Abstract
Reviewed are the phosphorylation events reporting activation of protein kinases and the key substrates critical for the DNA damage signaling (DDS). These DDS events are detected immunocytochemically using phospho-specific Abs; flow cytometry or image-assisted cytometry provide the means to quantitatively assess them on a cell by cell basis. The multiparameter analysis of the data is used to correlate these events with each other and relate to the cell cycle phase, DNA replication and induction of apoptosis. Expression of γH2AX as a possible marker of induction of DNA double strand breaks is the most widely studied event of DDS. Reviewed are applications of this multiparameter approach to investigate constitutive DDS reporting DNA damage by endogenous oxidants byproducts of oxidative phosphorylation. Also reviewed are its applications to detect and explore mechanisms of DDS induced by variety of exogenous agents targeting DNA such as exogenous oxidants, ionizing radiation, radiomimetic drugs, UV light, DNA topoisomerase I and II inhibitors, DNA crosslinking drugs and variety of environmental genotoxins. Analysis of DDS induced by these agents provides often a wealth of information about mechanism of induction and the type of DNA damage (lesion) and is reviewed in the context of cell cycle phase specificity, DNA replication, and induction of apoptosis or cell senescence. Critically assessed is interpretation of the data as to whether the observed DDS events report induction of a particular type of DNA lesion.
Keywords: ATM, γH2AX, Chk2, DNA double stand breaks, chromatin relaxation, DNA replication, DNA topoisomerase inhibitors, UV, cytometry, confocal microscopy
I. DNA damage signaling
Damage to DNA triggers a cascade of multifaceted molecular changes collectively defined as the DNA damage signaling (DDS) or DNA damage response (DDR). These DDS events comprise of a multitude of protein post-translational modifications and have four critical goals: (i) enabling accessibility of the DNA repair machinery to the damage site; (ii) activating and engaging repair machinery; (iii) halting cell cycle progression and division to prevent transfer of damaged DNA to the progeny cells, and (iv) mobilizing mechanisms that prepare cell to undergo either apoptosis or senescence, the means to eradicate cells with extensive DNA damage that cannot be fully repaired (reviews1–7). Flow cytometry and imaging-assisted cytometry are the major methodologies used to assess DDS in individual cells and correlate particular DDS events with the cell cycle phase or induction of apoptosis. This review outlines molecular changes occurring in the course of DDS, describes cytometric approaches detecting these changes, provides examples of their application for mechanistic studies of genotoxic effects of various agents and lists applications of cytometric DDS assays for genotoxicity screening. It updates and expands our prior review which was focused on technical aspects of the methodology.8
A. Chromatin relaxation (decondensation)
The chromatin compaction, particularly within regions of heterochromatin, restricts the accessibility of DNA damage sites to DNA damage response proteins making DNA in these regions refractory to repair. Remodeling of chromatin structure that leads to its decondensation (relaxation) is one of the earliest events of the DDR (reviews9–13). This event increases accessibility of the damage sites to the repair machinery. The signal to initiate chromatin decondensation appears to be triggered by decline of torsional strain of the DNA double helical structure resulting from DNA damage, particularly when DNA double-strand breaks (DSBs) are being formed (Fig. 1). In the absence of DNA damage the DNA torsional strain (topological stress) is being maintained by its winding onto the nucleosome core composed of histone octamers causing supercoiling to maintain the supra-nucleosomal chromatin structure.14 Following DNA damage high mobility group proteins (HMGs) and histone H1 play major role in providing the mechanism for local chromatin decondensation.15–18 Compared with other nuclear proteins HMGs are more extensively and rapidly modified by phosphorylation, acetylation, methylation, ribosylation and/or sumoylation in response to a change in the physiological state of the cell, induction of stress or cell cycle perturbation.19 This set of proteins provides a continuous dynamic interplay modulating molecular structure of chromatin.20,21 Of particular significance is the binding of the HMGN1 protein to the nucleosome, which alters the architecture of chromatin and affects the levels of post-transcriptional modifications of the tails of nucleosomal histones. Specifically, upon HMGN1 binding to nucleosomes, phosphorylation of histone H3 on Ser10 is reduced.20 Because Ser10 histone H3 phosphorylation is required to maintain chromatin in a condensed state such as seen during mitosis22 or premature chromosome condensation,23 its low level of phosphorylation facilitates chromatin decondensation (relaxation). Prevention of histone H3 phosphorylation by activation of HMGN1 and its binding to nucleosomes may thus provide the mechanism for chromatin decondensation in response DNA damage.
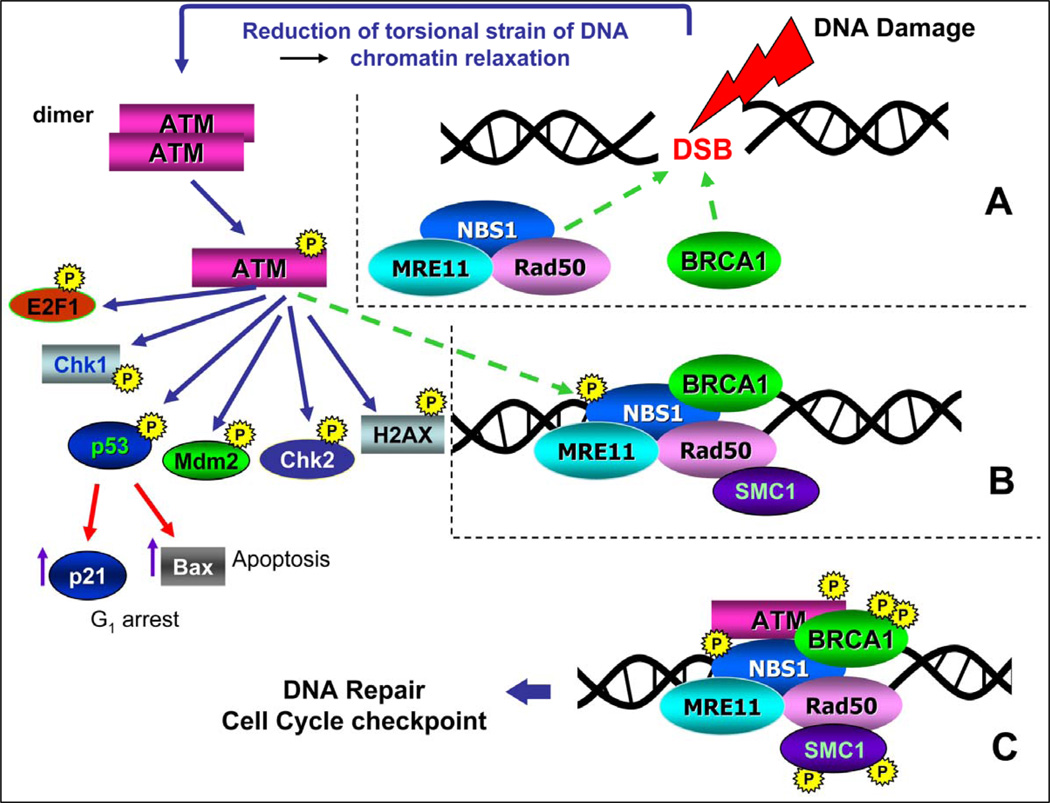
Fig. 1. The ATM signaling pathway triggered by induction of DSBs.1–7.
Induction of DSB leads to lessening of torsional strain and unwinding of DNA superhelical structure which triggers local decondensation of chromatin and recruits the MRE11, RAD50 and NBS1 proteins (MRN complex), as well as BRCA1 to the DSB site (A, dashed arrows). These events activate ATM which occurs by autophosphorylation of Ser1981 and leads to dissociation of the ATM dimer onto two monomers that have enzymatic activity. Activated ATM is then recruited to the site of the DSB (B, dashed arrow) where it phosphorylates several substrates including NBS1, BRCA1 and SMC1 (C). NBS1 phosphorylation is required for targeting ATM to phosphorylate Chk1 and Chk2. Phosphorylation of SMC1 activates S-phase checkpoints whereas BRCA1 phosphorylation engages this protein in the DSB repair pathway. ATM also phosphorylates E2F1, Chk1, p53, Mdm2, Chk2, and H2AX and several other substrates. Activated p53 (phosphorylated on Ser 15) induces transcription of p21WAF1 and/or Bax genes whose protein products arrest cells in G1 or promote apoptosis, respectively.
Histone acetyltransferase TIP60 also plays significant role in modulation of chromatin spatio-temporal dynamics triggered by DNA damage. It acetylates histone H2AX, which is a prerequisite for its phosphorylation on Ser139. Furthermore, TIP60 regulates the ubiquitination of H2AX via the ubiquitin-conjugating enzyme UBC.24,25 The presence of wt p53 appears to be essential for the induction of chromatin decondensation after DNA damage9, 26 likely through its effect on the tumor suppressor p33ING2.27 Chromatin relaxation not only augments accessibility of the repair proteins to DNA damage sites but also provides the signal for activation of Ataxia Telangiectasia Mutated (ATM) protein kinase through its phosphorylation on Ser1981.
At the moment of chromatin relaxation and ATM activation the MRN complex consisting of Meiotic Recombination 11 Homolog A (Mre11), Rad50 homolog and Nijmegen Breakage Syndrome 1 (NMR1) proteins undergoes translocation into the site of DNA damage.28–31 Relaxation of chromatin and translocation of MRN to the damage site are considered both to contribute to ATM activation. Whereas ATM activation takes place in chromatin at some distance from the DNA break site the activated ATM undergoes rapid translocation to the site.
B. Activation of phosphatidyl inositol 3' kinase-related kinases (PIKKs)
The DDR is regulated by three PIKKs: ATM, ATM and Rad3-related (ATR), and DNA dependent protein kinase (DNA-PKcs).4,32–34 These kinases phosphorylate multitude of proteins whose ultimate purpose is to preserve integrity of the genome. The substrates phosphorylated by these PIKKs are implicated in regulation of DNA damage repair, cell cycle progression, apoptosis and cell senescene. In many instances these PIKKs can have redundant activities and back-up each other in terms of phosphorylation of the same proteins.
Among the PIKKs activated in response to DNA damage the most extensively studied was ATM, which is the key component of the signal transduction pathways mobilized by the induction of DSBs.35,36 Activation of ATM is provided through its autophosphorylation on Ser1981 within the ATM dimer and is preceded by its prior acetylation that is mediated by the Tip60 histone acetyltransferase37. Following ATM phosphorylation the inactive dimers dissociate onto monomers having kinase enzymatic activity.2,3 (Fig. 1) The MRN protein complex plays a key role in the process of ATM activation by detecting DSBs, recruiting ATM to the damage site and targeting ATM to the respective substrates to carry on their phosphorylation38. Activation of ATM is also being promoted by p53-binding protein 1 (53BP1)39. The catalytic domain of ATM is outside of the Ser1981 prosphorylated site and becomes accessible to the kinase substrates when ATM is in its monomeric conformation.1,2
It should be stressed that in addition to its essential role in signaling DNA damage ATM can be directly activated by oxidation and plays an important function in oxidative stress response.40–42 In this capacity ATM senses redox potential, controls the level of reactive oxygen species within the cell and interacts with the AMP-activated protein kinase (AMPK), mammalian target of rapamycin (mTOR) and hypoxia-inducible factor 1 (HIF1)42. Interestingly, unlike in the case of activation by DNA damage which induces monomerization, ATM activated by the oxidative stress remains as a covalently linked dimer.42
As presented in Fig. 1 ATM phosphorylates several substrates at the site of the DSB, including NBS1, Structural Maintenance of Chromosomes 1 (SMC1) and Breast Cancer 1 (BRCA1) proteins. Phosphorylated NBS1 targets ATM towards Chk1, phosphorylated SMC1 engages the S-phase checkpoints halting DNA replication43 and BRCA1 phosphorylation is required to activate this protein along the DNA repair pathway. The BRCA1 (E3-ubiquitin ligase) is involved in several biochemical processes related to DNA repair.5,30 BRCA2 is essential for locating Rad51 to the sites of DNA damage and both BRCA proteins are involved in DNA repair by homologous recombination (HR).44 Also recruited to the DSB site is the mediator of DNA damage checkpoint 1 (MDC1).45 This nuclear protein activates the S phase- and G2/M phase- cell cycle checkpoints and interacts with phosphorylated histone H2AX near sites of DSB facilitating recruitment of the ATM and other repair factors to the damage site.
