Abstract
Recent studies of mammalian genomes have uncovered the vast extent of copy number variations (CNVs) that contribute to phenotypic diversity. Compared to SNP, a CNV can cover a wider chromosome region, which may potentially incur substantial sequence changes and induce more significant effects on phenotypes. CNV has been becoming an alternative promising genetic marker in the field of genetic analyses. Here we firstly report an account of CNV regions in the cattle genome in Chinese Holstein population. The Illumina Bovine SNP50K Beadchips were used for screening 2047 Holstein individuals. Three different programes (PennCNV, cnvPartition and GADA) were implemented to detect potential CNVs. After a strict CNV calling pipeline, a total of 99 CNV regions were identified in cattle genome. These CNV regions cover 23.24 Mb in total with an average size of 151.69 Kb. 52 out of these CNV regions have frequencies of above 1%. 51 out of these CNV regions completely or partially overlap with 138 cattle genes, which are significantly enriched for specific biological functions, such as signaling pathway, sensory perception response and cellular processes. The results provide valuable information for constructing a more comprehensive CNV map in the cattle genome and offer an important resource for investigation of genome structure and genomic variation underlying traits of interest in cattle.
Introduction
With the rapid progress of genome sequencing, the abundance of single nucleotide polymorphisms (SNPs) as a major source of genetic variation has been widely recognized. As a result, great efforts were made to develop high-throughput SNP genotyping platforms, and genome-wide high density SNP chips have been designed for many species including human and major farm animal species, such as cattle, swine, sheep, and chicken. Using these SNP chips, a large number of genome-wide association studies (GWAS) have been carried out in milking and meat production as well as diseases in cattle [1], [2], [3]. Over last few years, a few of crucial SNPs have been identified and confirmed with effects on milk production traits, e.g., the K232A mutation in DGAT1 [4].
In addition to SNPs, another form of genetic variation, i.e., copy number variation (CNV), has been identified in many species, including human [5], [6], [7], [8], [9], [10], mouse [11], [12], [13], [14], [15], rat [16], fruit fly [17], [18], dog [19], pig [20], [21], and cattle [22], [23], [24], [25], [26]. CNV is defined as a variable copy number of DNA segments ranging from 1 kilobase (Kb) to several megabases (Mb) compared with a reference genome [27]. CNVs take several forms, including deletions, duplications, insertions and complex rearrangements in the genome. So far, there are 179,450 CNVs identified in human genome which cover more than 53% of the human genome according to the Database of Genomic Variants (DGV) (http://dgvbeta.tcag,ca/dgv/app/home?ref=NCBI36/hg18, Apr, 2012). Thousands of genes, regulation elements and segmental duplications are harbored within these CNV regions [27], [28]. CNVs can potentially influence phenotypes or lead to diseases by altering gene dosage and/or disrupting genes in the form of deletion or duplication [29], [30], [31]. Furthermore, CNVs can modulate gene expression indirectly through disturbing the regulation regions of genes [5]. It has been found that many CNVs contribute to phenotypic variation in animals [32], [33], [34] as well as in humans [35], [36], [37], [38], [39].
Currently, CNVs can been identified using different technological approaches. Two major platforms are commonly used. One is the comparative genomic hybridization (CGH) array based approach [9], [25], [40], [41], [42], in which signal intensities of reference and target DNA samples labeled with different fluorescent tags are compared. The other is the SNP array based approach [24], [43], [44], in which intensity values of SNPs derived from each sample are used to estimate copy numbers in each individual. In comparison between these two existing panels, CGH array based approach has excellent performance in signal-to-noise ratios, while the SNP array based approach is more convenient for high-throughput analyses and follow-up association studies [45]. With the development of high density SNP arrays, higher resolution of genomic regions can be achieved [46]. Furthermore, recent advances in next-generation sequencing technology allow for more detailed characterization of CNVs [47], [48], [49], [50] and detect CNVs with a higher effective resolution and sensitivity and become more and more popular due to the cost decreases for sequencing. Therefore, many studies pay more attention to efficient algorithms to detect reliable CNVs via SNP array data [51], [52] and sequence data [53], [54].
So far, only a few CNV studies in cattle have been performed and relative few CNVs were detected or confirmed. Using CGH array, Fadista et al. [23] reported 304 CNV regions (CNVRs) from 20 bovine samples derived from 4 dairy and beef breeds, and Liu et al. [25] identified over 200 CNVRs from 90 animals of several different cattle breeds. By SNP array, Bae et al. [22] identified 368 CNVRs from 265 samples, Hou et al. [24], [26] reported 682 candidate CNVRs in 521 animals of 21 cattle breeds and 811 CNVRs in 472 Angus cattle. Using sequencing platform, Bickhart et al. [48], Zhan et al. [49] and Stothard et al. [50] reported 1265, 520 and 790 CNVRs from one, two and five individuals, respectively. Although some novel CNVRs were found by sequencing platform in these studies, it is limited by using very limited numbers of tested animals. Compared with the coverage of CNVRs detected in the human genome, the total length of CNVRs reported in cattle only cover 0.13% (3.3 Mb) to 5.57% (141.8 Mb) of the cattle genome. It can be envisaged that there are still a large number of CNVs undetected. Considering potential significance of CNV contributing to complex traits, further efforts should be made to obtain a more comprehensive CNV map in cattle genome.
In this study, we investigated genome-wide CNVRs in a Chinese Holstein population with a larger sample size of 2047 individuals. To pursue convincing results, we employed three programs (PennCNV, GADA and cnvPartition) based on different algorithms to analyze Bovine SNP50 genotyping data along with very strict quality control. Consequentially, we identified 99 candidate CNV regions. Our study provides useful addition to the cattle CNV map.
