Abstract
The accessibility of DNA in chromatin is an essential factor in regulating its activities. We studied the accessibility of the DNA in a ∼170 kb circular minichromosome to DNA-cleaving reagents using pulsed-field gel electrophoresis and fibre-fluorescence in situ hybridization on combed DNA molecules. Only one of several potential sites in the minichromosome DNA was accessible to restriction enzymes in permeabilized cells, and in growing cells only a single site at an essentially random position was cut by poisoned topoisomerase II, neocarzinostatin and γ-radiation, which have multiple potential cleavage sites; further sites were then inaccessible in the linearized minichromosomes. Sequential exposure to combinations of these reagents also resulted in cleavage at only a single site. Minichromosome DNA containing single-strand breaks created by a nicking endonuclease to relax any unconstrained superhelicity was also cut at only a single position by a restriction enzyme. Further sites became accessible after ≥95% of histones H2A, H2B and H1, and most non-histone proteins were extracted. These observations suggest that a global rearrangement of the three-dimensional packing and interactions of nucleosomes occurs when a circular minichromosome is linearized and results in its DNA becoming inaccessible to probes.
INTRODUCTION
Understanding how the accessibility of DNA in chromatin is regulated is central to models of DNA transcription and replication and their control (1), and factors that influence accessibility in vivo include nucleosomal structure (2–7), compaction of chromatin (8) and unrestrained superhelicity in DNA (9–11). As a model system to explore these and other features of chromatin we are studying a minichromosome that contains a ∼172 kb DNA with a canonical nucleosomal structure, the Epstein–Barr virus (EBV) episome, which is formed by circularization of the DNA of EBV through its cohesive termini and maintained at ∼50 copies per nucleus in Raji (human lymphoblastoid) cells (12–15). The only protein other than nucleosomes which is known to be associated with DNA in this minichromosome is EBNA-1, which is bound to a ∼1.7 kb region at oriP (14) (Figure 1A). The length of the minichromosome is in the range of those of the topologically closed loops in genomic chromatin (9–11,16,17).
Figure 1.
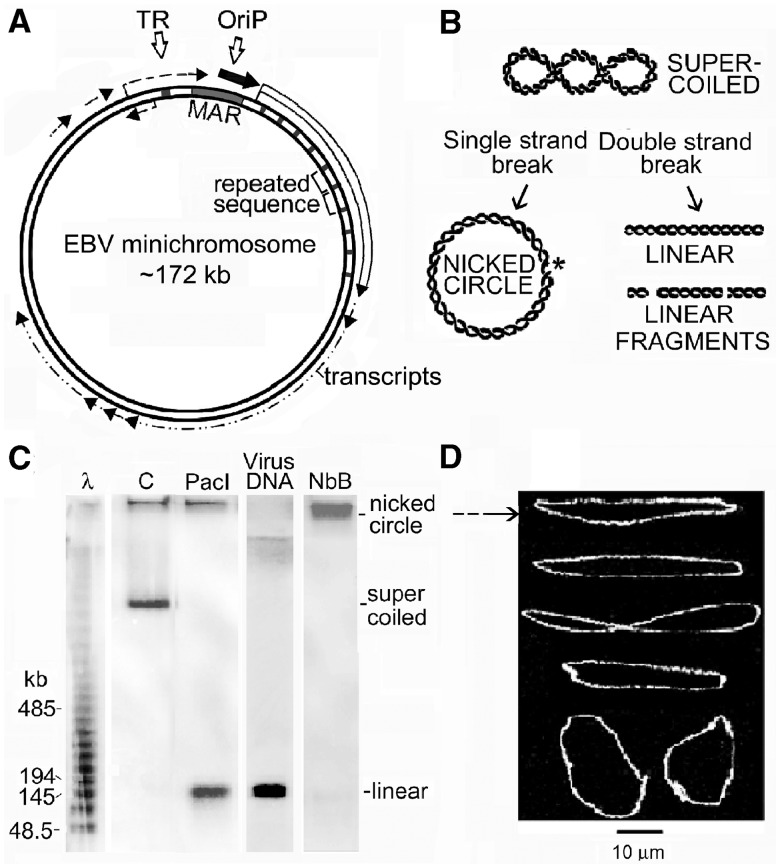
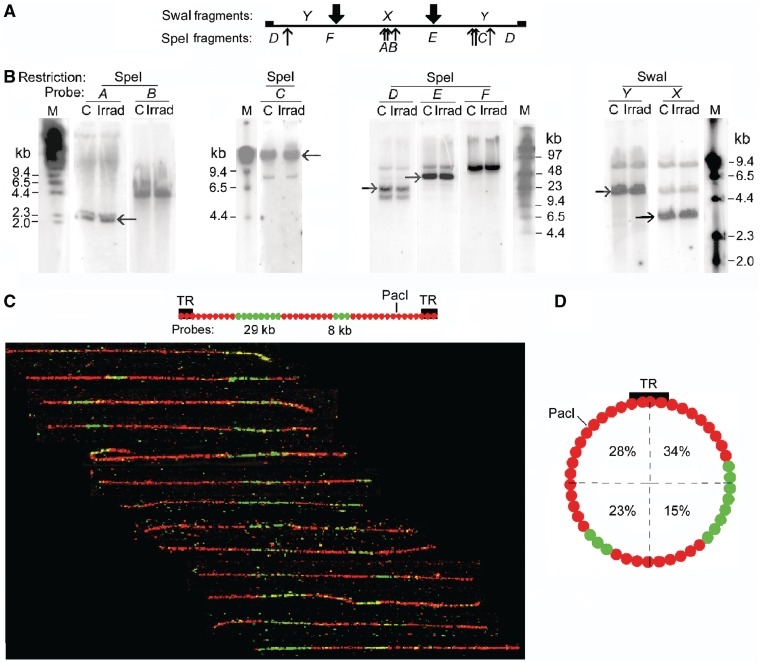
(A) The circular DNA of the EBV minichromosome. TR is the terminal repeated sequence through which linear EBV DNA is circularized to form the minichromosome, oriP is the preferred but not unique origin of replication, MAR is the nuclear matrix attachment region which coincides with a micrococcal nuclease-hypersensitive region. Approximate positions of regions transcribed in Raji cells are shown by arrows; black, highest level; white, intermediate; dashed, lowest (22). (B) Forms of minichromosome DNA considered in this work and (C) their migration in a PFGE gel of total cell DNA shown by hybridizing with an EBV DNA probe. In all gel images, the top coincides with the sample wells and each panel shows lanes from the same gel. Cells encapsulated in agarose beads and deproteinized were incubated with: C, no addition (2 h); PacI (100 U/ml, 2 h) which cuts minichromosome DNA at a single site; NbB, nicking endonuclease Nb.BbvCI (100 U/ml, 1 h). Lane virus DNA, linear EBV DNA; lane λ, oligomers of λ DNA (48.5 kb). (D) Representative DNA molecules extracted from the region close to the origin after incubating cells with Nb.BbvCI, stained with YOYO-1 and spread by molecular combing; these are believed to be nicked circular minichromosome DNA (see text).
The objective of this study was to understand our unexpected observation that the minichromosome DNA was converted quantitatively to full-length linear when permeabilized cells were incubated with a restriction enzyme with multiple cutting sites, showing that in the circular minichromosome the accessibility of DNA to the enzyme was limited to only a single site (the term site is used here for simplicity and does not imply a precise nucleotide position). Here, we report these experiments and show that other agents which create double-strand DNA breaks also cleave the minichromosome DNA at only one site. These findings suggest the existence of a previously unidentified influence of the topology of chromatin on the accessibility of DNA.
MATERIALS AND METHODS
Cells and exposure to DNA-cleaving reagents
Raji cells were maintained in Roswell Park Memorial Institute (RPMI)-1640 medium with 2 mM L-glutamine and 10% heat-inactivated foetal bovine serum and harvested during exponential growth. When indicated, the medium was supplemented for 18–24 h with 5-bromo-2′-deoxyuridine (BrdU) (30 μM) (Sigma-Aldrich) or a [35S] protein labelling mix (4 MBq/litre) (NEN). Cells were incubated with a nuclease (New England Biolabs) after encapsulation in beads of 1% low melting point (LMP) agarose (18), permeabilization in 10 mM Tris–HCl, 140 mM NaCl, 1 mM MgCl2, pH 7.6 containing 0.5% v/v Triton X-100 (Sigma-Aldrich) and washing 3× for 30 min in the same buffer without Triton X-100. Beads were equilibrated for 30 min on ice in the appropriate buffer, re-suspended in fresh buffer and incubated with an enzyme at 25°C for SwaI or at 37°C for others. Etoposide and neocarzinostatin (NCS; Sigma-Aldrich) dissolved in dimethyl sulfoxide (DMSO) were added to cultures as detailed in Figure legends; control cultures were incubated with DMSO alone at the highest concentration used. For pulsed-field gel electrophoresis (PFGE), cells washed in phosphate buffered saline (PBS) were embedded in 1% LMP agarose blocks (∼0.5–2 × 106 cells/block). For irradiation, the blocks were permeabilized and washed as described above, immersed in growth medium in closed 2-ml microtubes on ice and irradiated with 60Co γ photons at 4.3 Gy/min in a Teratron (Atomic Energy of Canada) calibrated as described in (19).