Whereas induction of DSBs such as caused by ionizing radiation triggers activation of ATM, DNA damage can also lead to activation of ATR and DNA-PKcs32–34. ATR becomes activated in response to UV radiation and to replication stress e.g. after exposure of cells to inhibitors of DNA polymerase.46–48 However, activation of DNA-PKcs takes place during repair of DSBs where it is an essential factor for the non-homologous end-joining (NHEJ) mechanism of DNA repair.33,49 Because the NHEJ mechanism also operates during V(D)J recombination and is responsible for antibody diversity DNA-PKcs is a critical element for normal immune development.50 DNA-PKcs is also strongly implicated in telomere maintenance.51 The process of activation and inactivation of DNA-PKcs is mediated by its extensive post-translational modification33. Among several sites of its phosphorylation Thr2609, which becomes autophosphorylated in response to DNA damage by ionizing radiation, has been the most extensively studied.52
C. Activation of checkpoint kinases
p53 (TP53), checkpoint kinase 2 (Chk2) and histone H2AX are among the most important downstream substrates phosphorylated by PIKKs.2,53 The purpose of checkpoint pathways activation is to halt cell cycle progression until integrity to DNA is restored by the repair mechanisms.54–57 In response to induction of DSBs ATM activates Chk2 via phosphorylation of Thr68 of this kinase which leads to Chk2 dimerization and acquirement of the catalytic activity.54,55 (Fig. 2) Phosphorylation of Chk2 on Thr68 can also be mediated by ATR, which takes place in response to DNA replication stress.56 Further steps in activation of Chk2 involve intermolecular phosphorylation on Thr383, Thr387 and Ser516 within the Chk2 dimers, which leads to dissociation of the dimers. Both the monomers and the multi-phosphorylated dimers are enzymatically active (Fig. 2). The DNA damage-activated Chk2 undergoes dissociation from chromatin which facilitates further signal amplification and translocation to soluble substrates.58 Activated Chk2 phosphorylates numerous downstream substrates including Cdc25A and Cdc25C phosphatases which after activation induce cell arrest at the G1 or at the transition from G2 to M, respectively (Fig. 2). Chk2 plays also a role in mediating the response to DNA damage by promoting apoptosis. For example after DNA damage induced by the DNA topoisomerase II (Top2) inhibitor etoposide, Chk2 phosphorylates and activates the E2F-1 transcription factor that activates apoptotic pathways.59 Phosphorylation of p53 by Chk2 may lead to upregulation of Bax, an event also promoting apoptosis. Phosphorylation of p53 may also lead to upregulation of p21Waf1 which in addition to activation of Cdc25A provides another mechanism to stop cell progression through G1.60 Still other substrates phosphorylated by Chk2 are BRCA1 and Promyelocytic Leukemia (PML) proteins.61,62 Phosphorylation of BRCA1 engages this protein in the DNA repair pathway while of PML increases cells proclivity to undergo apoptosis.55 Activated Chk2 also stabilizes the FoxM1 transcription factor thereby enhancing expression of DNA repair genes.63 There is strong redundancy between Chk1 and Chk2 as well as among all three isoforms of Cdc25 (Cdc25A, Cdc25B and Cdc25C) in their enzymatic activities of phosphorylation (Chk1, Chk2) or dephosphorylation (Cdc25A, Cdc25B, Cdc25C), of the respective substrates.64–66 This leads to redundancy and possibility of use different mechanisms to halt cell progression through G1 or G2 in response to DNA damage.
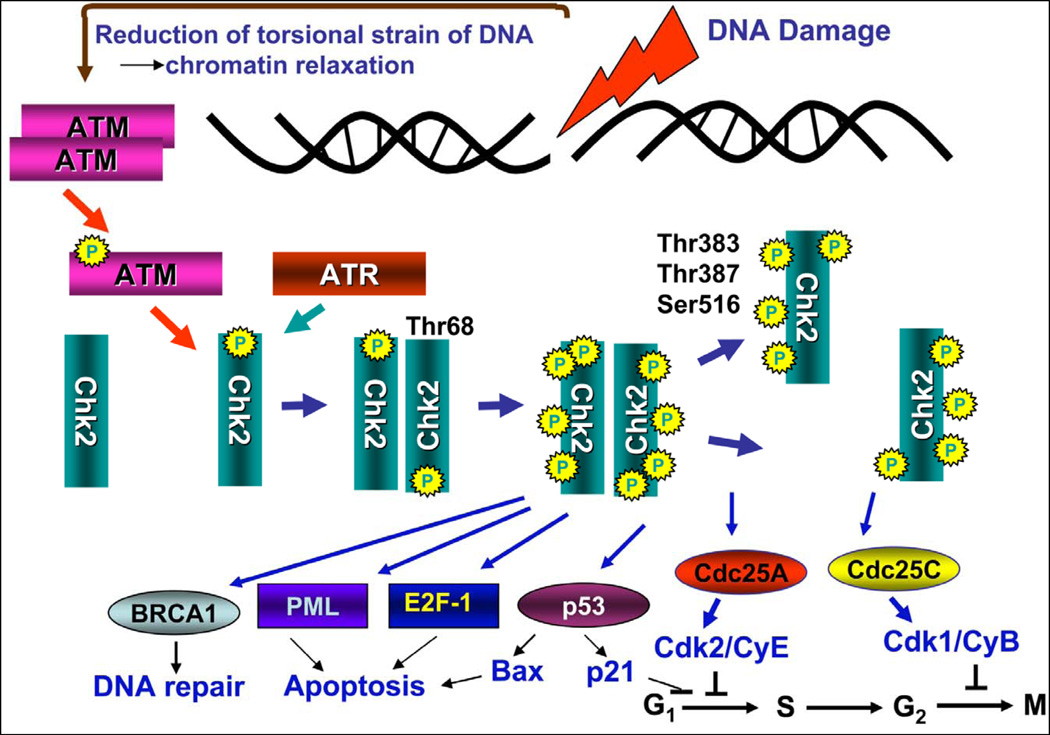
Fig. 2. Activation of Chk2 and Chk2’s major substrates.54–66.
DNA damage (induction of DSBs) triggers activation of ATM (Fig 1) which in turn phosphorylates Chk2 on Thr68 causing its dimerization. In response to replication stress rather than DSB Chk2 can be phosphorylated by ATR. Within the dimer Chk2 is further phosphorylated at Thr383, Thr387 and Ser516 which leads to its dissociation onto monomers. Both multi-phosphorylated dimers and monomers are enzymatically active and able to phosphorylate the downstream substrates. Among these substrates are the Cdc25C and Cdc25A phosphatases whose phosphorylation by Chk2 promotes binding to a 14-3-3 protein thereby preventing translocation into the nucleus and dephosphorylation of inhibitory phosphorylation at Thr14 and Tyr15 on cyclin/CDK complexes. This halts cell cycle transitions from G2 to M (Cdc25C) and G1 to S, (Cdc25A) respectively. Phosphorylation of Cdc25 phosphatases also accelerates their proteasomal degradation. A redundant mechanism of cell arrest in G1 involves phosphorylation of p53 by Chk2 which may lead to upregulation of the cdk2 inhibitor p21CIP1/WAF1. Phosphorylation of p53 may also result in upregulation of the pro-apoptotic protein Bax. Apoptosis may additionally be promoted by phosphorylation of PML and E2F-1. Phosphorylation of BRCA1 engages it in the DNA repair pathway.
D. Histone H2AX phosphorylation
Histone H2AX, one of the variants of histone H2A67 plays a critical role in surveillance of genome integrity.68–70 In response to DNA damage, particularly when the damage involves induction of DSBs, H2AX becomes phosphorylated on Ser139; so modified H2AX is defined as γH2AX.71–73 The phosphorylation can be mediated by ATM-74,75, ATR-76, and/or DNA-PKcs77 and takes place on nucleosomes located on both sides flanking DSBs along a megabase domain of DNA.71 H2AX is also phosphorylated during induction of DSBs in physiological processes such as DNA recombination in V(D)J class-switch during the process of immune system development and in meiosis.50,78 DNA fragmentation in cells undergoing apoptosis also induces extensive H2AX phosphorylation.79–81
II. DNA damage signaling (DDS) detected and measured in individual cells
A. Constitutive DDS
In live cells DNA is continuously exposed to reactive oxidants, by-products of aerobic respiration generated in mitochondria, as well as to exogenous oxidants or oxidant-inducers. Such exposure leads to a progressive oxidative DNA damage. According to one estimate during a single cell cycle of 24 h duration in the average cell in the human body the oxidants generate approximately 5,000 DNA single-strand lesions (SSLs).82 These lesions are single-strand breaks, apurinic/apyrimidinic sites, oxidation products such as 8-oxoguanine and thymine glycol, and some alkylation products83. Whereas about 99% of SSLs are repaired by essentially error-free mechanisms, one percent of them (~50) become converted to DSBs, predominantly during DNA replication. Recombinatorial repair (homologous recombination repair) and nonhomologous DNA-end joining (NHEJ) are the major pathways for repair of DSBs. The NHEJ pathway is error-prone, often resulting in deletion of a few base pairs.84 This leads to an accumulation of DNA damage with each cell division. Such permanent damage if occurs at sites of oncogenes or tumor suppressor genes promotes development of pre-neoplastic changes.85 It is also considered to contribute to cell aging and senescence.
The presence of oxidants and certain DNA primary lesions resulting from oxidative damage such as 8-oxoguanine, can be detected in individual cells by cytometry.86,87 However, because of the scarcity of DSBs generated by endogenous oxidants their detection is difficult. The comet methodology88 lacks the desired sensitivity and it also cannot provide information on the subnuclear location, relationship between the presence of DSBs and the cell cycle phase, or DNA ploidy, of the examined cells.
While studying a variety of cell lines during their unperturbed growth, as well as mitogenically stimulated normal lymphocytes, in the absence of any added exogenous genotoxic agents, we noticed the background level of H2AX phosphorylation and activation of ATM. This background level of γH2AX expression and ATM phosphorylation was defined as constitutive DDS.89–93 The level of constitutive DDS was shown to vary between different cell types (cell line) and was higher for cells having wt p53 compared to cells with null- or mutated- p53.93 It also was dissimilar for cells in different phases of the cell cycle being higher for S- and G2M- compared with G1- phase cells. Constitutive DNA damage response was markedly attenuated by exposure of cells to N-acetyl-L-cysteine (NAC), the scavenger of reactive oxidative species, and the attenuation was NAC-concentration dependent.89 Expression of γH2AX was also markedly decreased in cells growing at hypoxic (3% O2) conditions (Fig. 3).
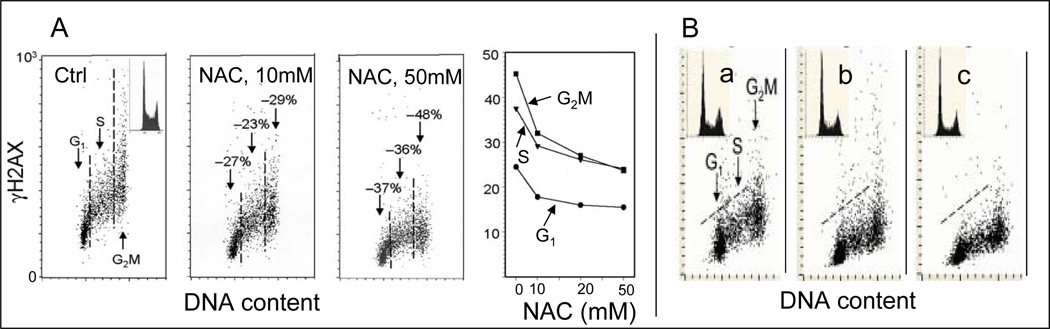
Fig. 3. Attenuation of constitutive expression of γH2AX by treatment with the ROS scavenger N-acetyl-L-cysteine (NAC) (A) and by hypoxia (B).
A. The bivariate (DNA content versus γH2AX IF) distributions show a decrease in the level of constitutive expression of γH2AX in TK6 lymphoblastoid cells growing in the presence of 10 or 50 mM NAC, added into cultures for 1 h prior to cell harvesting, compared to the untreated cells (Ctrl). The percent decline in mean values of γH2AX IF of G1, S and G2M phase cell subpopulations in cultures treated with NAC in relation to the respective subpopulations of the untreated (Ctrl) cells are shown above the respective arrows. The inset in the left panel shows the DNA content frequency histogram representative of the cells in these cultures. The right panel shows the plot of the mean values of γH2AX IF for G1, S and G2M cells, established by gating analysis, in relation to NAC concentration90. Because the level of constitutive expression of γH2AX (or ATM-S1981P) is relatively low, compared with the level seen after treatment with DNA damaging agents (e.g. Fig. 4) the γH2AX IF measurements have been carried out at higher sensitivity (higher voltage) of the photomultiplier.
B. Constitutive expression of γH2AX in WI-38 cells growing in 21% concentration of O2 (a), and 3% concentration of O2 for 24 (b) and 48 h (c).92.
Further studies demonstrated that the decreased metabolic activity such as induced by cell treatment with 2-deoxy-D-glucose or 3-bromopyruvate, exposure to the antioxidants ascorbate or celecoxib, confluency in cultures, as well as growth at low serum concentrations, all reduced the constitutive level of expression of γH2AX and of activated ATM. Conversely, the elevated level of constitutive expression of γH2AX and activated ATM was seen to be correlated with the increased metabolic activity and oxidative phosphorylation following mitogenic stimulation of lymphocytes or treatment with dichloroacetate, an agent known to shift metabolism from anaerobic to oxidative glycolysis through its effect on pyruvate dehydrogenase kinase.92 It also correlated well with the intracellular level of reactive oxygen species (ROS).91 Collectively, these data provide strong evidence that analysis of constitutive levels of H2AX phosphorylation and ATM activation in individual cells by multiparameter cytometry offers the means of measuring the effectiveness of factors such as antioxidants, ROS scavengers, caloric restriction mimetics or geroprotecting agents on protection of DNA from the damage caused by endogenous oxidants. Using this cytometric approach the effectiveness of hyaluronate, vitamin D3 and metformin, the agents considered to have geroporotective properties,94–96 in reducing the level of constitutive DNA damage, has been clearly demonstrated.97–99
Interestingly, a strong correlation between the level of constitutive expression of γH2AX and human age was observed for both males and females, as older individuals had markedly elevated expression of γH2AX.100 An increase in constitutive DDS is observed during tumor progression where it appears to provide clinically useful prognostic biomarker of the progression.101–105
B. DDS induced by exogenous oxidants
Exposure of cells to exogenous oxidants triggers DDS. The earliest event of DDS, relaxation (decondensation) of chromatin, can be detected by as early as 10 min after exposure to H2O2.106 The relaxation is evident in across the cell cycle being approximately of similar degree in G1, as in S or G2M cells. The subsequent events of DDS, however, such as activation of ATM and H2AX phosphorylation, are much more pronounced in S- compared to G1- or G2M- cells (Fig. 4).107,108
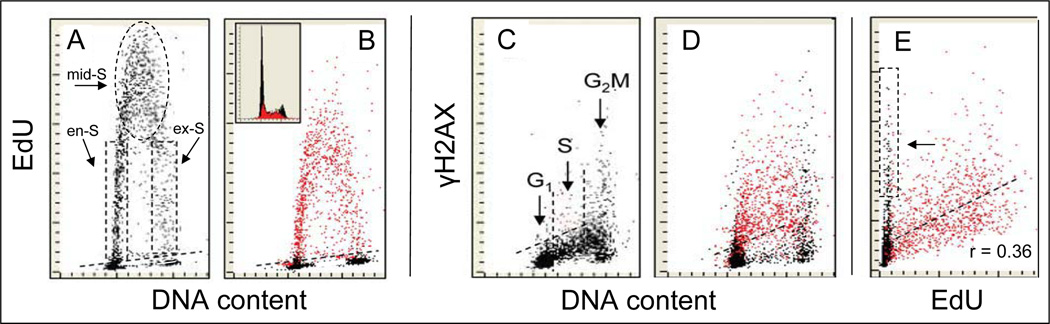
Fig. 4. Correlation between DNA replication and induction of γH2AX in A549 cells treated with H2O2.