Materials and Methods
Sample Collection and Genotyping
The study population consisted of 2047 Chinese Holstein cattle, including 1960 cows and 87 sires (of which 14 are sires of these cows, each has 83 to 358 daughters) with unknown relationship. The Chinese Holstein originated from crosses of European Holstein-Friesian with Chinese Yellow cattle about 70 yr ago. Since then, continuous introgression of foreign Holstein genes (live bulls, semen, and embryos), mainly from North America, have been conducted. Therefore, the current population has a close relationship with the North American Holstein.
DNA was extracted from blood samples of cows and semen samples of bulls. The concentration and the purity of genomic DNA were assessed on the Nanovue Spectrophotometer. All samples were genotyped with the Illumina BovineSNP50 BeadChip. All the markers were clustered and genotyped using the BEADSTUDIO software.
The blood samples were collected along with the regular quarantine inspection of the farms, so no ethical approval was required for this study.
CNV Detection
In order to increase the confidence in CNV detection and decrease the rate of false discoveries, we used three programs to infer CNVs: PennCNV [55], cnvPartition v2.4.4 Plug-in (http://www.illumina.com) and GADA (Genome Alteration Detection Algorithm, [56]). The required data on signal intensities (Log R ratio, LRR) and allelic intensity ratios (B allele frequency, BAF) of all SNPs for all samples were generated from the Illumina BeadStudio 3.5 software (Illumina). For PennCNV, which is the most widely used program for inferring CNV based on SNP data [21], [24], the analysis of the X chromosome and autosomes were separately performed. PennCNV was run using the –test option without considering pedigree information since the cows in our study population merely have sire information and the relationship of these bulls is unknown. The PFB (population frequency of B allele) file was generated based on the BAF of each marker in this population. The signal intensity of each SNP which is subject to genomic waves was adjusted for the GC content of the 500 Kb genomic region of its both sides. The parameters involved were defined as 0.24 for standard deviation of LRR, 0.01 for BAF_DRIFT and 0.05 for waviness factor. For cnvPartition v2.4.4 Plug-in, the default parameters set by Illumina were used. For GADA, the parameters involved were defined as 0.8 for sparseness hypeparameter (aα) and 8 for critical value of the backward elimination (BE).
For each program, we employed the following criteria to define a potential CNV: its size was less than 1 Mb; it contained three or more consecutive SNPs; and it was detected in at least two animals (the overlapped region detected in different animals was defined as a CNV). In addition, to minimize the false positive rate, the union region of overlapping CNVs detected by different programs was defined as a CNV region (CNVR).
Information on gene annotations within the CNVRs was retrieved from the NCBI Gene Database based on Btau_4.0 genome assembly (The Bovine Genome Sequencing and Analysis Consortium, 2009).
qPCR Validation
Quantitative real time PCR (qPCR) was used to validate CNVRs or CNVs detected in the study. The relative comparative threshold cycle (2−△△CT) method was used to quantify copy number changes by comparing the ΔCT [cycle threshold (CT) of target region minus CT of control region] value of samples with CNV to the ΔCT of a calibrator without CNV [57], [58], [59]. CNVRs (CNVs) were tested by using SYBR Green chemistry as recommended by the manufacturers. We designed the PCR primers using Primer 3 web tool (http://frodo.wi.mit.edu/primer3/). For each target CNVR, two pairs of primers were designed considering the uncertainty of the CNV boundaries. Moreover, In-Silico PCR program from the UCSC browser (http://genome.ucsc.edu/) was used for in silico specificity analysis to ensure the primers only matching with the sequence of interest. We generated standard curves for each primer of target and control regions in order to ensure approximately equal PCR efficiencies between them. A serial diluted genomic DNA samples from a common cattle was used as template for creating a standard curve of each primer. Amplification efficiencies of all primers were calculated based on the standard curves. The copy number of each CNVR (CNV) was compared with a region in the control gene Basic transcription factor 3 (BTF3) as done in previous studies [22]. All PCR primers were designed based on its reference sequence in NCBI. PCR amplifications were performed in a total volume of 20 µL consisting of the following reagents: 1 µL DNA (around 50 ng), 1 µL(20 pM/µL) of both forward primer and reverse primer, 10 µL of Master Mix (2×) and water (Roche Applied Science). All RT-PCRs were run in triplicate. PCRs were run as follows: 5 min at 95°C followed by 40 cycles at 95°C for 10sec and 60°C for 10 sec. All PCRs were performed in 96-well clear reaction plates (Roche Applied Science). The average CT value of three replications for each sample was calculated and normalized against the control gene with the assumption of existing two copies of DNA segment in the control region. For each CNVR (CNV) to be validated, a value from the formula 2×2−△△CT was calculated for each individual. The value obtained was used to judge if an individual is in normal status without CNV (if the value was around 2), in gain status (if the value was around 3 or above), or in loss status (if the value was around 0 or 1).