PFGE, probes and hybridization
Agarose beads or blocks were deproteinized for PFGE in 1 ml of 0.2 M ethylenediaminetetraacetic acid, 1% sodium dodecyl sulphate (SDS) and 1 mg/ml proteinase K (Invitrogen) for 48 h at ∼18°C with rocking and stored in 10 mM Tris-Cl, pH 7.5, 1 mM ethylenediaminetetraacetic acid (TE) at 4°C. PFGE was in 1% agarose in 0.5× 89 mM Tris base, 89 mM boric acid, 2 mM ethylenediaminetetraacetic acid at 14°C and 190 v for 20 h with pulse time increasing linearly from 50 to 90 s, or as described in the Figure legends. For hybridization, gels placed on 3 MM paper and covered with Saran Wrap were vacuum-dried at 60°C for 1 h and incubated successively in 0.5 M NaOH, 1.5 M NaCl (30 min), H2O, 0.5 M Tris, pH 8.0, 1.5 M NaCl (30 min) and 6× SSC (150 mM NaCl, 15 mM Na3citrate) (20 min), all at room temperature. Minichromosome DNA was detected by hybridizing gels with EBV DNA (GenBank AJ507799.2) isolated by PFGE from virus from B95-8 cells (20). Restriction fragments of this DNA cut with SpeI or SwaI (100 U/ml) were separated by PFGE in 1% LMP agarose at 190 V/cm for 7 or 20 h and switch time ramped linearly from 0.4 to 6 or 0.3 to 3 s, respectively, excised from gels and purified on Ultrafree columns (Millipore). Gel lanes containing length markers were hybridized with an appropriate probe. Probes were labelled with [α-32P]dCTP (111 TBq/mmol) using Megaprime kits (Amersham). Gels were pre-hybridized (30 min) and hybridized (18 h) in 6× SSC, 5× Denhardt’s solution, 0.5% SDS, 0.5 μg/ml human Cot-1 DNA (Invitrogen) at 68°C, washed 3× 30 min in 0.1× SSC, 0.5% SDS at 68°C, sealed in Saran Wrap and exposed to PhosphorImager screens. Signals imaged using ImageQuant (Molecular Dynamics) are shown (10−7 × measured arbitrary units) after subtracting the mean background in identical areas below and above the region of interest and are from at least triplicate independent experiments; P-values were calculated by the unpaired t-test.
Molecular combing and hybridization of minichromosome DNA
DNA of cells grown with BrdU was separated by PFGE and the region containing linear minichromosome DNA was excised, incubated with YOYO-1 (5 μM; Molecular Probes) for 30 min, washed in TE, incubated in β-agarase buffer for 30 min on ice, melted in 50 mM 2-(N-morpholino)ethanesulfonic acid, pH 5.7 at 65°C for 10 min and solubilized by β-agarose (New England Biolabs) at 42°C for 4 h. Four microlitres of DNA in the same buffer (2 μg/ml) was placed on a 3-aminopropyltriethoxysilane-coated microscope slide (Sigma-Aldrich) and covered with a standard cover glass, which was pulled horizontally across the slide at a constant speed of ∼300 μm/s after 2 min. Slides with well-spread DNA molecules as seen by fluorescence microscopy (Nikon E800,100× objective) were dried (5 min, room temperature and overnight, 60°C), incubated in 0.6× SSC, 70% formamide (3 min, 95°C) and in cold 70%, 85% and 95% ethanol (2 min each). Fluorescence in situ hybridization (FISH) probes were an 8.1-kb BamHI–SalI fragment of cosmid cM301-99 and a 29-kb HindIII fragment of cosmid cMB-14 (21) (gifts from G. Bornkamm) excised from gels, purified on a Microcon YM-100 (Qiagen) and labelled with biotin-11-dUTP (Fermentas) by nick translation. Hybridization was at 37°C in a humidified chamber for up to 48 h. Probes were detected with fluorescein isothiocyanate-goat anti-biotin (Sigma-Aldrich) (1:50, 20 min) followed by Alexa 488-rabbit anti-goat antibody (Invitrogen; 1:50, 20 min) and DNA by subsequent incubation with rat anti-BrdU (Abcam) (1:30, 20 min) followed by Alexa 594-goat anti-rat antibody (Invitrogen; 1:50, 20 min). Antibody dilutions and washing were in PBS and 0.05% Tween-20, and slides were mounted in SlowFade Gold (Invitrogen). Minichromosome DNA molecules which showed hybridization signals from both probes were imaged by confocal microscopy (Bio-Rad MRC1024). Lengths of molecules were calculated using a factor of 2.2 kb DNA/µm after minor adjustment of images to normalize the distance between the two probes (22,23).
Extraction of chromatin proteins
Cells in agarose beads were permeabilized as described above; washed 2× in 10 mM Tris–HCl, 140 mM NaCl, 1 mM MgCl2, pH 7.6 and protease inhibitor cocktail (Sigma-Aldrich; 1/200) and incubated in this buffer supplemented with NaCl at 0.14, 0.35, 0.6, 1.2 or 2 M for 1 h on ice with gentle agitation; the buffer was replaced and incubation continued for 18 h at 4°C. Beads were washed in cold PBS, boiled for 5 min in 2× SDS–polyacrylamide gel electrophoresis (PAGE) buffer, and proteins were separated on denaturing 12% gels with size markers (Bio-Rad) and calf thymus histone markers (gift of W. T. Garrard) and stained with Coomassie blue. Histones were quantitated by blotting replicate gels onto nitrocellulose, detection with rabbit primary antibodies against histone H1 (GeneTex, 1:1500), H2A (Upstate Biotechnology, 1:3000), H2B (Active Motif, 1:3000), H3 (Abcam, 1:3000) or H4 (Abcam, 1:3000) followed by horseradish peroxidase-conjugated secondary antibody, and signals were developed by enhanced chemiluminescence (Perkin-Elmer) and quantitated on a Phosphorimager.
RESULTS
Minichromosome DNA and forms produced by single- and double-strand breaks
The EBV minichromosome (Figure 1A) is formed by circularization of the linear DNA of EBV through its terminal repeated sequences (TR); it contains a major but not unique origin of replication (oriP) (23), a region more sensitive to micrococcal nuclease ascribed to a non-canonical arrangement (24) or absence (13) of nucleosomes and which contains a nuclear matrix attachment region (25,26); only 10 of ∼100 genes are transcribed in the Raji cells used here (23,27). Genome-wide mapping of histone modifications has not been reported, but the markers H3K9me2, H3K9me3, H3K27me3 and H4K20me3 are present throughout the minichromosome DNA (28). Commonly present in the nuclei of human B lymphocytes (14), this minichromosome has proved a particularly valuable experimental system to study origins of DNA replication (22,23).
The minichromosome DNA was detected by hybridizing PFGE gels of total cell DNA with an EBV DNA probe (Figure 1C). For PFGE, DNA was deproteinized at room temperature (∼18°C) because extra strand breaks can be formed at temperatures >20°C (29); >99% of the 10% trichloroacetic acid-precipitable radioactivity was extracted from cells containing 35S-labelled proteins (unpublished results). Dried gels were used for hybridization because transfer of large DNA fragments by blotting is not quantitative (30,31). In the gel shown in Figure 1C, some hybridizing material remained in the sample wells, but in most experiments this was less evident or not seen and is believed to reflect trapping of minichromosome DNA by the abundant high-molecular-weight genomic DNA in this region due to slight overloading of gels (32). The linear form of minichromosome DNA, produced by cutting at the single PacI site, migrated with the same mobility as EBV DNA (Figure 1C, lanes PacI and viral DNA). Circular minichromosome DNA containing single-strand breaks migrated close to the sample well (Figure 1C, lane NbB) like other large nicked circular DNAs (33,34), and molecules with the appropriate length (181 ± 11 kb, SEM from 30 molecules) and conformation were seen when DNA from this region was spread by molecular combing after permeabilized cells were incubated with the nicking endonuclease Nb.BbvCI (Figure 1D); these molecules were not seen in gels of DNA from control cells and supercoiled DNA does not bind to slides in these conditions (unpublished results; see also 35), and they did not have the theta conformation characteristic of replicating minichromosome DNA (15).
