Abstract
Spatial control of chemical reactions, with micro- and nanometer scale resolution, has important consequences for one pot synthesis, engineering complex reactions, developmental biology, cellular biochemistry and emergent behavior. We review synthetic methods to engineer this spatial control using chemical diffusion from spherical particles, shells and polyhedra. We discuss systems that enable both isotropic and anisotropic chemical release from isolated and arrayed particles to create inhomogeneous and spatially patterned chemical fields. In addition to such finite chemical sources, we also discuss spatial control enabled with laminar flow in 2D and 3D microfluidic networks. Throughout the paper, we highlight applications of spatially controlled chemistry in chemical kinetics, reaction-diffusion systems, chemotaxis and morphogenesis.
1. Introduction
Chemical reactions are often discussed in the context of well-mixed reactants in homogeneous media. The outcome of such reactions is described by spatially homogeneous reaction rates and free energy differences between reactants and products; microscopic mass transport and concentration gradients are often not taken into account. However, reactions occurring in a spatially heterogeneous chemical medium can feature novel properties and outcomes. For example, in multiple-step chemical synthesis, spatial isolation of reactants can reduce interference between simultaneously occurring chemical reactions and allow sequential or hierarchical reactions within the same medium. Further, products can be synthesized using high local reactant concentrations which might otherwise be too dilute when spread across the entire medium. Additionally, three-dimensional heterogeneous chemical patterns and gradients are critical in a variety of inter and intracellular processes. Spatially encoded chemical patterns are important for defining cellular polarity and transmitting information in the biological world, and the function of many biochemical pathways require chemicals to be spatially separated in three dimensional spaces.
In order to engineer spatial control over chemical reactions, there is a need to accurately and reproducibly generate spatially patterned chemical fields (Fig. 1). As compared to a spatially homogeneous chemical field (Fig. 1a) that is typically achieved using well mixed reactants, the precise positioning of discrete chemical sources can be utilized to generate a spatially heterogeneous chemical field (Fig. 1b–f). The spatial characteristics of the field can be tuned by varying its constituent local and global features such as the shape, size, permeability, chemical concentration and spatial distribution of chemical sources in both stationary and flowing media. We review methods to generate a variety of chemical fields by confining and releasing chemical reactants from containers or fluidic channels. The containers can be in the form of vesicles, microspheres, fibers, polyhedra or other well-shaped particles or capsules. We discuss methods for synthesis of symmetrical and asymmetrical chemical-releasing containers and survey both passively diffusing and stimuli-responsive release mechanisms. Further, in addition to tuning the chemical release characteristics of individual containers, we discuss the manipulation of chemical fields by arranging the sources in 2D and 3D arrays. Examples of chemical patterning generated by flowing media in microfluidic devices can enable dynamic fields, and these are highlighted. We conclude by comparing the advantages and limitations of chemical pattern generation by discrete particles and by microfluidic devices.
Fig. 1.
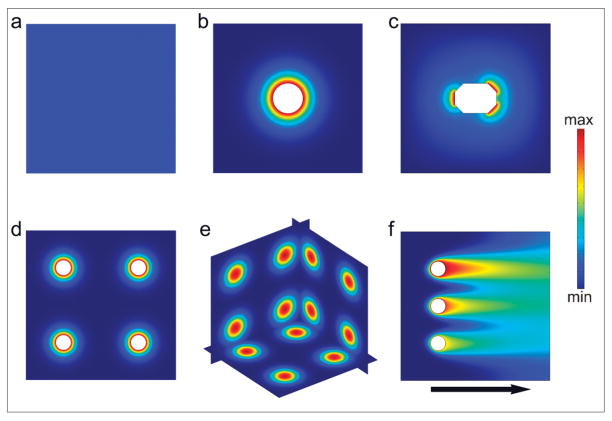
Schematic illustration of chemical fields generated by particles and streams. (a) Homogeneous chemical field. (b) Radially symmetric chemical field generated by a single particle source. (c) Asymmetric chemical field generated by a single particle source. (d) 2D array of radially-symmetric particulate sources. (e) 3D array of spherically symmetric chemical sources. (f) Chemical field generated by three sources releasing the same chemical with different concentration in a flowing stream.
2. Motivation
The need for engineering chemical fields is largely motivated by observations that it can be used to significantly enhance functionality as is evident in a variety of cellular processes. Further, a few synthetic and spatially patterned chemical systems have been shown to display emergent behaviors such as periodic oscillations. We highlight important examples of chemical patterning that have been observed within individual or multiple cells and reaction-diffusion systems.
2.1. Chemical patterning within individual cells
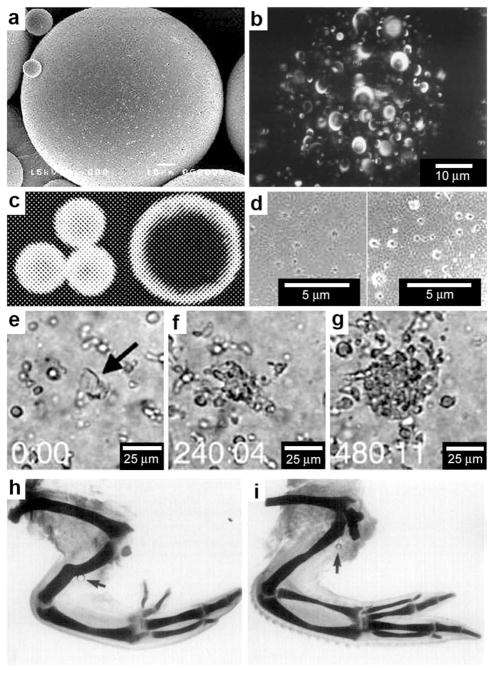
The development and application of advanced optical imaging methods and genetically encoded fluorescent reporters have revealed that critical cellular functions require precise spatial patterning of chemicals both within and around cells.1,2 Recent studies have convincingly demonstrated that spatial localization of biomolecules such as proteins and DNA occurs even in prokaryotic cells which do not contain any membrane bound intracellular organelles.3,4 These studies have challenged traditional views, which describe prokaryotic cells as being non-compartmentalized with a homogeneous distribution of biomolecules within the cell. For example, in Caulobacter crescentus, intracellular localization of histidine kinases and proteins control important functions such as cell division (Fig. 2a). A striking example of intercellular spatial localization of protein secretion has been observed in gram-positive bacteria such as Streptococcus pyogenes (Fig. 2b).5 As compared to gram-negative bacteria, which possess a periplasmic space that can be utilized to fold proteins prior to secretion, gram-positive bacteria must rely on mechanisms such as spatially controlled secretion of chaperones at the same site of protein secretion to assist in site-localized folding and maturation of important functional proteins such as the cysteine protease SpeB (Note: a list of all abbreviations and acronyms used in the paper is included in the ESI†). Spatially directed chemical secretion of virulence factors by gram-negative bacteria such as Vibrio cholerae are important during cholera pathogenesis. It has been hypothesized that the localized bacterial secretion of hemagglutinin/protease at its site of attachment would likely result in rapid detachment and would reduce the amount of protein required for detachment from the intestinal epithelium (Fig. 2c).6 Flow in the vicinity of isotropically secreting chemical sources can also induce anisotropy (Fig. 2d,e) and it has been hypothesized that this anisotropy may have an important role in cancer metastasis. For example, Swartz et al. have suggested that autologous chemotaxis may be an important mechanism in the migration of individual cancer cells to the lymphatics.7,8 They argue that the flow generated due to lymphatic drainage spatially polarizes transcellular chemokine gradients and these anisotropic gradients allow individual cells to sense direction and move into the lymph vessels.7,8 The above examples are intended to highlight the importance of spatial patterning of biochemicals to a variety of important single cellular functions in both prokaryotes and eukaryotes.
Fig. 2.
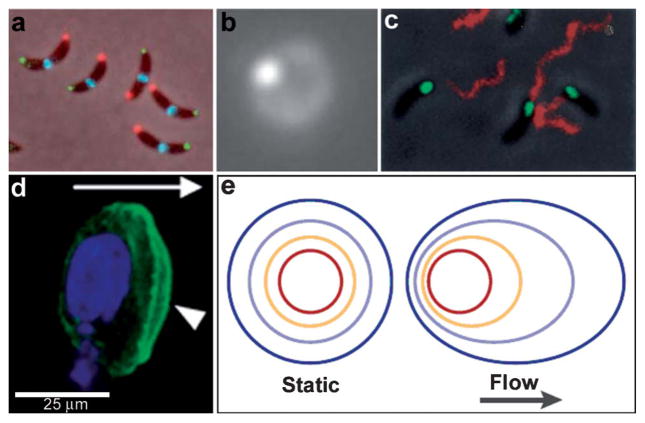
Examples of chemical patterning at the single cell level. (a) An anterior-posterior cellular localization axis is exhibited by the Caulobacter histidine kinases PleC (red) and DivJ (green) that dynamically and selectively localize to specific cell poles. The ZapA cell division protein (blue) localizes to the FtsZ ring. (Reprinted with permission from ref. 2. Copyright (2009) by The American Association for the Advancement of Science). (b) Live streptococcal cells (seen in the image as a diffuse ring) secrete enzymatically active SpeB at a single locus (brighter spot in the image). Representative micrograph shows that each cell displays a single punctate fluorescent locus. (Reprinted with permission from ref. 5. Copyright (2004) by The American Association for the Advancement of Science). (c) Co-localization of expression of Eps apparatus (green) and flagellum (red) at the same pole in the cells of Vibrio cholerae. The bacteria were visualized by phase-contrast microscopy. (Reprinted with permission from ref. 6. Copyright (2001) by The National Academy of Sciences, USA). (d) Staining of fixed samples with phalloidin demonstrates directional reorganization and membrane localization (arrowheads) of the actin cytoskeleton in a flow-dependent manner. (Reprinted with permission from ref. 8. Copyright (2007) by Elsevier.) (e) An idealized steady-state concentration profile of a secreted chemokine surrounding a cell (centered in the red ring) under interstitial flow. The color indicates the concentration range of the secreted molecule, from 100% of the secreted concentration (red) to 0% (dark blue). (Reprinted with permission from ref. 233. Copyright (2005) by The Nature Publishing Group).
2.2. Chemical patterning in multi-cellular systems
In addition to spatial patterning within and around single cells, spatial patterning of chemicals within multi-cellular systems is critical to the function of organs and organisms. Here, chemical fields generated by a multitude of cellular sources orchestrate inter-cellular signaling and embryonic development. For example, during the development of the central nervous system (CNS), gradients of diffusible chemoattractants direct the migration of neuroblasts to their final positions.9 Similarly, in the vertebrate nervous system, distinct neurogenic and non-neurogenic regions form, each providing a uniquely patterned chemical environment.10 Numerous experiments have shown that individual neurons and their growth cones are affected and can be guided by chemical gradients.11–16 Angiogenesis, the process of blood vessel growth, also critically depends on spatially patterned chemical gradients of oxygen and proteins such as vascular endothelial growth factor (VEGF).17–20
Striking examples of the influence of the spatial patterning of chemicals in multi-cellular organisms can be seen in the shaping of the developing embryo due to the patterning of signaling molecules or morphogens.21–25 For example, gradients of morphogens such as transcription factors can alter gene expression and cellular differentiation. In embryonic development, chemical gradients play a critical role in transmitting positional and polarity information.26 Experiments with blastula cells of Xenopus, a genus of aquatic frogs, have shown that single cells respond to different concentrations of activin, irrespective of their contact with other cells.27 Further, the specification of the pattern of digits in chick limb morphogenesis has been observed to depend on their distance from the zone of polarizing activity along the anterior-posterior axis.28 Consequently, it has been suggested that diffusion distances from morphogen sources could provide a means to define spatial positioning.29
It is also important to note that morphogen gradients need to be robust and precise to generate reproducible outcomes from embryo to embryo. For example, Gregor et al. have investigated the precision required in gradients of the Bicoid morphogen in the Drosophila embryo (Fig. 3a).30,31 They argue that in order to adopt distinct fates, adjacent nuclei in the developing embryo must acquire positional information with a remarkable accuracy of 1–2%. Their calculation is based on the fact that adjacent nuclei are separated by a distance of approximately 8 μm within a developing embryo of approximately 500 μm in length. The reproducibility and precision of the generation of Bicoid morphogen gradients is orchestrated by a complex set of input/output mechanisms. In fact, immunostaining experiments have revealed an exponential gradient with a length constant of approximately 100 μm.32
Fig. 3.
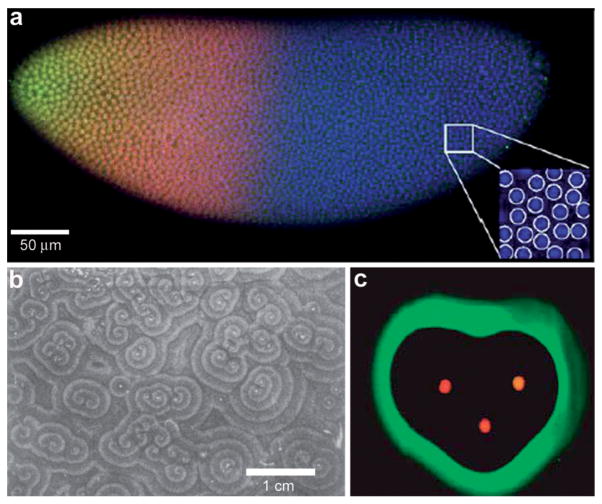
Examples of chemical patterning at the multi-cellular level. (a) Scanning confocal microscope image of a Drosophila embryo in early nuclear cycle, stained for DNA (blue), Hb (red), and Bcd (green); scale bar 50 μm. Inset (28 × 28 μm2) shows how DNA staining allows for automatic detection of nuclei. (Reprinted with permision from ref. 30. Copyright (2007) by Elsevier.) (b) Organized waves of cell movement during aggregation in Dictyostelium discoideum. (Reprinted with permission from ref. 33. Copyright (1981) by The American Association for the Advancement of Science). (c) Experimental result showing heart-shaped GFP patterns formed based on the placement and initial concentrations of sender cells (three sender disks are shown in the image in red color) expressing DsRed-Express. (Reprinted with permission from ref. 35. Copyright (2005) by The Nature Publishing Group).
In addition to exponential gradients, a variety of other spatial distributions of chemicals are important in multicellular behavior. For example, waves of chemicals such as adenosine 3′,5′-monophosphate (cyclic AMP) have been observed during aggregation of the slime mold Dictyostelium discoideum (Fig. 3b).33,34 Here, the spatial distribution of high concentrations of c-AMP was observed to form in spiral or concentric patterns and this had important consequences for determining cell shape; cells in the vicinity of high c-AMP concentrations were elongated, whereas cells were randomly shaped in low concentrations.33 Recently, programmed chemical patterning has been demonstrated in a genetically engineered, multicellular E. coli system.35 Here, patterns of green fluorescent protein (GFP) in a variety of 2D shapes, such as an ellipse or a heart (Fig. 3c) were created by positioning discs of so called sender cells that synthesize chemicals and release gradients of acyl-homoserine lactone (AHL), which represses GFP production.35 These experiments suggest the important link between spatial patterning of chemicals and its outcome on multi-cellular organization and behavior.
2.3. Chemical patterning in reaction-diffusion systems
Chemical reaction-diffusion systems represent one of nature’s most commonly used strategies to generate patterns.36–38 In these systems, the concentration of one or more species ci varies when subject to both chemical reactions (represented by the reaction rates Ri) and diffusion (with diffusion coefficients Di); such systems in general can be represented by systems of partial differential equations for concentrations of the form,
The solutions to these partial differential equations display a wide range of behaviors, including self-organized patterns and wave-like phenomena. As first shown by Turing, two interacting chemicals can generate an inhomogeneous pattern if one of the reactants diffuses much faster than the other so as to generate short range reaction activation and long range inhibition.36 To date, most studies of synthetically generated chemical reaction diffusion systems, such as oscillating reactions, have been carried out in homogeneous media. In these experiments, the chemicals are typically added with no imposed spatial organization39,40 (Fig. 4a) and with or without convection.41
Fig. 4.