Results
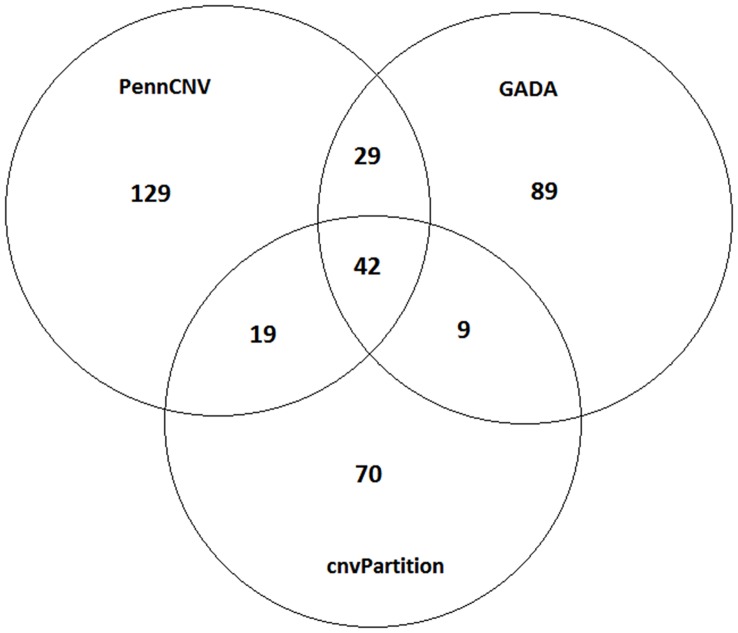
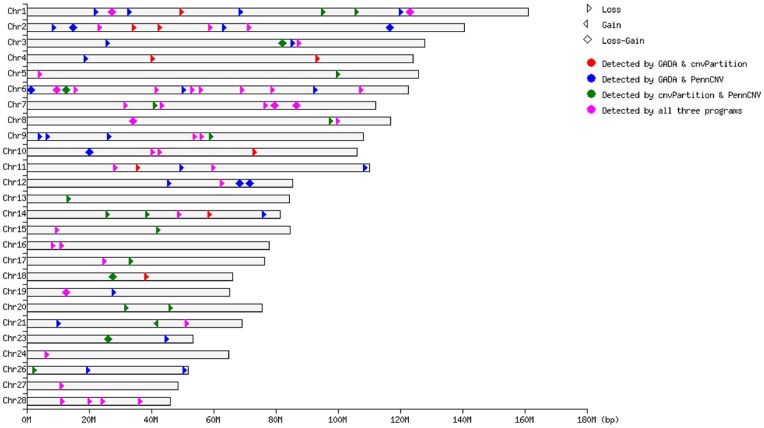
The numbers of CNVs called by PennCNV, GADA and cnvPartition were 219, 169 and 140, respectively (Fig. 1). Among these CNVs, 71 were commonly called by both PennCNV and GADA, 61 by both PennCNV and cnvPartition, 51 by both GADA and cnvPartition, and 42 by all of the three programs. A total of 99 CNVRs (union region of overlapping CNVs called by two or three programs) across genome were identified. The lengths of these CNVRs range from 27.01 Kb to 1.31 Mb, with an average size of 234.76 Kb and a median size of 151.69 Kb. The total length of all CNVRs is 23.24 Mb and covers 0.91% of the whole bovine genome. These CNVRs are located on all chromosomes except BTAs 22, 25, 29 and X. The numbers of CNVRs vary across different chromosomes, with BTA6 having the largest proportion (Fig. 2). Among the 99 CNVRs, 81 are in loss status, one in gain status and 17 in loss-gain status. The frequencies of these CNVRs in the study population are quite different. Specifically, 14 (14.1%), 17 (17.2%), 22 (22.2%), 34 (34.3%) and 52 (52.5%) CNVRs have frequencies of above 5%, 4%, 3%, 2% and 1%, respectively. Furthermore, 11 CNVRs were identified in more than 100 individuals, 63 CNVRs in more than 10 animals and the rest in more than 3 individuals. The CNVR with the highest frequency is on BTA10, reaching 27.09% in the population. The detailed description of each CNVR identified is given in Table S1.
Figure 1. Numbers of CNVs identified by three programs and numbers of CNVs overlapped between different programs.
Figure 2. The distribution and status of detected CNVRs across the bovine genome (based on the Btau_4.0 assembly).
These identified CNVRs contain abundant annotated genes. A total of 138 genes are harbored within 51 CNVRs (Table S2), of which 20 contain two or more annotated genes. On the other hand, there are 48 CNVRs without any known genes.
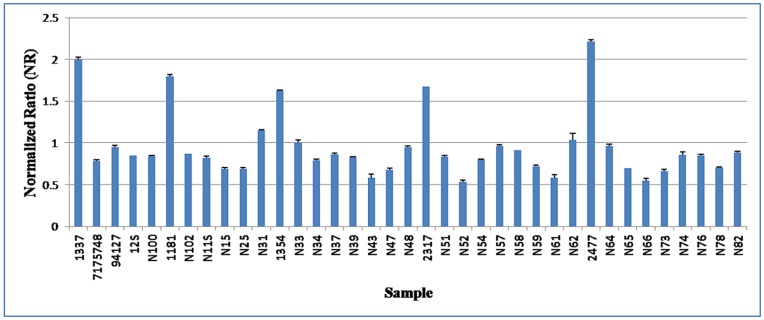
To further convince our results, we selected 6 CNVRs and 6 CNVs (detected only by PennCNV) to be validated by qPCR. These CNVRs or CNVs represent different status of copy numbers variation (i.e., loss and both) and different CNVR frequencies (varied from 0.19 to 6.3%) (See Table S1). In summary, of the 6 CNVRs, 4 (IDs = 43, 78, 80, and 84) were confirmed by qPCR, while of the 6 CNVs only 2 (CNV IDs = 100 and 101) were confirmed. Figure 3 illustrates the qPCR results for one confirmed CNVR (ID = 43). The full results for all of the 6 CNVRs are given in Figure S1. The detailed information of the validated CNVRs or CNVs and the primers used in qPCR analyses is given in Table S3.
Figure 3. Results of qPCR validation for one CNVR (No. 43).
NR around 2 indicates normal status (no CNV) and NR around 1 indicates one copy loss. The error bars represent the standard error among three technical replicates.