Minichromosome DNA is resistant to further breakage after one double-strand cut by a restriction enzyme
To cut minichromosome DNA by restriction enzymes, cells were encapsulated in agarose beads and permeabilized (18); in these conditions the synthesis of RNA (18) and DNA (Kumala et al., manuscript submitted) are maintained at essentially the same rates as in growing cultures and the minichromosome DNA conserves its native supercoiled conformation (Figure 2B and C, lanes no enzyme). Incubation with Spe1 or SwaI, which have, respectively, seven or two cutting sites and are not affected by methylated sequences which occur in some regions of minichromosome DNA (36), produced only full-length linear minichromosome DNA (Figure 2B and C), whereas in deproteinized cells SwaI cut minichromosome DNA at both its sites to produce the expected fragments (Figure 2C, lane DNA).
Figure 2.
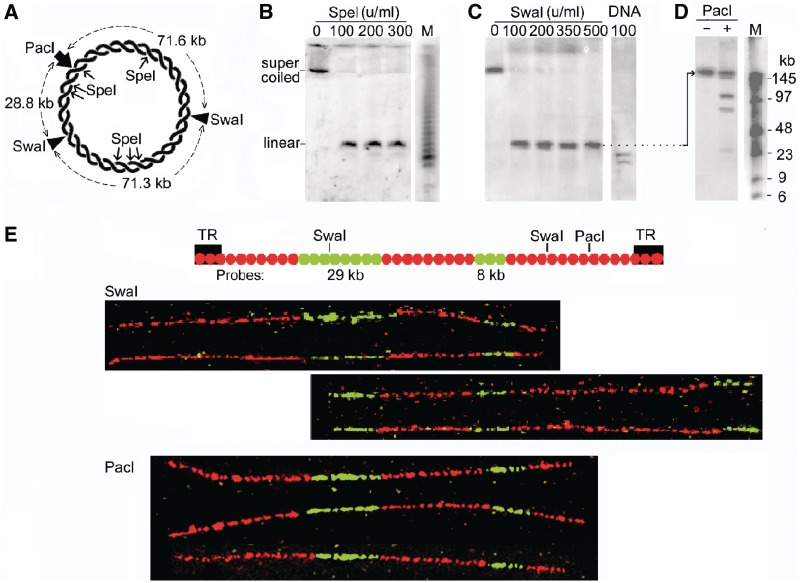
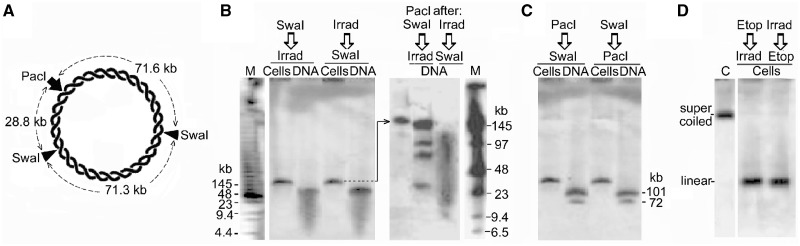
Minichromosome DNA is cut at only one site by SpeI or SwaI, which have seven and two cutting sites, respectively. (A) Circular minichromosome DNA showing SpeI, SwaI and PacI sites; SpeI fragment lengths are not shown for clarity. (B) Minichromosome DNA from permeabilized cells incubated with SpeI or (C) with SwaI, and deproteinized. Lane DNA, SwaI fragments produced in deproteinized cells. (D) The SwaI cleavage site mapped by gel hybridization. DNA linearized by SwaI (200 U/ml, 2 h) was isolated from a PFGE gel, cut by PacI (100 U/ml, 18 h), and the products were separated by PFGE. Lanes M, oligomers with HindIII fragments of λ DNA. The four fragments produced (∼140, 100, 72 and 29 kb) represent a mixture of the pairs produced after SwaI had cut at only one of its two sites in different minichromosomes. (E) The SwaI cleavage site mapped by fibre-FISH. The positions of the SwaI sites and hybridization probes (green) are shown above; the biotin-labelled probes were detected with anti-biotin antibodies (green) and BrdU-labelled DNA with anti-BrdU antibodies (red). Images show representative linear molecules from the two classes observed, which had been cut by SwaI (shown by the extremities of the molecule) at either the left (upper panel) or the right (lower panel) site on the map. The probe positions were aligned approximately considering the variable stretching during combing (22,23). Below, linear molecules cleaved at the single PacI site for comparison.
To distinguish if all minichromosome DNA molecules were cut at one particular site or at any one of their potential sites, we mapped the sites cut by SwaI whose fragments were fewer and, therefore, more easily identified. Minichromosome DNA linearized by SwaI was extracted from a gel and digested with PacI, which has a single cutting site; if SwaI had cut all minichromosomes at the same site, PacI would produce a pair of fragments of either 71.6 and 100.1 kb or 28.8 and 142.9 kb (Figure 2A). Instead, four fragments of ∼140, 100, 72 and 29 kb were produced (Figure 2D) representing a mixture of these pairs, showing that in different minichromosomes SwaI had cut at either of its two sites. Fibre-FISH mapping of SwaI-linearized minichromosome DNA confirmed this conclusion (Figure 2E). Linear DNA from BrdU-labelled cells was extracted from gels, combed and hybridized with two biotin-labelled probes to identify minichromosome DNA molecules and orient their images, and DNA was labelled with anti-BrdU antibodies. The length of these molecules was measured using the factor of 2.2 kb DNA/µm determined experimentally for combed linear DNA of this minichromosome (22). The molecules detected by the probes were 167 ± 10 kb in length (SEM from 100 molecules in five independent experiments), corresponding closely to full-length minichromosome DNA (∼172 kb). They fell into two classes; 28 of a total of 50 molecules imaged had been cut by SwaI (represented by their extremity) at the left site on the map in Figure 2E and 22 at the right site. As a control, PacI-linearized minichromosome DNA showed cleavage at the expected single position (Figure 2E, lower panel).
Other DNA cleavage agents create only one double-strand break in minichromosome DNA
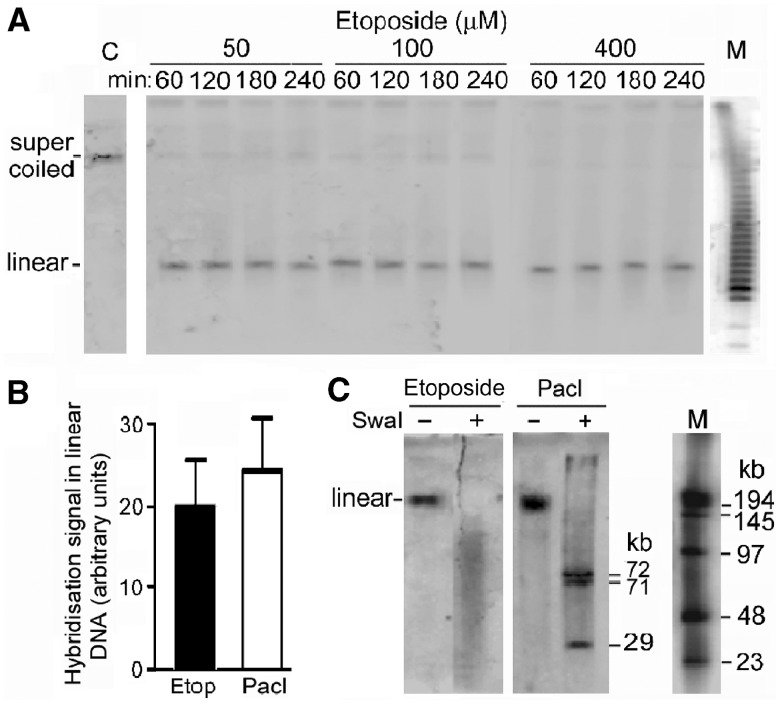
To explore if the inaccessibility of minichromosome DNA to restriction enzymes at all but one site was an example of a more general phenomenon, growing cells were incubated with the DNA-cleaving agent etoposide (37–40) or NCS (41–43), or were exposed to γ-radiation (44). Etoposide, termed a topoisomerase II poison, arrests the re-ligation step of topoisomerase II covalently integrated into DNA, resulting in double-strand breaks after deproteinization (37–40). Linear minichromosome DNA was formed when growing cells were incubated with etoposide (Figure 3A), and after the maximum extent of cleavage its amount was not significantly different from that when minichromosome DNA was linearized at the single PacI site (P = 0.42 from three replicate experiments; Figure 3B). The site of the single cleavage varied in different minichromosomes, since cutting the linearized DNA with SwaI produced a smear of fragments whose length extended down to ∼10 kb (Figure 3C, lane etoposide + SwaI). As a control, PacI-linearized minichromosome DNA showed the expected SwaI fragments (Figure 3C, lane PacI + SwaI).
Figure 3.