A: Incorporation of EdU by untreated cells during 120 min exposure to EdU. The cells which during the duration of 120 EdU pulse were entering (en-S) or exiting (ex-S) S phase show variable level of EdU incorporation as the interval of their exposure to the precursor at the time of DNA replication varied; the mid-S phase cells show maximal EdU incorporation. .B. The cells were initially treated with EdU for 60 min and then, still in the presence of EdU, were exposed to 200 µM H2O2 for an additional 60 min. Note a decline in EdU incorporation compared to A. For the “paint-a-gate” multiparameter analysis the cells incorporating EdU (above the threshold marked by the dashed line), were electronically colored red. C. Constitutive expression of γH2AX in untreated (control) cells; the skewed dashed line shows the upper threshold of γH2AX expression for 97% of G1 and S-phase cells. D. Cells were treated as in B, the EdU incorporating cells are colored red. Note the increased expression of γH2AX (compared with C) predominantly in DNA replicating cells. E. Bivariate (EdU vs γH2AX) distribution shows a correlation between incorporation of EdU and expression of γH2AX. The EdU negative cells expressing γH2AX (arrow) are predominantly G2M cells that show increased level of constitutive DNA damage signaling.98
As is evident in Fig. 4 the DNA replicating cells marked by the pulse-labeling them with EdU (identified by the red dots in panel D) show considerably higher expression of γH2AX than the EdU unlabeled cells. A modest degree of correlation (r = 0.36) between extent of EdU incorporation and expression of γH2AX is apparent (Fig. 4D).108 The extent of EdU incorporation varies between the cells predominantly due to variability in the interval of DNA replication during duration of exposure to EdU (Fig. 4A, en-S and ex-S cells). The correlation thus indicates that the longer the cells were replicating DNA at the time of exposure to EdU (which was constant, 2 h) the greater was the intensity of DDS manifesting as the induction of γH2AX. These data are consistent with the mechanism in which the elongating replication forks stalled or collapsed upon encountering the primary DNA lesions induced by H2O2.provide signal for H2AX phosphorylation.109
C. DDS induced by ionizing radiation (IR), radiomimetic drugs and UV
The early studies on the detection of DDR in individual cells exposed to IR or radiomimetic drugs were focused on the assessment of a possible relationship between the extent IR-induced γH2AX, considered to report the presence of DSBs, and cell survival.110–116 The induction of γH2AX in several cell types was detected by flow cytometry already 1 h after exposure to the dose of X-rays as low as a 20 cGy.111 While expression of γH2AX in non-irradiated cells was higher in S-phase the X-ray induced increase in γH2AX was distinctly more pronounced in G1 than in S-phase cells.113 A strong correlation was observed between the induced expression of γH2AX and cell survival as measured by clonogenicity assays. The half-times of the radiation-induced γH2AX foci, ranged from 1.6 to 7.2 h, correlated well with clonogenic survival for 10 cell lines.110 Also strongly correlated with cell survival was expression of γH2AX measured 1 h after treatment of Chinese hamster V79 cells with several radiomimetic drugs examined within a range of over two decades of cell kill.112 This strong correlation led the authors to postulate that the assessment of H2AX phosphorylation by flow cytometry can be used as a surrogate of the cell kill estimate.112 Interestingly, the cell line-dependent differences in the rate of disappearance of IR induced γH2AX were associated with the status of p53 (wt vs p53 deficient) as well as was related to intrinsic radiosensitivity of the lines.110 The treatment of human and rodent DNA-repair proficient and deficient cell lines with cisplatin revealed that the level of the retention of γH2AX foci 24 h after the treatment was much more correlated with the fraction of cells that lost their clonogenic potential than the initial intensity of γH2AX expression following the treatment.116
Several subsequent studies provided further evidence that both, the extent of induction γH2AX by IR, and perhaps to an even greater degree, the duration (persistence) of the induced γH2AX, likely reporting the presence of the unrepaired DSBs, are strong biomarkers of cytotoxic effects of IR, predictive of elimination of cells’ proliferation capability.117–121
Unlike in the case of IR, the energy of UV light is absorbed by thymine and cytosine (pyrimidine bases) which leads to formation of the four-membered cyclobutane ring of the pyrimidine dimer.122 In addition, a single bond may form between two carbon atoms on the cyclobutane ring resulting in the so called ‘6–4 (T-C) photoproduct.122 Nucleotide excision repair (NER) is the primary mechanism of repair of DNA damage induced by UV.123 One of its earliest events of the DDR is chromatin relaxation. This event, detected by flow cytometry, has been seen in all cells, regardless of the cell cycle phase, already 10 min after exposure to UV.106
Phosphorylation of H2AX induced by UV is primarily mediated by ATR.124,125 However, there is a redundancy and activation of ATM as well as DNA PKcs can also be seen in UV treated cells, contributing to phosphorylation of H2AX and other downstream protein substrates.124 Unlike chromatin relaxation which takes place in all phases of the cell cycle,106 the activation of ATM and induction of γH2AX by UV was evident exclusively in DNA replicating cells and inhibition of DNA replication by the DNA polymerase inhibitor aphidicolin prevented the induction of γH2AX.126,127 It should be noted, however, that aphidicolin as well other inhibitors of DNA replication such hydroxyurea or excess of thymidine also induce γH2AX.46,47 The cell cycle-related pattern of γH2AX induction by these inhibitors, however, is very much different than that of the pattern induced by UV. Namely, unlike UV which affects cells uniformly across the S-phase, the most affected by the replication inhibitors were cells initiating DNA replication, at the very-early section of S-phase.46,47
Further evidence linking DNA replication with the induction of γH2AX by UV was obtained from the studies designed to correlate incorporation the DNA precursor 5-ethynyl-2'-deoxyuridine (EdU) with the extent of induction of γH2AX in the same cells. Exposure of cells to EdU to mark the cells replicating DNA, followed by exposure to UV revealed that γH2AX was induced exclusively in EdU-positive cells, and the extent of EdU incorporation versus the level of UV-induced γH2AX were strongly correlated (R2 = 0.98).127 There was no evidence of H2AX phosphorylation in the non-replicating cells.127 The data are consistent with the mechanism in which the UV-induced primary DNA lesions cause an initial total block to replication forks elongation.128–130 However DNA replication may still continue leaving unreplicated gaps behind the replication forks. It is not entirely clear to what extent the formation of DSBs130 versus the block in forks elongation128,129 contribute to activation of ATR, ATM and DNA-PKcs that leads to H2AX phosphorylation. It should be noted that the H2AX phosphorylation may occur during replication stress in the absence of DSBs formation,81 and thus the γH2AX may not be induced exclusively in response to DSBs caused by UV. Definitely, however, the induction of γH2X takes place only during replication.
D. DDS induced by anticancer drugs targeting DNA
1. DNA topoisomerase inhibitors
DNA topoisomerase I and II (Top1, Top2) inhibitors are among the most widely used anticancer drugs. Their mechanism of action is considered to involve stabilization of Top-DNA complexes (“cleavable complexes”) which are transiently formed to cut DNA in order to untangle double helix which becomes entangled or over-wound during replication or transcription. Collisions of elongating replication forks, or moving RNA polymerase molecules, with these complexes result in formation of DSBs, the potentially lethal lesions.131–133 Extensive studies were carried out to compare the cells cycle-specific effects of Top1 inhibitor topotecan (Tpt) with the Top2 inhibitors mitoxantrone (Mxt) and etoposide (Etp).134–142 Tpt is the analog of camptothecin (Cpt), binds to DNA by intercalation and stabilizes the Top1-DNA complex.143 The Top2 inhibitor Mxt which is anthracenedione analog also intercalates into DNA144,145 and stabilizes Top2-DNA complex.133 Etp (also named VP-16), one of the podophyllotoxins family of drugs, binds to the Top2-DNA complex stabilizing it.146 The pattern of DDS as related to the cell cycle phase specificity, induction of apoptosis and involvement of ROS was very much different for each drug.134–140
Specifically, the treatment of human lymphoblastoid TK6 or pulmonary adenocarcinoma A549 cells with Tpt (or Cpt) led to rapid induction of phosphorylation of ATM and H2AX exclusively in S phase cells. In contrast, their exposure to Mxt or Etp induced ATM and H2AX phosphorylation in all phases of the cycle.134,136,138 Moreover, whereas DDS induced by Mxt or Etp was significantly attenuated by the reactive oxygen species (ROS) scavenger N-acetyl-L-cysteine (NAC), no attenuation at all was seen after induction of DNA damage by Tpt or Cpt.135,138 The induction of apoptosis, seen subsequently to ATM activation and H2AX phosphorylation, was also cell cycle-phase specific. The S-phase cells were selectively undergoing apoptosis in cultures treated either with Tpt/Cpt or Mxt whereas apoptosis of the cells treated with Etp was seen in all phases of the cell cycle and actually was more pronounced in G1 phase.134
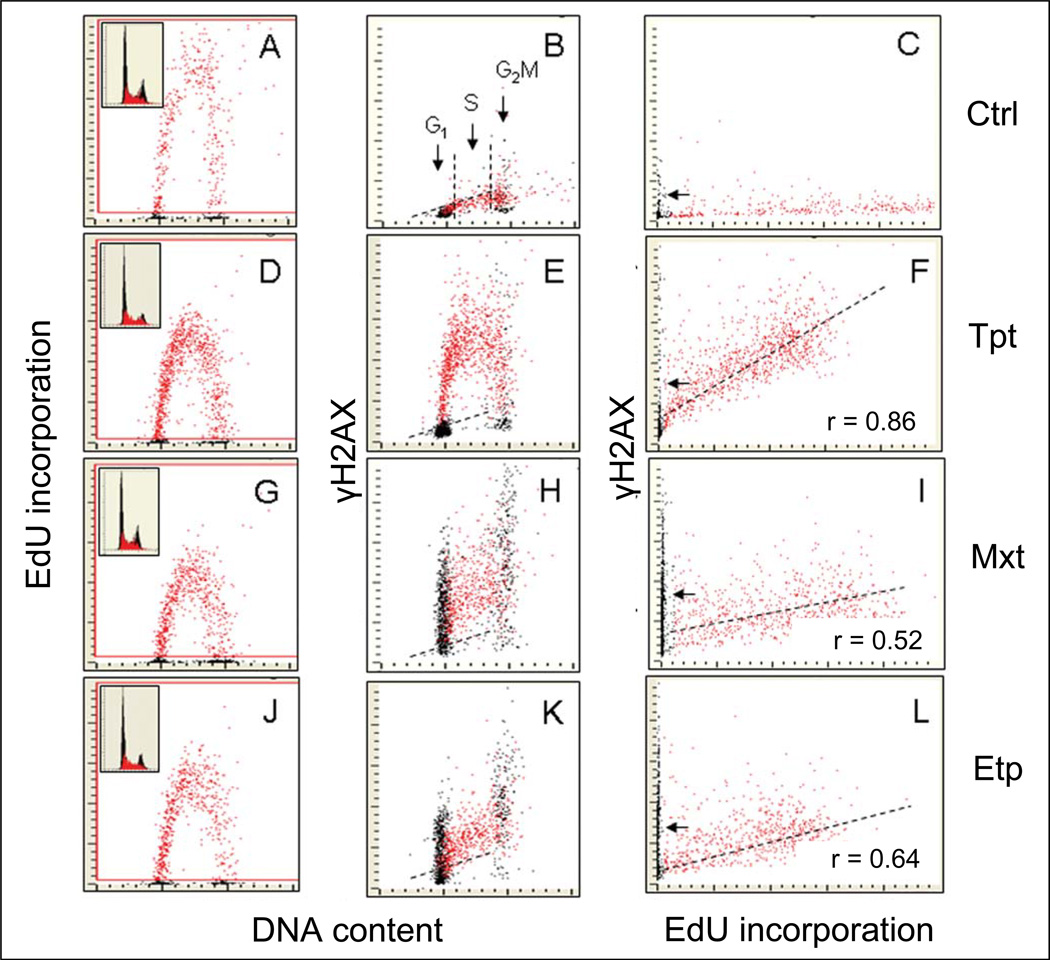
A relationship between the induction of DDR by Tpt, Mxt and Etp and actual DNA replication was explored by multiparameter cytometry, by labeling DNA replicating cells with EdU and measuring the induction of γH2AX, differentially labeled, in the same cells (Fig. 5).147 Toward this end the topoisomerase inhibitors were included into the cultures at the time when the cells were incorporating EdU. The multiparameter cytometric analysis made it possible to identify EdU-incorporating cells and using a “paint-a-gate” algorithm to directly correlate the expression of γH2AX with EdU incorporation, on cell by cell basis.147As is evident in Fig. 5 the induction of γH2AX after exposure to Cpt is seen exclusively in DNA replicating cells. Also, a relatively strong correlation between EdU incorporation and induction of γH2AX is evident in cells treated with Cpt (r = 0.86). In cells treated with Mxt or Etp, on the other hand, both DNA replicating as well as non-replicating cells demonstrate distinctly elevated level of γH2AX. The correlation between induction of γH2AX and EdU incorporation is also less apparent (Mxt, r = 0.52; Etp, r = 0.64). The data shown in this figure also indicate that, as would be expected, the treatment with Cpt, Mxt, or Etp decreased DNA replication rate revealed by lowered intensity of labeling with EdU seen in the D, G, and J panels compared with A (Fig. 5).
Fig. 5. Correlation between induction of H2AX by Cpt, Mxt, and Etp and DNA replication in A549 cells.