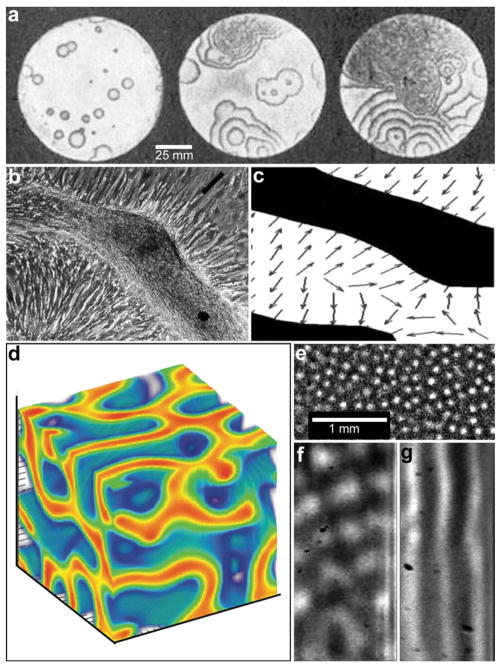
Reaction-diffusion chemical patterns in 2D and 3D. (a) Photographs of a concentration wave propagation in two-dimensional self-oscillatory chemical system. The images were taken at 4 min intervals. Ring diameter equals 100 mm. (Reprinted with permission from ref. 39. Copyright (1970) by The Nature Publishing Group). (b) Pattern formation in cultured Vascular Mesenchymal Cells (VMCs) in vitro. Over 20 days, VMCs develop from a monolayer of randomly oriented cells (not shown in the figure) of nearly uniform density to a ridge with the perpendicular orientation of cells in the monolayer relative to the edges of the multicellular ridge. The black bar in (b) shows the approximate size, shape, and orientation of a single cell. (Reprinted with permission from ref. 51. Copyright (2004) by The National Academy of Sciences, USA). (c) Numerical solutions corresponding to the experimental image shown in (b). Model results are displayed as levels of one of the chemicals involved (the activator) with black representing high and white representing low levels. Gray arrows depict the direction field of gradients of the activator concentration, which corresponds to the perpendicular orientation of cells in culture. (Reprinted with permission from ref. 51. Copyright (2004) by The National Academy of Sciences, USA). (d) 3D steady state patterns of VMCs obtained in simulations of pattern formation of the cells arising from their interaction with Bone Morphogenic Protein-2 (BMP-2) and its inhibitor, Matrix Gla Protein (MGP) in three dimensions. In 3D, the steady state patterns produced are highly interconnected tubes which have planar surfaces. (Reprinted with permission from ref. 52). (e–g) Tomographic study of the Belousov-Zhabotinsky reaction: snapshots of stationary 2D spots (e) in a thin layer and 2D images of the corresponding 3D structures (f) and (g). Bright regions correspond to high concentrations of the oxidized form of the catalyst. (Reprinted with permission from ref. 53. Copyright (2011) by The American Association for the Advancement of Science).
Living systems, in contrast tend to be cellular with well-defined spatial structure. Hence, the study of coupled cellular spatiotemporal systems is also important; a few seminal studies such as those involving coupled linear reaction oscillators operating with the bistable chlorite-iodide reactions,42,43 diffusive coupling of coupled spatio-temporal patterns across a membrane44 and excitable catalyst loaded particles45 have shown rich emergent behavior including quorum sensing.46 Computationally, the concept of interacting cellular automata with coupled chemical reactions is well developed.47,48 However, it is also known that the number of limit cycles for a coupled system tends to grow exponentially with the number of nonlinear oscillator subsystems,49 and new statistical methods are needed to capture the behavior of such systems.
These oscillating chemical reactions are interesting not just for their own sake but also as models of biological processes. By a hierarchical coupling of such systems, highly complex patterns can be generated with one pattern directing a subsequent pattern and so on.50 For example, recent experimental and theoretical work has shown that multipotent adult vascular mesenchymal cells self-organize in vitro into 2D patterns that are predicted by a mathematical model based on molecular morphogens interacting in a reaction-diffusion process (Fig. 4b, c).51 The authors have identified activator and inhibitor morphogens for stripe, spot, and labyrinthine patterns and confirm the model predictions in vitro.
While the bulk of the research on reaction-diffusion pattern formation has been done in 2D, recent simulation (Fig. 4d)52 and experimental (Fig. 4e–g)53 studies on reaction diffusion pattern formation in 3D highlight that dimensionality of the pattern plays an important role. For example, Bansagi et al. demonstrated the existence of Turing patterns including curved surfaces, hexagonally packed cylinders, spots, labyrinthine and lamellar patterns that can exist only in three dimensions (Fig. 4e–g).53 They used a Belousov–Zhabotinsky (BZ) reaction incorporated into a water-in-oil aerosol and studied 3D patterns generated in a cylindrical quartz capillary. Simulations of 3D patterns of vascular mesenchymal cells arising from their interaction with Bone Morphogenic Protein-2 (BMP-2) and its inhibitor, Matrix Gla Protein (MGP) also show a wide variety of three-dimensional cellular patterns such as spheres, tubes, and sheets that are not observed in a two-dimensional analysis.52
The aforementioned studies suggest unusual properties and outcomes when chemistry is carried out in heterogeneous media. The examples discussed highlight the importance of the ability to generate reproducible and precise chemical sources and fields in both two and three dimensions.54 In the following sections, we review the existing methodologies including particle-based and fluidic techniques that can be used to generate such heterogeneous and cellular media in both two and three dimensions and at sub-mm length scales.
3. Chemical patterning using synthetic particles
Synthetic point-like chemical sources can restrict chemical reactions to a particular region of space. Hence, an important synthetic strategy to locally alter the chemical properties of a uniform medium is to carry out reactions with small solid particles or liquid drops of reactants or catalysts.55–58 However, in the absence of any secondary surface stabilizing molecules or synthetic package, there is limited control over the time scale of dissolution of the solid particulates or liquid drops in the medium, and consequently the chemical release rate. This control can be achieved by utilizing particles in a variety of shapes, sizes and molecular compositions that delay the release of reactants to different extents. Many of these particles have been developed for alternate applications such as drug delivery but could as well be utilized to generate 3D spatial control in chemistry. The important particle classes that can be utilized for spatially engineered chemical release are reviewed below.
3.1. Vesicles and capsules
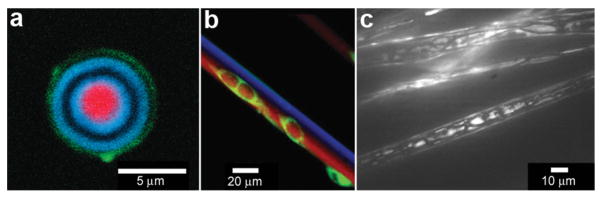
In order to provide additional control over the rate of reactant release and to provide protection from undesirable interactions, chemicals can be confined within thin shells or molecular membranes composed of a variety of short and long chain molecules such as surfactants, lipid bilayers (liposomes; Fig. 5a, b) and polymers.59–62 Since the permeability of these membranes can vary widely for different chemicals, these particles facilitate spatial separation of reactant molecules based on the shell porosity and the solubility and diffusion rate of the encapsulated molecules through the shell. In addition, these particles can be made very small so that they encapsulate single molecules (Fig. 5a) and extremely small volumes, on the order of zeptoliters (10−21 liters; approximately 10 nm diameter scale particles) to femtoliters (10−15 liters; approximately 1 μm diameter scale particles). As pointed out by Chiu et al., at these small sizes, the surface-to-volume ratio is very high and the contained molecules frequently collide with the shell.63 Their Brownian motion simulations indicate that, in a 170-nm-diameter vesicle, the frequency of substrate–phospholipid wall collisions are almost three orders of magnitude more frequent than collisions of a single contained enzyme and substrate.63 Thus, the reaction kinetics within these core-shelled particles would be dominated by surface collisions. Additionally, diffusion based mixing is very fast in these small volumes, typically on the order of micro to milliseconds allowing reactions to be carried out under conditions far different than can be achieved by convective mixing in the bulk. In addition to showing that chemical reactions could be carried out within a single vesicle, the authors describe chemical reactions by initiating electrofusion between two vesicles loaded with fluo-3 (10 μM) and Ca2+ (10 μM). The two chemicals reacted locally as indicated by a 40 fold enhancement in fluorescence (Fig. 5c–f).
Fig. 5.
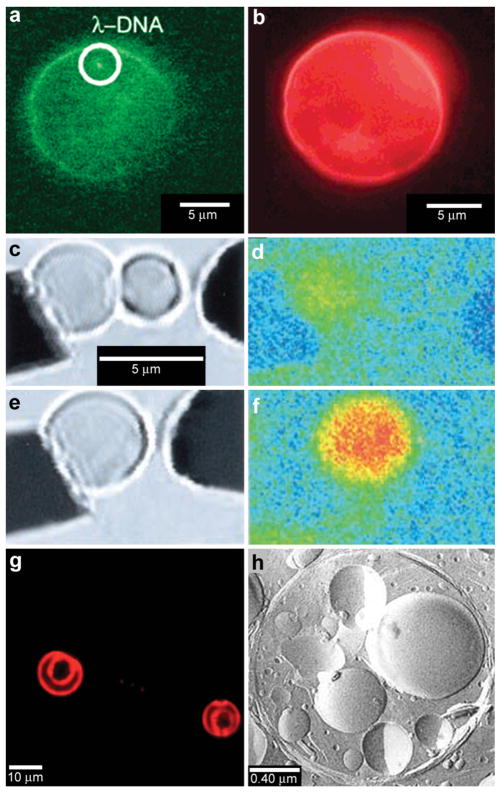
Chemical reactions with vesicles and spherical capsules. (a,b) Fluorescence photographs of λ-DNA-loaded (green, shown in (a)) liposome (red, shown in (b)). (Reprinted with permission from ref. 59. Copyright (2005) by The American Chemical Society). (c–f) Chemical reactions in merging lipid vesicles. (c, e) Electrofusion (about 75 kV cm−1; 10 μs) of a 10 μM fluo-3 containing vesicle (left) and a 10 μM Ca2+ containing vesicle (right) under bright-field illumination. Corresponding fluorescence images are shown in (d) and (f). Binding of Ca2+ by fluo-3 increases the fluorescence quantum yield of this chelator by about 40-fold as was indeed observed. (Reprinted with permission from ref. 63. Copyright (1999) by The American Association for the Advancement of Science). (g) A light scanning confocal microscope (LSCM) image of multivesicle assembly. (Reprinted with permission from ref. 234. Copyright (2008) by John Wiley and Sons). (h) A transmission electron microscopy (TEM) image of a multicompartment structure, the vesosome, formed from a lipid mixture. Multiple small vesicle compartments are visible inside one of more exterior bilayers. (Reprinted with permission from ref. 73. Copyright (2004) by Bentham Science Publishers Ltd.).
Vesicles can be created in a variety of sizes and configurations (e.g. unilamellar and multilamellar) and with a variety of phospholipids and polymers (such as polymersomes).64,65 The choice of material and size often determines durability of the vesicles, the release rate of the chemical into the external medium as well as allows for location-based targeting when vesicles are used in drug delivery applications. Indeed, it has been found that polymer vesicles are more durable than phospholipids.66 Polymer vesicles are also capable of slower release of their content. Here the release rate f can be estimated by the following relation: , where P is the permeation coefficient, l is the membrane thickness and R is the vesicle radius. Since polymer bilayers are generally thicker (5–20 nm) than lipid bilayers (3–5 nm) permeation of chemicals tends to be slower in the case of polymers.66
Vesicles with multiple compartments, such as vesosomes, have also been synthesized (Fig. 5g, h).67–69,73,234 Compartmentalization provides a means to encapsulate different chemical reagents70,71 and delay the release of chemicals to the external medium by providing additional barriers against degradative enzymes and/or proteins.72,73 Individual compartments within the compartmentalized structures can be individually designed to meet specific requirements.
Compartmentalization can also be achieved by spatially localizing chemicals within a core shelled structure and thus provides a means to achieve sequential chemical reactions. For example, van Dongen et al. have designed a three enzyme cascade reaction by positioning three enzymes: glucose oxidase (GOx), Candida antarctica lipase B (CalB) and horseradish peroxidase (HRP), in the lumen, membrane and surface of a polymersome.62 They observed overall kinetics of a two enzyme system instead of the expected three enzyme system due to a high activity of HRP which they attribute to a consequence of spatial positioning, allowing it to be inconsequential in the overall rate equation. Enzyme encapsulation has also been achieved in layer-by-layer polymer multilayer capsules. The encapsulated enzymes remain accessible to their substrates but are protected from high molecular weight inhibitors, and the products of the reaction are able to diffuse to the exterior.74–76
3.2. Polymer microspheres
Polymer microspheres are particles typically synthesized by emulsion methods and are often utilized for localized and sustained release of drugs.77–79 The polymeric particles often adopt spherical shapes due to minimization of surface energy (Fig. 6a) and the chemicals soaked within the polymer microsphere are released during polymer degradation and microsphere erosion. The chemicals encapsulated in polymer microspheres on the average end up being distributed with radial symmetry within the microsphere but the distribution can be varied based on the details of the fabrication process and material properties (Fig. 6b).80 In addition, it has been reported that for the same polymer–drug combination, the microsphere size can have a pronounced effect on the spatial distribution of the encapsulated chemicals within the polymer matrix (Fig. 6c).81
Fig. 6.
Chemical patterning with polymer microspheres. (a) Surface scanning electron (SEM) micrograph of a polymer microsphere. (Reprinted with permission from ref. 83. Copyright (2000) by Elsevier). (b) Three-dimensional confocal microscopy image depicting the spherical-occlusion structure of a single PLGA microsphere and localization of (fluorescently labeled) drug along the periphery of the occlusions. (Reprinted with permission from ref. 80. Copyright (1997) by John Wiley and Sons). (c) Laser scanning confocal microscopy cross-sections through the midline of 20 μm and 40 μm rhodamine-loaded microspheres, revealing increasing surface distribution of rhodamine as microsphere diameter increases. (Reprinted with permission from ref. 81. Copyright (2003) by Springer). (d) Surface scanning electronic micrographs of microspheres prepared at 38 °C and 4 °C respectively. (Reprinted with permission from ref. 83. Copyright (2000) by Elsevier). (e–g) Three frames from a time-lapse imaging experiment showing chemotaxis of dendritic cells into direct contact with one isolated large PLGA microsphere (denoted in the first frame by black arrow) filled with a chemoattractant. Elapsed times (min:s) are shown in the lower left of each frame. (Reprinted with permission from ref. 82. Copyright (2005) by Elsevier). (h,i) Chick wings that developed following the application of a range of doses (0.5 μg ml−1 for (h) and 0.1 mg ml−1 for (i)) of all-trans-retinoic acid from AGl-X2 beads (beads are indicated by arrows). Wing digit patterns vary depending on the concentration of all-trans-retinoic acid. (Reprinted with permission from ref. 84. Copyright (1985) by Academic Press).
A consequence of the spherical shape of these polymeric particles and the spherically-symmetric distribution of encapsulated chemicals is that chemicals are released primarily in spherically-symmetric spatial patterns.82 However, within this spherically-symmetric pattern, it is possible to control the radial concentration gradient as well as the chemical release rate using a variety of microsphere synthesis parameters that control the porosity and degradation rate (Fig. 6d).83 A major advantage of the synthesis methods for polymer microspheres is that large numbers of particles with a variety of material constituents including biodegradable ones can be created in a cost-effective manner.
Spherical patterns of released chemicals have been used to guide motile cells to self-organize around polymer microspheres.82 For example, Fig. 6e–g shows the migration of dendritic cells as they chemotax into direct contact with chemoattractant loaded controlled release microspheres; the chemoattractant chosen in this case was macrophage inflammatory protein-3α, MIP-3α. Three frames from a time-lapse imaging experiment near one isolated large PLGA microsphere (the microsphere is denoted in the first frame of the figure by a black arrow) show that the dendritic cells continuously accumulated around this particle over eight hours of observation.82 Another example of an application of spherically-symmetric chemical release involves the release of a morphogen, specifically all-trans-retinoic acid from 200 μm polymer beads (Fig. 6h, i). Following implantation of beads containing all-trans-retinoic acid in the chick embryo wings, the wing digit patterns varied depending on the local chemical concentration.84–86 In addition to beads, 500 μm long and 100 μm wide, chemically loaded paper strips were also tried,84,87 but it was found that beads led to more reproducible digit patterns.
Just as in the case of vesicles, polymer microspheres with multiple compartments88 can be used for chemical reactions. Baeumler et al. used coprecipitation and crosslinking to fabricate spherical multicompartment biopolymer particles (Fig. 7a).89 Their goal was to understand the influence of compartmentalization on coupled reactions inside one particle.89,90 Each of the resulting concentric compartments could be independently loaded with one of the three enzymes, β-glucosidase, glucose oxidase, and horseradish peroxidase. The distance between the enzyme containing compartments was varied using bovine serum albumin (BSA) spacer layers. When fluorogenic substrates for β-glucosidase and horseradish peroxidase were used, the beginning and the end of the coupled enzyme reaction were visualized and recorded inside individual particles, using confocal laser scanning microscopy. A strong influence of the BSA compartment spacing on the reaction kinetics of the last enzyme was observed, suggesting an impaired diffusion of the intermediate products of the chain reaction through the compartments.89
Fig. 7.
Chemical patterning with complex and non-spherical polymer particles and fibers. (a) A CLSM image of a single five-compartment particle which contained three enzymes placed in separated compartments. The image shows the particle after the coupled reaction has completed. Reprinted with permission from ref. 89. Copyright (2010) by The American Chemical Society). (b) A CLSM image of a bicompartmental microfiber. Acetylene-PLGA was incorporated in the red compartment only followed by selective peptide conjugation resulting in cell (shown in green) adhesion alongside the red compartment only. (Reprinted with permission from ref. 98. Copyright (2009) by John Wiley and Sons). (c) FITC-BSA-encapsulated polymer (PCLEEP) electrospun fibers. (Reprinted with permission from ref. 92. Copyright (2005) by The American Chemical Society).