Minichromosome DNA is cut at a single but variable site in growing cells incubated with the topoisomerase II poison etoposide. (A) PFGE gel of DNA from cells incubated with etoposide. (B) Quantitation of linear minichromosome DNA after etoposide cleavage (Etop) (400 μM, 240 min), compared with that after PacI cleavage (100 U/ml, 1 h) in permeabilized cells; error bars show SEM from three independent experiments. (C) Sites of cleavage by etoposide (100 μM, 60 min) compared with those of PacI (100 U/ml, 3 h). Linearized minichromosome DNA from a PFGE gel was digested with SwaI (100 U/ml, 18 h), and the products were separated by PFGE (10 h, pulse time 10 to 70 s). Lanes M, oligomers with HindIII fragments of λ DNA. The smear of SwaI fragments from cells incubated with etoposide shows that the DNA cleavage position varied in different minichromosomes, while DNA cut at the single PacI site showed the expected SwaI fragments.
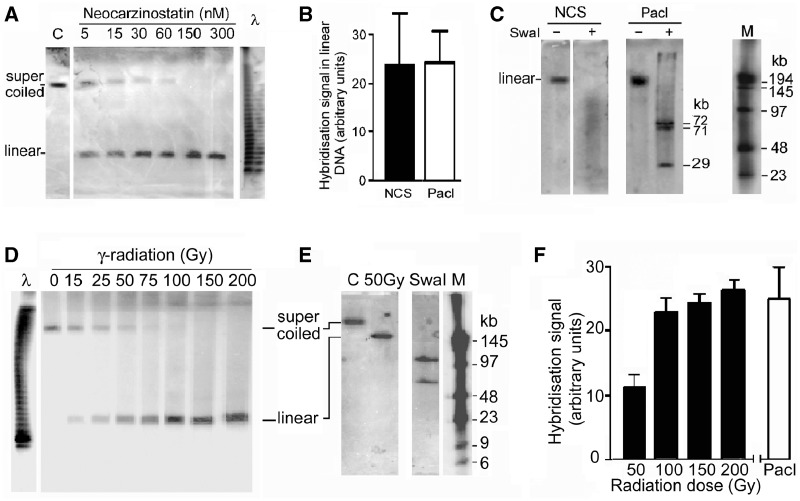
NCS, an enediyne chromophore bound to a small protein, creates DNA strand breaks in vivo and in vitro (41–43) and has numerous preferred cleavage sequences (41). Incubation of cells with NCS resulted in conversion of the supercoiled minichromosome DNA to the linear form (Figure 4A) whose amount was not significantly different from that after linearization at the single PacI site (P = 0.94 from three replicate experiments; Figure 4B). The linear molecules were 170 ± 10 kb in length (SEM from five replicate experiments) and the cleavage position varied in different minichromosomes, since a smear of fragments of length down to ∼10 kb was produced when the linear DNA was cut with SwaI (Figure 4C, lane NCS+Swa1). NCS, therefore, cleaved minichromosome DNA at a single but variable site.
Figure 4.
Cleavage of minichromosome DNA by neocarzinostatin (NCS) or γ-irradiation produces full-length linear DNA. (A) Minichromosome DNA from growing cells incubated with NCS for 1 h. (B) Quantitation of the linear DNA produced by NCS (300 nM) compared with that produced by PacI cleavage (100 U/ml, 1 h) in permeabilized cells; error bars show SEM from three independent experiments. (C) Sites of cleavage by NCS (300 nM); linearized minichromosome DNA isolated from a PFGE gel was incubated without or with SwaI (100 U/ml, 18 h), and the products were separated by PFGE (10 h, pulse time 10 to 70 s). The smear of SwaI fragments from cells incubated with NCS shows that the position of cleavage varied in different minichromosomes, whereas PacI produced the expected fragments. Lane M, oligomers with HindIII fragments of λ DNA. (D) Minichromosome DNA from γ-irradiated cells separated in standard PFGE conditions or (E) to detect fragments shorter than full-length linear DNA. Lane C, un-irradiated cells; lane SwaI, SwaI fragments (100 U/ml, 18 h) from control cells demonstrating the separation obtained in these conditions; lane M, oligomers with HindIII fragments of λ DNA. (F) Quantitation of linear minichromosome DNA produced by γ-irradiation compared with that produced by PacI (100 U/ml, 3 h); error bars show SEM from three independent experiments.
Minichromosome DNA was also linearized in an essentially quantitative manner in cells exposed to γ-radiation (Figure 4D), which causes DNA breaks mainly due to OH radicals and other reactive species produced from H2O (44,45). The amount of linear DNA formed was not significantly different from that after cleavage at the single PacI site (P = 0.45 from three independent experiments; Figure 4F), and no shorter fragments were detected using PFGE conditions which separated DNA of length down to ∼5 kb (Figure 4E). The break was mapped by restricting the linearized DNA, separating the fragments by PFGE and hybridizing with the same restriction fragments of EBV DNA; if all minichromosomes were broken at a specific site the fragment containing this site would be truncated, allowing the site’s position to be deduced (39). Overlapping probe sets of SwaI and SpeI fragments (Figure 5A) were used to ensure detection of truncation sites close to fragment ends. Some probes detected weaker bands in addition to those predicted from the minichromosome DNA sequence (46) (Figure 5B), which may reflect polymorphisms (47) and inhibition of SpeI cleavage due to rare N6-methyladenines (48). For all the probes, the pattern of hybridizing DNA fragments was identical in irradiated and control cells and no truncated fragments were detected (Figure 5B), showing that the single cleavage of minichromosome DNA in irradiated cells was at an essentially random position. This was confirmed by fibre-FISH; the extremities of the linearized DNA were in variable positions with respect to the probes (Figure 5C), and the breakage site was localized within any of four quadrants of the circular DNA (Figure 5D).
Figure 5.
Sites of breakage of minichromosome DNA in γ-irradiated cells. (A) SpeI and SwaI EBV DNA fragments used as probes for PFGE gels. (B) Hybridization of these probes to gels of DNA from control cells (C) or cells irradiated with 100 Gy (Irrad) restricted by the same enzyme; the PFGE conditions differed according to the length of fragments to be detected. Arrows show fragments predicted from the minichromosome DNA sequence; the origin of the weakly-hybridizing fragments is discussed in the text. (C) Fibre-FISH; the hybridization probes and procedure were as described in Figure 2E. Images show representative linear molecules from irradiated cells (100 Gy); probes are green and DNA is red, and the extremities of the molecule represent the site of breakage. (D) Distribution of the breakage site expressed as the % of 55 circular DNA molecules in which the break occurred in one of four quadrants.
Linearization of minichromosome DNA confers resistance to further DNA cleavage by other reagents
Three possible mechanisms were considered to understand why only one site, which was not the same in all minichromosomes, was accessible to the reagents tested earlier. The conformation of the minichromosome could be altered following rapid phosphorylation of histone H2AX (49) or other histone modifications triggered by the first strand break, but this appeared to be excluded since adenosine triphosphate (ATP) and other factors had been extracted by permeabilizing the cells. Alternatively, all potential sites, except one, could be masked by nucleosomes or other proteins; this is perhaps plausible for restriction enzymes whose sites can be masked when they are on the nucleosome surface (3–7) but is improbable for NCS and γ-radiation which have many potential cleavage sites, predominantly in internucleosomal linker DNA (41–43,50–52). In this model, minichromosome DNA linearized by one agent would be still accessible at other positions to reagents with different cleavage sites. Finally, upon linearization of the minichromosome a global rearrangement of its chromatin structure could occur, which made DNA inaccessible to all the reagents; in this case, after linearization by one reagent, the DNA would not be cleaved further by any of them.
To distinguish the two latter scenarios, cells were exposed sequentially to two different conditions which created a double-strand break. For example, they were incubated first with SwaI, which cut minichromosome DNA at only one of its two sites, and then irradiated, which produced a single break at an essentially random site. The circular minichromosome DNA was cut at only one site to produce linear DNA (Figure 6B, left panel, lane SwaI ⇒ Irrad, cells). Restriction of this linear DNA by PacI produced only the fragments characteristic of cutting by SwaI at either of its two sites, showing that no further breaks were made by radiation after the DNA had been cut by SwaI (Figure 6B, right panel, lane PacI after SwaI ⇒ Irrad). Similarly, only linear DNA was produced when this sequence was reversed and cells were first irradiated and then incubated with SwaI, showing that the DNA was not cut further by SwaI after it had been linearized by radiation (Figure 6B, left panel, lane Irrad ⇒ SwaI, cells). In this case, PacI restriction of the linearized DNA produced a smear of shorter fragments (Figure 6B, right panel, lane PacI after Irrad ⇒ SwaI), confirming that the single break was located randomly and therefore must have been produced by the initial irradiation. Similarly, cells were first incubated with etoposide and then irradiated, both of which alone create a single break at a random site (Figures 3, 5) or this sequence was inversed. Only linear minichromosome DNA was produced in both cases, showing that the circular DNA had been cleaved at only a single site (Figure 6D). Cleavage with PacI followed by SwaI or the inverse sequence produced only linear DNA (Figure 6C, lanes cells), whereas three fragments of 71.6, 71.3 and 28.8 kb would have been produced if they could access all their sites (Figure 6A).
