acid (6) when the fraction of 'B' form conformers is approximately 25% and approximately 91%. The t*,g+ conformer is the second lowest energy form in the C-(2)-endo puckering domain, following the 'B' form.
Full text
PDF







Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Achter E. K., Felsenfeld G. The conformation of single-strand polynucleotides in solution: sedimentation studies of apurinic acid. Biopolymers. 1971;10(9):1625–1634. doi: 10.1002/bip.360100916. [DOI] [PubMed] [Google Scholar]
- Altona C., Sundaralingam M. Conformational analysis of the sugar ring in nucleosides and nucleotides. A new description using the concept of pseudorotation. J Am Chem Soc. 1972 Nov 15;94(23):8205–8212. doi: 10.1021/ja00778a043. [DOI] [PubMed] [Google Scholar]
- Arnott S., Selsing E. Structures for the polynucleotide complexes poly(dA) with poly (dT) and poly(dT) with poly(dA) with poly (dT). J Mol Biol. 1974 Sep 15;88(2):509–521. doi: 10.1016/0022-2836(74)90498-7. [DOI] [PubMed] [Google Scholar]
- Broyde S. B., Wartell R. M., Stellman S. D., Hingerty B., Langridge R. Classical potential energy calculations for ApA, CpC, GpG, and UpU. The influence of the bases on RNA subunit conformations. Biopolymers. 1975 Aug;14(8):1597–1613. doi: 10.1002/bip.1975.360140805. [DOI] [PubMed] [Google Scholar]
- Cheng D. M., Sarma R. H. Intimate details of the conformational characteristics of deoxyribodinucleoside monophosphates in aqueous solution. J Am Chem Soc. 1977 Oct 26;99(22):7333–7348. doi: 10.1021/ja00464a038. [DOI] [PubMed] [Google Scholar]
- Eisenberg H., Felsenfeld G. Studies of the temperature-dependent conformation and phase separation of polyriboadenylic acid solutions at neutral pH. J Mol Biol. 1967 Nov 28;30(1):17–37. doi: 10.1016/0022-2836(67)90240-9. [DOI] [PubMed] [Google Scholar]
- Fang K. N., Kondo N. S., Miller P. S., Ts'o P. O. Proton magnetic resonance study on adenine dideoxynucleoside monophosphate with emphasis on the furanose conformation. J Am Chem Soc. 1971 Dec;93(24):6647–6656. doi: 10.1021/ja00753a053. [DOI] [PubMed] [Google Scholar]
- Gorenstein D. G., Findlay J. B., Momii R. K., Luxon B. A., Kar D. Temperature dependence of the 31P chemical shifts of nucleic acids. A prode of phosphate ester torsional conformations. Biochemistry. 1976 Aug 24;15(17):3796–3803. doi: 10.1021/bi00662a023. [DOI] [PubMed] [Google Scholar]
- Hingerty B., Broyde S. Helix geometry of single stranded DNA 'A' and 'B' forms from minimum energy conformations of dimeric subunits. Nucleic Acids Res. 1978 Jan;5(1):127–137. doi: 10.1093/nar/5.1.127. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Inners L. D., Felsenfeld G. Conformation of polyribouridylic acid in solution. J Mol Biol. 1970 Jun 14;50(2):373–389. doi: 10.1016/0022-2836(70)90199-3. [DOI] [PubMed] [Google Scholar]
- Miller P. S., Fang K. N., Kondo N. S., Ts'o P. O. Syntheses and properties of adenine and thymine nucleoside alkyl phosphotriesters, the neutral analogs of dinucleoside monophosphates. J Am Chem Soc. 1971 Dec;93(24):6657–6665. doi: 10.1021/ja00753a054. [DOI] [PubMed] [Google Scholar]
- Olson W. K. Configurational statistics of polynucleotide chains. A single virtual bond treatment. Macromolecules. 1975 May-Jun;8(3):272–275. doi: 10.1021/ma60045a006. [DOI] [PubMed] [Google Scholar]
- Olson W. K., Flory P. J. Spatial configuration of polynucleotide chains. II. Conformational energies and the average dimensions of polyribonucleotides. Biopolymers. 1972 Jan;11(1):25–56. doi: 10.1002/bip.1972.360110103. [DOI] [PubMed] [Google Scholar]
- Olson W. K., Flory P. J. Spatial configurations of polynucleotide chains. 3. Polydeoxyribonucleotides. Biopolymers. 1972 Jan;11(1):57–66. doi: 10.1002/bip.1972.360110104. [DOI] [PubMed] [Google Scholar]
- Olson W. K., Flory P. J. Spatial configurations of polynucleotide chains. I. Steric interactions in polyribonucleotides: a virtual bond model. Biopolymers. 1972 Jan;11(1):1–23. doi: 10.1002/bip.1972.360110102. [DOI] [PubMed] [Google Scholar]
- Olson W. K. The spatial configuration of ordered polynucleotide chains. I. Helix formation and base stacking. Biopolymers. 1976 May;15(5):859–878. doi: 10.1002/bip.1976.360150505. [DOI] [PubMed] [Google Scholar]
- Olson W. K. The spatial configuration of ordered polynucleotide chains. II. The poly(rA) helix. Nucleic Acids Res. 1975 Nov;2(11):2055–2068. doi: 10.1093/nar/2.11.2055. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stannard B. S., Felsenfeld G. The conformation of polyriboadenylic acid at low temperature and neutral pH. A single-stranded rodlike structure. Biopolymers. 1975 Feb;14(2):299–307. doi: 10.1002/bip.1975.360140205. [DOI] [PubMed] [Google Scholar]
- Tewari R., Nanda R. K., Govil G. Spatial configuration of single-stranded polynucleotides. Calculations of average dimensions and Nmr coupling constants. Biopolymers. 1974;13(10):2015–2035. doi: 10.1002/bip.1974.360131007. [DOI] [PubMed] [Google Scholar]