In addition to spherical particles, liquefied or heated polymer microspheres embedded in thin films have been deformed to create more complicated shapes. Notably, Champion et al. have created polystyrene particles in a variety of shapes including disk, rods, and oblate ellipsoids using such an approach.91 Cylindrically shaped polymer fibers have been synthesized by electrospinning and the chemical field generated from single fibers also has cylindrical symmetry (Fig. 7b, c),92–95 while release from a bundle of fibers is dependent on the overall geometry.96,97 Electrohydrodynamic co-spinning can also create dual-compartment fibers with micrometer patterns for cell guidance.98
3.3. Lithographically structured containers
While the aforementioned vesicles and polymeric particles are inherently spherical resulting in spherical spatial control, lithographic approaches99 enable the size and shape of the devices to be varied. Additionally, these highly precise methods can result in better reproducibility and accuracy over chemical release. As an example of utilization of these advantages Ainslie et al. describe the development of multi-layered polymeric cubic containers with an opening in one face (Fig. 8a, b) developed specifically for unidirectional release of chemicals.100 A reservoir is patterned into the microdevice that allows for asymmetric delivery of chemicals, such as therapeutics. Asymmetric delivery was shown to concentrate drug at the device/cell interface, wherein ten times more drug permeated an intestinal epithelial cell monolayer as compared to unprotected drug-loaded hydrogels.100–102 The device also allowed for release of multiple chemical patterns at the same time.
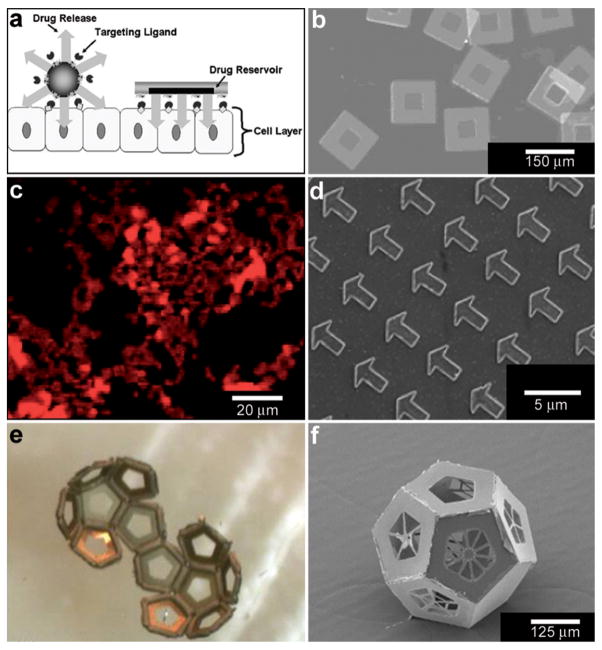
Fig. 8.
Lithographically-structured and self-assembled containers. (a) Schematic representation of a spherical particle and lithographically-fabricated microdevice interface with intestinal epithelial cell surface. This illustration displays the advantages of a microfabricated drug delivery particle over traditional spherical particles: asymmetric release of drug, multi-site targeting for flow stability, and drug reservoir protection can be engineered into the design of the microdevice. (Reprinted from ref. 100 with permission from The Royal Society of Chemistry). (b) Light micrograph of detached SU-8 microdevices without hydrogel. (Reprinted from ref. 100 with permission from The Royal Society of Chemistry). (c) Fluorescence images of cubic (side length 2 μm), Doxorubicin-loaded Trojan horse particles produced using the PRINT technique. (Reprinted with permission from ref. 106. Copyright (2008) by The American Chemical Society). (d) Manipulation of shape using PRINT: 3 μm arrow PEG particles. (Reprinted with permission from ref. 104. Copyright (2005) by The American Chemical Society). (e) Video snapshots featuring the hierarchical self-assembly of a 500 μm dodecahedron. (Reprinted with permission from ref. 111. Copyright (2009) by IOP Publishing Ltd.). (f) An SEM image of a folded 3D dodecahedron container featuring anisotropic surface patterning (i.e., different specific desired patterns on each panel). (Reprinted with permission from ref. 111. Copyright (2009) by IOP Publishing Ltd.).
Molding of pre-polymers in microfabricated molds, channels and capillaries can be utilized to create polymeric particles with a variety of sizes and shapes.103 One notable strategy is a versatile top-down methodology for fabricating monodisperse particles in a variety of sizes, shapes and material composition. PRINT or Particle Replication In Nonwetting Templates utilizes molds microfabricated with non-adherent fluorinated polymers such as photocurable perfluoropolyether to confine liquid precursors within the features of the mold.104 The non-stick nature of the mold allows particles to be well formed and easily released. PRINT particles loaded with a variety of chemicals such as doxorubicin (Fig. 8c) or vaccines have been developed and investigated primarily as an alternative to spherically shaped particles (Fig. 8d) in drug delivery.105,106
3.4. Self-assembled patterned polyhedral containers
When lithographic fabrication is combined with self-assembly, precisely patterned three-dimensional hollow polyhedra can be formed. For example, Leong et al., have developed the process of surface-tension107 and stress-driven108 self-assembly to create such three-dimensional patterned porous structures (Fig. 8e, f). The process utilizes hinge-based self-folding methods which are compatible with planar lithographic methods such as photo, e-beam and nano-imprint lithography. It has been shown that it is possible to use a single self-assembly methodology, which is compatible with the techniques used to pattern microelectronic devices, to create hollow and well-sealed polyhedral particles of different materials (metals, semiconductors and polymers),109 size (ranging from 100 nm to 2 cm),110 shapes (cube, tetrahedron, pyramid, dodecahedron and truncated octahedron),111 and precise sub-100 nm sized pores.112 One challenge in this approach is that while particles with 10 μm sizes and larger can be patterned using photolithography, smaller sized particles require the use of nanopatterning techniques such as electron beam lithography which are serial and expensive processes. Nevertheless, these hollow polyhedra have already been utilized in a range of applications that require high precision in size, shape and side-wall porosity.
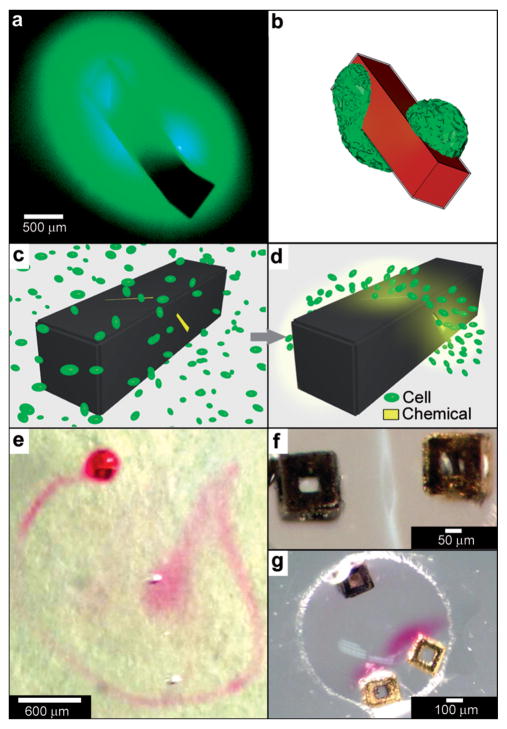
Kalinin et al. utilized the flexibility of the process in terms of creating sealed and precisely patterned three-dimensional containers to enable chemical release with unprecedented 3D spatial control. Following fabrication, the containers were filled with chemicals by submerging them into aqueous solutions of the chemicals. When these containers were submerged into porous media such as gels, the loaded chemical emerged from the pores and diffused into the medium. The three-dimensional shape of the emerging chemical pattern could be independently controlled by the overall shape of the container as well as the pattern of pores on its surface.113 For example, Fig. 9a, b show how a three-dimensional chemical pattern in the shape of a helix could be obtained by defining slanted slits at incrementally increasing positions along the long axis of the container. Consequently, when the containers were filled with chemoattractants such as L-serine, it was possible to guide microorganisms to self-organize into spatial patterns that resemble the chemical pattern created by the patterned container. By selecting the appropriate chemoattractant concentration and time scale of the experiment, the authors showed that it was possible to cause motile E. coli bacteria to self-organize in a helical pattern (Fig. 9c, d).113
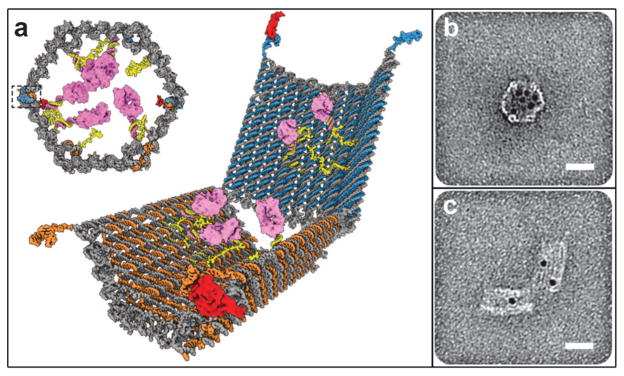
Fig. 9.
Curved, anisotropic and dynamic chemical patterns created by self-assembled microcontainers. (a, b) Generation of 3D spatial patterns by varying pore placement on self-assembled microcontainers. Experimental optical image (a) and numerical simulation (b) of the helical spatial pattern of fluorescein emerging from the container. (Reprinted with permission from ref. 113. Copyright (2011) by John Wiley and Sons). (c, d) Conceptual representation of chemotactic self-organization of motile cells in the shape of the underlying chemical pattern. At the start of the experiment, the chemoattractant is confined to the container, and the cells (represented by green ellipsoids) are distributed uniformly throughout the medium (c). The cells then self-organize in a helical pattern based on the chemical pattern once the chemoattractant (yellow) is allowed to diffuse out of the container (d). (Reprinted with permission from ref. 113. Copyright (2011) by John Wiley and Sons). (e) Optical images showing remotely guided spatially controlled chemical pattern. In this case the container was remotely guided using magnetic fields. The letter G was formed by the direct writing of phenolphthalein in an alkaline water-glycerol medium. (Reprinted with permission from ref. 114. Copyright (2006) by The American Chemical Society). (f) Spatially controlled chemical reactions between multiple containers: reaction of copper sulfate and potassium hydroxide in an aqueous medium resulting in the formation of copper hydroxide along the central line between the containers. (Reprinted with permission from ref. 114. Copyright (2006) by The American Chemical Society). (g) Reaction of phenolphthalein (diffusing out of the two bottom containers) and potassium hydroxide (diffusing out of the top container) in an aqueous medium. (Reprinted with permission from ref. 114. Copyright (2006) by The American Chemical Society).
Further, these polyhedral containers can be fabricated with a variety of materials, including nickel (Ni), which is ferromagnetic. Hence, it is possible to remotely manipulate the position and orientation of the container while it is releasing a chemical, using an external magnetic field. For instance, chemistry could be carried out in the shape of the letter G (Fig. 9e), highlighting the feasibility to spatially control chemistry in any desired 2D or 3D trajectory.114 It was also possible to spatially localize chemical reactions by positioning multiple containers relative to each other. For example, Fig. 9f, g shows two examples of spatially-confined chemical reactions. When chemical reactants diffuse out from these containers, the reaction occurs only where the reactants meet. Variations in the container position, chemical diffusion coefficients and container geometry can be used to further control chemical reactions in different regions of space.114
In addition to controlling the diffusion of chemicals outside the containers, it is also possible to control chemical fields inside the containers in 3D. For example, Randall et al. cultured mammalian cells inside these microcontainers (Fig. 10)115 and observed that the shape of the viable cellular distribution depended on the diffusion pattern of nutrients into the containers. This pattern in turn could be varied by manipulating the pore placement and density on the faces of the containers. Numerical simulations of oxygen diffusion within the containers confirmed the experimentally-observed dependence of the fraction of viable cells on the container face porosity. In the future, this method might be used to create precisely shaped viable 3D cell aggregates by altering the nutrient diffusion patterns around the cells. In addition to manipulating cell viability Randall et al. have also utilized such polyhedral containers for size-exclusion based separations116,117 as well as the generation of cell immunoisolation devices.118,119
Fig. 10.
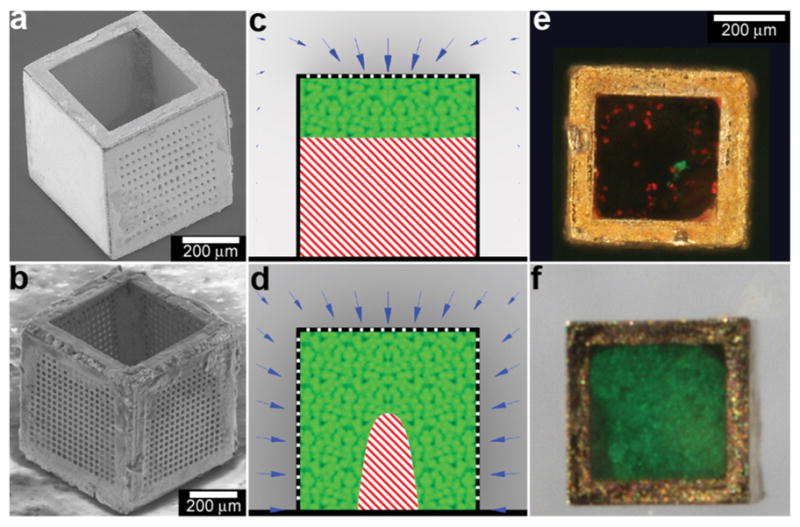
Three dimensional nutrient patterning inside polyhedral micro-containers. (a, b) Electron microscopy images of a self-assembled one porous-faced (a) and five porous-faced (b) microcontainers. The open face at the top of the containers is used for cell loading and it is sealed during the experiment. (Reprinted from ref. 115 with permission from The Royal Society of Chemistry). (c, d) Numerical simulations of spatial variation of viable (green) and necrotic (red) cells within a micro-container with (c) one porous face and (d) a microcontainer with porosity on all faces except the one at the bottom (similar to the containers shown in (a, b)). The O2 concentration outside the microcontainers is color coded with darker gray colors indicating lower O2 concentrations. The arrows represent the diffusive flux of O2 in the medium surrounding the microcontainer. (Reprinted from ref. 115 with permission from The Royal Society of Chemistry). (e, f) Representative images of 500 μm sized microcontainers with one porous face (e) and five porous faces (f) removed from the cell culture medium and opened for inspection after 7 days. Cells were stained using the live/dead (green/red) assay. Microcontainer with one porous face showed significant numbers of dead cells (e) while those with five porous faces (f) showed high cell viability. (Reprinted from ref. 115 with permission from The Royal Society of Chemistry).
3.5. Matrices with immobilized chemicals
Laser-based methods can be used to immobilize chemicals within hydrogels or polymer matrices. The key idea relies on activation of specific chemical reactions with a laser beam. By restricting the beam’s location to specific spatial regions, it is possible to restrict these reactions to only occur within these regions. For example, Luo et al. used a focused laser to immobilize the adhesive fibronectin peptide fragment, glycine–arginine–glycine–aspartic acid–serine (GRGDS), in selected volumes of agarose hydrogel matrices (Fig. 11a–c). They then verified the influence of GRGDS oligopeptide-modified channels on the 3D cell migration and neurite outgrowth in vitro. Their method for immobilizing biomolecules in 3D matrices could be applied to any optically clear hydrogel, offering a methodology to construct scaffolds with programmed spatial features for tissue engineering applications.120,121
Fig. 11.
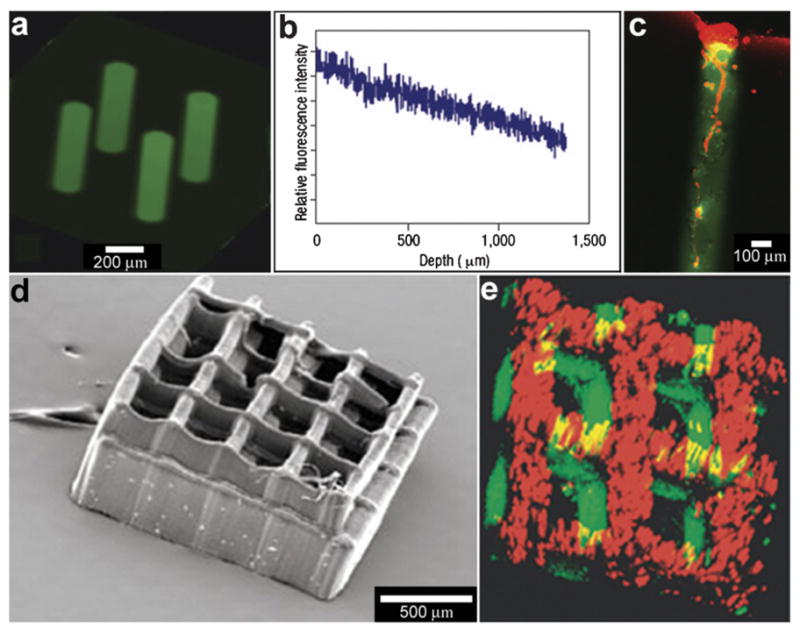
Laser-assisted immobilization of chemicals within polymer matrices. (a) Biochemical channels synthesized in agarose hydrogels and characterized with a fluorescein-tagged GRGDS peptide. (Reprinted with permission from ref. 120. Copyright (2004) by The Nature Publishing Group). (b) The longitudinal fluorescence intensity profile along the central axis of the channel shows a decrease in fluorescent intensity with depth, indicating a concentration gradient of oligopeptide. (Reprinted with permission from ref. 120. Copyright (2004) by The Nature Publishing Group). (c) Primary rat dorsal root ganglia cells were plated on 3D patterned GRGDS oligopeptide-modified agarose gels. Three days after plating, DRG cells grew within GRGDS-oligopeptide-modified agarose channels only, and not in surrounding volumes. A cell cluster on top of a GRGDS channel shows cell migration into the channel and extension of a process into the oligopeptide-modified channel as viewed by confocal fluorescent microscopy, where the channel is green (due to a fluorescein-labeled oligopeptide) and the cells are red (due to the cytoskeletal F-actin rhodamine-phalloidin stain). (Reprinted with permission from ref. 120. Copyright (2004) by The Nature Publishing Group). (d, e) Laser-fabricated 3D polymer structure (d) with two chemicals incorporated into it (e). (Reprinted with permission from ref. 122. Copyright (2005) by John Wiley and Sons).