Figure 6.
Minichromosome DNA is cut at only a single site when cells are exposed sequentially to two different cleavage reagents. (A) Cutting sites for SwaI and PacI in minichromosome DNA. (B) Left panel, permeabilized cells (lanes cells) or deproteinized cells (lanes DNA) were incubated with SwaI (100 U/ml, 2 h) and then irradiated (200 Gy) (Irrad) or this sequence was reversed. Right panel, the linearized minichromosome DNA in (B) was isolated and cut with PacI; after the sequence SwaI ⇒ irradiation only the predicted PacI fragments were produced, showing that irradiation had produced no further breaks after the DNA had been linearized by SwaI. In contrast, a smear of shorter fragments was seen after the sequence irradiation ⇒ SwaI, confirming that the initial break caused by radiation had occurred at a variable site in different minichromosomes. Lane M, oligomers with HindIII fragments of λ DNA. (C) Identical samples were incubated with PacI followed by SwaI (both 100 U/ml, 2 h), or this sequence was reversed. In deproteinized DNA, either sequence resulted in fragments cut by both enzymes (the ∼72 kb band contains two fragments of similar length, see panel A). (D) Identical samples were incubated alone (C) or with etoposide (Etop) (100 μM) and then irradiated (Irrad) (200 Gy) or this sequence was reversed.
Single-strand breaks in minichromosome DNA do not affect the single-site mode of cutting by a restriction enzyme
In the preceding experiments, only the conversion of circular to linear DNA was assayed, but the event that caused minichromosome DNA to become inaccessible could be a single-strand break since NCS and γ-radiation (41–43,50–52), and also some restriction enzymes (53) and possibly topoisomerase II poisons (54), can create single- as well as double-strand breaks. To create only single-strand breaks in minichromosome DNA, permeabilized cells were incubated with the nicking endonuclease Nb.BsmI or Nb.BbvCI. About 80% of the circular DNA was converted to the nicked form by Nb.BsmI in optimum conditions (Figure 7); some linear DNA was also produced, probably due to single-strand cuts at close sites on opposite strands. The nicked circular minichromosomes were cut by SwaI in exactly the same manner as unnicked minichromosomes, producing only full-length linear DNA (Figure 7). Cutting by SwaI of minichromosome DNA nicked by Nb.BbvCI also produced only full-length linear DNA (unpublished results). We conclude that even if the reagents tested earlier created single-strand breaks in minichromosome DNA, these breaks would not be sufficient to allow SwaI to cut at all of its potential sites.
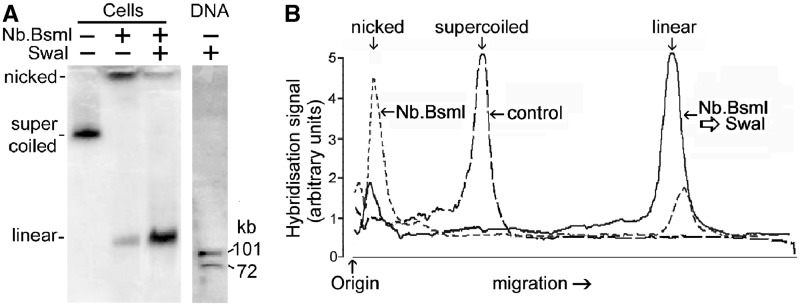
Figure 7.
Circular minichromosome DNA containing single-strand breaks is cut by SwaI at only one of its two potential sites. (A) Minichromosome DNA was incubated with endonuclease Nb.BsmI (100 U/ml, 2 h) in permeabilized cells to produce predominantly nicked circular DNA (cells, centre lane). Subsequent cutting by SwaI (100 U/ml, 2 h) produced only full-length linear DNA (cells, right lane) showing that only one of the two SwaI sites was cut in nicked circular DNA. Lane DNA shows SwaI fragments of deproteinized DNA; the ∼72 kb band contains two fragments of similar length (Figure 2A). (B) Scans of lanes in panel A.
Proteins associated with inaccessibility of DNA in linearized minichromosomes
Quite different patterns of DNA cleavage were seen when minichromosomes had been deproteinized. SwaI followed by irradiation or the reverse sequence produced a smear of fragments of length down to ∼6 kb (Figure 6B, left panel, lanes DNA), and SwaI was able to cut at both of its potential sites as shown by subsequent restriction by PacI (Figure 6C, lanes DNA) (the ∼72 kb band contains two fragments of similar length, Figure 2A).
To identify the class of proteins which modulated the accessibility of minichromosome DNA, permeabilized cells were extracted with NaCl at different concentrations (Figure 8A). After the majority of non-histone proteins and 87% of histone H1, but no core histones, had been extracted at NaCl concentrations ≤0.6 M, SwaI or irradiation still cut only a single site in minichromosome DNA to produce full-length linear DNA (Figure 8C). In contrast, after ≥90% of H2A and H2B but ≤5% of histones H3 and H4 had been extracted by 1.2 M NaCl, both SwaI sites could be cut (Figure 8C) and irradiation created multiple breaks producing fragments of length down to ∼10 kb (Figure 8D, E). Thus, the access of cleaving agents to linearized minichromosome DNA was no longer impaired after H2A and H2B had been detached from nucleosomes.
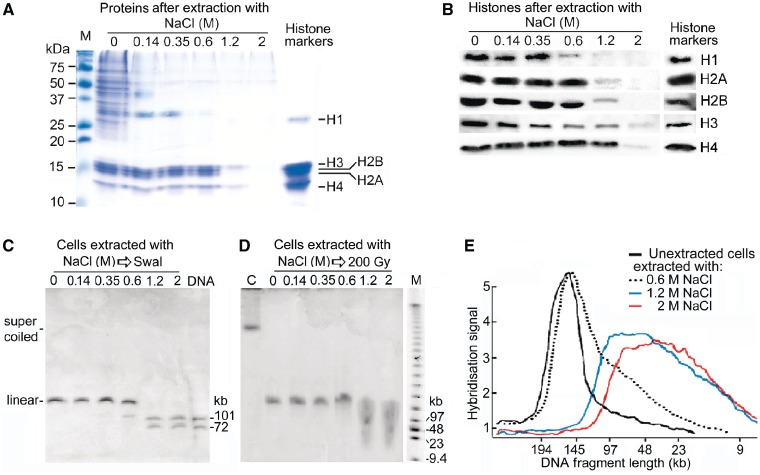
Figure 8.
Cutting of minichromosome DNA by SwaI or γ-radiation after extraction of proteins. (A) Proteins remaining in permeabilized cells after extraction with NaCl, shown by denaturing SDS–PAGE and Coomassie blue staining; histone markers are calf thymus histones. (B) Quantitation of histones in extracted cells; proteins transferred to membranes from a gel like that in panel A were probed with antibodies specific for each histone and the signals were quantitated. (C) Cutting of minichromosome DNA by SwaI (100 U/ml, 2 h) in extracted cells; lane DNA shows cutting in deproteinized cells. Only one of the two SwaI sites was accessible in cells extracted with ≤0.35 M NaCl producing linear DNA, but both sites had become accessible after extraction with ≥1.2 M NaCl as shown by the production of the three predicted SwaI fragments (the ∼72 kb band contains two fragments of similar length). (D) Minichromosome DNA from cells irradiated (200 Gy) after extracting proteins; lane C, un-irradiated cells. Only a single site was cut in cells extracted with ≤0.35 M NaCl producing linear DNA, but many further sites were cut after extraction with ≥1.2 M NaCl as shown by the smear of shorter DNA fragments. Lane M, oligomers with HindIII fragments of λ DNA. (E) Minichromosome DNA fragment lengths in irradiated cells without or after extraction with NaCl, calculated from scans of a gel like that in panel D and interpolation from the positions of the λ DNA markers.
DISCUSSION
The essential conclusion of these experiments is that several enzymatic and chemical agents which produce double-strand DNA breaks cut the DNA of the circular EBV minichromosome within cells at only one of their multiple potential sites, and once the DNA has been linearized other sites are inaccessible. Accessibility of only one site to a restriction enzyme could result from masking of other sites by nucleosomes, but it is implausible that access to NCS and for OH radicals generated by γ-radiation could be limited in this way in view of the multiplicity of their potential cleavage sites (41–43,50–52). We note that the radiation doses used here are 10-fold lower than those which create sufficient strand breaks to footprint DNA-bound proteins in vivo (55). The production of only one double-strand break in all the minichromosomes in γ-irradiated cells was particularly unexpected, since in genomic DNA breaks are assumed to follow a Poisson distribution (e.g., 56,57), which would result in conversion of only 38% (1/e) of the minichromosome DNA to full-length linear DNA, with the remainder as shorter fragments.