Discussion
CNVs contribute greatly to the genomic structure variation. In the past few years, CNVs have been explored extensively in the human genome and some of them were found to play important roles in disease susceptibility [52], [60], [61]. In animals, CNVs also contribute to the variation of phenotypes or some common diseases. For instance, the Pea-comb phenotype in chicken is caused by the duplication of the first intron of the Sox5 gene [32]. The white coat phenotype in pigs is caused by the copy number variation of the KIT gene [10]. Copy number variation of the ASIP (agouti signaling protein) gene in goat leads to different coat colors [62]. It is also reported that CNVs may be associated with cattle health and adaptive traits [26], [34]. These demonstrate that CNVs can be considered as promising markers for some traits or diseases in domestic animals. Our results illustrate the widespread of CNVR in Chinese Holstein genome. In total, 387 CNVs were detected by either of the three programs (PennCNV, GADA and cnvPartition), 99 out of which were called by at least two programs.
So far, a number of algorithms have been developed to infer CNVs based on SNP data. In order to minimize the false positive rate, we used three programs to detect CNV. It can be found that different methods lead to different results. 219, 169 and 140 CNVs were detected by PennCNV, GADA, and cnvPartition, respectively, and only 42 were commonly detected by the three programs. The inconsistence between different programs was also reported in other CNV studies. For example, using data from Porcine SNP60 BeadChip and also the same three programs as we used here, Ramayo-Caldas et al. [21] reported 94, 84, and 200 CNVs called by cnvPartition, PennCNV and GADA, respectively, in pigs from an Iberian x Landrace cross and only 26 were overlapped among them. Winchester et al. [63] compared 7 programs (including PennCNV, GADA and cnvPartition) using a common data set from the HapMap collection and found a large variation in numbers of copy number events among these algorithms. The inconsistence among different programs should be mainly due to the different algorithms implemented in these programs. In particular, PennCNV is based on the Hidden Markov model, GADA uses sparse Bayesian learning algorithms, while cnvPartition is a plug-in software within BeadsStudio (illumina) which uses the log R ratio and BAF and compares the data to 14 different Gaussian distribution models to assess copy number level. Each algorithm has its strengths and weaknesses as summarized by Winchester et al [63]. Therefore, Winchester et al. [63] recommended using multiple algorithms on a single dataset to produce the most informative results and also utilize the different advantages of each software.
Although the Bovine 50 K Beadchip is feasible for CNV detection, SNP probes on the chip are neither dense enough nor uniformly distributed to achieve an unbiased and high-resolution cattle CNV map. The average interval between adjacent SNPs on the Bovine 50 K Beadchip is 51.5 Kb. In addition, this chip was originally developed for SNP genotyping in association studies, and a large proportion of probes may be positioned beyond CNVRs. Hence, only the CNVRs with sufficient length were expected to be discovered. Some studies in humans suggested that smaller CNVs are much more frequent than larger ones [10], [64]. With application of the Bovine high-density 800 K chip or next generation sequencing methods, it can be expected that CNV resources across genome can be increasingly identified.
It is notable that these 99 CNVRs include 81 loss, 1 gain and 17 both (loss and gain) events in our study, i.e., loss-type CNVs are much more common than gain-type ones. Similar results have been reported in other studies [22], [23], [24]. But this is different from the results reported in the human genome studies and in the porcine genome studies [21]. This may be because that some CNVRs are not discovered in our study due to the limitation of the Bovine SNP 50 K Beadchip and the strict quality control criteria.
CNV content varies significantly among different chromosomes. The proportion of the total CNVR length on different chromosomes to the length of the chromosome ranged from 0.19% to 3.90% (see Table S7). Chromosomes 6, 1 and 2 show the greatest enrichment of CNVRs with two-fold of the average CNVR content across the whole genome. Compared with the reported CNVRs of bovine genome based on SNP array [22], [23], [24], [25], [26], our results are largely consistent with them (see Table 1). Specifically, 70 CNVRs (70.7%) in our results are overlapped with those reported by Hou et al. [24] and the total length of overlapped regions is 12.2 Mb (52.6%), 42 CNVRs (42.4%) overlapped with those reported by Bae et al. [22] and the length of overlapped region is 5.4 Mb (23.3%). In comparison with the CNV findings based on CGH-array, only 11 CNVRs (11.1%) with the total length of 0.8 Mb (3.4%) and six CNVRs (6.1%) with the total length of 0.7 Mb (3%) identified in our study are overlapped with those reported by Fadista et al. [23] and Liu et al. [25], respectively. In addition, we compared our results with the CNVRs detected based on sequence data [48], [49], [50]. The number of overlapped CNVRs varies from 10 to 77. The total length of 0.2 Mb (9%), 0.1 Mb (5%) and 0.36 Mb (16%) in our study are overlapped with those reported by Bickhart et al. [48], Zhan et al. [49] and Stothard et al. [50], respectively (Table 1). This demonstrates that different technology platforms for genome-wide CNV surveys can lead to different results, and it also illustrates that even using the same platform and program, different sets of CNVs can be inferred in different populations due to differences in population genetic background, sample size, CNV and CNVR definition, and technical errors. Since the identified CNVRs by different studies do not completely overlap, a great amount of CNVRs are still undiscovered in cattle genomes.