Layer-by-layer stereolithography can also precisely pattern ligands and growth factors within a polymeric scaffold. Mapili et al. used photocrosslinkable poly(ethylene glycol) dimethacrylate as the basic structural component of these microfabricated scaffolds.122 Fabrication was accomplished by placing the solutions for polymerization on a micromanipulator stage and then slowly (at a scanning speed of approximately 50 μm s−1) translating the stage in the direction perpendicular to the laser beam. Chemicals and polymer microparticles could be incorporated into the polymer structure during polymerization and the process could be repeated allowing multiple chemicals to be incorporated into the scaffold. The method allows for fabrication of complex geometric architectures with precisely incorporated distributions of single or multiple chemical factors (Fig. 11d, e).122
Alternatively, multiphoton excitation can be used to activate chemical reactions in specific spatial locations. In this case, the reaction requires multiple photons to proceed,123,124 opening up the possibility of defining sub-micron resolved spatial regions at the intersection of two laser beams. Multiphoton excitation has been used to photocrosslink protein microstructures within three-dimensional, optically transparent hydrogel materials, such as those based on hyaluronic acid.125–128 Submicron resolution allowed the creation of structures of relevance to in vivo studies. In particular, 3D structures could be fabricated within cellular microenvironments. The experiments showed that three-dimensional chemical patterning in combination with topographic cues was sufficient to guide dorsal root ganglion cells (DRGs) and hippocampal neural progenitor cells (NPCs) along a specified 3D path.125
In summary, the techniques of laser fabrication are versatile and can aid in understanding cell behavior inside complex 3D microenvironments with controlled spatiotemporal patterning of physical and biochemical factors. The primary areas of interest are studies of basic biology of organ development and tissue functions, in particular studies of embryonic development.120 A particular advantage of these fabrication techniques is the ability to incorporate multiple chemicals while at the same time permitting control over topographic features of the fabricated structures. Both features are important in simulating spatially heterogeneous natural microenvironments as is required in tissue and organ regeneration (e.g., bone, blood vessels) that require the controlled delivery of multiple growth factors.129
4. Chemical patterning using stimuli-responsive particles
Particles with stimuli-responsive behaviors enable on-demand and environmental control over chemistry. A variety of physical (e.g. optical) and chemical (e.g. antigen) stimuli responsive particles are available.
4.1. Optical, radio-frequency (RF) and thermal stimulation
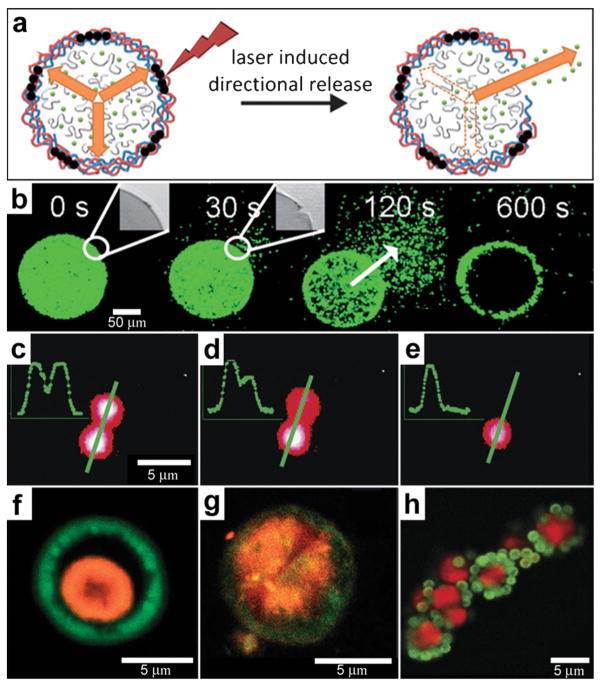
In addition to the fabrication of 3D structures, spatial localization of a laser beam can also be used to induce chemical release. For example, lasers can be used to rupture the wall of spherically-symmetric capsules in order to release chemicals in an asymmetric manner (Fig. 12a). Bedard and coworkers utilized polyelectrolyte capsules for this purpose.130 To make their 100 μm sized capsules respond to near-infrared light, the polyelectrolyte membranes were functionalized with gold nano-particles. The incident laser light leads to plasmon excitation and a local temperature increase which is localized to the nanoparticles and consequently results in localized damage to the polyelectrolyte membrane creating an opening in the capsule. Once the pore has been created, osmotic pressure drives chemical release from the capsule. Fluorescent microscopy snapshots (Fig. 12b) show site-specific opening of a giant polyelectrolyte capsule by near-infrared laser activation. This experimental approach allows release of the chemical in a spatial direction specified by the laser. These devices operate in a manner similar to drug delivery systems where the drug is pumped by osmotic pressure through a laser-drilled hole.77,131
Fig. 12.
Chemical patterning with optically responsive particles. (a) Schematic representation of the laser-induced opening of a capsule at a desired area and the release of encapsulated material in a pre-selected direction. The degradation products of the dex-HEMA/DMAEMA hydrogel onto the polyelectrolyte membrane exert an osmotic pressure against the capsule wall (orange arrows). The large size of the capsules and the presence of IR-sensitive gold nanoparticles (black dots) make it easy to open the shells at a desired site using an IR laser. Once an incision is made, the content of the capsules is released. (Reprinted from ref. 130 with permission from The Royal Society of Chemistry). (b) Fluorescence microscopy snapshots of the site-specific opening of a giant polyelectrolyte capsule by IR laser activation. The inset shows the pore in the polyelectrolyte shell. The arrow indicates the direction of release as osmotic pressure drives encapsulated material out of the capsule. (Reprinted from ref. 130 with permission from The Royal Society of Chemistry). (c–e) A polymeric microcapsule shell acts as a reversible nanomembrane. Upon laser light illumination one of the microcapsules (top) partially releases encapsulated polymers (c) and reseals (d). After the second illumination the microcapsule completely releases its content (e). Profiles in the left upper corner are drawn along the green line. (Reprinted with permission from ref. 132. Copyright (2008) by The American Chemical Society). (f, g) CLSM images taken before (f) and after (g) laser-illumination of the inner shell (orange) doped with gold particles in the course of laser stimulated mixing of both compartments (outer compartment shown in green). (Reprinted with permission from ref. 134. Copyright (2007) by John Wiley and Sons). (h) Fluorescence CLSM images of pericentric multicompartment structures based on the CaCO3 inner core (red) and PS nanoparticles in the outer (green). (Reprinted with permission from ref. 133. Copyright (2010) by John Wiley and Sons).
The technique of laser excitation is not limited to directional release from single capsules. Several capsules can be positioned right next to each other (Fig. 12c–e) and it is possible to selectively release chemicals from a capsule while keeping the neighboring capsules intact.132 Further, the laser damage is reversible and experiments indicate that no content is lost in the absence of laser irradiation even in the capsules which were previously broken by the laser (Fig. 12c–e). The complexity of multi-capsule structures can be further increased by creating multicompartment capsules in a variety of geometries, such as concentric, pericentric, innercentric and acentric.133 The use of these complex capsules in enabling spatially controlled chemistry has been demonstrated by the incorporation of IR-light absorbing gold nanoparticles into the inner shell or a two-layer capsule. The incorporation allowed selective rupture of only the inner membrane while preserving an intact outer membrane. Rupture of the inner shell on illumination of the capsules with near-infrared laser light caused intermixing of the two shells (Fig. 12f, g) providing a route to conduct chemical reactions in confined volumes.134 This strategy could be extended to more complex shaped pericentric capsules, which are multicompartment capsules where smaller capsules are adsorbed onto larger inner capsules (Fig. 12h).133 In fabricating such capsules, Delcea et al. have described how the ratio of the capsule numbers in the assembled capsule structure could be varied by adjusting the concentration of small and large particles prior to electrostatic self-assembly. The highlight of such microcompartment capsules is that they can be synthesized with a range of materials and make it possible to release different chemicals.
Light activation can also be used to perform photolytic conversion of molecules essentially allowing for instantaneous “creation” of molecules of a certain type. Specifically, light can be used to break chemical bonds in “caged” compounds.135 Upon illumination, established spatial gradients of a molecule and its container molecule (“the cage”) can be converted into spatial gradients of the caged compound alone. Examples of applications of such photosensitive caged compounds include studies of spatial patterns of Ca2+ ions in Xenopus oocytes,136 generation of local increases in Ca2+ concentrations inside certain cell regions137 and the release of proteins to study their interactions inside E. coli bacteria.138 Temperature changes can also be used to control the positions and to conduct chemical reactions at the level of individual molecules. For example, Bolinger et al. described a system that consists of small unilamellar vesicles (SUV) trapped inside a larger lipid-based nanoreactor (Fig. 13a).139,140 By using different lipids with different ordered-fluid phase transition temperatures, they showed that it was possible to precisely control the release of chemicals individually from each of the SUVs allowing for single-molecule control.
Fig. 13.
Chemical patterning with temperature and RF responsive particles. (a) Two consecutive reactions in a single large unilamellar vesicle (LUV) nanoreactor. The nanoreactor was loaded with two kinds of small unilamellar vesicles (SUVs), the first kind with a phase transition temperature Tt = 23 °C and encapsulating dichlorodimethylacridinone (DDAO) phosphate (dark red), the other with Tt = 41 °C and containing fluorescein diphosphate (FDP, dark green). (Reprinted with permission from ref. 139. Copyright (2008) by John Wiley and Sons). (b–d) Schematic illustration (b) and TEM images (c, d) of the controlled-release Au-nanocages-based system. (b) On exposure to a near-infrared laser, the light is absorbed by the nanocage and converted into heat, triggering the smart polymer to collapse and thus release the pre-loaded effector. When the laser is turned off, the polymer chains will relax back to the extended conformation and terminate the release. (Reprinted with permission from ref. 141. Copyright (2009) by The Nature Publishing Group). (c) TEM images of Au nanocages for which the surface was covered by a pNIPAAm-co-pAAm copolymer with the lower critical solution temperature at 39 °C. The inset shows a magnified TEM image of the corner of such a nanocage. (Reprinted with permission from ref. 141. Copyright (2009) by The Nature Publishing Group). (d) TEM of multiple-walled Au/Ag nanoshells. (Reprinted with permission from ref. 143. Copyright (2004) by The American Chemical Society). (e–g) Optical images showing the remote controlled, spatially localized microfabrication within a capillary. Two microwires (1 and 2) were embedded within a microfabricated capillary (ca. 1 mm in diameter and 1.5 cm in length) and the capillary was aligned on top of a 2D microcoil. The microcoil was used to remotely increase the temperature of the container. Separate containers filled with pluronic and soaked with the chemical sensitizer and activator were guided into the capillary to the site of the gap within wire 1 using a magnetic stylus (e, f). The capillary was then flushed with a commercial electroless copper-plating solution; chemical reduction (bubbles of the hydrogen gas, a byproduct in the reaction, can be seen) of copper sulfate to metallic copper, occurred at the gap within microwire 1 (g). As a result copper was deposited only in the gap within wire 1 (not shown in the figure). (Reprinted with permission from ref. 145. Copyright (2007) by John Wiley and Sons).
While the method of creating spatial gradients via laser irradiation of caged compounds has been effectively used to study biological systems, their synthesis can be challenging. Further, the intensity and wavelength of light utilized may also be detrimental to the surrounding medium, especially for biological samples. To address these challenges, Yavuz et al. developed a platform where chemicals are encapsulated inside smart polymer-covered gold nanocages.141 Gold nanocages are nanometer scale hollow structures with porous walls (Fig. 13b–d) capable of encapsulating numerous chemicals in their hollow interior. The nanocages have strong absorption in the near-infrared region, a feature that was utilized to locally heat poly(N-isopropylacrylamide) pNIPAAm and a pNIPAAm-co-pAAm copolymer above their low critical solution temperature (LCST) of 32 and 39 °C respectively. This heating lead to reversible changes in the polymer permeability and resulted in chemical release from the nanocage.141–143
While thermal changes can be achieved using laser irradiation, they can also be effected using inductively coupled radio frequency (RF) fields. Hamad-Schifferli et al. have even claimed to control DNA hybridization by localized heating using RF fields. Here, RF fields were coupled to a metal nanocrystal covalently linked to DNA.144 RF fields can also be used to heat larger metallic containers for on-demand chemistry. For example, Ye et al. have shown that by combining external magnetic guiding with such RF heating, it is possible to release chemicals locally and accomplish local catalyst deposition and consequently spatially controlled electroless deposition within thin capillaries.145 In this case, two self-assembled porous microcontainers both filled with a pluronic gel with one containing a sensitizer and the other an activator were guided sequentially using a magnetic stylus into a glass capillary (Fig. 13e). The containers were sequentially localized to a gap between two microwires (Fig. 13f) and the sensitizer and activator were released by heating the gel using an RF source. Subsequent flushing of the capillary with copper-plating solution (Fig. 13g) allowed for electroless copper deposition in the gap. RF heating is attractive since it can be made frequency selective, and it can enable localized heating. Additionally, RF fields can be transmitted through a variety of materials with relatively low loss.
4.2. Chemical stimulation
Most biodegradable polymer particles rely on chemical degradation at a specific pH or in the presence of specific enzymes to release chemicals over extended periods of times. However, a number of chemical release systems have also been developed where release is induced in the presence of chemical stimuli that effect a rapid structural change. For example, stimuli responsive hydrogels have been shown to be responsive to pH,146,147 ionic strength,148 saccharides149 and even antigens.150 Since the response of these materials is often accompanied by a large change in volume, capsules or particles composed of these materials can be triggered to release chemicals in the presence of these chemical stimuli. More recently, Douglas et al. created 35 nm sized DNA-based capsules which are capable of recognizing specific targets on cell surfaces (Fig. 14).151 Their capsule was synthesized using DNA origami techniques,152,153 and consists of two parts, each of which is loaded with antibodies. The parts are held closed by a DNA-aptamer-based lock. Binding of the lock to its leukemic cell marker antigen triggers opening of the capsule making it possible to bring the antibodies loaded inside the capsule to the cell surface which can induce cell death.
Fig. 14.
Antigen-responsive capsules. (a) Design of an aptamer-gated DNA nanorobot. The device transitions from its closed state to open when aptamer-based locks are displaced by binding to an antigen key. Payloads such as gold nanoparticles and antibody fragments (shown in pink) can be loaded. (b, c) TEM images of robots loaded with 5-nm gold nanoparticles in closed and open conformations. Scale bars, 20 nm. (Reprinted with permission from ref. 151. Copyright (2012) by The American Association for the Advancement of Science).
5. Chemical patterning using arrays of chemical sources
In order to heterogeneously pattern chemicals over large distances and to study the interplay between chemicals released from different sources, it is necessary to create arrays of chemical releasing particles. Here, the spacing between the particles and consequently the density can have important consequences for chemical reactions. Tinsley et al. confined the ferroin catalysts of the BZ reaction to microporous cation-exchange beads and submerged large numbers of these beads in a catalyst-free medium (Fig. 15a).45,154 They observed different oscillatory behaviors depending on the stirring rate and the density of the particles within the medium. The authors argue that their observations may serve as a model of quorum sensing where microorganisms switch from individual to collective behavior155 as their concentration changes.
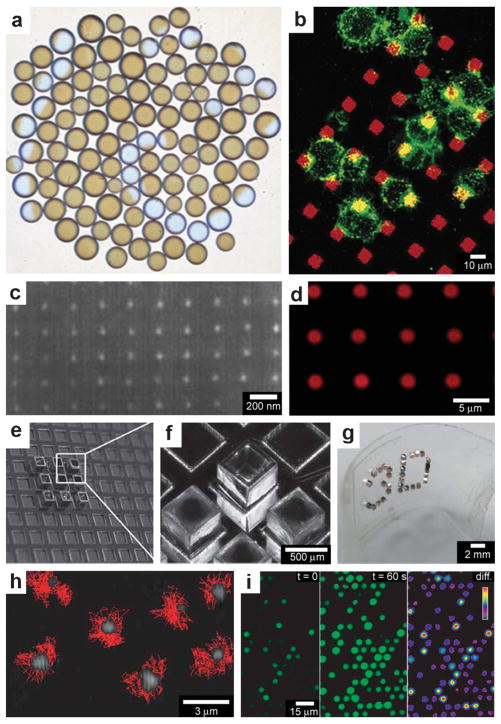
Fig. 15.