The position of the initial break produced by NCS and γ-radiation varied in different minichromosomes and therefore cannot be determined by a feature of the DNA sequence, for example the region with a non-canonical nucleosomal structure (24) or the transcribed regions (27) (Figure 1A). Molecules that are replicating during a 1 h incubation would not exceed ∼5% of the total number since the average duplication time for minichromosome DNA is ∼1 h (22) and therefore should not contribute significantly to the observed responses. It is improbable that the one-site mode of cleavage results from a conformational change of the minichromosome caused by rapid modification of histones in response to a first break, because ATP and other factors had been depleted by permeabilizing the cells and no formation of γ-H2AX was detected by immunofluorescence after irradiation (unpublished results). As discussed below, a further argument against a response of this type is that a similar one-site accessibility to probes been observed in vitro for other minichromosomes.
Whether the DNA of this minichromosome contains unconstrained superhelicity has not been explored, but this is plausible in view of the examples of other circular minichromosomes (58,59); nevertheless, the one-site mode of cleavage cannot be associated with unrestrained superhelicity since it was not abrogated when single-strand nicks were created in the circular form. The one-site mode is, however, determined by the structure and/or interactions of intact nucleosomes; the accessibility of multiple sites after histones H2A and H2B are removed is likely to reflect the reduced length of DNA bound to the remaining (H3-H4)2 tetramers (60). It is intuitively probable that nucleosome orientation and contacts will undergo considerable adjustment when a circular chain of polynucleosomes is converted to the linear form. Models of circular chromatin (61–64) are not yet as detailed as those of linear chromatin (63–68), but they predict multiple and dynamic conformations and emphasize the crucial influence of DNA helicity (60,69,70) and torsional forces (60–62,69–71) with the possible formation of plectonemes depending on the twist and writhe of the DNA (71). All the DNA-cleaving agents tested here, restriction enzymes (3–5,7), topoisomerase II poisons (38,68), NCS (41–43) and γ-radiation (52,72), cut DNA in chromatin preferentially in internucleosomal linkers and the simplest assumption is that the breaks observed here are in linker DNA, although fluctuations of nucleosome conformation may allow transient access to sites on their surface (3–7). Studies of circular chromatin suggest how linker DNA could be exposed to probes; it has been proposed that nucleosomes are distorted (63–65) and/or re-orientated (71,73) and histone–DNA contacts loosened with transfer of some linker onto their surface (74). In contrast, in compact linear chromatin, linker DNA is poorly accessible (75); most evidence indicates that it lies in the interior in vitro (75–78) and in vivo (79), and its accessibility could be limited further by self-association and interdigitation (80) and hairpin formation (81) due to internucleosomal attractive forces.
Properties of other circular minichromosomes that probably have a similar mechanistic origin were observed in earlier studies, but were not further explored or interpreted. DNA of circular bovine papilloma virus minichromosomes in isolated nuclei could be methylated by the DNA-(cytosine-5)-methyltransferase Hhal, but it was inaccessible after the minichromosome was linearized (59). DNA of circular SV40 minichromosomes was cut in vitro at a single but not unique site by multiple-site restriction enzymes (61). Similarly, DNA of a chimeric variant of this minichromosome (37) and EBV minichromosomes (52) was converted essentially quantitatively to the full-length linear form in cells exposed to a topoisomerase II poison. Understanding these switches in DNA accessibility depends on more detailed models of the nucleosomal conformation and topology of linear and circular chromatin, particularly in the crowded environment within the nucleus, and could be relevant to the accessibility of DNA in closed loops of genomic chromatin in vivo, which are topologically analogous to a circular minichromosome.
FUNDING
The Polish Ministry of Education and Science [Grant N N518 497639 to J.R-W.]; the Medical Faculty and Cancer Research Centre of Laval University. Funding for open access charge: Polish Ministry of Education and Science [Grant NN518 497639].
Conflict of interest statement. None declared.
ACKNOWLEDGEMENTS
We thank L. Frappier for Raji cells, P. de Campos-Lima for B95-8 cells, G. Bornkamm for cosmids, W. T. Garrard for calf thymus histones, J. St-Hilaire and N. Octave for help with irradiation, A. Prunell and W. K. Olson for helpful discussions and anonymous reviewers for constructive comments.
REFERENCES
- 1.Bell O, Tiwari VK, Thomä NH, Schübeler D. Determinants and dynamics of genome accessibility. Nat. Rev. Genet. 2011;12:554–564. doi: 10.1038/nrg3017. [DOI] [PubMed] [Google Scholar]
- 2.Morse RH. Nucleosomes inhibit both transcriptional initiation and elongation by RNA polymerase Ill in vitro. EMBO J. 1989;8:2343–2351. doi: 10.1002/j.1460-2075.1989.tb08362.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Anderson JD, Thåström A, Widom J. Spontaneous access of proteins to buried nucleosomal DNA target sites occurs via a mechanism that is distinct from nucleosome translocation. Mol. Cell. Biol. 2002;22:7147–7157. doi: 10.1128/MCB.22.20.7147-7157.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Poirier MG, Bussiek M, Langowski J, Widom J. Spontaneous access to DNA target sites in folded chromatin fibers. J. Mol. Biol. 2008;379:772–786. doi: 10.1016/j.jmb.2008.04.025. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Prinsen P, Schiessel H. Nucleosome stability and accessibility of its DNA to proteins. Biochimie. 2010;92:1722–1728. doi: 10.1016/j.biochi.2010.08.008. [DOI] [PubMed] [Google Scholar]
- 6.Böhm V, Hieb AR, Andrews AJ, Gansen A, Rocker A, Tóth K, Luger K, Langowski J. Nucleosome accessibility governed by the dimer/tetramer interface. Nucleic Acids Res. 2011;39:3093–3102. doi: 10.1093/nar/gkq1279. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Tims HS, Gurunathan K, Levitus M, Widom J. Dynamics of nucleosome invasion by DNA binding proteins. J. Mol. Biol. 2011;411:430–448. doi: 10.1016/j.jmb.2011.05.044. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.van Steensel B. Chromatin: constructing the big picture. EMBO J. 2011;30:1885–1895. doi: 10.1038/emboj.2011.135. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Luchnik AN, Hisamutdinov TA, Georgiev GP. Inhibition of transcription in eukaryotic cells by X-irradiation: relation to the loss of topological constraint in closed DNA loops. Nucleic Acids Res. 1988;16:5175–5190. doi: 10.1093/nar/16.11.5175. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Rodi CP, Sauerbier W. Structure of transcriptionally active chromatin: radiological evidence for requirement of torsionally constrained DNA. J. Cell Physiol. 1989;141:346–352. doi: 10.1002/jcp.1041410216. [DOI] [PubMed] [Google Scholar]
- 11.Villeponteau B, Martinson HG. Gamma rays and bleomycin nick DNA and reverse the DNase I sensitivity of ß-globin gene chromatin in vivo. Mol. Cell. Biol. 1987;7:1917–1924. doi: 10.1128/mcb.7.5.1917. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Shaw JE, Levinger LFC, Carter W. Nucleosomal structure of EB virus DNA in transformed cell lines. J. Virol. 1979;29:657–665. doi: 10.1128/jvi.29.2.657-665.1979. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Dyson PJ, Farrell PJ. Chromatin structure of Epstein-Barr virus. J. Gen. Virol. 1985;66:1931–1940. doi: 10.1099/0022-1317-66-9-1931. [DOI] [PubMed] [Google Scholar]
- 14.Sugden B, Leight ER. EBV's plasmid replicon: an enigma in cis and trans. Curr. Top. Microbiol. Immunol. 2001;258:3–11. doi: 10.1007/978-3-642-56515-1_1. [DOI] [PubMed] [Google Scholar]