Table 1. Comparison between results of the current study and results from other studies.
| Findings from different studies | Overlapped CNVRs with this Study | ||||||
| Study | Count | Total Length(Mb) | Count | Percentage ofcount | Total length(Mb) | Percentage oflength | |
| CGH-based Studies | Fadista et al. [23] a | 266 | 16.6 | 11 | 11.1% | 0.8 | 3.4% |
| Liu et al. [25] b | 177 | 28.1 | 6 | 6.1% | 0.7 | 3% | |
| SNP-based Studies | Hou et al. [24] | 682 | 139.8 | 70 | 70.7% | 12.2 | 52.6% |
| Bae et al. [22] | 368 | 63.1 | 42 | 42.4% | 5.4 | 23.3% | |
| Hou et al. [26] | 811 | 141.8 | 59 | 59.6% | 10.6 | 45.7% | |
| Resequencing-based | Bickhart et al. [48] | 1265 | 55.6 | 10 | 10.1% | 0.202 | 0.9% |
| studies | Zhan et al. [49] | 520 | 3.6 | 16 | 16.2% | 0.112 | 0.5% |
| Stothard et al. [50] | 790 | 3.3 | 77 | 77.8% | 0.367 | 1.6% | |
| This study | 99 | 23.2 | |||||
: CNVRs on Chr Un and mitochondrial sequence are excluded;
: CNVRs on Chr Un are excluded.
Previous studies have shown that CNVs play an important role in phenotypic variation and are often related with disease susceptibility [5], [65], [66]. We compared the 99 CNVRs identified in this study with the reported QTL regions collected in the cattle QTL database (http://www.animalgenome.org/cgi-bin/QTLdb/BT/index). Since some QTLs have too large confidence interval and some QTLs reported by different studies are overlapped, we focused on QTL with confidence interval less than 30cM and considered those QTLs with overlapped confidence intervals greater than 50% as the same QTL. In this way, we identified 402 QTLs in total. 95 out of the 99 CNVRs harbor or partially overlap with 69 (17%) such QTL (Table S4). Since the total length of the 99 CNVRs covers only 0.91% of the whole bovine genome, there is a much greater QTL density coinciding with the CNVRs than we see in the genome as a whole. These QTLs are involved in many disease susceptibility traits, such as clinical mastitis, somatic cell score, bovine spongiform encephalopathy and gastrointestinal nematode burden (see Table S4). There are also CNVRs harboring QTLs which are associated with feed conversion, milk production and reproduction traits, such as calving ease, gestation length, birth body weight and non-return rate (Table S4). We also performed highly conserved elements (HECs) analysis and found 5,660 conserved elements in the CNVRs. The number of HECs in each CNVR is given in Table S5.
Furthermore, 51 of the identified CNVRs are completely or partially overlap with regions of bovine genes and encompass 138 known genes in total. The DAVID Bioinformatics Resources v6.7 [http://david.abcc.ncifcrf.gov/summary.jsp] [67] was used for gene ontology (GO) [68] and KEGG (Kyoto Encyclopedia of Genes and Genomes) [69] pathway analysis. Because some genes in the bovine genome do not have known function, the GO analysis was performed with the orthologous human genes of these bovine genes. As a result, we found that the functions of these genes are enriched in multiple categories of molecular functions, including sensory perception activity, regulation of biosynthetic process and cellular processes. Some genes in common GO terms among mammals (human, mouse) were also observed in cattle, e.g., the olfactory receptors gene families [13], [70], [71], [72], [73]. Besides, the KEGG analysis revealed a significant pathway, i.e., the Notch signaling pathway, which has been demonstrated to be very important in cell development in human and mouse [74], [75].We also compared the genes in the CNVRs detected in this study with those harbored in CNVRs of the human genome. As a result, 59 genes in CNVRs in the cattle genome also exist in CNVRs in the human genome (see Table S6).
In order to confirm these potential CNVRs, we performed quantitative PCR for 12 randomly selected CNVs, of which 6 were identified by two or three programs and 6 detected only by one program (PennCNV). From the former, 4 were confirmed successfully. This is similar to some previous reports in animals [21], [24]. From the later, only 2 were confirmed successfully. This suggests that using multiple CNV detection algorithms simultaneously can reduce the false positives, but it can also lead to some false negative results. It should be pointed out that those CNVs which are not confirmed by qPCR may not be really false positive. Three potential factors may contribute to this: First, SNP probes on the BovineSNP50 platform are neither dense enough nor uniformly distributed to achieve an unbiased CNVR map. Second, it is difficult to establish the exact boundaries of CNVRs. The breakpoint estimation of a CNVR may not be correct, leading to the designed primers outside the structural polymorphic region. Finally, the true CNVR boundaries may be also diverse among different animals.
In summary, we identified 99 CNVRs in Chinese Holstein by three different programs based on whole genome SNP genotyping data. These CNVRs covered 26 autosomes. Six of them were validated by qPCR successfully. Although the number of detected CNVRs here is probably an underestimate given the wide interval between SNPs in the Bovine 50 K BeadChip, the results provide a more comprehensive map of copy number variation in the cattle genome and it is an important resource for investigation of genome structure and cattle disease in the future studies.
Supporting Information
The file contains one figure with six subfigures. The figures display the detailed information of outcomes of qPCR validation for 6 detected CNVRs.
(DOC)
The detailed feature of each CNVR identified in this study.
(XLS)
Annotation of genes in CNVRs detected in this study.
(XLS)
Information of the validated CNVRs or CNVs and the primers used in quantitative PCR analyses.
(XLS)
Annotation of QTLs harbored within or partially overlapped with identified CNVRs across the bovine genome.
(XLS)
High Conservation Elements (HCE) in CNVRs detected in this study.
(XLS)
Gene content in the CNVRs and comparison with genes involved in Human Database of Genomic Variants.
(XLS)
The proportion of total CNVRs length on each chromosome.