Chemistry using arrays of chemical sources. (a) An array of 106 catalyst-loaded excitable particles (brown spheres) with a spiral Belousov–Zhabotinsky wave behavior (blue spiral). (Reprinted with permission from ref. 45. Copyright (2009) by The American Physical Society). (b) Confocal images of RBL cells (green) on a patterned supported liquid bilayer membrane (red) showing interaction between RBL cells and the patterned lipid bilayer. (Reprinted with permission from ref. 166. Copyright (2003) by The American Chemical Society). (c) SEM micrograph of a platinum nanocluster array fabricated on an oxidized silicon surface by electron beam lithography. (Reprinted with permission from ref. 161. Copyright (1998) by The American Chemical Society). (d) CLSM imaging in situ of resorufin formation in an array of polyelectrolyte multilayer microcapsules. Fluorescence occurs in the interior of the patterned microcapsules (capsules shells did not contain fluorescently labeled polymer). (Reprinted from ref. 174 with permission from The Royal Society of Chemistry). (e, f) Optical images of a 65 μm thick SU-8 holder with recessed slots and a 3 × 3 array of self-assembled microcontainers positioned in it. (Reprinted from ref. 115 with permission from The Royal Society of Chemistry). (g) Optical image of an ordered 3D microcontainer array on a curved surface. (Reprinted from ref. 115 with permission from The Royal Society of Chemistry). (g) Trajectories recorded over 2 min time interval of single λ-DNA chains undergoing Brownian motion inside micrometer-sized chambers fabricated in silicone. (Reprinted with permission from ref. 175. Copyright (2005) by The Nature Publishing Group). (h) Fluorescent images of enzymatic activity in micrometer-sized chambers. (Reprinted with permission from ref. 175. Copyright (2005) by The Nature Publishing Group).
In addition to doing chemistry with randomly positioned chemical sources, precise spacing of chemical reagents allows further control over chemical reactions since the size of each source as well as its spacing can be precisely controlled. There are a multitude of methods to achieve this control by forming arrays in 2D including top-down micro and nanofabrication, micro-contact printing and dippen lithography; these methods have been utilized to spatially control chemistry on 2D substrates.156–161 The ability to create these arrays of chemicals can be used to investigate fundamental aspects of cell behavior such as the influence of cell shape and spreading on cell growth, protein secretion and apoptosis,162–164 chemotactic responses of mammalian carcinoma cells,165 and morphological changes and receptor clustering on patterned lipid rafts with embedded antigens166 (Fig. 15b). Furthermore, lithographic technologies such as electron beam writing can be used to create arrays of catalytic particles (Fig. 15c). Such arrays have been used to study reactivity of metal catalysts when the particle size is reduced to nanometer-scale dimensions.161
In addition to biological applications, arrays of catalytic particles allow multiple catalysts to be combined at the same location, making it possible to perform chemical reactions with multiple steps. Such multi-step chemical reactions are particularly advantageous if the intermediate products of the reaction are unstable. By positioning multiple catalysts at the same site, the intermediate products can react locally without having to diffuse very far.167,168 Such closely coupled individual reactions often yield a product that is difficult to obtain by a single process169,170 and these catalytic cascades mimic tandem reactions in biosynthesis.171–173
Studies that require patterned arrays of chemicals may be further advanced by developing methods to create well-ordered and well-characterized arrays of capsules. Antipina et al., have developed a technique for making arrays of highly ordered patterns of polyelectrolyte multilayer microcapsules.174 The authors used imprint molding based on embossing to pattern a substrate composed of an inert ethylene and tetrafluoroethylene copolymer with micrometer-sized cavities. Once the cavities were fabricated, a drop of a colloidal dispersion containing core-shell microparticles covered with polyelectrolyte membranes was deposited on the surface and allowed to dry. In the process of drying, the particles were driven into the cavities where they self-assembled due to a combination of gravitational and capillary forces. By selectively dissolving the core of the particles with hydrochloric acid, without disturbing their spatial positions, it was possible to transform the array of microparticles into an array of hollow polyelectrolyte microcapsules (Fig. 15d). The authors then loaded their microcapsules with enzymes such as glucose oxidase (GOD) and horseradish peroxidase (horseradish POD, HRP) by altering the capsule membrane permeability with pH. They demonstrated site specific and enzymatic chemical reactions using these arrays of capsules. Similarly Randall et al., have shown how arrays of polyhedral self-assembled microcontainers can be formed by positioning cubic containers in lithographically patterned SU-8 crates.115,117 These crates feature indents slightly larger than the container size (600 μm openings for 500 μm sized cubes, Fig. 15e, f). By using adhesive substrates it was also possible to transfer the arrays of the containers to the flexible films (Fig. 15g). Such arrays of flexible substrates can be deformed into a variety of 3D geometries and thus used to mimic biologically relevant structures.
Arrays of femtoliter-sized cavities can aid in studies of single biological molecules and reveal trends and phenomena not observed when experiments are carried in the bulk. For example, Rondelez et al. developed a silicone device containing a large array of micrometer-sized cavities and used the device to measure enzymatic activities of single molecules.175 They found that the PDMS chambers were capable of confining single molecules, such as λ-DNA (Fig. 15h), as well as individual beads and quantum dots. The enzyme β-galactosidase was then dispensed at low concentrations and enclosed inside chambers containing fluorescein-di-β-D-galactopyranoside. This led to a hydrolysis reaction with fluorescein being one of the end products. They observed that differences in their fluorescent light intensities were quantized (Fig. 15i) indicating that it was possible to localize a very small number (0 to 3) of enzymes inside each individual chamber.
6. Chemical patterning using microfluidics
Microfluidics provides a set of techniques to manipulate fluid flow at very small scales. These techniques can be utilized to create static and dynamic droplets and streams for carrying out chemical reactions with spatial and often real-time control.
6.1. Droplet microfluidics
Droplet microfluidics is a set of techniques to generate and manipulate large numbers of droplets.176–182 Multiple identical droplets can be generated simply by controllably flowing immiscible liquids in different geometries, such as a T-junction (Fig. 16a);183–185 minimization of interfacial energy leads to the spontaneous formation of multiple identical droplets. The shape is not restricted to a sphere and alternate shapes such as discs, rods and ellipsoids, with one or multiple compartments, have been realized (Fig. 16b–d).177,186–190 Chemical localization is accomplished when liquids with similar hydrophobicity or hydrophilicity intermix. Droplet microfluidic methods can thus be used to synthesize a multitude of particles for use in the applications described in the preceding sections of this paper. In addition, the ability to generate and manipulate multiple droplets in real time provides further capabilities; multiple droplets can be merged and their contents mixed together in a controllable manner (Fig. 16e).191 For example, Tresset et al. have accomplished controlled electrofusion of liposomes using a microfluidic device they designed.192 They have also shown that their device can be used to deliver artificial microstructures into living cells when these microstructures are encapsulated in liposomes. Controllable mixing of the contents of the droplets can also be achieved by their contact with the microchannel walls.193 This controlled mixing facilitates tunability over the starting point and duration of chemical reactions; additionally, small volumes are utilized.193 Indeed, Song et al.194 have carried out chemical reactions inside droplets with millisecond time resolution (Fig. 16f). Furthermore, droplet microfluidic systems have been used for encapsulation (Fig. 16g, h) of both prokaryotic and eukaryotic cells.178,195 It has also been shown that cells confined to droplets behave in a manner similar to multiple coexisting cells (Fig. 16g).196 Future advances in this area may help further elucidate the complex reaction networks of relevance to synthetic biology.
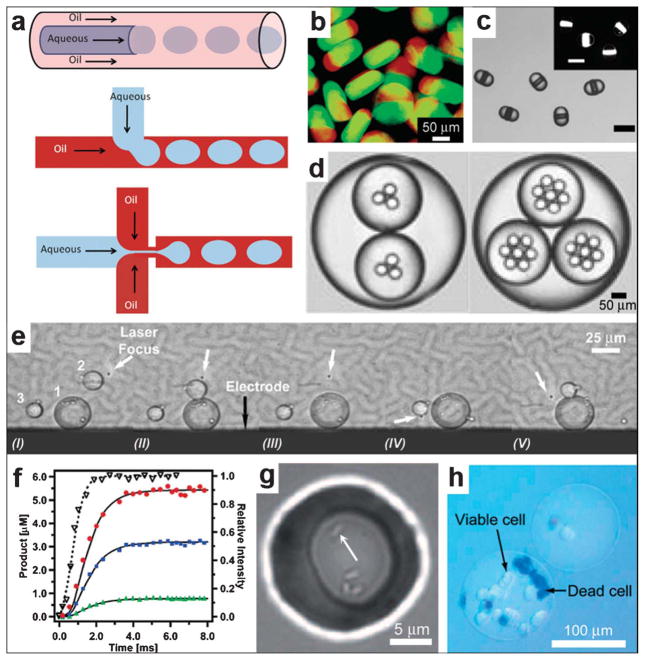
Fig. 16.
Spatially controlled chemistry with droplet microfluidics. (a) A schematic of three different approaches, namely co-flow, T-junction and flow focusing, conventionally used for droplet generation. (Reprinted from ref. 183 with permission from The Royal Society of Chemistry). (b–d) Types of droplets/particles created in microfluidic devices. The devices are capable of creating disc-shaped Janus droplets which can be polymerized with UV into Janus particles, ternary droplets as well as emulsions with a hierarchy of droplets confined within one another (i.e. multiple emulsions). (Reprinted with permission from ref. 187, 190, 188. Copyright (2006), (2007) by The American Chemical Society and (2006) by John Wiley and Sons). (e) Sequential fusion of droplets in a microfluidic device. The laser (at the position indicated with the white arrow) is used to position the droplets while the electrodes (the black line at the bottom of the figure shows one of the electrodes, the top electrode is not shown) are used to induce coalescence of the droplets. The Roman numerals below the figure indicate different stages of the fusion process. (Reprinted from ref. 191 with permission from the Royal Society of Chemistry). (f) Chemical reaction product formation as a function of time for a reaction proceeding inside droplets as observed in a microfluidic device. The experimental setup allows for millisecond resolution of the reaction progress. (Reprinted with permission from ref. 194. Copyright (2003) by The American Chemical Society). (g) Small group of bacteria confined to a picoliter droplet. The white arrow points to one of the bacteria. (Reprinted with permission from ref. 196. Copyright (2009) by John Wiley and Sons). (h) An alginate bead encapsulating mammalian cells. The cells have been stained with trypan blue to differentiate the living cells from the dead ones. (Reprinted with permission from ref. 195. Copyright (2007) by John Wiley and Sons).
6.2. Two dimensional microfluidic networks
Microfluidic devices that rely on both diffusion and advection have evolved into a highly versatile technique for controlled chemical gradient creation. Laminar flows of fluids are of particular interest since they permit creation of stable chemical patterns. In this section, we focus on devices which allow creation of chemical gradients on two-dimensional surfaces197 or in general where the presence of the third dimension is not important for the phenomena studied. Several excellent reviews have been written describing the design of microdevices198 and microfluidic networks199,200 as well as their use for chemical gradient creation.201–203 Here, we highlight some recently developed microfluidic devices mainly to compare and contrast their capabilities with those of particles.
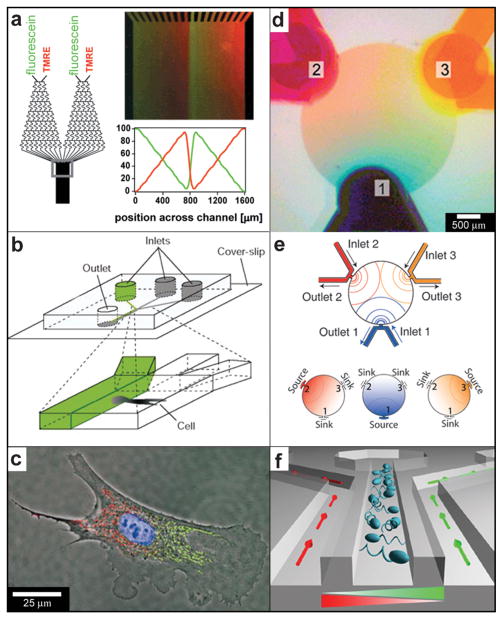
A well-known device for 2D gradient generation was described by Dertinger et al. (Fig. 17a) and relies on merging together and flowing multiple streams of fluid side-by-side.204 Each stream typically carries a different chemical or the same chemical at a different concentration. The downstream concentration of the chemical is determined by the network of channels located upstream and can be altered by varying the network geometry. Merging of multiple streams results in concentration gradients at the point at which they merge and these step gradients are subsequently smoothed out by diffusion (see Fig. 1f). Dertinger et al. have shown that their device is capable of generating a wide range of chemical gradients, including linear and periodic chemical gradients of both one and multiple chemicals (Fig. 17a). By relying on the slowness of diffusion or by increasing the speed of the flow, it is possible to decrease the width of the transition regions at any given position along the channel. Takayama et al. utilized this feature to achieve “subcellular positioning of small molecules”, i.e. they were able to expose a single cell to two different chemical concentrations (Fig. 17b).205 In order to position the interface region over the cell, they varied the relative amounts of fluid in each of the two merging streams. The technique allowed localized labeling of mitochondria in two spatially separated regions of a single cell (shown with red and green in Fig. 17c). Such labeling has been used to study mitochondrial movement and to deliver small molecules to selected regions inside single cells.205,206
Fig. 17.
Chemical patterning with 2D microfluidic devices. (a) A schematic of the microfluidic network and fluorescence micrographs of a periodic overlapping sawtooth gradient of fluorescein and TMRE in ethanol. The plot below the micrograph shows the numerically calculated fluorescence intensity profile across the broad channel. (Reprinted with permission from ref. 204. Copyright (2001) by The American Chemical Society). (b) Experimental set-up for differential manipulation of regions of a single bovine capillary endothelial cell using multiple laminar flows. Lower panel shows a close-up of the point at which the inlet channels combine into one main channel. (Reprinted with permission from ref. 205. Copyright (200) by The Nature Publishing Group). (c) Fluorescence images of a single cell after treatment of its right pole with Mitotracker Green FM and its left pole with Mitotracker Red CM-H2XRos. The entire cell is treated with the DNA-binding dye Hoechst 33342. (Reprinted with permission from ref. 205. Copyright (2001) by The Nature Publishing Group). (d, e) Overlapping diffusive gradients. Color dyes were introduced through different access ports and their concentration was maintained with a constant flow rate. For each dye an independent gradient formed with 120° angular displacement. Overlapping the three gradients results into a blend of dye concentrations where each spatial location has different combinations of dye concentrations. Scale bar 500 μm. (Reprinted from ref. 209 with permission from The Royal Society of Chemistry). (f) A microfluidic chemotaxis device design: details of a three channel unit. Motile cells are injected into the center channel. Dual chemical gradients (schematically illustrated with gradient-colored triangles) are generated in the center channel by pumping media containing different concentrations of chemoattractants through the two side channels. (Relevant reference: 210).
Microfluidic devices that rely on merging of multiple streams of fluid, as described above, represent a major class of 2D chemical pattern generators. The other major type of micro-fluidic gradient generators relies on diffusion of chemicals through pores at the end or in the side walls of fluidic channels. As compared to particle sources, the notable advantage is that the chemicals can be replenished or varied during chemical reactions. Since, multiple non-interacting chemicals can often diffuse through the same region, these microfluidic gradient generators can be used to create gradients with multiple chemicals within the same spatial region.207,208 For example, Atencia et al. describe a microfluidic device they called the “microfluidic palette”, which is capable of simultaneously generating multiple spatial chemical gradients inside a millimeter-sized chamber (Fig. 17d). In their case, three channels have openings into this chamber and chemicals flowing through each of the three channels diffuse into the chamber through the openings resulting in a complex three-component gradient (Fig. 17e). They demonstrate a variety of overlapping gradients and a dynamic diffusive gradient that rotates around its center.209 Such gradients are useful to study the behavior of multicellular systems in synthetic environments that are intended to simulate their natural habitats. Kalinin et al. exposed motile bacterial cells to two spatially opposing, but equally potent chemoattractant gradients (Fig. 17f) and observed that chemotactic decisions of E. coli cells depended on expression levels of Tar and Tsr methyl-accepting chemotaxis protein receptors.210 In addition to creating gradients of soluble factors for chemotaxis studies, the 2D microfluidic devices have been used to create gradients of bound chemicals,197 for example, for studies of cell migration, such as neurons and neutrophils,12,211–213 on gradients of substrate-bound chemicals as well as for chemical reactions.214
6.3. Three dimensional microfluidic networks
The design and fabrication of 3D microfluidic networks has been largely motivated by the need to replicate vascular channels and capillaries in vivo,215–217 but could also be used to engineer chemical reactions with spatial control in all three dimensions. A number of recent methods have been developed to create three-dimensional fluidic networks, including direct write methods218 the use of electric discharge (Fig. 18a),219 layered extensions of planar microfluidic devices220 and threading PDMS channels221 (Fig. 18b). These networks have only recently begun to be used to release chemicals and create patterns or gradients with spatial control in 3D media. Most of the studies have utilized inherently linear gradients but these have been established to a finite thickness in 3D media.222–228 Notably, Choi et al. have designed channels within an agarose gel and utilized them to established gradients using a variety of small molecules such as fluorescein (Fig. 18c, d).229,230 Similarly, both linear and non-linear gradients across two dimensional surfaces embedded within 3D gels have been developed by Mosadegh et al. (Fig. 18e).231 Recently, Jamal et al. have shown that polymers photopatterned with differential stress can enable curved and flexible micro-fluidic devices (Fig. 18f).232 These SU-8/PDMS devices can be utilized to dispense chemicals through channels and pores in a variety of cylindrical and undulatory geometries and suggest a means to take two dimensional microfluidic chemical release into curved and 3D geometries, with unprecedented spatial control.