- 15.Gussander E, Adam A. Electron microscopic evidence for replication of circular Epstein-Barr virus genomes in latently infected Raji cells. J. Virol. 1984;52:549–556. doi: 10.1128/jvi.52.2.549-556.1984. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Benyajati C, Worcel A. Isolation, characterization, and structure of the folded interphase genome of Drosophila melanogaster. Cell. 1976;9:393–407. doi: 10.1016/0092-8674(76)90084-2. [DOI] [PubMed] [Google Scholar]
- 17.Jackson DA, Dickinson P, Cook PR. The size of chromatin loops in HeLa cells. EMBO J. 1990;9:567–571. doi: 10.1002/j.1460-2075.1990.tb08144.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Jackson DA, Cook PR. A general method for preparing chromatin containing intact DNA. EMBO J. 1985;4:913–918. doi: 10.1002/j.1460-2075.1985.tb03718.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Almond PR, Biggs PJ, Coursey BM, Hanson WF, Huq MS, Nath R, Rogers DW. AAPM's TG-51 protocol for clinical reference dosimetry of high-energy photon and electron beams. Med. Phys. 1999;26:1847–1870. doi: 10.1118/1.598691. [DOI] [PubMed] [Google Scholar]
- 20.zur Hausen H, Bornkamm GW, Schmidt R, Hecker E. Tumor initiators and promoters in the induction of Epstein-Barr virus. Proc. Natl. Acad. Sci. USA. 1979;76:782–785. doi: 10.1073/pnas.76.2.782. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Polack A, Hartl G, Zimber U, Freese UK, Laux G, Takaki K, Hohn B, Gissmann L, Bornkamm GW. A complete set of overlapping cosmid clones of M-ABA virus derived from nasopharyngeal carcinoma and its similarity to other Epstein-Barr virus isolates. Gene. 1984;27:279–288. doi: 10.1016/0378-1119(84)90072-6. [DOI] [PubMed] [Google Scholar]
- 22.Norio P, Schildkraut CL. Visualization of DNA replication on individual Epstein-Barr virus episomes. Science. 2001;294:2361–2364. doi: 10.1126/science.1064603. [DOI] [PubMed] [Google Scholar]
- 23.Little RD, Schildkraut CL. Initiation of latent DNA replication in the Epstein-Barr virus genome can occur at sites other than the genetically defined origin. Mol. Cell. Biol. 1995;15:2893–2903. doi: 10.1128/mcb.15.5.2893. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Wensing B, Stuhler A, Jenkins P, Hollyoake M, Karstegl CE, Farrell J. Variant chromatin structure of the oriP region of Epstein-Barr virus and regulation of EBER1 expression by upstream sequences and oriP. J. Virol. 2001;75:6235–6241. doi: 10.1128/JVI.75.13.6235-6241.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Jankelevich S, Kolman JL, Bodnar JW, Miller G. A nuclear matrix attachment region organizes the Epstein-Barr viral plasmid in Raji cells into a single DNA domain. EMBO J. 1992;11:1165–1176. doi: 10.1002/j.1460-2075.1992.tb05157.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Mearini G, Chichiarelli S, Zampieri M, Masciarelli S, D’Erme M, Ferraro A, Mattia E. Interaction of EBV latent origin of replication with the nuclear matrix: identification of S/MAR sequences and protein components. FEBS Lett. 2003;547:119–124. doi: 10.1016/s0014-5793(03)00690-2. [DOI] [PubMed] [Google Scholar]
- 27.Bernasconi M, Berger C, Sigrist JA, Bonanomi A, Sobek J, Niggli FK, Nadal D. Quantitative profiling of housekeeping and Epstein-Barr virus gene transcription in Burkitt lymphoma cell lines using an oligonucleotide microarray. Virol. J. 2006;3:43. doi: 10.1186/1743-422X-3-43. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Murata T, Kondo Y, Sugimoto A, Kawashima D, Saito S, Isomura H, Kanda T, Tsurumia T. Epigenetic histone modification of Epstein-Barr Virus BZLF1 promoter during latency and reactivation in Raji Cells. J. Virol. 2012;86:4752–4761. doi: 10.1128/JVI.06768-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Stenerlöw B, Karlsson KH, Cooper B, Rydberg B. Measurement of prompt DNA double-strand breaks in mammalian cells without including heat-labile sites: results for cells deficient in nonhomologous end joining. Radiat. Res. 2003;159:502–510. doi: 10.1667/0033-7587(2003)159[0502:mopdds]2.0.co;2. [DOI] [PubMed] [Google Scholar]
- 30.Wohl T, Brecht M, Lottspeich F, Ammer H. The use of genomic DNA probes for in-gel hybridization. Electrophoresis. 1995;16:739–741. doi: 10.1002/elps.11501601120. [DOI] [PubMed] [Google Scholar]
- 31.Leach TJ, Glaser R. Quantitative hybridization to genomic DNA fractionated by pulsed-field gel electrophoresis. Nucl. Acids Res. 1998;26:4787–4789. doi: 10.1093/nar/26.20.4787. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Gurrieri S, Smith B, Bustamente C. Trapping of megabase-sized DNA molecules during agarose gel electrophoresis. Proc. Natl. Acad. Sci. USA. 1999;96:453–458. doi: 10.1073/pnas.96.2.453. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Maleszka R. Single-stranded regions in yeast mitochondrial DNA revealed by pulsed-field gel electrophoresis. Appl. Theor. Electrophor. 1993;3:259–263. [PubMed] [Google Scholar]
- 34.Beverley SM. Estimation of Circular DNA Size Using γ-Irradiation and Pulsed-Field Gel Electrophoresis. Anal. Biochem. 1989;177:110–114. doi: 10.1016/0003-2697(89)90023-7. [DOI] [PubMed] [Google Scholar]
- 35.Allemand JF, Bensimon D, Jullien L, Bensimon A, Croquette V. pH-dependent specific binding and combing of DNA. Biophys. J. 1997;73:2064–2070. doi: 10.1016/S0006-3495(97)78236-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Ernberg I, Falk K, Minarovits J, Busson P, Tursz T, Masucci MG, Klein G. The role of methylation in the phenotype-dependent modulation of Epstein-Barr nuclear antigen 2 and latent membrane protein genes in cells latently infected with Epstein-Barr virus. J. Gen. Virol. 1989;70:2989–3002. doi: 10.1099/0022-1317-70-11-2989. [DOI] [PubMed] [Google Scholar]
- 37.Cullinan EB, Beerman TA. A study of drug-induced topoisomerase II-mediated DNA lesions on episomal chromatin. J. Biol. Chem. 1989;264:16268–16275. [PubMed] [Google Scholar]
- 38.Udvardy A, Schedl P. Chromatin structure, not DNA sequence specificity, is the primary determinant of topoisomerase II sites of action in vivo. Mol. Cell. Biol. 1991;11:4973–4984. doi: 10.1128/mcb.11.10.4973. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Razin SV, Petrov P, Hancock R. Precise localization of the α-globin gene cluster within one of the 20- to 300-kb DNA fragments released by cleavage of chicken chromosomal DNA at topoisomerase II sites in vivo: evidence that the fragments are DNA loops or domains. Proc. Natl. Acad. Sci. USA. 1991;88:8515–8519. doi: 10.1073/pnas.88.19.8515. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.D'Arpa P, Liu LF. Topoisomerase-targeting antitumor drugs. Biochim. Biophys. Acta. 1989;989:163–177. doi: 10.1016/0304-419x(89)90041-3. [DOI] [PubMed] [Google Scholar]
- 41.D'Andrea AD, Haseltine WA. Sequence specific cleavage of DNA by the antitumor antibiotics neocarzinostatin and bleomycin. Proc. Natl. Acad. Sci. USA. 1978;75:3608–3612. doi: 10.1073/pnas.75.8.3608. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Kuo MT, Samy TS. Effects of neocarzinostatin on mammalian nuclei: release of nucleosomes. Biochim. Biophys. Acta. 1978;518:186–190. doi: 10.1016/0005-2787(78)90129-6. [DOI] [PubMed] [Google Scholar]
- 43.McHugh MM, Woynarowski J, Beerman T. Degradation of HeLa cell chromatin by neocarzinostatin and its chromophore. Biochim. Biophys. Acta. 1982;696:7–14. doi: 10.1016/0167-4781(82)90003-3. [DOI] [PubMed] [Google Scholar]
- 44.Halliwell B, Gutteridge JM. Free Radicals in Biology and Medicine. Oxford: Oxford University Press; 1999. [Google Scholar]
- 45.Ward JF. DNA damage produced by ionizing radiation in mammalian cells: identities, mechanisms of formation, and reparability. Prog. Nucleic Acid Res. Mol. Biol. 1988;35:95–125. doi: 10.1016/s0079-6603(08)60611-x. [DOI] [PubMed] [Google Scholar]
- 46.de Jesus O, Smith PR, Spender L, Karstegl CE, Niller HH, Huang D, Farrell PJ. Updated Epstein-Barr virus DNA sequence and analysis of a promoter for the BART (CST, BARF0) RNAs of Epstein-Barr virus. J. Gen. Virol. 2003;84:1443–1450. doi: 10.1099/vir.0.19054-0. [DOI] [PubMed] [Google Scholar]