Exponentially growing A549 cells were exposed to EdU for 30 min and then (still in the presence of EdU) were treated either with Cpt (D, E, F), Mxt (G, H, I), or Etp (J, K, L) for 2 h. Control cells (A—C) were exposed only to EdU for 2.5 h. The cells were then fixed and their blue (DAPI), green (EdU), and red (γH2AX) fluorescence were measured by LSC.147 The cells that incorporated EdU were “paint-gated’ (red) and the data were plotted as bivariate distributions representing γH2AX versus DNA content (B, E, H, K) or γH2AX versus EdU incorporation (C, F, I, L). Note that Cpt induced γH2AX only in EdU incorporating cells. The very few EdU-negative cells expressing γH2AX, marked by the arrows, (F) are predominantly G2M cells constitutively expressing γH2AX (C). In contrast, in response to treatment with Mxt or Etp, H2AX was phosphorylated in both EdU incorporating and non-incorporating (G1 and G2M) cells. A correlation between EdU incorporation and induction of cH2AX by the Top inhibitors is shown in Panels F, I, and L; the correlation coefficient refers only to DNA replicating cells. Unlike in the case of Cpt-treatment, there are numerous EdU-negative cells expressing γH2AX following exposure to Mxt or Etp (I, L, marked by arrows). Insets in A, D, G, and J panels present DNA content frequency histograms from the respective cultures, with EdU incorporating cells marked in red. The dashed skewed lines in B, E, H, and K show the upper threshold for 97% of G1 and S-phase cells expressing γH2AX in the untreated culture (Ctrl).147
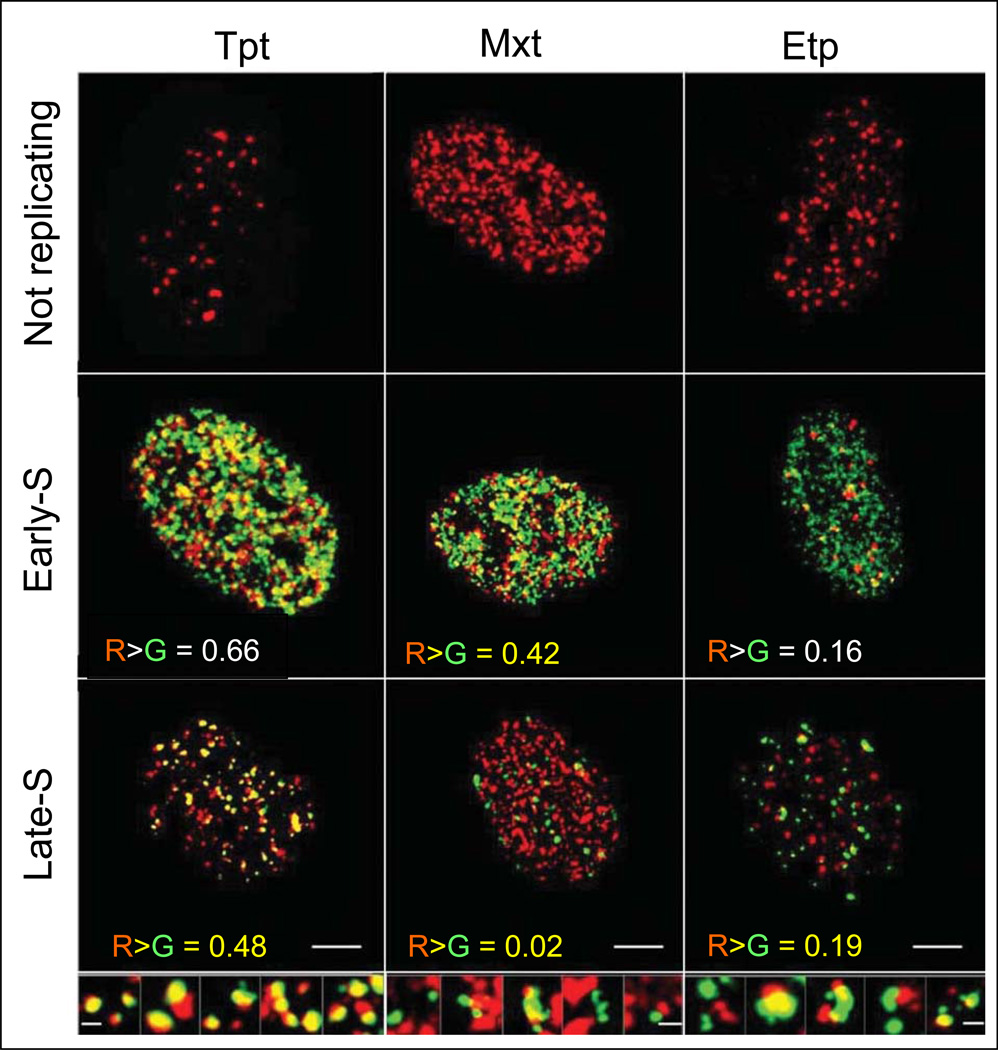
The approach to correlate actual DNA replication sites with the induction of γH2AX, has been further advanced using confocal microscopy to measure the spatial relationship in chromatin between the sites of DNA replication (“replication factories”) and the sites of γH2AX foci, the latter considered to be reporters of the induction of DSBs.72,73 Toward this end the cells were shortly exposed to EdU and subsequently treated with Tpt, Mxt, or Etp (Fig. 6).147 The incorporated EdU was detected using the green-fluorochrome (AlexaFluor 488)-tagged azide, whereas the Tpt-, Mxt-, or Etp- induced γH2AX foci were detected immunocytochemically using Ab conjugated to a red fluorescing dye (AlexaFluor 568). Among the DNA replicating cells the greatest degree of co-localization of γH2AX foci with DNA replication sites was seen in Tpt-treated cells, particularly during early-S (r>G = 0.66). The co-localization was also quite evident in the early-S phase Mxt-treated cells (R > G = 0.42) but was very low in late-S phase cells (R>G = 0.02). Etp-treated cells showed relatively low level of co-localization in both, early- (R>G = 0.16) and late-S phase (R>G = 0.19) phase cells.
Fig. 6. Spatial relationship in chromatin between the sites of EdU incorporation (“replication factories”) and the induction of γH2AX foci in A549 cells treated with Tpt, Mxt or Etp.
Confocal images of A549 nuclei that were briefly (30 min) exposed to EdU and then for additional 60 min treated with Tpt, Mxt, or Etp as described in detail elsewhere.147 The incorporation of EdU was detected utilizing AlexaFluor 488-tagged azide (green fluorescence) while γH2AX was detected using a secondary Ab labeled with AlexaFluor 568 (red fluorescence), as in Fig. 5.The size marker = 10µm. The top row shows nuclei of cells not incorporating EdU, the mid row - nuclei of cells in early-S phase, the bottom - nuclei of late-S phase. The distinction between early- and late-S phases is based on the characteristic differences in the pattern of DNA replication sites148. Enlarged images of selected foci/sites of DNA replication (late-S phase) are shown at the bottom of the respective columns; the size marker = 0.5 µM. The fraction of DNA replication sites that co-localize with γH2AX foci (yellow fluorescence) is shown in each panel.147
All these studies employing cytometry and confocal microscopy provided wealth of information regarding cell cycle phase differences in the mechanism by which these DNA topoisomerase inhibitors induce DDS and subsequently apoptosis. Specifically, the effects of Tpt/Cpt can be explained exclusively by stabilization of DNA-Top1 cleavable complexes. As originally postulated, the collisions of replication forks with these complexes generate DSBs.131–133 Rapid (10–30 min) ATM phosphorylation and induction of γH2AX exclusively in S-phase cells, at the sites of DNA replication factories (Fig. 6), is followed 2 h by Chk2 activation and 4 h by p53-Ser15 phosphorylation, all preferentially in S-phase cells.141,142 Apoptosis is also induced exclusively in S-phase cells.79,80 All these findings are consistent with the mechanism in which collisions of replication forks with the stabilized DNA-Top1 complexes are the sole mechanism of induction of DDS and subsequent apoptosis. The lack of evidence of DDS in G1 or G2M cells points out that collisions of the moving RNA polymerase with DNA-Top1 complexes during translation131 play no role in the possible induction of DSBs. Also, there is no involvement of ROS.135,138.
More complex is cell response to Mxt. While ATM activation and H2AX phosphorylation was seen in all phases of the cell cycle, the S-phase cells were preferentially undergoing apoptosis.80,141. Thus, collisions of replication forks with the stabilized DNA-Top2 complexes, which as evident from colocalization of replications sites and γH2AX foci (Fig. 6) appear to be much more extensive during the early– than the late- S phase (Fig. 6), seem to be responsible for induction of γH2AX and apoptosis. However, significant involvement of ROS in the induction of DDS (ATM, Chk2, p53 and H2AX phosphorylation) was also apparent in as much as the ROS scavenger NAC markedly suppressed DDS events, in all phases of the cell cycle.134,135 In the case of Etp, the collisions of replicating forks with the DNA-Top2 complexes seem to play relatively minor role, as evident with low level of co-localization of γH2AX (DSBs) and DNA replication sites (Fig. 6). Similar to Mxt, the involvement of ROS contributes to the induction of DDS in all phases of the cell cycle in cells treated with Etp. Because of DDS seen in G1 and G2M cells the possible role of RNA polymerase molecules colliding during translation with the stabilized DNA-Top2 complexes cannot be excluded as an additional factor contributing to the induction of DDS in cells treated with Mxt or Etp.
The induction of γH2AX and activation of ATM was detected in peripheral blood blasts of the patients with different types of acute leukemia treated with topoisomerase inhibitors, one hour after drug infusion. The number of the studied patients (20) and the number of the clinically non-responding ones (2) was too low to draw a conclusion whether the extent of treatment-induced DDS can be clinically prognostic.149 However, the findings demonstrate the feasibility of assessment of DDS during treatment of leukemia with drugs targeting DNA and its possible use as biomarker predictive of treatment effectiveness.149
2. DNA crosslinking agents
Exposure of cells to DNA crosslinking agents such as cisplatin or mitomycin C triggers DDS, revealed as induction of γH2AX, predominantly in S-phase cells.80,116,150–152 Compared to DNA topoisomerase inhibitors the kinetics of γH2AX induction by these agents is different. Unlike in the case of Mxt or Tpt which induce γH2AX already 10 min after cell exposure, the rise in expression of γH2AX occurs with a 2–3 h delay and is seen in all phases of the cell cycle.80 The repair process of DNA damage by cisplatin involves the NER mechanism, known to generate ssDNA breaks. A fraction of ssDNA lesions is known to be converted to DSBs. Furthermore, repair of cisplatin-induced damage also involves NHEJ.80 This mechanism may additionally contribute to the formation of ds DNA breaks and to the observed H2AX phosphorylation. Thus, when the primary cisplatin-induced lesions do not involve ds DNA breaks, but ds DNA breaks are formed during DNA repair, H2AX phosphorylation may indicate efficiency of the repair process. This mechanism is consistent with the observation that persistence of γH2AX foci, most likely reporting unrepaired DNA, is predictive of the cell-lethal effects of the treatment.152 It has been postulated that assessment of frequency of cells that retain γH2AX foci 24 h after treatment may predict tumor cell killing.152
Exposure of cells to very low concentrations of mitomycin C for a prolonged time period (6 days) was shown to trigger moderate level of DDS manifesting as elevation of γH2AX expression.150 Such treatment induced cell senescence and irreversible loss of reproductive capacity. Mutations of oncogenes and tumor suppressor genes that lead to activation of mTOR signaling are common to cancer. Because mTOR activation predisposes cells to undergo senescence153 it was postulated that cancers characterized by such mutations may be particularly sensitive to treatment with low doses of DNA damaging drugs (“metronomic chemotherapy”) that cause persistent replication stress.150 Analysis of DDS in response to persistent replication stress if found to be predictive of cell senescence my provide the means to identify tumors prone to undergo senescence when treated with low drug doses.
E. DDS as a marker of genotoxcicity. Screening for environmental genotoxins
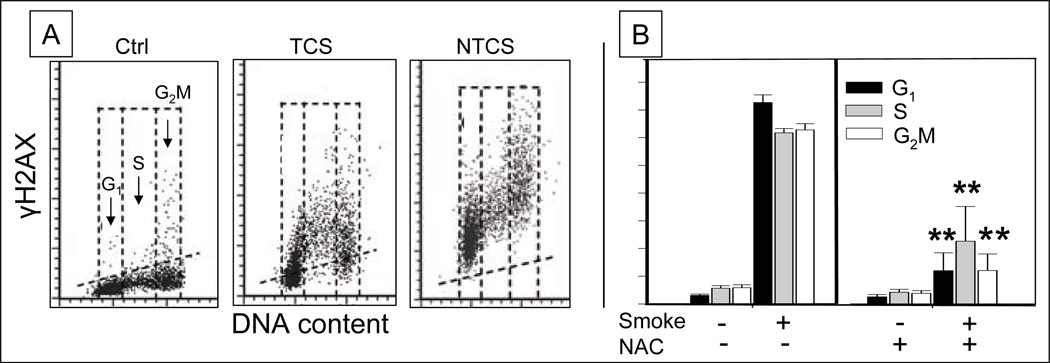
High sensitivity in detection of DNA damage combined with a possibility to correlate it with the cell cycle phase and/or apoptosis as offered by flow or imaging cytometry, contributed to wide application of this approach for screening of a variety of agents with respect to their genotoxic properties. One of first such applications was screening for genotoxic effects of cigarette smoke154–160. The induction of γH2AX foci, considered to be markers of DSBs, after exposure of normal human bronchial epithelial cells in vitro to cigarette smoke or smoke condensate, seen already 10 min after the exposure, was proposed as a new assay for carcinogens in tobacco.154 The induction γH2AX, measured by laser scanning cytometry, did not show significant differences vis-à-vis cell cycle phase.157(Fig. 7) Activation of ATM and other events of DDS paralleled H2AX phosphorylation.156 Human lung adenocarcinoma A549 cells, responded to cigarette smoke similarly as normal bronchial epithelial cells thereby providing an alternative experimental model for screening tobacco products.155–157 Interestingly, the smoke from tobacco-free cigarettes, which are commercially available as a substitute for tobacco cigarettes (with the aim to curtail smoking) were more potent in inducing γH2AX and ATM-S1981P, and thus potentially more carcinogenic, than tobacco cigarettes.159 (Fig.7). The induction of DDS by cigarettes smoke is prevented to a large degree by cells’ exposure to the ROS scavenger NAC.161 (Fig. 7) It is likely thus, that ROS mediate some of the genotoxic effects of tobacco smoke. Cigarette side-stream smoke was reported to induce DDS158 thereby confirming reports on genotoxic properties of secondhand smoke.162
Fig. 7. Induction of γH2AX in A549 cells after their exposure to cigarette smoke.