(XLS)
Funding Statement
This study was supported by the 948 Program of the Ministry of Agriculture of China (2011-G2A(2)), the National High Technology Research and Development Program of China (863 Program 2011AA100302), the National Natural Science Foundations of China (30972092), the Natural Science Foundations of Beijing (6102016), the New-Century Training Program for Talents of the Ministry of Education of China (NETC-10-0783) and the China Postdoctoral Science Foundation (2011M500037). The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
References
- 1. Huang W, Maltecca C, Khatib H (2008) A proline-to-histidine mutation in POU1F1 is associated with production traits in dairy cattle. Anim Genet 39: 554–557. [DOI] [PubMed] [Google Scholar]
- 2. Khatib H, Monson RL, Schutzkus V, Kohl DM, Rosa GJ, et al. (2008) Mutations in the STAT5A gene are associated with embryonic survival and milk composition in cattle. J Dairy Sci 91: 784–793. [DOI] [PubMed] [Google Scholar]
- 3. Jiang L, Liu J, Sun D, Ma P, Ding X, et al. (2010) Genome wide association studies for milk production traits in Chinese Holstein population. PLoS One 5: e13661. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4. Kuhn C, Thaller G, Winter A, Bininda-Emonds OR, Kaupe B, et al. (2004) Evidence for multiple alleles at the DGAT1 locus better explains a quantitative trait locus with major effect on milk fat content in cattle. Genetics 167: 1873–1881. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5. Freeman JL, Perry GH, Feuk L, Redon R, McCarroll SA, et al. (2006) Copy number variation: new insights in genome diversity. Genome Res 16: 949–961. [DOI] [PubMed] [Google Scholar]
- 6. Fiegler H, Redon R, Andrews D, Scott C, Andrews R, et al. (2006) Accurate and reliable high-throughput detection of copy number variation in the human genome. Genome Res 16: 1566–1574. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7. Schaschl H, Aitman TJ, Vyse TJ (2009) Copy number variation in the human genome and its implication in autoimmunity. Clin Exp Immunol 156: 12–16. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8. Springfield CL, Sebat F, Johnson D, Lengle S, Sebat C (2004) Utility of impedance cardiography to determine cardiac vs. noncardiac cause of dyspnea in the emergency department. Congest Heart Fail 10: 14–16. [DOI] [PubMed] [Google Scholar]
- 9. Komura D, Shen F, Ishikawa S, Fitch KR, Chen W, et al. (2006) Genome-wide detection of human copy number variations using high-density DNA oligonucleotide arrays. Genome Res 16: 1575–1584. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10. Perry GH, Ben-Dor A, Tsalenko A, Sampas N, Rodriguez-Revenga L, et al. (2008) The fine-scale and complex architecture of human copy-number variation. Am J Hum Genet 82: 685–695. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11. Adams DJ, Dermitzakis ET, Cox T, Smith J, Davies R, et al. (2005) Complex haplotypes, copy number polymorphisms and coding variation in two recently divergent mouse strains. Nat Genet 37: 532–536. [DOI] [PubMed] [Google Scholar]
- 12. Kohler JR, Cutler DJ (2007) Simultaneous discovery and testing of deletions for disease association in SNP genotyping studies. Am J Hum Genet 81: 684–699. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13. Graubert TA, Cahan P, Edwin D, Selzer RR, Richmond TA, et al. (2007) A high-resolution map of segmental DNA copy number variation in the mouse genome. PLoS Genet 3: e3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14. She X, Cheng Z, Zollner S, Church DM, Eichler EE (2008) Mouse segmental duplication and copy number variation. Nat Genet 40: 909–914. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15. Watkins-Chow DE, Pavan WJ (2008) Genomic copy number and expression variation within the C57BL/6J inbred mouse strain. Genome Res 18: 60–66. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16. Guryev V, Saar K, Adamovic T, Verheul M, van Heesch SA, et al. (2008) Distribution and functional impact of DNA copy number variation in the rat. Nat Genet 40: 538–545. [DOI] [PubMed] [Google Scholar]
- 17. Aguade M (2009) Nucleotide and copy-number polymorphism at the odorant receptor genes Or22a and Or22b in Drosophila melanogaster. Mol Biol Evol 26: 61–70. [DOI] [PubMed] [Google Scholar]
- 18. Emerson JJ, Cardoso-Moreira M, Borevitz JO, Long M (2008) Natural selection shapes genome-wide patterns of copy-number polymorphism in Drosophila melanogaster. Science 320: 1629–1631. [DOI] [PubMed] [Google Scholar]
- 19. Chen WK, Swartz JD, Rush LJ, Alvarez CE (2009) Mapping DNA structural variation in dogs. Genome Res 19: 500–509. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20. Fadista J, Nygaard M, Holm LE, Thomsen B, Bendixen C (2008) A snapshot of CNVs in the pig genome. PLoS One 3: e3916. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21. Ramayo-Caldas Y, Castello A, Pena RN, Alves E, Mercade A, et al. (2010) Copy number variation in the porcine genome inferred from a 60 k SNP BeadChip. BMC Genomics 11: 593. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22. Bae JS, Cheong HS, Kim LH, NamGung S, Park TJ, et al. (2010) Identification of copy number variations and common deletion polymorphisms in cattle. BMC Genomics 11: 232. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23. Fadista J, Thomsen B, Holm LE, Bendixen C (2010) Copy number variation in the bovine genome. BMC Genomics 11: 284. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24. Hou Y, Liu GE, Bickhart DM, Cardone MF, Wang K, et al. (2011) Genomic characteristics of cattle copy number variations. BMC Genomics 12: 127. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25. Liu GE, Hou Y, Zhu B, Cardone MF, Jiang L, et al. (2010) Analysis of copy number variations among diverse cattle breeds. Genome Res 20: 693–703. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26. Hou Y, Liu GE, Bickhart DM, Matukumalli LK, Li C, et al. (2011) Genomic regions showing copy number variations associate with resistance or susceptibility to gastrointestinal nematodes in Angus cattle. Funct Integr Genomics 12: 81–92. [DOI] [PubMed] [Google Scholar]