Fig. 18.
Chemical patterning with 3D microfluidic devices. (a) Branched microvascular network embedded in PLA substrates which incorporates a hierarchy of microchannel diameters. An aqueous solution of blue food dye was injected into the interconnected microchannel array for visualization purposes. (Reprinted with permission from ref. 219. Copyright (2009) by John Wiley and Sons). (b) A microfluidic network having the geometry of a basketweave. The channels were filled with an aqueous solution of fluorescein (green) or Cascade Blue (blue) and illuminated with UV light. (Reprinted with permission from ref. 221. Copyright (2003) by The American Chemical Society). (c, d) Cross-sectional views of cell-seeded microfluidic scaffolds. Dispersed cells are shown as double circles. Microchannels are shown as squares. The pink shading represents steady-state 3D distributions of solutes: in (c), reactive solute is delivered via the channels and is consumed by cells as it diffuses into the matrix; in (d), non-reactive solute is delivered via the two channels on the left and extracted by the channels on the right. (Reprinted with permission from ref. 229. Copyright (2007) by The Nature Publishing Group). (e) The gradient-generating region of a microfluidic device features a tapered microchamber to produce a nonlinear gradient. By changing the shape of the gradient-generating region it is possible to change the shape of the chemical gradient. (Reprinted with permission from ref. 231. Copyright (2007) by The American Chemical Society). (f) A self-assembling microfluidic device with PDMS inlets/outlets attached to a Si substrate and with PDMS channels integrated with a differentially crosslinked SU-8 film. Fluorescence images showing the flow of fluorescein (green)/rhodamine B (red) through a dual channel device. (Reprinted with permision from ref. 232. Copyright (2011) by The Nature Publishing Group).
7. Concluding remarks
Recent advances in colloidal science, self-assembly, microfluidics and nanofabrication have enabled a range of discrete particles and fluidic devices that can be used to release chemicals with unprecedented spatial control. These advances suggest that it is now possible to engineer chemical reactions with spatial control in both two and three dimensions. The methodologies for enabling spatial control are summarized in Table 1. It can be seen that for the most part particle-based technologies are capable of operating independently of external equipment while microfluidic devices have advantages in terms of precision and real-time external control. Accordingly, microfluidic devices can be used to elucidate cell motion and responses in well-controlled chemical environments while the particle based technologies, due to their ability to function in the absence of any external equipment, can be used in confined and hard to reach spaces, such as in vivo.
Table 1.
Characteristics of particles and streams for chemistry with spatial control
| Class | Typical size utilized | Spatial characteristic | Applications demonstrated |
|---|---|---|---|
| Molecular cages | 1 nm–20 nm | Release by bond rupture | Instantaneous chemical gradients, studies with single molecules |
| Vesicles and liposomes | 100 nm–100 μm | High concentration, release by diffusion or rupture | Chemistry within confined spaces, low numbers of molecules, rapid mixing by diffusion |
| Polyelectrolyte capsules and their assemblies | 10 μm–1 mm | 3D spherically symmetric diffusion, 1D ejection, confinement | Studies of spatially-confined chemical reactions, 1D chemical delivery |
| Polymer microspheres | 1 μm–100 μm | Spherically-symmetric release by diffusion or degradation | Drug delivery, cell taxis and embryonic studies |
| Lithographically-structured containers | 100 nm–1 cm | Symmetric and asymmetric diffusion and/or degradation | Drug delivery and cell encapsulation therapy |
| Self-assembled polyhedra | 100 nm–1 cm | Arbitrary space curve by diffusion through precisely positioned pores | Bacterial chemotaxis, drug delivery, cell encapsulation therapy |
| Matrices with immobilized chemicals | 1 μm–10 cm | 3D bound | Engineering cellular microenvironments for cell and tissue engineering |
| Arrays | 100 nm–1 cm | 2D/3D diffusion, 2D/3D bound | Periodic patterns in reaction-diffusion and biological systems |
| Droplet microfluidics | 1 μm–1 mm | Confinement, 3D diffusion | Chemistry within confined spaces, control over reaction rates, cell encapsulation |
| 2D microfluidic networks | 10 μm–10 cm | 2D flow and diffusion | Cellular chemotaxis, chemistry in laminar flows, substrate patterning |
| 3D microfluidic networks | 100 μm–10 cm | 3D flow and diffusion | 3D gradients in gels |
Spatial control over chemistry with micro- and nanoscale resolution offers many opportunities, such as, (a) the use of small volumes of chemicals; (b) the localization of specific reactions while limiting interference from others; (c) rapid mixing; and (d) high local concentrations of chemicals with even single molecule precision. Additionally, the spatial control of reactions using synthetic chemical releasing particles and streams provides a means to mimic biological control of chemistry within and in between cells. As in biological networks, this spatial control can be used to, (1) conserve chemical reactants by secreting chemicals only at the location where they are needed; (2) enable adaptive behaviors such as sensing and orientation; (3) communicate and convey positional information as is achieved in an embryo in the process of embryonic development; (4) direct movement as occurs in angiogenesis and neural development; and (5) orchestrate collective behavior. The realization of these capabilities in synthetic systems necessitates further development and application of methods to create and control spatial patterns of chemicals, especially in three dimensions.
Supplementary Material
Acknowledgments
The authors would like to thank Svetlana Ratner for providing the illustration of an embryo used in the TOC graphic, Shivendra Pandey for the TOC images of the BZ reaction, Andrew Darling for suggesting the idea of Fig. 17f and Mustapha Jamal for discussions. This work was supported by the NSF EFRI-1022730 and the NIH Director’s New Innovator Award Program, part of the NIH Roadmap for Medical Research, through grant number 1-DP2-OD004346-01. Information about the NIH Roadmap can be found at http://nihroadmap.nih.gov.
Footnotes
Electronic Supplementary Information (ESI) available. See DOI: 10.1039/c2ra20337e
References
- 1.Shapiro L, McAdams HH, Losick R. Science. 2002;298:1942–1946. doi: 10.1126/science.1072163. [DOI] [PubMed] [Google Scholar]
- 2.Shapiro L, McAdams HH, Losick R. Science. 2009;326:1225–1228. doi: 10.1126/science.1175685. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Shapiro L, Losick R. Cell. 2000;100:89–98. doi: 10.1016/s0092-8674(00)81686-4. [DOI] [PubMed] [Google Scholar]
- 4.Goley ED, Iniesta AA, Shapiro L. J Cell Sci. 2007;120:3501–3507. doi: 10.1242/jcs.005967. [DOI] [PubMed] [Google Scholar]
- 5.Rosch J, Caparon M. Science. 2004;304:1513–1515. doi: 10.1126/science.1097404. [DOI] [PubMed] [Google Scholar]
- 6.Scott ME, Dossani ZY, Sandkvist M. Proc Natl Acad Sci U S A. 2001;98:13978–13983. doi: 10.1073/pnas.241411198. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Swartz MA. Curr Opin Biotechnol. 2003;14:547–550. doi: 10.1016/j.copbio.2003.09.003. [DOI] [PubMed] [Google Scholar]
- 8.Shields JD, Fleury ME, Yong C, Tomei AA, Randolph GJ, Swartz MA. Cancer Cell. 2007;11:526–538. doi: 10.1016/j.ccr.2007.04.020. [DOI] [PubMed] [Google Scholar]
- 9.Behar TN, Schaffner AE, Colton CA, Somogyi R, Olah Z, Lehel C, Barker JL. J Neurosci. 1994;14:29–38. doi: 10.1523/JNEUROSCI.14-01-00029.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Gonzalez-Quevedo R, Lee Y, Poss KD, Wilkinson DG. Dev Cell. 2010;18:136–147. doi: 10.1016/j.devcel.2009.11.010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Isbister CM, Mackenzie PJ, To KCW, O’Connor TP. J Neurosci. 2003;23:193–202. doi: 10.1523/JNEUROSCI.23-01-00193.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Baier H, Bonhoeffer F. Science. 1992;255:472–475. doi: 10.1126/science.1734526. [DOI] [PubMed] [Google Scholar]
- 13.Tessier-Lavigne M, Goodman CS. Science. 1996;274:1123–1133. doi: 10.1126/science.274.5290.1123. [DOI] [PubMed] [Google Scholar]
- 14.Goodman CS. Annu Rev Neurosci. 1996;19:341–377. doi: 10.1146/annurev.ne.19.030196.002013. [DOI] [PubMed] [Google Scholar]
- 15.Mueller BK. Annu Rev Neurosci. 1999;22:351–388. doi: 10.1146/annurev.neuro.22.1.351. [DOI] [PubMed] [Google Scholar]
- 16.Song HJ, Poo MM. Curr Opin Neurobiol. 1999;9:355–363. doi: 10.1016/s0959-4388(99)80052-x. [DOI] [PubMed] [Google Scholar]
- 17.Conway EM, Collen D, Carmeliet P. Cardiovasc Res. 2001;49:507–521. doi: 10.1016/s0008-6363(00)00281-9. [DOI] [PubMed] [Google Scholar]
- 18.Noden DM. Am Rev Respir Dis. 1989;140:1097–1103. doi: 10.1164/ajrccm/140.4.1097. [DOI] [PubMed] [Google Scholar]
- 19.Stone J, Dreher Z. J Comp Neurol. 1987;255:35–49. doi: 10.1002/cne.902550104. [DOI] [PubMed] [Google Scholar]
- 20.Gerhardt H, Golding M, Fruttiger M, Ruhrberg C, Lundkvist A, Abramsson A, Jeltsch M, Mitchell C, Alitalo K, Shima D, Betsholtz C. J Cell Biol. 2003;161:1163–1177. doi: 10.1083/jcb.200302047. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Entchev EV, Schwabedissen A, Gonzalez-Gaitan M. Cell. 2000;103:981–991. doi: 10.1016/s0092-8674(00)00200-2. [DOI] [PubMed] [Google Scholar]
- 22.Gurdon JB, Bourillot PY. Nature. 2001;413:797–803. doi: 10.1038/35101500. [DOI] [PubMed] [Google Scholar]
- 23.Teleman AA, Cohen SM. Cell. 2000;103:971–980. doi: 10.1016/s0092-8674(00)00199-9. [DOI] [PubMed] [Google Scholar]
- 24.Dubois L, Lecourtois M, Alexandre C, Hirst E, Vincent JP. Cell. 2001;105:613–624. doi: 10.1016/s0092-8674(01)00375-0. [DOI] [PubMed] [Google Scholar]
- 25.Neumann C, Cohen S. BioEssays. 1997;19:721–729. doi: 10.1002/bies.950190813. [DOI] [PubMed] [Google Scholar]
- 26.Wolpert L. J Theor Biol. 1969;25:1–47. doi: 10.1016/s0022-5193(69)80016-0. [DOI] [PubMed] [Google Scholar]
- 27.Gurdon JB, Standley H, Dyson S, Butler K, Langon T, Ryan K, Stennard F, Shimizu K, Zorn A. Development. 1999;126:5309–5317. doi: 10.1242/dev.126.23.5309. [DOI] [PubMed] [Google Scholar]
- 28.Tickle C, Summerbell D, Wolpert L. Nature. 1975;254:199–202. doi: 10.1038/254199a0. [DOI] [PubMed] [Google Scholar]
- 29.Crick F. Nature. 1970;225:420–422. doi: 10.1038/225420a0. [DOI] [PubMed] [Google Scholar]
- 30.Gregor T, Wieschaus EF, McGregor AP, Bialek W, Tank DW. Cell. 2007;130:141–152. doi: 10.1016/j.cell.2007.05.026. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Gregor T, Tank DW, Wieschaus EF, Bialek W. Cell. 2007;130:153–164. doi: 10.1016/j.cell.2007.05.025. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Houchmandzadeh B, Wieschaus E, Leibler S. Nature. 2002;415:798–802. doi: 10.1038/415798a. [DOI] [PubMed] [Google Scholar]
- 33.Tomchik KJ, Devreotes PN. Science. 1981;212:443–446. doi: 10.1126/science.6259734. [DOI] [PubMed] [Google Scholar]
- 34.Lee KJ, Cox EC, Goldstein RE. Phys Rev Lett. 1996;76:1174–1177. doi: 10.1103/PhysRevLett.76.1174. [DOI] [PubMed] [Google Scholar]
- 35.Basu S, Gerchman Y, Collins CH, Arnold FH, Weiss R. Nature. 2005;434:1130–1134. doi: 10.1038/nature03461. [DOI] [PubMed] [Google Scholar]
- 36.Turing AM. Philos Trans R Soc London, Ser B. 1952;237:37–72. [Google Scholar]
- 37.Murray JD. Sci Am. 1988;258:80–87. [Google Scholar]
- 38.Murray JD. Mathematical Biology. 3. Springer; 2007. [Google Scholar]
- 39.Zaikin AN, Zhabotinsky AM. Nature. 1970;225:535–537. doi: 10.1038/225535b0. [DOI] [PubMed] [Google Scholar]
- 40.Belousovin BP. In: Oscillations and traveling waves in chemical systems. Field RJ, Burger M, editors. Wiley; New York: 1985. [DOI] [PubMed] [Google Scholar]
- 41.Stern KH. Chem Rev. 1954;54:79–99. [Google Scholar]
- 42.Laplante JP, Erneux T. J Phys Chem. 1992;96:4931–4934. [Google Scholar]
- 43.Epstein IR, Golubitsky M. Chaos. 1993;3:1–5. doi: 10.1063/1.165974. [DOI] [PubMed] [Google Scholar]
- 44.Winston D, Arora M, Maselko J, Gaspar V, Showalter K. Nature. 1991;351:132–135. [Google Scholar]
- 45.Tinsley MR, Taylor AF, Huang ZY, Showalter K. Phys Rev Lett. 2009;102:158301. doi: 10.1103/PhysRevLett.102.158301. [DOI] [PubMed] [Google Scholar]
- 46.Taylor AF, Tinsley MR, Wang F, Huang ZY, Showalter K. Science. 2009;323:614–617. doi: 10.1126/science.1166253. [DOI] [PubMed] [Google Scholar]
- 47.Markus M, Hess B. Nature. 1990;347:56–58. [Google Scholar]
- 48.Madore BF, Freedman WL. Science. 1983;222:615–616. doi: 10.1126/science.222.4624.615. [DOI] [PubMed] [Google Scholar]
- 49.Wiesenfeld K, Hadley P. Phys Rev Lett. 1989;62:1335–1338. doi: 10.1103/PhysRevLett.62.1335. [DOI] [PubMed] [Google Scholar]