A. Bivariate (DNA content versus γH2AX) distributions representing mock-treated (Ctrl), cells exposed to smoke from standard tobacco cigarette (2R4F) (TCS), or smoke from tobacco-free (Bravo) cigarette (NTCS). A549 cells were exposed to the smoke for 10 min and the induction of γH2AX was assessed 1 h after the exposure159. The skewed dashed line shows the top level of γH2AX expression for 97% cells from Ctrl culture.
B. Reduction of the cigarette smoke induced γH2AX in A549 cells by co-treatment with the radicals’ scavenger n-acetyl-L-cysteine (NAC); p<0.01 marked by asterisks.161
As exemplified by analysis of genotoxicity of cigarette smoke, the cytometric analysis of DDS, particularly of γH2AX, provides high sensitivity, convenience and rapidity in detecting the genotoxic potential of different agents. This approach was recently used to assess genotoxicity of different types of nanoparticles.163–166 Of interest was the observation that the orthopedic particles made of metal (CoCr alloy) induced high level of γH2AX whereas the ceramic particles had no such effect.166 This observation provides further evidence of the advantage of the ceramic over metal alloys implants. Possible involvement of reactive oxidants in the induction of DDS by different types of nano- or micro-particles was a subject of particular inquiry in several studies.163–166
Utility of the γH2AX assay has been also tested in the assessment of genotoxic properties of agents used to manufacture plastics and epoxy resins such as bisphenol A and bisphenol F,167 ubiquitous pollutants such as polycyclic aromatic hydrocarbons,168 the non-ionic surfactans nonylphenol polyethoxylates that are widely used for industrial and household purposes,169 potential genotoxins in cosmetic preservatives,170 or several different pharmaceuticals frequently detected in freshwater environment.171
The finding that the nitrogen oxide-releasing aspirin induces γH2AX, activates ATM and leads to apoptosis of TK6 cells provides another example of detection of genotoxic effects by assessing induction of DDS.172 This is an important observation since so modified aspirin, which as it appears from these findings may have genotoxic properties, is being promoted as a novel anti-inflammatory agent. Attenuation of genotoxicity of the acridine orange mutagen 191 by chlorophyllin, likely via the mutagen-interceptor mechanism173 was also detected using the γH2AX assay.174 In a yet another application of the DDS assay it was observed that exposure of cells to Hoechst 33342 or DRAQ5 fluorochromes, the widely used supravital DNA probes, led to activation of ATM and Chk2 as well as phosphorylation of p53 and H2AX.175 This observation cautions on use of these probes, which themselves appear to have DNA damaging properties, in long-term cell viability studies.
The successful use of DDS assays for screening and examining genotoxic properties of different agents prompted several authors to propose this approach as the end-point for the assessment of genotoxicity176–178. Developments and applications of the γH2AX assay in the areas of radiation bio-dosimetry, evaluation or validation of new cancer drugs in clinical studies, chronic inflammation and environmental genotoxicity, have been recently highlighted in comprehensive and authoritative reviews.178,179
III. New cytometric techniques and approaches for analysis of DDS
Cytometry remains a vibrant and dynamic field of engineering where new technological advancements often lead to development of new bioassays probing molecular mechanisms in individual cells.180,181 In the recent years in particular we have seen many spectacular developments in the field of cytometry.180,181 The latest innovative platforms such as multispectral imaging cytometry182–184, mass cytometry185–188, Raman cytometry189–191, and microfluidic Lab-on-a-Chip192–195 solutions rapidly emerge as the next generation of tools for the analysis of DDS.
The multispectral imaging cytometry (Image Stream Flow Cytometry) revolutionized the conventional flow cytometry by integrating a fast in-flow imaging with the conventional layout of the flow cytometer.182–183 Such hybrid cytometric system is capable of simultaneously acquiring six spectrally decomposed images from each cell providing a valuable morphometric analysis.182–183 Multispectral cytometry has recently revealed distinct frequencies of γ-H2AX foci induction in DNA double strand break repair defective human cell lines.184
Mass cytometry is yet another proverbial quantum leap in expansion of analytical capabilities probing individual cells.185–187 Instead of using fluorescent probes cell staining is performed with stable isotopes of rare earth metals. Mass cytometry evades limitations of conventional flow cytometry such as the need for compensation of the emission spectra and offers a possibility for multiparametric analysis supreme to any other methodology currently available on the market.185,186 It has already been shown in the analysis of the cell cycle with the pRb phosphorylation [p-Rb(S807/S811)] as a biomarker.187 Moreover, mass cytometry it has been also shown capable of discriminating live vs. dying cells using platinum-containing chemotherapy drug cisplatin.188
Raman Spectral Flow Cytometry is another example of enabling bioanalytical technology of recent years.189–190 Non-resonant Raman spectroscopy is one of the most dynamically developing technologies based on inelastic scattering of incident photons and resulting vibrational excitation of the biomolecules.181–182 Non-invasive methods of detecting necrotic cell death using Raman Cytometry has recently been reported. Also the Raman micro-spectroscopy to non-invasively track cell cycle dynamics in living cells has also been recently investigated.189 The advent of surface enhanced Raman spectroscopy (SERS), which features signal enhancement of over 14 orders of magnitude over non-enhanced Raman scattering, opens up new avenues for real-time dynamic spectroscopy.180,190 Nanostructures within a hybrid SERS probe provide sufficient enhancement in the electromagnetic field to analyze single biomolecular species while using only low laser powers and short data acquisition times.180,190,191 The SERS flow cytometry was recently developed providing prospects of a massively multiplexed chemical cytometry on living cells.191
Capacitance cytometry is also a radical and pioneering approach to develop label-less flow cytometry.196 This technology is based on a linear relationship between the DNA content of eukaryotic cell and the change in capacitance that is evoked by the passage of individual cell across a 1-kHz electric field.196 It has been capable of rapid analysis of cell cycle and fractional DNA content (sub-G1) without a necessity to label or process cell samples.196 A capacitance flow cytometry can be particularly useful in the real-time monitoring of changes in cell cycle distribution offering non-exciting analytical advantages.196
The image-based cytometry represents a useful alternative to flow cytometry as it combines advantages of flow cytometry (FCM) and fluorescence image analysis (FIA).181,197 Automated fluorescence mosaic microscopy as a high-content imaging cytometry has recently been proven in systematic analysis of cells and subcellular protein locations such as dose-dependent changes in γH2AX response.197 Combined with automated machine learning algorithms it is reportedly a powerful tool for multivariate phenotypic profiling of individual cell nuclei and quantification of subnuclear spots using, optimized image processing tools, and supervised classification.197
Finally a number of emerging, microfluidic technologies for cell-based assays have recently been reported.192–195 The advent of so-called Lab-on-a-Chip technologies (LOC) is one of the most innovative cytometric approaches to the analysis of rare cells and organelles.193–194 It also provides exceptional evolutionary avenues for microfluidic flow cytometry (µFACS), cell sorting and revolutionary cell microarrays.181,193,195 Microfluidic flow cytometers and cell sorters require a greatly reduced number of cells per sample when compared with conventional analyzers.195 A great number of unconventional technologies have recently been proposed for a non-invasive and real-time cell analysis on microfluidic chips. These include real-time studies on a single cell level such as time-of-flight (TOF) optophoresis198, scanning thermal lens microscopy (TLM)199 and microfluidic living cell arrays192,200. The microfluidic platforms that can track single cell responses, multiparametically on a large scale are so far the only tool that can support personalized treatment in oncology and systems biology efforts and provide new vistas for a new generation of rationally designed anti-cancer drugs.193,194
Advances are also being made in analytical chemistry of bioassays. In this regard a high-throughput flow cytometric assays has recently been developed to measure the RPA-ssDNA formation during exposure to DNA damaging agents.201 Moreover, a new class of probes for cell imaging, such as functionalized gold or silver nanoparticles has recently been created.180 The binding of proteins to such nanoparticles changes the wavelength of plasmon resonance, which can be followed by dark-field microscopy.180 Moreover, the light-scattering intensity of a pair of nanoparticles is greater then for two separate particles, and increases even further for multi-particle assemblies.180 This feature has been recently utilized to study activation of caspase-3 using crown nanoparticle probes composed of a NeutrAvidin-coated gold-core nanoparticle with biotinylated gold satellite nanoparticles.202 It is postulated that small-cleaved satellite nanoparticles can move and allow analysis of single molecule trajectories.180 This could be beneficial to track movements of protein complexes involved in DDS.
IV. Interpretation of cytometric data
Detection and evaluation of the extent of DNA damage caused by different genotoxins that would be predictive of potential mutagenic and cytotoxic consequences is the primary goal of most applications of the DDS assays utilizing cytometry. Because DSBs represent the most deleterious DNA lesions, both in terms of their mutagenic potential as well as their role in signaling cell death pathways, it is of importance to correlate the particular events of the DDS with formation of DSBs. Initially, phosphorylation of H2AX on Ser139 was considered to be the specific marker of induction of DSBs. Indeed, in instances of ionizing radiation and radiomimetic agents that directly generate DSBs the intensity of expression of γH2AX, measured as the integrated γH2AX phospho-specific immunofluorescence per nucleus correlated well with the extent of DSBs and was prognostic of cell viability.
It should be cautioned, however, that expression of γH2AX after induction of DSBs is a kinetic event of rather short and often of variable duration. The rate of γH2AX dephosphorylation varies markedly depending the cell cycle phase, nature of the inducer of DSBs and rate of DNA repair.203 Expression of γH2AX however is not measured dynamically, in real time, but at the end-point at the time of cell harvest/fixation. Thus, there is an uncertainty as to whether the expression of γH2AX is being measured at the peak of response and thus whether intensity of the measured γH2AX immunofluorescence or frequency of γH2AX foci correlates with number of DSBs induced by DNA damaging agent.
There is strong evidence that H2AX phosphorylation, triggered by formation of DSBs, is mediated by ATM.73,75 One would expect therefore that concurrent ATM activation and induction of γH2AX may provide more definite evidence of the presence of DSBs than γH2AX alone. It should be stressed, however, that there are the instances when DDS, including concurrent phosphorylation of both H2AX and ATM, is triggered by DNA lesions other than DSBs. This has been seen to occur at mitosis204 or during premature chromosome condensation.23 Likewise, ATM and H2AX can be phosphorylated in the absence of DSBs during replication stress.46,47,81,204,205 Because of redundancy of PIKKs in phosphorylation of different substrates when one of these kinases is initially activated, activation of the other PIKK often follows. This was observed after induction of DNA damage by UV when activation of ATM was seen to be followed shortly (30 min) after activation of ATR.127 Thus, the expression of γH2AX per se or even when accompanied by ATM activation is not a conclusive proof of the presence of DSBs.
The presence of distinct γH2AX IF foci, each focus reported to represent a single DSB.71–73 appears to be more reliable reporter of DSBs. The number of individual IF foci per nucleus, especially if double-differentially labeled with ATM-S1981P and γH2AX, is thus likely to correspond to the count of DSBs.138,206 Quantification of the foci can be accomplished by image-assisted cytometric analysis utilizing, for example, a laser scanning cytometer (LSC)207 or similar instruments.207,209 Multiparameter analysis of such data make it possible to correlate the number of individual foci per nucleus with the cell cycle phase or with a marker of apoptosis.8,208 Image analysis was shown to be able to detect very low level of DNA damage, for example, as occurs in vivo in lymphocytes of patients subjected to computed tomography examinations.210 It should be noted, however, that accurate count of individual foci is limited to their relatively small number (<20) per nucleus. With greater numbers, proximity of foci to each other and possibility of their spatial overlap prevent accurate contouring and counting. Quantification of foci by LSC or other imaging instruments can be accurate, therefore, during analysis of constitutive DDS or for assessing environmental genotoxins at their low concentration, when relatively few foci are present per nucleus.
There are other issues that should also be considered when using image-assisted cytometry for foci quantification. Namely, the rates of phosphorylation and de-phosphorylation of H2AX and/or ATM in individual foci in the same cell may vary depending on the rate of DNA repair at each particular DSB site. Furthermore, when induction of DSBs are asynchronous within the cell (for example as is the case of progressive DNA damage following treatment with low doses of genotoxins) the “age” and thus the IF intensity of individual foci may vary. This variability in intensity of fluorescence of individual foci may complicate the setting optimal thresholds for contouring all foci.