- 27. Feuk L, Carson AR, Scherer SW (2006) Structural variation in the human genome. Nat Rev Genet 7: 85–97. [DOI] [PubMed] [Google Scholar]
- 28. Redon R, Ishikawa S, Fitch KR, Feuk L, Perry GH, et al. (2006) Global variation in copy number in the human genome. Nature 444: 444–454. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29. Margareto J, Leis O, Larrarte E, Pomposo IC, Garibi JM, et al. (2009) DNA copy number variation and gene expression analyses reveal the implication of specific oncogenes and genes in GBM. Cancer Invest 27: 541–548. [DOI] [PubMed] [Google Scholar]
- 30. Mileyko Y, Joh RI, Weitz JS (2008) Small-scale copy number variation and large-scale changes in gene expression. Proc Natl Acad Sci U S A 105: 16659–16664. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31. Stranger BE, Forrest MS, Dunning M, Ingle CE, Beazley C, et al. (2007) Relative impact of nucleotide and copy number variation on gene expression phenotypes. Science 315: 848–853. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32. Wright D, Boije H, Meadows JR, Bed’hom B, Gourichon D, et al. (2009) Copy number variation in intron 1 of SOX5 causes the Pea-comb phenotype in chickens. PLoS Genet 5: e1000512. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33. Norris BJ, Whan VA (2008) A gene duplication affecting expression of the ovine ASIP gene is responsible for white and black sheep. Genome Res 18: 1282–1293. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Clop A, Vidal O, Amills M (2011) Copy number variation in the genomes of domestic animals. Anim Genet. [DOI] [PubMed]
- 35. Cusco I, Corominas R, Bayes M, Flores R, Rivera-Brugues N, et al. (2008) Copy number variation at the 7q11.23 segmental duplications is a susceptibility factor for the Williams-Beuren syndrome deletion. Genome Res 18: 683–694. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36. Frank B, Bermejo JL, Hemminki K, Sutter C, Wappenschmidt B, et al. (2007) Copy number variant in the candidate tumor suppressor gene MTUS1 and familial breast cancer risk. Carcinogenesis 28: 1442–1445. [DOI] [PubMed] [Google Scholar]
- 37. Wang J, Hegele RA (2007) Homozygous missense mutation (G56R) in glycosylphosphatidylinositol-anchored high-density lipoprotein-binding protein 1 (GPI-HBP1) in two siblings with fasting chylomicronemia (MIM 144650). Lipids Health Dis 6: 23. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38. McKinney C, Merriman ME, Chapman PT, Gow PJ, Harrison AA, et al. (2008) Evidence for an influence of chemokine ligand 3-like 1 (CCL3L1) gene copy number on susceptibility to rheumatoid arthritis. Ann Rheum Dis 67: 409–413. [DOI] [PubMed] [Google Scholar]
- 39. Shlien A, Malkin D (2009) Copy number variations and cancer. Genome Med 1: 62. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40. de Smith AJ, Tsalenko A, Sampas N, Scheffer A, Yamada NA, et al. (2007) Array CGH analysis of copy number variation identifies 1284 new genes variant in healthy white males: implications for association studies of complex diseases. Hum Mol Genet 16: 2783–2794. [DOI] [PubMed] [Google Scholar]
- 41. de Bustos C, Diaz de Stahl T, Piotrowski A, Mantripragada KK, Buckley PG, et al. (2006) Analysis of copy number variation in the normal human population within a region containing complex segmental duplications on 22q11 using high-resolution array-CGH. Genomics 88: 152–162. [DOI] [PubMed] [Google Scholar]
- 42.Tchinda J, Lee C (2006) Detecting copy number variation in the human genome using comparative genomic hybridization. Biotechniques 41: 385, 387, 389 passim. [DOI] [PubMed]
- 43. Kamath BM, Thiel BD, Gai X, Conlin LK, Munoz PS, et al. (2009) SNP array mapping of chromosome 20p deletions: genotypes, phenotypes, and copy number variation. Hum Mutat 30: 371–378. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44. Yau C, Holmes CC (2008) CNV discovery using SNP genotyping arrays. Cytogenet Genome Res 123: 307–312. [DOI] [PubMed] [Google Scholar]
- 45. Ionita-Laza I, Rogers AJ, Lange C, Raby BA, Lee C (2009) Genetic association analysis of copy-number variation (CNV) in human disease pathogenesis. Genomics 93: 22–26. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46. Rincon G, Weber KL, Eenennaam AL, Golden BL, Medrano JF (2012) Hot topic: performance of bovine high-density genotyping platforms in Holsteins and Jerseys. J Dairy Sci 94: 6116–6121. [DOI] [PubMed] [Google Scholar]
- 47. Sudmant PH, Kitzman JO, Antonacci F, Alkan C, Malig M, et al. (2010) Diversity of human copy number variation and multicopy genes. Science 330: 641–646. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48. Bickhart DM, Hou Y, Schroeder SG, Alkan C, Cardone MF, et al. (2012) Copy number variation of individual cattle genomes using next-generation sequencing. Genome Res 22: 778–790. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49. Zhan B, Fadista J, Thomsen B, Hedegaard J, Panitz F, et al. (2011) Global assessment of genomic variation in cattle by genome resequencing and high-throughput genotyping. BMC Genomics 12: 557. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50. Stothard P, Choi JW, Basu U, Sumner-Thomson JM, Meng Y, et al. (2011) Whole genome resequencing of black Angus and Holstein cattle for SNP and CNV discovery. BMC Genomics 12: 559. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51. Zollner S, Su G, Stewart WC, Chen Y, McInnis MG, et al. (2009) Bayesian EM algorithm for scoring polymorphic deletions from SNP data and application to a common CNV on 8q24. Genet Epidemiol 33: 357–368. [DOI] [PubMed] [Google Scholar]