- 47.Fruscalzo A, Marsili G, Busiello V, Bertolini L, Frezza D. DNA sequence heterogeneity within the Epstein–Barr virus family of repeats in the latent origin of replication. Gene. 2001;265:165–173. doi: 10.1016/s0378-1119(01)00345-6. [DOI] [PubMed] [Google Scholar]
- 48.Hofer B. The sensitivity of DNA cleavage by SpeI and ApaLI to methylation by M.EcoK. Nucleic Acids Res. 1988;16:5206. doi: 10.1093/nar/16.11.5206. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Rogakou EP, Pilch DR, Orr AH, Ivanova VS, Bonner WM. DNA double-stranded breaks induce histone H2AX phosphorylation on serine 139. J. Biol. Chem. 1998;273:5858–5868. doi: 10.1074/jbc.273.10.5858. [DOI] [PubMed] [Google Scholar]
- 50.Spotheim-Maurizot M, Charlier M. Sequencing gel electrophoresis: a tool for studying radiation damage to DNA. Radiat. Res. 1997;48:501–502. [Google Scholar]
- 51.Johnson PG, Beerman TA. Damage induced in episomal EBV DNA in Raji cells by antitumor drugs as measured by pulsed field gel electrophoresis. Anal. Biochem. 1994;220:103–114. doi: 10.1006/abio.1994.1305. [DOI] [PubMed] [Google Scholar]
- 52.Barone F, Belli M, Rongoni E, Sapora O, Tabocchini MA. X-ray induced DNA double-strand breaks in polynucleosomes. In: Bums FJ, Upton AC, Silini G, editors. Radiation Carcinogenesis and DNA Alterations. Plenum. New York, NY: 1986. pp. 293–296. [Google Scholar]
- 53.Nishigaki K, Kaneko Y, Wakuda H, Husimi Y, Tanaka T. Type II restriction endonucleases cleave single-stranded DNAs in general. Nucleic Acids Res. 1985;13:5747–5760. doi: 10.1093/nar/13.16.5747. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Muslimovic A, Nyström S, Gao Y, Hammarsten O. Numerical analysis of etoposide induced DNA breaks. PLoS One. 2009;4:e5859. doi: 10.1371/journal.pone.0005859. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Ottinger LM, Tullius TD. High-resolution in vivo footprinting of a protein-DNA complex using γ-radiation. J. Am. Chem. Soc. 2000;122:5901–5902. [Google Scholar]
- 56.Löbrich M, Ikpeme S, Kiefer J. Measurement of DNA double-strand breaks in mammalian cells by pulsed-field gel electrophoresis: a new approach using rarely cutting restriction enzymes. Radiat. Res. 1994;138:186–192. [PubMed] [Google Scholar]
- 57.Cedervall B, Radivoyevitch T. Methods for analysis of DNA fragment distributions on pulsed field gel electrophoretic gels. Electrophoresis. 1996;17:1080–1086. doi: 10.1002/elps.1150170617. [DOI] [PubMed] [Google Scholar]
- 58.Barsoum J, Berg P. Simian virus 40 minichromosomes contain torsionally strained DNA molecules. Mol. Cell. Biol. 1985;5:3048–3057. doi: 10.1128/mcb.5.11.3048. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Rösl F, Waldeck W. Topological properties of bovine papillomavirus type 1 (BPV-1) DNA in episomal nucleoprotein complexes: a model system for chromatin organization in higher eukaryotes. Mol. Carcinog. 1991;4:248–256. doi: 10.1002/mc.2940040312. [DOI] [PubMed] [Google Scholar]
- 60.Sivolob A, Prunell A. Nucleosome conformational flexibility and implications for chromatin dynamics. Phil. Trans. R. Soc. Lond. A. 2004;362:1519–1547. doi: 10.1098/rsta.2004.1387. [DOI] [PubMed] [Google Scholar]
- 61.Liggins GL, English M, Goldstein DA. Structural changes in simian virus 40 chromatin as probed by restriction endonucleases. J. Virol. 1979;31:718–732. doi: 10.1128/jvi.31.3.718-732.1979. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Bancaud A, Conde e Silva N, Barbi M, Wagner G, Allemand J-F, Mozziconacci J, Lavelle C, Croquette V, Victor J-M, Prunell A, et al. Structural plasticity of single chromatin fibers revealed by torsional manipulation. Nat. Struct. Mol. Biol. 2006;13:444–450. doi: 10.1038/nsmb1087. [DOI] [PubMed] [Google Scholar]
- 63.Engelhardt M. Choreography for nucleosomes: the conformational freedom of the nucleosomal filament and its limitations. Nucleic Acids Res. 2007;35:e106. doi: 10.1093/nar/gkm560. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Wong H, Victor J-M, Mozziconacci J. An all-atom model of the chromatin fiber containing linker histones reveals a versatile structure tuned by the nucleosomal repeat length. PLoS ONE. 2007;2:e877. doi: 10.1371/journal.pone.0000877. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Cinacchi G, La Penna G, Perico A. Anisotropic internucleosome interactions and geometrical constraints in the organization of chromatin. Macromolecules. 2007;40:9603–9613. [Google Scholar]
- 66.Depken M, Schiessel H. Nucleosome shape dictates chromatin fiber structure. Biophys. J. 2009;96:777–784. doi: 10.1016/j.bpj.2008.09.055. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Scipioni A, Turchetti G, Morosetti S, De Santis P. Geometrical, conformational and topological restraints in regular nucleosome compaction in chromatin. Biophys. Chem. 2010;148:56–67. doi: 10.1016/j.bpc.2010.02.010. [DOI] [PubMed] [Google Scholar]
- 68.Capranico G, Jaxel C, Roberge M, Kohn KW, Pommier Y. Nucleosome positioning as a critical determinant for the DNA cleavage sites of mammalian DNA topoisomerase II in reconstituted simian virus 40 chromatin. Nucleic Acids Res. 1990;18:4553–4559. doi: 10.1093/nar/18.15.4553. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Thoma F, Zatchej M. Chromatin folding modulates nucleosome positioning in yeast minichromosomes. Cell. 1988;55:945–953. doi: 10.1016/0092-8674(88)90240-1. [DOI] [PubMed] [Google Scholar]
- 70.White JH, Gallo R, Bauer WR. Dependence of the linking deficiency of supercoiled minichromosomes upon nucleosome distortion. Nucleic Acids Res. 1989;17:5827–5835. doi: 10.1093/nar/17.14.5827. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Martino JA, Katritch V, Olson WK. Influence of nucleosome structure on the three-dimensional folding of idealized minichromosomes. Structure. 1999;7:1009–1022. doi: 10.1016/s0969-2126(99)80127-0. [DOI] [PubMed] [Google Scholar]
- 72.Enright HU, Miller WJ, Hebbel RP. Nucleosomal histone protein protects DNA from iron-mediated damage. Nucleic Acids Res. 1992;20:3341–3346. doi: 10.1093/nar/20.13.3341. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Dobrovolskaia IV, Kenward M, Arya G. Twist propagation in dinucleosome arrays. Biophys. J. 2010;99:3355–3364. doi: 10.1016/j.bpj.2010.09.055. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Garner MM, Felsenfeld G, O'Dea MH, Gellert M. Effects of DNA supercoiling on the topological properties of nucleosomes. Proc. Natl. Acad. Sci. USA. 1987;84:2620–2623. doi: 10.1073/pnas.84.9.2620. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Zlatanova J, Leuba SH, Yang G, Bustamente C, Van Holde K. Linker DNA accessibility in chromatin fibers of different conformations: a reevaluation. Proc. Natl. Acad. Sci. USA. 1994;91:5277–5280. doi: 10.1073/pnas.91.12.5277. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Labarbe R, Mignon S, Flock S, Houssier C. Diffusion-enhanced resonance energy transfer shows that linker-DNA accessibility decreases during salt-induced chromatin condensation. J. Fluorescence. 1996;6:107–118. doi: 10.1007/BF00732050. [DOI] [PubMed] [Google Scholar]
- 77.Staynov DZ. DNase I footprinting of the nucleosome in whole nuclei. Biochem. Biophys. Res. Comm. 2008;372:226–229. doi: 10.1016/j.bbrc.2008.05.024. [DOI] [PubMed] [Google Scholar]
- 78.Schiessel H, Gelbart WM, Bruinsma R. DNA folding: structural and mechanical properties of the two-angle model for chromatin. Biophys. J. 2001;80:1940–1956. doi: 10.1016/S0006-3495(01)76164-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Scheffer MP, Eltsov M, Frangakisa AS. Evidence for short-range helical order in the 30-nm chromatin fibers of erythrocyte nuclei. Proc. Natl. Acad. Sci. USA. 2011;108:16992–16997. doi: 10.1073/pnas.1108268108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Stehr R, Kepper N, Rippe K, Wedemann G. The effect of the internucleosomal interaction on the folding of the chromatin fiber. Biophys. J. 2008;95:3677–3691. doi: 10.1529/biophysj.107.120543. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Mergell B, Everaers R, Schiessel H. Nucleosome interactions in chromatin: fiber stiffening and hairpin formation. Phys. Rev. E. Stat. Nonlin. Soft Matter Phys. 2004;70:011915. doi: 10.1103/PhysRevE.70.011915. [DOI] [PubMed] [Google Scholar]