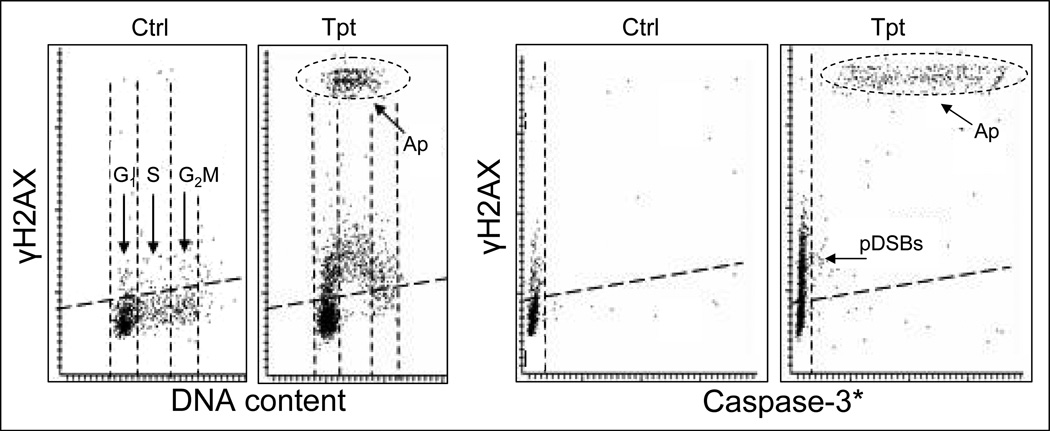
DNA fragmentation mediated by the caspase-activated DNase (CAD) is one of the hallmarks of apoptosis.211,212 Fragmentation of DNA triggers massive DDS, reflected by particularly extensive phosphorylation of H2AX.79,80 The question may be posed that if apoptosis is induced by agents damaging DNA how DDS triggered by these agents can be distinguished from the secondary DDS triggered by DNA fragmentation induced by CAD. The data shown in Fig. 8 illustrate such an instance, when cells were treated with Tpt which causes DSBs predominantly in S-phase cells, and also induces apoptosis.80 As is evident, the level of induction of γH2AX in apoptotic cells is by an order of magnitude greater than that of γH2AX induced in response to the primary DNA damage by Tpt. The high intensity of IF of the phospho-specific Ab recognizing phosphorylated H2AX is therefore one of the markers that can be used to identify apoptotic cells and distinguish primary DSBs from apoptotic DNA fragmentation. The other marker is the presence of activated caspase-3 (Fig. 8). It should be noted, however that at late stages of apoptosis γH2X is being degraded and the intensity of its IF drops.80 In still another approach that allows one to identify DDS triggered by the primary DNA damage and exclude apoptosis as the trigger of DNA fragmentation, a caspase inhibitor such as z-VAD-FMK can be used to selectively prevent DNA fragmentation.79,80
Fig. 8. Correlation between induction of γH2AX, activation of caspase-3, and cell cycle phase of HL-60 cells treated with Tpt for 3 h.
After 3 h of treatment with Tpt some cells (predominantly in S phase) undergo apoptosis. Apoptotic cells (Ap) show an order of magnitude higher intensity of γH2AX phospho-specific IF compared with non-apoptotic cells. The caspase-3 versus γH2AX bivariate distributions (right panels) reveal that some non-apoptotic (caspase-3 negative) cells have elevated expression of γH2AX reflecting the presence to the Tpt-induced primary DSBs (pDSBs).80
Conclusion
The major advantage of cytometry stems from the possibility to correlate expression of particular cell attributes with each other, on cell by cell basis. The correlation analysis is highly informative, often revealing mechanisms of interactions between the measured cell constituents. For many years cytometry has been successfully used for quantitative assessment of different cellular constituents, among them discrete parts of cell signaling pathways. By complementing findings obtained by the methods that rely on data obtained on cells measured in bulk, cytometry contributed a great deal towards revealing molecular mechanisms operating in normal and diseased cells, particularly in areas of cell growth, cell cycle progression and programmed cell death. The introduction of Abs that detect posttranslational modification of proteins, such as phosphorylation, opened an opportunity to explore actual cell signaling along different pathways in individual cells.213,214 High sensitivity in detection of DNA damage offered by analysis of γH2AX foci, approaching the limit of revealing of a single DSB per cell, with a possibility of a measurement of tens of thousands of cells per sample, is one of the major assets of such an approach. As is evident in this review, the multivariate analysis of different cell signaling events along DDS pathway, combined with probes of the cell cycle and apoptosis, yields a wealth of information pertinent to mechanism of action of a variety of genotoxic agents. Further information can be obtained by a combination of high resolution imaging offered by confocal microscopy with multivariate analysis by laser scanning cytometry. This is exemplified by analysis of spatio-temporal relationship between DNA replication and DNA damage sites in revealing mechanism of action of Top1 and Top2 inhibitors (Fig. 5 and 6). It can be expected that approaches along this way will yield further information leading to expansion of our understanding of the mechanisms of DNA damage and DNA repair.
Acknowledgement
Supported by NCI RO1 28 704 and by the Robert Andrew Welke Foundation for Cancer Research.(ZD, HDH, HZ); National Committee for Science (UMO-2011/01/B/NZ3/00609) and EU structural funds grant BMZ No, POIG.02.01.00-12-064/08 (JD, PR); Faculty Research Development Fund, University of Auckland (DW); Early Career Research Excellence Award, University of Auckland (DW); Ministry of Science & Innovation (MSI) New Zealand (DW); Biotechnology and Biological Sciences Research Council (BBSRC), Engineering and Physical Sciences Research Council (EPSRC) and Scottish Funding Council UK, funded under the RASOR (Radical Solutions for Researching the proteome) Program (DW).
Abbreviations
- 53BP1
p53-binding protein 1
- Ab
antibody
- AMPK
AMP-activated protein kinase
- ATM
Ataxia telangiectasia mutated
- ATR
ATM and Rad3-related protein kinase
- BRCA1
Breast Cancer 1
- CAD
caspase-activated DNase
- Chk1
checkpoint kinase 1
- Chk2
checkpoint kinase 2
- Cpt
camptothecin
- DDR
DNA damage response
- DDS
DNA damage signaling
- DSBs
DNA double-strand breaks
- DNA-PKcs
DNA dependent protein kinase
- EdU
5-ethynyl-2'-deoxyuridine
- Etp
etoposide
- FCM
flow cytometry
- FIA
Fluorescence image analysis
- µFACS
microfluidic flow cytometry
- HIF1
hypoxia-inducible factor 1
- HMG
high mobility group proteins
- HR
homologous recombination
- IF
immunofluorescence
- IR
ionizing radiation
- LSC
laser scanning cytometry
- MDC1
mediator of DNA damage checkpoint 1
- Mre11
Meiotic Recombination 11 Homolog A
- mTOR
mammalian target of rapamycin
- Mxt
mitoxantrone
- NAC
N-acetyl-L-cysteine
- NER
Nucleotide excision repair
- NHEJ
non-homologous end-joining
- NMR1
Nijmegen Breakage Syndrome 1
- PIKKs
phosphatidyl inositol 3' kinase-related kinases
- PML
Promyelocytic Leukemia protein
- ROS
reactive oxygen species
- SERS
surface enhanced Raman spectroscopy
- SMC1
Structural Maintenance of Chromosomes 1
- SSLs
DNA single-strand lesions
- TLM
thermal lens microscopy
- TOF
time-of-flight
- Top1
DNA topoisomerase I
- Top2
DNA topoisomerase II
- Tpt
topotecan
- UBC
ubiquitin-conjugating enzyme
REFERENCES
- 1.Bakkenist CJ, Kastan MB. DNA damage activates ATM through intermolecular autophosphorylation and dimer dissociation. Nature. 2003;421:499–506. doi: 10.1038/nature01368. [DOI] [PubMed] [Google Scholar]
- 2.Bakkenist CJ, Kastan MB. Initiating cellular stress responses. Cell. 2004;118:9–17. doi: 10.1016/j.cell.2004.06.023. [DOI] [PubMed] [Google Scholar]
- 3.Bonner WM, Redon CE, Dickey JS, Nakamura AJ, Sedelnikova OA, Solier S, Pommier Y. γH2AX and cancer. Nat. Rev. Cancer. 2008;8:957–966. doi: 10.1038/nrc2523. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Helt CE, Cliby WA, Keng PC, Bambara RA, O’Reilly MA. Ataxia telangiectasia mutated (ATM) and ATM and Rad3-related protein exhibit selective target specificities in response to different forms of DNA damage. J. Biol. Chem. 2005;280:1186–1192. doi: 10.1074/jbc.M410873200. [DOI] [PubMed] [Google Scholar]
- 5.Kastan MB. DNA damage responses: Mechanisms and roles in human disease. 2007 G.H.A. Cloves Memorial Award Lecture. Mol Cancer Res. 2008;6:517–524. doi: 10.1158/1541-7786.MCR-08-0020. [DOI] [PubMed] [Google Scholar]
- 6.Nakamura AJ, Rao VA, Pommier Y, Bonner WM. The complexity of phosphorylated H2AX foci formation and DNA repair assembly at DNA double-strand breaks. Cell Cycle. 2010;9:389–398. doi: 10.4161/cc.9.2.10475. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Lord CJ, Ashworth A. The DNA damage response and cancer therapy. Nature. 2012;481:287–294. doi: 10.1038/nature10760. [DOI] [PubMed] [Google Scholar]
- 8.Darzynkiewicz Z, Traganos F, Zhao H, Halicka HD, Skommer J, Wlodkowic D. Analysis of individual molecular events of DNA damage response by flow and image-assisted cytometry. Meth Cell Biol. 2011;103:15–148. doi: 10.1016/B978-0-12-385493-3.00006-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Murga M, Jaco I, Soria R, Martinez-Pastor B, Cuadrado M, Yang S-M, Blasco MA, Skoultchi AI, Fernandez-Capetillo O. Global chromatin compaction limits the strength of the DNA damage response. J Cell Biol. 2007;178:1101–1108. doi: 10.1083/jcb.200704140. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Pandita TK, Richardson C. Chromatin remodeling finds its place in the DNA double-strand breaks. Nucleic Acids Res. 2009;37:1363–1377. doi: 10.1093/nar/gkn1071. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Rouleau M, Aubin RA, Poirier GG. Poly(ADP-ribosyl)ated chromatin domains: access granted. J. Cell Sci. 2004;117:815–825. doi: 10.1242/jcs.01080. [DOI] [PubMed] [Google Scholar]
- 12.Ziv Y, Bielopolski D, Galanty Y, Lukas C, Taya Y, Schultz DC, Lukas J, Bekker-Jensen S, Bartek J, Shiloh Y. Chromatin relaxation in response to DNA double-strand breaks is modulated by a novel ATM- and KAP-1 dependent pathway. Na. Cell Biol. 2006;8:870–876. doi: 10.1038/ncb1446. [DOI] [PubMed] [Google Scholar]
- 13.Cann KL, Dellaire G. Heterochromatin and the DNA damage response: the need to relax. Biochem Cell Biol. 2011;89:45–60. doi: 10.1139/O10-113. [DOI] [PubMed] [Google Scholar]
- 14.Marko JF. Linking topology to large DNA molecules. Physica A. 2010;389:2997–3001. doi: 10.1016/j.physa.2010.01.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Gerlitz G, Bustin M. Nucleosome binding proteins potentiate ATM activation and DNA damage response by modifying chromatin. Cell Cycle. 2009;8:1641–1644. doi: 10.4161/cc.8.11.8637. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Kim YC, Gerlitz G, Furusawa T, Catez F, Nussenzweig A, Oh KS, Kraemer KH, Shiloh Y, Bustin M. Activation of ATM depends on chromatin interactions occurring before induction of DNA damage. Nat Cell Biol. 2009;11:92–96. doi: 10.1038/ncb1817. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Sinha M, Peterson CL. Chromatin dynamics during repair of chromosomal DNA double-strand breaks. Epigenomics. 2009;1:371–385. doi: 10.2217/epi.09.22. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Shiloh Y, Shema E, Moyal L, Oren M. RNF20-RNF40: A ubiquitin-driven link between gene expression and the DNA damage response. FEBS Lett. 2011;585:2795–2802. doi: 10.1016/j.febslet.2011.07.034. [DOI] [PubMed] [Google Scholar]
- 19.Zhang Q, Wang Y. High mobility group proteins and their post-transcriptional modifications. Biochim Biophys Acta. 2008;1794:1159–1166. doi: 10.1016/j.bbapap.2008.04.028. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Lim J, Catez F, Birger Y, West K, Prymakowska-Bosak M, Postnikov MV, Bustin M. Chromosomal protein HMGN1 modulates histone H3 phosphorylation. Mol Cell. 2004;15:573–584. doi: 10.1016/j.molcel.2004.08.006. [DOI] [PubMed] [Google Scholar]
- 21.Misteli T, Soutoglou E. The emerging role of nuclear architecture in DNA repair and genome maintenance. Mol Cell Biol. 2009;10:243–254. doi: 10.1038/nrm2651. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Juan G, Traganos F, James WM, Ray JM, Roberge M, Sauve DM, Darzynkiewicz Z. Histone H3 phosphorylation and expression of cyclins A and B1 measured in individual cells during their progression through G2 and mitosis. Cytometry. 1998;32:71–77. doi: 10.1002/(sici)1097-0320(19980601)32:2<71::aid-cyto1>3.0.co;2-h. [DOI] [PubMed] [Google Scholar]
- 23.Huang X, Kurose A, Tanaka T, Traganos F, Dai W, Darzynkiewicz Z. Sequential phosphorylation of Ser-10 on histone H3 and Ser-139 on histone H2AX and ATM activation during premature chromosome condensation: Relationship to cell-cycle and apoptosis. Cytometry A. 2006;69A:222–229. doi: 10.1002/cyto.a.20257. [DOI] [PubMed] [Google Scholar]