- 50.Koch AJ, Meinhardt H. Rev Mod Phys. 1994;66:1481–1507. [Google Scholar]
- 51.Garfinkel A, Tintut Y, Petrasek D, Bostrom K, Demer LL. Proc Natl Acad Sci U S A. 2004;101:9247–9250. doi: 10.1073/pnas.0308436101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Danino T, Volfson D, Bhatia SN, Tsimring L, Hasty J. PLoS One. 2011;6:e20182. doi: 10.1371/journal.pone.0020182. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Bansagi T, Vanag VK, Epstein IR. Science. 2011;331:1309–1312. doi: 10.1126/science.1200815. [DOI] [PubMed] [Google Scholar]
- 54.Edelman DB, Keefer EW. Exp Neurol. 2005;192:1–6. doi: 10.1016/j.expneurol.2004.10.005. [DOI] [PubMed] [Google Scholar]
- 55.Pardo L, Wilson WC, Boland T. Langmuir. 2003;19:1462–1466. [Google Scholar]
- 56.Rosoff WJ, McAllister R, Esrick MA, Goodhill GJ, Urbach JS. Biotechnol Bioeng. 2005;91:754–759. doi: 10.1002/bit.20564. [DOI] [PubMed] [Google Scholar]
- 57.Rosoff WJ, Urbach JS, Esrick MA, McAllister RG, Richards LJ, Goodhill GJ. Nat Neurosci. 2004;7:678–682. doi: 10.1038/nn1259. [DOI] [PubMed] [Google Scholar]
- 58.Bachmatiuk A, Boerrnert F, Grobosch M, Schaeffel F, Wolff U, Scott A, Zaka M, Warner JH, Klingeler R, Knupfer M, Buechner B, Ruemmeli MH. ACS Nano. 2009;3:4098–4104. doi: 10.1021/nn9009278. [DOI] [PubMed] [Google Scholar]
- 59.Tresset G, Takeuchi S. Anal Chem. 2005;77:2795–2801. doi: 10.1021/ac048207o. [DOI] [PubMed] [Google Scholar]
- 60.Lin L, Beyer S, Wohland T, Trau D, Lubrich D. Angew Chem, Int Ed. 2010;49:9773–9776. doi: 10.1002/anie.200907321. [DOI] [PubMed] [Google Scholar]
- 61.Gregoriadis G. Trends Biotechnol. 1995;13:527–537. doi: 10.1016/S0167-7799(00)89017-4. [DOI] [PubMed] [Google Scholar]
- 62.van Dongen SFM, Nallani M, Cornelissen J, Nolte RJM, van Hest JCM. Chem–Eur J. 2009;15:1107–1114. doi: 10.1002/chem.200802114. [DOI] [PubMed] [Google Scholar]
- 63.Chiu DT, Wilson CF, Ryttsen F, Stromberg A, Farre C, Karlsson A, Nordholm S, Gaggar A, Modi BP, Moscho A, Garza-Lopez RA, Orwar O, Zare RN. Science. 1999;283:1892–1895. doi: 10.1126/science.283.5409.1892. [DOI] [PubMed] [Google Scholar]
- 64.Kataoka K, Harada A, Nagasaki Y. Adv Drug Delivery Rev. 2001;47:113–131. doi: 10.1016/s0169-409x(00)00124-1. [DOI] [PubMed] [Google Scholar]
- 65.Richmond DL, Schmid EM, Martens S, Stachowiak JC, Liska N, Fletcher DA. Proc Natl Acad Sci U S A. 2011;108:9431–9436. doi: 10.1073/pnas.1016410108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Antonietti M, Forster S. Adv Mater. 2003;15:1323–1333. [Google Scholar]
- 67.Al-Jamal WT, Kostarelos K. Int J Pharm. 2007;331:182–185. doi: 10.1016/j.ijpharm.2006.11.020. [DOI] [PubMed] [Google Scholar]
- 68.Kim S, Turker MS, Chi EY, Sela S, Martin GM. Biochim Biophys Acta, Biomembr. 1983;728:339–348. doi: 10.1016/0005-2736(83)90504-7. [DOI] [PubMed] [Google Scholar]
- 69.Thünemann AF, Kubowicz S, von Berlepsch H, Möhwald H. Langmuir. 2006;22:2506–2510. doi: 10.1021/la0533720. [DOI] [PubMed] [Google Scholar]
- 70.Sengupta S, Eavarone D, Capila I, Zhao GL, Watson N, Kiziltepe T, Sasisekharan R. Nature. 2005;436:568–572. doi: 10.1038/nature03794. [DOI] [PubMed] [Google Scholar]
- 71.Xiao CJ, Qi XR, Maitani Y, Nagai T. J Pharm Sci. 2004;93:1718–1724. doi: 10.1002/jps.20086. [DOI] [PubMed] [Google Scholar]
- 72.Boyer C, Zasadzinski JA. ACS Nano. 2007;1:176–182. doi: 10.1021/nn7002025. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Kisak ET, Coldren B, Evans CA, Boyer C, Zasadzinski JA. Curr Med Chem. 2004;11:199–219. doi: 10.2174/0929867043456197. [DOI] [PubMed] [Google Scholar]
- 74.Stadler B, Price AD, Chandrawati R, Hosta-Rigau L, Zelikin AN, Caruso F. Nanoscale. 2009;1:68–73. doi: 10.1039/b9nr00143c. [DOI] [PubMed] [Google Scholar]
- 75.Caruso F, Trau D, Möhwald H, Renneberg R. Langmuir. 2000;16:1485–1488. [Google Scholar]
- 76.Nardin C, Thoeni S, Widmer J, Winterhalter M, Meier W. Chem Commun. 2000:1433–1434. [Google Scholar]
- 77.Langer R. Nature. 1998;392:5–10. [PubMed] [Google Scholar]
- 78.Langer R, Folkman J. Nature. 1976;263:797–800. doi: 10.1038/263797a0. [DOI] [PubMed] [Google Scholar]
- 79.Cohen S, Yoshioka T, Lucarelli M, Hwang LH, Langer R. Pharm Res. 1991;8:713–720. doi: 10.1023/a:1015841715384. [DOI] [PubMed] [Google Scholar]
- 80.Batycky RP, Hanes J, Langer R, Edwards DA. J Pharm Sci. 1997;86:1464–1477. doi: 10.1021/js9604117. [DOI] [PubMed] [Google Scholar]
- 81.Berkland C, Kim K, Pack DW. Pharm Res. 2003;20:1055–1062. doi: 10.1023/a:1024466407849. [DOI] [PubMed] [Google Scholar]
- 82.Zhao XJ, Jain S, Larman HB, Gonzalez S, Irvine DJ. Biomaterials. 2005;26:5048–5063. doi: 10.1016/j.biomaterials.2004.12.003. [DOI] [PubMed] [Google Scholar]
- 83.Yang YY, Chung TS, Bai XL, Chan WK. Chem Eng Sci. 2000;55:2223–2236. [Google Scholar]
- 84.Tickle C, Lee J, Eichele G. Dev Biol. 1985;109:82–95. doi: 10.1016/0012-1606(85)90348-3. [DOI] [PubMed] [Google Scholar]
- 85.Thaller C, Eichele G. Nature. 1987;327:625–628. doi: 10.1038/327625a0. [DOI] [PubMed] [Google Scholar]
- 86.Thaller C, Eichele G. Development. 1988;103:473–483. doi: 10.1242/dev.103.3.473. [DOI] [PubMed] [Google Scholar]
- 87.Tickle C, Alberts B, Wolpert L, Lee J. Nature. 1982;296:564–566. doi: 10.1038/296564a0. [DOI] [PubMed] [Google Scholar]
- 88.Pekarek KJ, Jacob JS, Mathiowitz E. Nature. 1994;367:258–260. doi: 10.1038/367258a0. [DOI] [PubMed] [Google Scholar]
- 89.Baumler H, Georgieva R. Biomacromolecules. 2010;11:1480–1487. doi: 10.1021/bm1001125. [DOI] [PubMed] [Google Scholar]
- 90.Du JZ, O’Reilly RK. Chem Soc Rev. 2011;40:2402–2416. doi: 10.1039/c0cs00216j. [DOI] [PubMed] [Google Scholar]
- 91.Champion JA, Katare YK, Mitragotri S. Proc Natl Acad Sci U S A. 2007;104:11901–11904. doi: 10.1073/pnas.0705326104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92.Chew SY, Wen J, Yim EKF, Leong KW. Biomacromolecules. 2005;6:2017–2024. doi: 10.1021/bm0501149. [DOI] [PubMed] [Google Scholar]
- 93.Xu X, Chen X, Xu X, Lu T, Wang X, Yang L, Jing X. J Controlled Release. 2006;114:307–316. doi: 10.1016/j.jconrel.2006.05.031. [DOI] [PubMed] [Google Scholar]
- 94.Liang D, Hsiao BS, Chu B. Adv Drug Delivery Rev. 2007;59:1392–1412. doi: 10.1016/j.addr.2007.04.021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 95.Zeng J, Yang LX, Liang QZ, Zhang XF, Guan HL, Xu XL, Chen XS, Jing XB. J Controlled Release. 2005;105:43–51. doi: 10.1016/j.jconrel.2005.02.024. [DOI] [PubMed] [Google Scholar]
- 96.Lee KJ, Yoon J, Lahann J. Curr Opin Colloid Interface Sci. 2011;16:195–202. [Google Scholar]
- 97.Ling XY, Phang IY, Acikgoz C, Yilmaz MD, Hempenius MA, Vancso GJ, Huskens J. Angew Chem, Int Ed. 2009;48:7677–7682. doi: 10.1002/anie.200903579. [DOI] [PubMed] [Google Scholar]
- 98.Mandal S, Bhaskar S, Lahann J. Macromol Rapid Commun. 2009;30:1638–1644. doi: 10.1002/marc.200900340. [DOI] [PubMed] [Google Scholar]
- 99.Madou MJ. Fundamentals of Microfabrication. 2. CRC Press; 2002. [Google Scholar]
- 100.Ainslie KM, Kraning CM, Desai TA. Lab Chip. 2008;8:1042–1047. doi: 10.1039/b800604k. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 101.Tao SL, Desai TA. Adv Mater. 2005;17:1625–1630. [Google Scholar]
- 102.Foraker AB, Walczak RJ, Cohen MH, Boiarski TA, Grove CF, Swaan PW. Pharm Res. 2003;20:110–116. doi: 10.1023/a:1022211127890. [DOI] [PubMed] [Google Scholar]
- 103.Dendukuri D, Doyle PS. Adv Mater. 2009;21:4071–4086. [Google Scholar]
- 104.Rolland JP, Maynor BW, Euliss LE, Exner AE, Denison GM, DeSimone JM. J Am Chem Soc. 2005;127:10096–10100. doi: 10.1021/ja051977c. [DOI] [PubMed] [Google Scholar]
- 105.Canelas DA, Herlihy KP, DeSimone JM. Wiley Interdiscip Rev: Nanomed Nanobiotechnol. 2009;1:391–404. doi: 10.1002/wnan.40. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 106.Petros RA, Ropp PA, DeSimone JM. J Am Chem Soc. 2008;130:5008–5009. doi: 10.1021/ja801436j. [DOI] [PubMed] [Google Scholar]
- 107.Leong TG, Lester PA, Koh TL, Call EK, Gracias DH. Langmuir. 2007;23:8747–8751. doi: 10.1021/la700913m. [DOI] [PubMed] [Google Scholar]
- 108.Leong TG, Benson BR, Call EK, Gracias DH. Small. 2008;4:1605–1609. doi: 10.1002/smll.200800280. [DOI] [PubMed] [Google Scholar]
- 109.Azam A, Laflin KE, Jamal M, Fernandes R, Gracias DH. Biomed Microdevices. 2011;13:51–58. doi: 10.1007/s10544-010-9470-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 110.Cho JH, Gracias DH. Nano Lett. 2009;9:4049–4052. doi: 10.1021/nl9022176. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 111.Filipiak DJ, Azam A, Leong TG, Gracias DH. J Micromech Microeng. 2009;19:075012. doi: 10.1088/0960-1317/19/7/075012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 112.Randall CL, Gultepe E, Gracias DH. Trends Biotechnol. 2012;30:138–146. doi: 10.1016/j.tibtech.2011.06.013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 113.Kalinin YV, Randhawa JS, Gracias DH. Angew Chem, Int Ed. 2011;50:2549–2553. doi: 10.1002/anie.201007107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 114.Leong T, Gu ZY, Koh T, Gracias DH. J Am Chem Soc. 2006;128:11336–11337. doi: 10.1021/ja063100z. [DOI] [PubMed] [Google Scholar]
- 115.Randall CL, Kalinin YV, Jamal M, Manohar T, Gracias DH. Lab Chip. 2011;11:127–131. doi: 10.1039/c0lc00368a. [DOI] [PubMed] [Google Scholar]
- 116.Randall CL, Gillespie A, Singh S, Leong TG, Gracias DH. Anal Bioanal Chem. 2009;393:1217–1224. doi: 10.1007/s00216-008-2538-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 117.Randall CL, Kalinin YV, Jamal M, Shah A, Gracias DH. Nanomed: Nanotechnol, Biol Med. 2011;7:686–689. doi: 10.1016/j.nano.2011.08.020. [DOI] [PubMed] [Google Scholar]
- 118.Desai TA, Chu WH, Tu JK, Beattie GM, Hayek A, Ferrari M. Biotechnol Bioeng. 1998;57:118–120. doi: 10.1002/(sici)1097-0290(19980105)57:1<118::aid-bit14>3.0.co;2-g. [DOI] [PubMed] [Google Scholar]
- 119.Gimi B, Kwon J, Liu L, Su Y, Nemani K, Trivedi K, Cui YH, Vachha B, Mason R, Hu WC, Lee JB. Biomed Microdevices. 2009;11:1205–1212. doi: 10.1007/s10544-009-9338-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 120.Luo Y, Shoichet MS. Nat Mater. 2004;3:249–253. doi: 10.1038/nmat1092. [DOI] [PubMed] [Google Scholar]
- 121.Wosnick JH, Shoichet MS. Chem Mater. 2008;20:55–60. [Google Scholar]
- 122.Mapili G, Lu Y, Chen SC, Roy K. J Biomed Mater Res, Part B. 2005;75B:414–424. doi: 10.1002/jbm.b.30325. [DOI] [PubMed] [Google Scholar]
- 123.Denk W, Strickler JH, Webb WW. Science. 1990;248:73–76. doi: 10.1126/science.2321027. [DOI] [PubMed] [Google Scholar]
- 124.Hill RT, Shear JB. Anal Chem. 2006;78:7022–7026. doi: 10.1021/ac061102w. [DOI] [PubMed] [Google Scholar]
- 125.Seidlits SK, Schmidt CE, Shear JB. Adv Funct Mater. 2009;19:3543–3551. [Google Scholar]
- 126.Kaehr B, Allen R, Javier DJ, Currie J, Shear JB. Proc Natl Acad Sci U S A. 2004;101:16104–16108. doi: 10.1073/pnas.0407204101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 127.Iosin M, Scheul T, Nizak C, Stephan O, Astilean S, Baldeck P. Microfluid Nanofluid. 2011;10:685–690. [Google Scholar]
- 128.Lee S-H, Moon JJ, West JL. Biomaterials. 2008;29:2962–2968. doi: 10.1016/j.biomaterials.2008.04.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 129.Richardson TP, Peters MC, Ennett AB, Mooney DJ. Nat Biotechnol. 2001;19:1029–1034. doi: 10.1038/nbt1101-1029. [DOI] [PubMed] [Google Scholar]
- 130.Bedard MF, De Geest BG, Moehwald H, Sukhorukov GB, Skirtach AG. Soft Matter. 2009;5:3927–3931. [Google Scholar]
- 131.Conley R, Gupta SK, Sathyan G. Curr Med Res Opin. 2006;22:1879–1892. doi: 10.1185/030079906x132613. [DOI] [PubMed] [Google Scholar]
- 132.Skirtach AG, Karageorgiev P, Bedard MF, Sukhorukov GB, Möhwald H. J Am Chem Soc. 2008;130:11572–11573. doi: 10.1021/ja8027636. [DOI] [PubMed] [Google Scholar]
- 133.Delcea M, Yashchenok A, Videnova K, Kreft O, Möhwald H, Skirtach AG. Macromol Biosci. 2010;10:465–474. doi: 10.1002/mabi.200900359. [DOI] [PubMed] [Google Scholar]
- 134.Kreft O, Skirtach AG, Sukhorukov GB, Möhwald H. Adv Mater. 2007;19:3142–3145. [Google Scholar]
- 135.Adams SR, Tsien RY. Annu Rev Physiol. 1993;55:755–784. doi: 10.1146/annurev.ph.55.030193.003543. [DOI] [PubMed] [Google Scholar]
- 136.Parker I, Yao Y. Philos T Roy Soc B. 1991;246:269–274. doi: 10.1098/rspb.1991.0154. [DOI] [PubMed] [Google Scholar]
- 137.O’Neill SC, Mill JG, Eisner DA. Am J Physiol. 1990;258:C1165–C1168. doi: 10.1152/ajpcell.1990.258.6.C1165. [DOI] [PubMed] [Google Scholar]
- 138.Sourjik V, Berg HC. Proc Natl Acad Sci U S A. 2002;99:12669–12674. doi: 10.1073/pnas.192463199. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 139.Bolinger PY, Stamou D, Vogel H. Angew Chem, Int Ed. 2008;47:5544–5549. doi: 10.1002/anie.200801606. [DOI] [PubMed] [Google Scholar]
- 140.Bolinger PY, Stamou D, Vogel H. J Am Chem Soc. 2004;126:8594–8595. doi: 10.1021/ja049023u. [DOI] [PubMed] [Google Scholar]