- 52. Diskin SJ, Hou C, Glessner JT, Attiyeh EF, Laudenslager M, et al. (2009) Copy number variation at 1q21.1 associated with neuroblastoma. Nature 459: 987–991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53. Mills RE, Walter K, Stewart C, Handsaker RE, Chen K, et al. (2011) Mapping copy number variation by population-scale genome sequencing. Nature 470: 59–65. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54. Hach F, Hormozdiari F, Alkan C, Birol I, Eichler EE, et al. (2010) mrsFAST: a cache-oblivious algorithm for short-read mapping. Nat Methods 7: 576–577. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55. Wang K, Li M, Hadley D, Liu R, Glessner J, et al. (2007) PennCNV: an integrated hidden Markov model designed for high-resolution copy number variation detection in whole-genome SNP genotyping data. Genome Res 17: 1665–1674. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56. Pique-Regi R, Monso-Varona J, Ortega A, Seeger RC, Triche TJ, et al. (2008) Sparse representation and Bayesian detection of genome copy number alterations from microarray data. Bioinformatics 24: 309–318. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57. Bodin L, Beaune PH, Loriot MA (2005) Determination of cytochrome P450 2D6 (CYP2D6) gene copy number by real-time quantitative PCR. J Biomed Biotechnol 2005: 248–253. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58. D’Haene B, Vandesompele J, Hellemans J (2010) Accurate and objective copy number profiling using real-time quantitative PCR. Methods 50: 262–270. [DOI] [PubMed] [Google Scholar]
- 59. Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 25: 402–408. [DOI] [PubMed] [Google Scholar]
- 60. Kirov G (2010) The role of copy number variation in schizophrenia. Expert Rev Neurother 10: 25–32. [DOI] [PubMed] [Google Scholar]
- 61. Ibanez P, Bonnet AM, Debarges B, Lohmann E, Tison F, et al. (2004) Causal relation between alpha-synuclein gene duplication and familial Parkinson’s disease. Lancet 364: 1169–1171. [DOI] [PubMed] [Google Scholar]
- 62. Fontanesi L, Beretti F, Riggio V, Gomez Gonzalez E, Dall’Olio S, et al. (2009) Copy number variation and missense mutations of the agouti signaling protein (ASIP) gene in goat breeds with different coat colors. Cytogenet Genome Res 126: 333–347. [DOI] [PubMed] [Google Scholar]
- 63. Winchester L, Yau C, Ragoussis J (2009) Comparing CNV detection methods for SNP arrays. Brief Funct Genomic Proteomic 8: 353–366. [DOI] [PubMed] [Google Scholar]
- 64. Conrad DF, Pinto D, Redon R, Feuk L, Gokcumen O, et al. (2010) Origins and functional impact of copy number variation in the human genome. Nature 464: 704–712. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65. McCarroll SA, Altshuler DM (2007) Copy-number variation and association studies of human disease. Nat Genet 39: S37–42. [DOI] [PubMed] [Google Scholar]
- 66. Shastry BS (2009) Copy number variation and susceptibility to human disorders (Review). Mol Med Report 2: 143–147. [DOI] [PubMed] [Google Scholar]
- 67. Huang da W, Sherman BT, Lempicki RA (2009) Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc 4: 44–57. [DOI] [PubMed] [Google Scholar]
- 68. Ashburner M, Ball CA, Blake JA, Botstein D, Butler H, et al. (2000) Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat Genet 25: 25–29. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69. Kanehisa M, Goto S, Furumichi M, Tanabe M, Hirakawa M (2010) KEGG for representation and analysis of molecular networks involving diseases and drugs. Nucleic Acids Res 38: D355–360. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70. Conrad DF, Andrews TD, Carter NP, Hurles ME, Pritchard JK (2006) A high-resolution survey of deletion polymorphism in the human genome. Nat Genet 38: 75–81. [DOI] [PubMed] [Google Scholar]
- 71. Cutler G, Marshall LA, Chin N, Baribault H, Kassner PD (2007) Significant gene content variation characterizes the genomes of inbred mouse strains. Genome Res 17: 1743–1754. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72. Hasin Y, Olender T, Khen M, Gonzaga-Jauregui C, Kim PM, et al. (2008) High-resolution copy-number variation map reflects human olfactory receptor diversity and evolution. PLoS Genet 4: e1000249. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73. Young JM, Endicott RM, Parghi SS, Walker M, Kidd JM, et al. (2008) Extensive copy-number variation of the human olfactory receptor gene family. Am J Hum Genet 83: 228–242. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74. Quillard T, Devalliere J, Chatelais M, Coulon F, Seveno C, et al. (2009) Notch2 signaling sensitizes endothelial cells to apoptosis by negatively regulating the key protective molecule survivin. PLoS One 4: e8244. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75. Tchorz JS, Kinter J, Muller M, Tornillo L, Heim MH, et al. (2009) Notch2 signaling promotes biliary epithelial cell fate specification and tubulogenesis during bile duct development in mice. Hepatology 50: 871–879. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
The file contains one figure with six subfigures. The figures display the detailed information of outcomes of qPCR validation for 6 detected CNVRs.
(DOC)
The detailed feature of each CNVR identified in this study.
(XLS)
Annotation of genes in CNVRs detected in this study.
(XLS)
Information of the validated CNVRs or CNVs and the primers used in quantitative PCR analyses.
(XLS)
Annotation of QTLs harbored within or partially overlapped with identified CNVRs across the bovine genome.
(XLS)
High Conservation Elements (HCE) in CNVRs detected in this study.
(XLS)
Gene content in the CNVRs and comparison with genes involved in Human Database of Genomic Variants.
(XLS)
The proportion of total CNVRs length on each chromosome.
(XLS)