- 24.Ikura T, Tashiro S, Kakino A, Shima H, Jacob N, Amunugama R, Yoder K, Izumi S, Kuraoka I, Tanaka K, Kimura H, Ikura M, Nishikubo S, Ito T, Muto A, Miyagawa K, Takeda S, Fishel R, Igarashi K, Kamiya K. DNA damage dependent acetylation and ubiquitination of H2AX enhances chromatin dynamics. Mol Cell Biol. 2007;27:7028–7040. doi: 10.1128/MCB.00579-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Kruhlak MJ, Celeste A, Nussenzweig A. Spatio-temporal dynamics of chromatin containing DNA breaks. Cell Cycle. 2006;5:1910–1912. doi: 10.4161/cc.5.17.3169. [DOI] [PubMed] [Google Scholar]
- 26.Rubi CP, Milner J. p53 is a chromatin accessibility factor for nucleotide excision repair of DNA damage. The EMBO J. 2003;22:975–986. doi: 10.1093/emboj/cdg082. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Wang J, Chin MY, Li G. The novel tumor suppressor p33ING2 enhances nucleotide excision repair via inducement of histone H4 acetylation and chromatin relaxation. Cancer Res. 2006;66:1906–1911. doi: 10.1158/0008-5472.CAN-05-3444. [DOI] [PubMed] [Google Scholar]
- 28.Abraham RT, Tibbetts RS. Guiding ATM to broken DNA. Science. 2005;308:510–511. doi: 10.1126/science.1112069. [DOI] [PubMed] [Google Scholar]
- 29.Downs JA, Cote J. Dynamics of chromatin during the repair of DNA double-strand breaks. Cell Cycle. 2005;4:1373–1376. doi: 10.4161/cc.4.10.2108. [DOI] [PubMed] [Google Scholar]
- 30.Kitagawa R, Kastan MB. The ATM-dependent DNA damage signaling pathway. Cold Spring Harb. Symp Quant Biol. 2005;70:99–109. doi: 10.1101/sqb.2005.70.002. [DOI] [PubMed] [Google Scholar]
- 31.Paull TT, Lee JH. The Mre11/Rad50/Nbs1 complex and its role as a DNA-double strand break sensor for ATM. Cell Cycle. 2005;4:737–740. doi: 10.4161/cc.4.6.1715. [DOI] [PubMed] [Google Scholar]
- 32.Cuadrado M, Martinez-Pastor B, Fernandez-Capetillo O. ATR activation in response to ionizing radiation: still ATM territory. Cell Div. 2006;1:7. doi: 10.1186/1747-1028-1-7. http://www.celldiv.com/content/1/1/7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Hill R, Lee PWK. The DNA-dependent protein kinase (DNA-PK). More than just a case of making ends meet? Cell Cycle. 2010;9:3460–3469. doi: 10.4161/cc.9.17.13043. [DOI] [PubMed] [Google Scholar]
- 34.Lovejoy CA, Cortez D. Common mechanisms of PIKK regulation. DNA Repair. 2009;8:1004–1008. doi: 10.1016/j.dnarep.2009.04.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Li L, Zou L. Sensing signaling and responding to DNA damage: organization of checkpoint pathways in mammalian cells. J. Cell Biochem. 2005;94:298–306. doi: 10.1002/jcb.20355. [DOI] [PubMed] [Google Scholar]
- 36.Shiloh Y. ATM and related protein kinases: safeguarding genome integrity. Nat Rev Cancer. 2003;22:5834–5868. doi: 10.1038/nrc1011. [DOI] [PubMed] [Google Scholar]
- 37.Sun Y, Jiang X, Chen S, Fernandes N, Price BD. A role for the Tip60 histone acetyltransferase in the acetylation and activation of ATM. Proc Natl Acad Sc. USA. 2005;102:3182–13187. doi: 10.1073/pnas.0504211102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Lee JH, Paull TT. ATM activation by DNA double-strand breaks through the Mre11-Rad50-Nbs1 complex. Science. 2005;308:551–554. doi: 10.1126/science.1108297. [DOI] [PubMed] [Google Scholar]
- 39.Lee JH, Gospodarzi AA, Jeggo PA, Paull TT. 53BP1 promotes ATP activity through direct interactions with the MRN complex. EMBO J. 2010;29:574–585. doi: 10.1038/emboj.2009.372. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Guo Z, Deshpande R, Paull TT. ATM activation in the presence of oxidative stress. Cell Cycle. 2010;9:4805–4811. doi: 10.4161/cc.9.24.14323. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Guo Z, Kozlov S, Lavin MF, Person MD, Paull TT. ATM activation by oxidative stress. Science. 2010;330:517–521. doi: 10.1126/science.1192912. [DOI] [PubMed] [Google Scholar]
- 42.Ditch S, Paull TT. The ATM protein kinase and cellular redox signaling: beyond the DNA damage response. Trends Biochem Sci. 2012;37:15–22. doi: 10.1016/j.tibs.2011.10.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Kitagawa R, Bakkenist CJ, McKinnon PJ, Kastan MB. Phosphorylation of SMC1 is a critical downstream event in the ATM-ANBS1-BRCA1 pathway. Genes Dev. 2004;18:1423–1438. doi: 10.1101/gad.1200304. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Yuan SS, Lee SE, Chen G, Song M, Tomlinson GE, Lee FY. BRCA2 is required for ionizing radiation-induced assembly of Rad51 complex in vivo. Cancer Res. 1999;59:3547–3551. [PubMed] [Google Scholar]
- 45.Stucki M, Jackson SP. MDC1/NFBD1: a key regulator of the DNA damage response in higher eukaryotes. DNA Repair (Amst.) 2005;3:953–957. doi: 10.1016/j.dnarep.2004.03.007. [DOI] [PubMed] [Google Scholar]
- 46.Kurose A, Tanaka T, Huang X, Traganos F, Dai W, Darzynkiewicz Z. Effects of hydroxyurea and aphidicolin on phosphorylation of ATM on Ser 1981 and histone H2AX on Ser 139 in relation to cell cycle phase and induction of apoptosis. Cytometry A. 2006;69A:212–221. doi: 10.1002/cyto.a.20241. [DOI] [PubMed] [Google Scholar]
- 47.Kurose A, Tanaka T, Huang X, Traganos F, Darzynkiewicz Z. Synchronization in the cell cycle by inhibitors of DNA replication induces histone H2AX phosphorylation, an indication of DNA damage. Cell Prolif. 2006;39:231–240. doi: 10.1111/j.1365-2184.2006.00380.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Ward M, Minn K, Chen J. UV-induced ataxia-telangiectasia-mutated and Rad3-related (ATR) activation requires replication stress. J Bio. Chem. 2004;279:9677–9680. doi: 10.1074/jbc.C300554200. [DOI] [PubMed] [Google Scholar]
- 49.Smith GC, Jackson SP. The DNA-dependent protein kinase. Genes Dev. 1999;13:916–934. doi: 10.1101/gad.13.8.916. [DOI] [PubMed] [Google Scholar]
- 50.Smith GR. How homologous recombination is initiated: unexpected evidence for single-strand nicks from v(d) site-specific recombination. Cell. 2004;17:146–148. doi: 10.1016/s0092-8674(04)00338-1. [DOI] [PubMed] [Google Scholar]
- 51.Samper E, Goytisolo EA, Slijepcevic P, van Buul PP, Blasco MA. Mammalinan Ku86 protein prevents telomeric fusions independently of the length of TTAGGG repeats and the G-strand overhang. EMBO Rep. 2000;1:244–252. doi: 10.1093/embo-reports/kvd051. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Chan SD, Luedke G, Valer M, Buhlmann C, Preckel T. Cytometric analysis of protein expression and apoptosis in human primary cells with a novel microfluidic chip-based system. Cytometry A. 2003;55A:119–125. doi: 10.1002/cyto.a.10070. [DOI] [PubMed] [Google Scholar]
- 53.Wakeman TP, Kim WJ, Callens S, Chu A, Brown KD, Xu B. The ATM-SMC1 pathway is essential for activation of chromium [VI]-induced S-phase checkpoint. Mutat Res. 2004;554:241–251. doi: 10.1016/j.mrfmmm.2004.05.006. [DOI] [PubMed] [Google Scholar]
- 54.Ahn JY, Li X, Davis HL, Canman CE. Phosphorylation of threonine 68 promotes oligomerization and autophosphorylation of Chk2 protein kinase via the forkhead-associated domain. J Biol Chem. 2002;277:19389–19395. doi: 10.1074/jbc.M200822200. [DOI] [PubMed] [Google Scholar]
- 55.Ahn J, Urist M, Prives C. The Chk2 protein kinase. DNA Repair. 2004;3:1039–1047. doi: 10.1016/j.dnarep.2004.03.033. [DOI] [PubMed] [Google Scholar]
- 56.Matsuoka S, Rotman G, Ogawa A, Shiloh K, Tamai SJ, Elledge SJ. Ataxia telangiectasia-mutated phosphorylates Chk2 in vivo and in vitro. Proc. Natl Acad Sci USA. 2000;97:10289–10394. doi: 10.1073/pnas.190030497. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Zhou BB, Elledge SJ. The DNA response: putting checkpoints in perspective. Nature. 2000;408:433–439. doi: 10.1038/35044005. [DOI] [PubMed] [Google Scholar]
- 58.Li J, Stern DF. DNA damage regulates Chk2 association with chromatin. J. Biol Chem. 2005;280:37948–37956. doi: 10.1074/jbc.M509299200. [DOI] [PubMed] [Google Scholar]
- 59.Stevens C, Smith L, La Thangue NB. Chk2 activates E2F-1 in response to DNA damage. Nat Cell Biol. 2003;5:4465–4479. doi: 10.1038/ncb974. [DOI] [PubMed] [Google Scholar]
- 60.Lin J, Reichner C, Wu X, Levine AJ. Analysis of wild-type and mutant p21WAF-1 gene activities. Mol Cell Biol. 1996;16:1786–1793. doi: 10.1128/mcb.16.4.1786. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Lee JS, Collins KM, Brown AL, Lee CH, Chung JH. hCds1-mediated phosphorylation of BRCA1 regulates the DNA damage response. Nature. 2000;404:201–204. doi: 10.1038/35004614. [DOI] [PubMed] [Google Scholar]
- 62.Yang S, Kuo C, Bisi JE, Kim MK. PML-dependent apoptosis after DNA damage is regulated by the checkpoint kinase hCdsa/Chk2. Nat Cell Biol. 2002;4:865–870. doi: 10.1038/ncb869. [DOI] [PubMed] [Google Scholar]
- 63.Tan Y, Raychaudhuri P, Costa RH. Chk2 mediates stabilization of the FoxM1 transcription factor to stimulate expression of DNA repair genes. Mol Cell Biol. 2007;27:1007–1016. doi: 10.1128/MCB.01068-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Boutros R, Dozier C, Ducommun B. The when and wheres of CDC25 phosphatases. Curr Opin Cell Biol. 2006;18:185–119. doi: 10.1016/j.ceb.2006.02.003. [DOI] [PubMed] [Google Scholar]
- 65.Boutros R, Lobjois V, Ducommun B. CDC25 phosphatases in cancer cells: key players? Good targets? Nat Rev Cancer. 2007;7:495–507. doi: 10.1038/nrc2169. [DOI] [PubMed] [Google Scholar]
- 66.Rudolph J. Cdc25 phosphatases: Structure, specificity and mechanism. Biochemistry. 2007;46:3595–3604. doi: 10.1021/bi700026j. [DOI] [PubMed] [Google Scholar]
- 67.Thatcher TH, Gorovsky MA. Phylogenetic analysis of the core histones H2A, H2B, H3 and H4. Nucleic Acids Res. 1994;22:174–183. doi: 10.1093/nar/22.2.174. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Bassing CH, Suh H, Ferguson DO, Chua KF, Manis J, Eckersdorff M, Gleason M, Bronson R, Lee C, Alt FW. (Histone H2AX: A dosage-dependent suppressor of oncogenic translocations in tumors. Cell. 2003;114:359–370. doi: 10.1016/s0092-8674(03)00566-x. [DOI] [PubMed] [Google Scholar]
- 69.Lukas J, Lukas C, Bartek J. More than just a focus: The chromatin response to DNA damage and its role in genome integrity maintenance. Nat Cell Biol. 2011;13:1161–1169. doi: 10.1038/ncb2344. [DOI] [PubMed] [Google Scholar]
- 70.Celeste A, Difilippantonio S, Fernandez-Capetillo O, Pilch DR, Sedelnikova O, Eckhaus M, Ried T, Bonner WM, Nussenzweig A. H2AX haploinsufficiency modifies genomic stability and tumor susceptibility. Cell. 2003;114:371–383. doi: 10.1016/s0092-8674(03)00567-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Rogakou EP, Boon C, Redon C, Bonner WM. Megabase chromatin domains involved in DNA double-strand breaks in vivo. J Cell Biol. 1999;146:905–916. doi: 10.1083/jcb.146.5.905. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Rogakou EP, Pilch DR, Orr AH, Ivanova VS, Bonner WM. DNA double-stranded breaks induce histone H2AX phosphorylation on serine 139. J. Biol. Chem. 1998;273:5858–5868. doi: 10.1074/jbc.273.10.5858. [DOI] [PubMed] [Google Scholar]
- 73.Sedelnikova OA, Rogakou EP, Panuytin IG, Bonner WM. Quantitive detection of 125IUdr-induced DNA double-strand breaks with γ-H2AX antibody. Ratiation Res. 2002;158:486–492. doi: 10.1667/0033-7587(2002)158[0486:qdoiid]2.0.co;2. [DOI] [PubMed] [Google Scholar]
- 74.Anderson L, Henderson C, Adachi Y. Phosphorylation and rapid relocalization of 53BP1 to nuclear foci upon DNA damage. Mol Cell Biol. 2001;21:1719–1729. doi: 10.1128/MCB.21.5.1719-1729.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Burma S, Chen BP, Murphy M, Kurimasa A, Chen DJ. ATM phosphorylates histone H2AX in response to DNA double-strand breaks. J Biol Chem. 2001;276:42462–42467. doi: 10.1074/jbc.C100466200. [DOI] [PubMed] [Google Scholar]
- 76.Furuta T, Takemura H, Liao Z-Y, Aune GJ, Redon C, Sedelnikova OA, Pilch DR, Rogakou EP, Celeste A, Chen HT, Nussenzweig A, Aladjem MI, Bonner WM, Pommier Y. Phosphorylation of histone H2AX and activation of Mre11, Rad50, and Nbs1 in response to replication-dependent DNA double-strand breaks induced by mammalian topoisomerase I cleavage complexes. J Biol Chem. 2003;278:20303–20312. doi: 10.1074/jbc.M300198200. [DOI] [PubMed] [Google Scholar]