- 141.Yavuz MS, Cheng YY, Chen JY, Cobley CM, Zhang Q, Rycenga M, Xie JW, Kim C, Song KH, Schwartz AG, Wang LHV, Xia YN. Nat Mater. 2009;8:935–939. doi: 10.1038/nmat2564. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 142.Skrabalak SE, Chen J, Sun Y, Lu X, Au L, Cobley CM, Xia Y. Acc Chem Res. 2008;41:1587–1595. doi: 10.1021/ar800018v. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 143.Sun YG, Wiley B, Li ZY, Xia YN. J Am Chem Soc. 2004;126:9399–9406. doi: 10.1021/ja048789r. [DOI] [PubMed] [Google Scholar]
- 144.Hamad-Schifferli K, Schwartz JJ, Santos AT, Zhang SG, Jacobson JM. Nature. 2002;415:152–155. doi: 10.1038/415152a. [DOI] [PubMed] [Google Scholar]
- 145.Ye HK, Randall CL, Leong TG, Slanac DA, Call EK, Gracias DH. Angew Chem, Int Ed. 2007;46:4991–4994. doi: 10.1002/anie.200604414. [DOI] [PubMed] [Google Scholar]
- 146.Tanaka T, Fillmore D, Sun ST, Nishio I, Swislow G, Shah A. Phys Rev Lett. 1980;45:1636–1639. [Google Scholar]
- 147.Siegel RA, Firestone BA. Macromolecules. 1988;21:3254–3259. [Google Scholar]
- 148.Bassik N, Abebe BT, Laflin KE, Gracias DH. Polymer. 2010;51:6093–6098. [Google Scholar]
- 149.Kokufata E, Zhang YQ, Tanaka T. Nature. 1991;351:302–304. [Google Scholar]
- 150.Miyata T, Asami N, Uragami T. Nature. 1999;399:766–769. doi: 10.1038/21619. [DOI] [PubMed] [Google Scholar]
- 151.Douglas SM, Bachelet I, Church GM. Science. 2012;335:831–834. doi: 10.1126/science.1214081. [DOI] [PubMed] [Google Scholar]
- 152.Andersen ES, Dong M, Nielsen MM, Jahn K, Subramani R, Mamdouh W, Golas MM, Sander B, Stark H, Oliveira CLP, Pedersen JS, Birkedal V, Besenbacher F, Gothelf KV, Kjems J. Nature. 2009;459:73–76. doi: 10.1038/nature07971. [DOI] [PubMed] [Google Scholar]
- 153.Rothemund PWK. Nature. 2006;440:297–302. doi: 10.1038/nature04586. [DOI] [PubMed] [Google Scholar]
- 154.Tinsley MR, Taylor AF, Huang Z, Showalter K. Phys Chem Chem Phys. 2011;13:17802–17808. doi: 10.1039/c1cp22109d. [DOI] [PubMed] [Google Scholar]
- 155.Miller MB, Bassler BL. Annu Rev Microbiol. 2001;55:165–199. doi: 10.1146/annurev.micro.55.1.165. [DOI] [PubMed] [Google Scholar]
- 156.Bernard A, Renault JP, Michel B, Bosshard HR, Delamarche E. Adv Mater. 2000;12:1067–1070. [Google Scholar]
- 157.Lee KB, Park SJ, Mirkin CA, Smith JC, Mrksich M. Science. 2002;295:1702–1705. doi: 10.1126/science.1067172. [DOI] [PubMed] [Google Scholar]
- 158.Mayer M, Yang J, Gitlin I, Gracias DH, Whitesides GM. Proteomics. 2004;4:2366–2376. doi: 10.1002/pmic.200300748. [DOI] [PubMed] [Google Scholar]
- 159.Piner RD, Zhu J, Xu F, Hong SH, Mirkin CA. Science. 1999;283:661–663. doi: 10.1126/science.283.5402.661. [DOI] [PubMed] [Google Scholar]
- 160.Wilbur JL, Kumar A, Kim E, Whitesides GM. Adv Mater. 1994;6:600–604. [Google Scholar]
- 161.Yang MX, Gracias DH, Jacobs PW, Somorjai GA. Langmuir. 1998;14:1458–1464. [Google Scholar]
- 162.Chen CS, Mrksich M, Huang S, Whitesides GM, Ingber DE. Science. 1997;276:1425–1428. doi: 10.1126/science.276.5317.1425. [DOI] [PubMed] [Google Scholar]
- 163.Singhvi R, Kumar A, Lopez GP, Stephanopoulos GN, Wang DIC, Whitesides GM, Ingber DE. Science. 1994;264:696–698. doi: 10.1126/science.8171320. [DOI] [PubMed] [Google Scholar]
- 164.Mossman KD, Campi G, Groves JT, Dustin ML. Science. 2005;310:1191–1193. doi: 10.1126/science.1119238. [DOI] [PubMed] [Google Scholar]
- 165.Bailly M, Yan L, Whitesides GM, Condeelis JS, Segall JE. Exp Cell Res. 1998;241:285–299. doi: 10.1006/excr.1998.4031. [DOI] [PubMed] [Google Scholar]
- 166.Orth RN, Wu M, Holowka DA, Craighead HG, Baird BA. Langmuir. 2003;19:1599–1605. [Google Scholar]
- 167.Broadwater SJ, Roth SL, Price KE, Kobaslija M, McQuade DT. Org Biomol Chem. 2005;3:2899–2906. doi: 10.1039/b506621m. [DOI] [PubMed] [Google Scholar]
- 168.Poe SL, Kobaslija M, McQuade DT. J Am Chem Soc. 2006;128:15586–15587. doi: 10.1021/ja066476l. [DOI] [PubMed] [Google Scholar]
- 169.Wasilke JC, Obrey SJ, Baker RT, Bazan GC. Chem Rev. 2005;105:1001–1020. doi: 10.1021/cr020018n. [DOI] [PubMed] [Google Scholar]
- 170.Tietze LF, Beifuss U. Angew Chem, Int Ed Engl. 1993;32:131–163. [Google Scholar]
- 171.Lehninger AL. Principles of biochemistry. Worth Publishers; New York, N. Y: 1982. [Google Scholar]
- 172.Enders D, Huttl MRM, Grondal C, Raabe G. Nature. 2006;441:861–863. doi: 10.1038/nature04820. [DOI] [PubMed] [Google Scholar]
- 173.Mann J. Chemical aspects of biosynthesis. 1. Oxford University Press; Oxford: 1994. [Google Scholar]
- 174.Antipina MN, Kiryukhin MV, Chong K, Low HY, Sukhorukov GB. Lab Chip. 2009;9:1472–1475. doi: 10.1039/b819725c. [DOI] [PubMed] [Google Scholar]
- 175.Rondelez Y, Tresset G, Tabata KV, Arata H, Fujita H, Takeuchi S, Noji H. Nat Biotechnol. 2005;23:361–365. doi: 10.1038/nbt1072. [DOI] [PubMed] [Google Scholar]
- 176.Steinbacher JL, McQuade DT. J Polym Sci, Part A: Polym Chem. 2006;44:6505–6533. [Google Scholar]
- 177.Seo M, Nie ZH, Xu SQ, Mok M, Lewis PC, Graham R, Kumacheva E. Langmuir. 2005;21:11614–11622. doi: 10.1021/la050519e. [DOI] [PubMed] [Google Scholar]
- 178.Kohler JM, Henkel T. Appl Microbiol Biotechnol. 2005;69:113–125. doi: 10.1007/s00253-005-0176-6. [DOI] [PubMed] [Google Scholar]
- 179.Gunther A, Jensen KF. Lab Chip. 2006;6:1487–1503. doi: 10.1039/b609851g. [DOI] [PubMed] [Google Scholar]
- 180.Kelly BT, Baret JC, Taly V, Griffiths AD. Chem Commun. 2007:1773–1788. doi: 10.1039/b616252e. [DOI] [PubMed] [Google Scholar]
- 181.Leamon JH, Link DR, Egholm M, Rothberg JM. Nat Methods. 2006;3:541–543. doi: 10.1038/nmeth0706-541. [DOI] [PubMed] [Google Scholar]
- 182.Breslauer DN, Lee PJ, Lee LP. Mol BioSyst. 2006;2:97–112. doi: 10.1039/b515632g. [DOI] [PubMed] [Google Scholar]
- 183.Solvas XCI, deMello A. Chem Commun. 2011;47:1936–1942. doi: 10.1039/c0cc02474k. [DOI] [PubMed] [Google Scholar]
- 184.Teh SY, Lin R, Hung LH, Lee AP. Lab Chip. 2008;8:198–220. doi: 10.1039/b715524g. [DOI] [PubMed] [Google Scholar]
- 185.Huebner A, Sharma S, Srisa-Art M, Hollfelder F, Edel JB, deMello AJ. Lab Chip. 2008;8:1244–1254. doi: 10.1039/b806405a. [DOI] [PubMed] [Google Scholar]
- 186.Zhang H, Tumarkin E, Peerani R, Nie Z, Sullan RMA, Walker GC, Kumacheva E. J Am Chem Soc. 2006;128:12205–12210. doi: 10.1021/ja0635682. [DOI] [PubMed] [Google Scholar]
- 187.Shepherd RF, Conrad JC, Rhodes SK, Link DR, Marquez M, Weitz DA, Lewis JA. Langmuir. 2006;22:8618–8622. doi: 10.1021/la060759+. [DOI] [PubMed] [Google Scholar]
- 188.Chu L-Y, Utada AS, Shah RK, Kim J-W, Weitz DA. Angew Chem, Int Ed. 2007;46:8970–8974. doi: 10.1002/anie.200701358. [DOI] [PubMed] [Google Scholar]
- 189.Dendukuri D, Tsoi K, Hatton TA, Doyle PS. Langmuir. 2005;21:2113–2116. doi: 10.1021/la047368k. [DOI] [PubMed] [Google Scholar]
- 190.Nie ZH, Li W, Seo M, Xu SQ, Kumacheva E. J Am Chem Soc. 2006;128:9408–9412. doi: 10.1021/ja060882n. [DOI] [PubMed] [Google Scholar]
- 191.Tan W-H, Takeuchi S. Lab Chip. 2006;6:757–763. doi: 10.1039/b517178d. [DOI] [PubMed] [Google Scholar]
- 192.Tresset G, Takeuchi S. Biomed Microdevices. 2004;6:213–218. doi: 10.1023/B:BMMD.0000042050.95246.af. [DOI] [PubMed] [Google Scholar]
- 193.Song H, Tice JD, Ismagilov RF. Angew Chem, Int Ed. 2003;42:768–772. doi: 10.1002/anie.200390203. [DOI] [PubMed] [Google Scholar]
- 194.Song H, Ismagilov RF. J Am Chem Soc. 2003;125:14613–14619. doi: 10.1021/ja0354566. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 195.Tan W-H, Takeuchi S. Adv Mater. 2007;19:2696. [Google Scholar]
- 196.Boedicker JQ, Vincent ME, Ismagilov RF. Angew Chem, Int Ed. 2009;48:5908–5911. doi: 10.1002/anie.200901550. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 197.Jiang XY, Xu QB, Dertinger SKW, Stroock AD, Fu TM, Whitesides GM. Anal Chem. 2005;77:2338–2347. doi: 10.1021/ac048440m. [DOI] [PubMed] [Google Scholar]
- 198.El-Ali J, Sorger PK, Jensen KF. Nature. 2006;442:403–411. doi: 10.1038/nature05063. [DOI] [PubMed] [Google Scholar]
- 199.Whitesides GM. Nature. 2006;442:368–373. doi: 10.1038/nature05058. [DOI] [PubMed] [Google Scholar]
- 200.Song H, Chen DL, Ismagilov RF. Angew Chem, Int Ed. 2006;45:7336–7356. doi: 10.1002/anie.200601554. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 201.Keenan TM, Folch A. Lab Chip. 2008;8:34–57. doi: 10.1039/b711887b. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 202.Kim S, Kim HJ, Jeon NL. Integr Biol. 2010;2:584–603. doi: 10.1039/c0ib00055h. [DOI] [PubMed] [Google Scholar]
- 203.Kim BJ, Wu M. Ann Biomed Eng. 2011 doi: 10.1007/s10439-10011-10489-10439. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 204.Dertinger SKW, Chiu DT, Jeon NL, Whitesides GM. Anal Chem. 2001;73:1240–1246. [Google Scholar]
- 205.Takayama S, Ostuni E, LeDuc P, Naruse K, Ingber DE, Whitesides GM. Nature. 2001;411:1016–1016. doi: 10.1038/35082637. [DOI] [PubMed] [Google Scholar]
- 206.Takayama S, McDonald JC, Ostuni E, Liang MN, Kenis PJA, Ismagilov RF, Whitesides GM. Proc Natl Acad Sci U S A. 1999;96:5545–5548. doi: 10.1073/pnas.96.10.5545. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 207.Cosson S, Kobel SA, Lutolf MP. Adv Funct Mater. 2009;19:3411–3419. [Google Scholar]
- 208.Georgescu W, Jourquin J, Estrada L, Anderson ARA, Quaranta V, Wikswo JP. Lab Chip. 2008;8:238–244. doi: 10.1039/b716203k. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 209.Atencia J, Morrow J, Locascio LE. Lab Chip. 2009;9:2707–2714. doi: 10.1039/b902113b. [DOI] [PubMed] [Google Scholar]
- 210.Kalinin Y, Neumann S, Sourjik V, Wu MM. J Bacteriol. 2010;192:1796–1800. doi: 10.1128/JB.01507-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 211.Jeon NL, Baskaran H, Dertinger SKW, Whitesides GM, Van de Water L, Toner M. Nat Biotechnol. 2002;20:826–830. doi: 10.1038/nbt712. [DOI] [PubMed] [Google Scholar]
- 212.Bagnard D, Thomasset N, Lohrum M, Püschel AW, Bolz J. J Neurosci. 2000;20:1030–1035. doi: 10.1523/JNEUROSCI.20-03-01030.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 213.Burden-Gulley SM, Payne HR, Lemmon V. J Neurosci. 1995;15:4370–4381. doi: 10.1523/JNEUROSCI.15-06-04370.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 214.Kenis PJA, Ismagilov RF, Whitesides GM. Science. 1999;285:83–85. doi: 10.1126/science.285.5424.83. [DOI] [PubMed] [Google Scholar]
- 215.Bettinger CJ, Weinberg EJ, Kulig KM, Vacanti JP, Wang YD, Borenstein JT, Langer R. Adv Mater. 2006;18:165–169. doi: 10.1002/adma.200500438. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 216.Labarbera M, Vogel S. Am Scientist. 1982;70:54–60. [Google Scholar]
- 217.Inamdar NK, Borenstein JT. Curr Opin Biotechnol. 2011;22:681–689. doi: 10.1016/j.copbio.2011.05.512. [DOI] [PubMed] [Google Scholar]
- 218.Therriault D, White SR, Lewis JA. Nat Mater. 2003;2:265–271. doi: 10.1038/nmat863. [DOI] [PubMed] [Google Scholar]
- 219.Huang J-H, Kim J, Agrawal N, Sudarsan AP, Maxim JE, Jayaraman A, Ugaz VM. Adv Mater. 2009;21:3567–3571. [Google Scholar]
- 220.Jo BH, Van Lerberghe LM, Motsegood KM, Beebe DJ. J Microelectromech Syst. 2000;9:76–81. [Google Scholar]
- 221.Wu H, Odom TW, Chiu DT, Whitesides GM. J Am Chem Soc. 2003;125:554–559. doi: 10.1021/ja021045y. [DOI] [PubMed] [Google Scholar]
- 222.Tan W, Desai TA. Biomed Microdevices. 2003;5:235–244. [Google Scholar]
- 223.Haessler U, Kalinin Y, Swartz MA, Wu MW. Biomed Microdevices. 2009;11:827–835. doi: 10.1007/s10544-009-9299-3. [DOI] [PubMed] [Google Scholar]
- 224.Vickerman V, Blundo J, Chung S, Kamm R. Lab Chip. 2008;8:1468–1477. doi: 10.1039/b802395f. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 225.Barkefors I, Thorslund S, Nikolajeff F, Kreuger J. Lab Chip. 2009;9:529–535. doi: 10.1039/b814691h. [DOI] [PubMed] [Google Scholar]
- 226.Saadi W, Rhee SW, Lin F, Vahidi B, Chung BG, Jeon NL. Biomed Microdevices. 2007;9:627–635. doi: 10.1007/s10544-007-9051-9. [DOI] [PubMed] [Google Scholar]
- 227.Cheung YK, Gillette BM, Zhong M, Ramcharan S, Sia SK. Lab Chip. 2007;7:574–579. doi: 10.1039/b700869d. [DOI] [PubMed] [Google Scholar]
- 228.Amadi OC, Steinhauser ML, Nishi Y, Chung S, Kamm RD, McMahon AP, Lee RT. Biomed Microdevices. 2010;12:1027–1041. doi: 10.1007/s10544-010-9457-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 229.Choi NW, Cabodi M, Held B, Gleghorn JP, Bonassar LJ, Stroock AD. Nat Mater. 2007;6:908–915. doi: 10.1038/nmat2022. [DOI] [PubMed] [Google Scholar]
- 230.Cabodi M, Choi NW, Gleghorn JP, Lee CSD, Bonassar LJ, Stroock AD. J Am Chem Soc. 2005;127:13788–13789. doi: 10.1021/ja054820t. [DOI] [PubMed] [Google Scholar]
- 231.Mosadegh B, Huang C, Park JW, Shin HS, Chung BG, Hwang SK, Lee KH, Kim HJ, Brody J, Jeon NL. Langmuir. 2007;23:10910–10912. doi: 10.1021/la7026835. [DOI] [PubMed] [Google Scholar]
- 232.Jamal M, Zarafshar AM, Gracias DH. Nat Commun. 2011;2:527. doi: 10.1038/ncomms1531. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 233.Randolph GJ, Angeli V, Swartz MA. Nat Rev Immunol. 2005;5:617–628. doi: 10.1038/nri1670. [DOI] [PubMed] [Google Scholar]
- 234.Chiu HC, Lin YW, Huang YF, Chuang CK, Chern CS. Angew Chem, Int Ed. 2008;47:1875–1878. doi: 10.1002/anie.200704078. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.