- 77.Park EJ, Chan DW, Park JH, Oettinger MA, Kwon J. DNA-PK is activated by nucleosomes and phosphorylated H2AX within the nucleosomes in an acetylation-dependent manner. Nucleic Acids Res. 2003;31:6819–6827. doi: 10.1093/nar/gkg921. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Modesti M, Kanaar R. DNA repair: spot(light)s on chromatin. Curr Biol. 2001;11:R229–R232. doi: 10.1016/s0960-9822(01)00112-9. [DOI] [PubMed] [Google Scholar]
- 79.Huang X, Traganos F, Darzynkiewicz Z. DNA damage induced by DNA topoisomerase I- and topoisomerase II- inhibitors detected by histone H2AX phosphorylation in relation to the cell cycle phase and apoptosis. Cell Cycle. 2003;2:614–619. [PubMed] [Google Scholar]
- 80.Huang X, Okafuji M, Traganos F, Luther E, Holden E, Darzynkiewicz Z. Assessment of histone H2AX phosphorylation induced by DNA topoisomerase I and II inhibitors topotecan and mitoxantrone and by DNA crosslinking agent cisplatin. Cytometry A. 2004;58A:99–110. doi: 10.1002/cyto.a.20018. [DOI] [PubMed] [Google Scholar]
- 81.Cleaver JE. γH2AX: biomarker of damage or functional participant in DNA repair “all that glitters is not gold”. Photochem Photobiol. 2011;87:1230–1239. doi: 10.1111/j.1751-1097.2011.00995.x. [DOI] [PubMed] [Google Scholar]
- 82.Vilenchik MM, Knudson AG. Endogenous DNA double-strand breaks: Production, fidelity of repair, and induction of cancer. Proc Natl Acad Sci USA. 2003;100:12871–12876. doi: 10.1073/pnas.2135498100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.Beckman KB, Ames BN. Oxidative decay of DNA. J Biol Chem. 1997;272:13300–13305. doi: 10.1074/jbc.272.32.19633. [DOI] [PubMed] [Google Scholar]
- 84.Pastwa E, Blasiak J. Nonhomologous DNA end joining. Acta Biochim Pol. 2003;50:891–908. [PubMed] [Google Scholar]
- 85.Gorbunova V, Seluanov A. Making ends meet in old age: DSB repair and aging. Mech Ageing Dev. 2005;126:621–628. doi: 10.1016/j.mad.2005.02.008. [DOI] [PubMed] [Google Scholar]
- 86.Pham NA, Robinson BH, Hedley DW. Simultaneous detection of mitochondrial respiratory chain activity and reactive oxygen in digitonin-permeabilized cells using flow cytometry. Cytometry. 2000;41:245–251. doi: 10.1002/1097-0320(20001201)41:4<245::aid-cyto2>3.0.co;2-#. [DOI] [PubMed] [Google Scholar]
- 87.Cheng TJ, Kao HP, Chan CC, Chang WP. Effects of ozone on DNA single-strand breaks and 8-oxoguanine formation in A549 cells. Environ Res. 2003;93:298–284. doi: 10.1016/s0013-9351(03)00041-0. [DOI] [PubMed] [Google Scholar]
- 88.Olive PL, Durand RE, Banath JP, Johnston PJ. Analysis of DNA damage in individual cells. Methods Cell Biol. 2001;64:235–249. doi: 10.1016/s0091-679x(01)64016-0. [DOI] [PubMed] [Google Scholar]
- 89.Huang X, Tanaka T, Kurose A, Traganos F, Darzynkiewicz Z. Constitutive histone H2AX phosphorylation on Ser-139 in cells untreated by genotoxic agents is cell-cycle phase specific and attenuated by scavenging reactive oxygen species. Int J Oncol. 2006;29:495–501. [PubMed] [Google Scholar]
- 90.Tanaka T, Halicka HD, Huang X, Traganos F, Darzynkiewicz Z. Constitutive histone H2AX phosphorylation and ATM activation, the reporters of DNA damage by endogenous oxidants. Cell Cycle. 2006;5:1940–1945. doi: 10.4161/cc.5.17.3191. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91.Tanaka T, Kajstura M, Halicka HD, Traganos F, Darzynkiewicz Z. Constitutive histone H2AX phosphorylation and ATM activation are strongly amplified during mitogenic stimulation of lymphocytes. Cell Prolif. 2007;40:1–13. doi: 10.1111/j.1365-2184.2007.00417.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92.Zhao H, Tanaka T, Halicka HD, Traganos F, Zarebski M, Dobrucki J, Darzynkiewicz Z. Cytometric assessment of DNA damage by exogenous and endogenous oxidants reports the aging-related processes. Cytometry A. 2007;71A:905–914. doi: 10.1002/cyto.a.20469. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 93.Tanaka T, Kurose A, Huang X, Traganos F, Dai W, Darzynkiewicz Z. Extent of constitutive histone H2AX phosphorylation on Ser-139 varies in cells with different TP53 status. Cell Prolif. 2006;39:313–323. doi: 10.1111/j.1365-2184.2006.00387.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 94.Wu H, Zhang H, Wang C, Wu Y, Xie J, Jin X, Yang J, Ye J. Genoprotective effect of hyaluronic acid against benzalkonium chloride-induced DNA damage in human corneal epithelial cells. Mol Vis. 2011;17:3364–3370. [PMC free article] [PubMed] [Google Scholar]
- 95.Bulterijs S. Metformin as a geroprotector. Rejuvenation Res. 2011;14:469–482. doi: 10.1089/rej.2011.1153. [DOI] [PubMed] [Google Scholar]
- 96.Nair-Shaliliker V, Armstrong BK, Fenech M. Does vitamin D protect against DNA damage? Mutat Res. 2012;733:50–57. doi: 10.1016/j.mrfmmm.2012.02.005. [DOI] [PubMed] [Google Scholar]
- 97.Zhao H, Tanaka T, Mitlitski V, Heeter J, Balazs EA, Darzynkiewicz Z. Protective effect of hyaluronate on oxidative DNA damage in WI-38 and A549 cells. Int J Oncol. 2008;32:1159–1169. doi: 10.3892/ijo_32_6_1159. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 98.Halicka HD, Zhao H, Li J, Traganos F, Zhang S, Lee M, Darzynkiewicz Z. Genome protective effect of metformin as revealed by reduced level of constitutive DNA damage signaling. Aging (Albany) 2011;3:1028–1038. doi: 10.18632/aging.100397. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 99.Halicka HD, Zhao H, Li J, Traganos DF, Studzinski G, Darzynkiewicz Z. Attenuation of constitutive DNA damage signaling by 1,25-dihydroxyvitamin D3. Aging (Albany) 2012;4:270–278. doi: 10.18632/aging.100450. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 100.Massudi H, Grant R, Braidy N, Guest J, Farnsworth B, Gullemin GJ. Age-associated changes in oxidative stress and NAD+ metabolism in human tissue. PLoS ONE. 2012;7:e42357. doi: 10.1371/journal.pone.0042357. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 101.Oka K, Tanaka T, Enoki T, Yoshimura K, Ohshima M, Kubo M, Murakami T, Gondou T, Minami Y, Takemoto Y, Harada E, Tsushimi T, Li T-S, Traganos F, Darzynkiewicz Z, Hamano K. DNA damage signaling is activated during cancer progression in human colorectal carcinoma. Cancer Biol Ther. 2010;9:246–252. doi: 10.4161/cbt.9.3.10751. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 102.Negelkerke A, van Kuijk SJ, Sweep FC, Nagtegaal ID, Hoogerbrugge N, Martens JW, Timmermans MA, van Laarhoven HW, Bussink J, Span PN. Constitutive expression of γ-H2AX has prognostic relevance in triple negative breast cancer. Radiother Oncol. 2011;101:39–45. doi: 10.1016/j.radonc.2011.07.009. [DOI] [PubMed] [Google Scholar]
- 103.Bartkova J, Hamerlik P, Stockhausen MT, Ehrmann J, Hlobilkova A, Laursen H, Kalita O, Kolar Z, Poulsen HS, Broholm H, Lukas J, Bartek J. Replication stress and oxidative damage contribute to aberrant constitutive activation of DNA damage signaling in human gliomas. Oncogene. 2010;29:5095–5102. doi: 10.1038/onc.2010.249. [DOI] [PubMed] [Google Scholar]
- 104.Walters DK, Tschumper RC, Arendt BK, Huddleston PM, Henderson KJ, Dispenzieri A, Jelinek DF. Evidence for ongoing DNA damage in multiple myeloma cells revealed by constitutive phosphorylation of H2AX. Leukemia. 2011;25:1344–1353. doi: 10.1038/leu.2011.94. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 105.Cavalier C, Didier C, Prade N, Mansat-De Mas V, Manenti S, Recher C, Demur C, Ducommun B. Constitutive activation of the DNA damage signaling pathway in acute myeloid leukemia with complex karyotype: potential importance for checkpoint targeting therapy. Cancer Res. 2009;69:8652–8661. doi: 10.1158/0008-5472.CAN-09-0939. [DOI] [PubMed] [Google Scholar]
- 106.Halicka HD, Zhao H, Podhorecka M, Traganos F, Darzynkiewicz Z. Cytometric detection of chromatin relaxation, an early reporter of DNA damage response. Cell Cycle. 2009;8:2233–2237. doi: 10.4161/cc.8.14.8984. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 107.Zhao H, Traganos F, Albino AP, Darzynkiewicz Z. Oxidative stress induces cell cycle-dependent Mre11 recruitment, ATM and Chk2 activation and histone H2AX phosphorylation. Cell Cycle. 2008;7:1490–1495. doi: 10.4161/cc.7.10.5963. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 108.Zhao H, Dobrucki J, Rybak P, Traganos F, Halicka HD, Darzynkiewicz Z. Induction of DNA damage signaling by oxidative stress in relation to DNA replication as detected using the “Click Chemistry”. Cytometry A. 2011;79A:897–902. doi: 10.1002/cyto.a.21137. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 109.Allen C, Ashley AK, Hromas R, Nickoloff JA. More forks on the road to replication stress recovery. J Mol Cell Biol. 2011;3:4–12. doi: 10.1093/jmcb/mjq049. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 110.Banath JP, MacPhail SH, Olive PL. Radiation sensitivity, H2AX phosphorylation, and kinetics of repair of DNA strand breaks in irradiated cervical cancer cells. Cancer Res. 2004;64:7144–7148. doi: 10.1158/0008-5472.CAN-04-1433. [DOI] [PubMed] [Google Scholar]
- 111.MacPhail SH, Banath JP, Yu TY, Chu EH, Lambur H, Olive PL. Expression of phosphorylated histone H2AX in cultured cell lines following exposure to X-rays. Int J Radiat Biol. 2003;79:351–358. doi: 10.1080/0955300032000093128. [DOI] [PubMed] [Google Scholar]
- 112.Banath JP, Olive PL. Expression of phosphorylated histone H2AX as a surrogate of cell killing by drugs that create DNA double-strand breaks. Cancer Res. 2003;63:4347–4350. [PubMed] [Google Scholar]
- 113.MacPhail SH, Banath JP, Yu Y, Chu E, Olive PL. Cell cycle-dependent expression of phosphorylated histone H2AX: reduced expression in unirradiated but not X-irradiated G1-phase cells. Radiat Res. 2003;159:759–767. doi: 10.1667/rr3003. [DOI] [PubMed] [Google Scholar]
- 114.Olive PL, Banath JP. Phosphorylation of histone H2AX as a measure of radiosensitivity. Int J Radiat Oncol Biol Phys. 2004;58:331–335. doi: 10.1016/j.ijrobp.2003.09.028. [DOI] [PubMed] [Google Scholar]
- 115.Olive PL, Banath JP, Sinnott LT. Phosphorylated histone H2AX in spheroids, tumors, and tissues of mice exposed to etoposide and 3-amino-1,2,4-benzotriazine-1,3-dioxide. Cancer Res. 2004;64:5363–5369. doi: 10.1158/0008-5472.CAN-04-0729. [DOI] [PubMed] [Google Scholar]
- 116.Olive PL, Banath JP. Kinetics of HAX phosphorylation after exposure to cisplatin. Cytometry B Clin Cytom. 2009;76:79–90. doi: 10.1002/cyto.b.20450. [DOI] [PubMed] [Google Scholar]
- 117.Fu S, Yang Y, Tirtha D, Yen Y, Zhou BS, Zhou MM, Ohimeyer M, Ko EC, Cagan R, Rosenstein BS, Chen SH, Kao J. γ-H2AX kinetics as a novel approach to high content screening for small molecule radiosensitizers. PloS One. 2012;7(6):e38465. doi: 10.1371/journal.pone.0038465. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 118.Ivashkevich A, Redon CE, Nakamura AJ, Martin RF, Martin OA. Use of the γH2AX assay to monitor DNA damage and repair in translational cancer research. Cancer Lett. 2011 Dec 21; doi: 10.1016/j.canlet.2011.12.025. (Epub) [DOI] [PMC free article] [PubMed] [Google Scholar]
- 119.Bourton EC, Plowman PN, Smith D, Arlett CF, Parris CN. Prolonged expression of the γ-H2AX DNA rep[air biomarker correlates with excess acute and chronic toxicity from radiotherapy treatment. Int J Cancer. 2011;129:2928–2934. doi: 10.1002/ijc.25953. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 120.Horn S, Barnard S, Rothkamm K. Gamma-H2AX-based dose estimation for whole and partial body radiation exposure. PloS One. 2011;6(9) doi: 10.1371/journal.pone.0025113. e225113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 121.Mah LJ, El-Osta A, Karagiannis TC. gammaH2AX: a sensitive molecular marker of DNA damage and repair. Leukemia. 2010;24:679–686. doi: 10.1038/leu.2010.6. [DOI] [PubMed] [Google Scholar]
- 122.Sinha RP, Häder D-P. UV-induced DNA damage and repair: A review. Photochem Photobiol Sci. 2002;1:225–236. doi: 10.1039/b201230h. [DOI] [PubMed] [Google Scholar]
- 123.Marti TM, Hefner E, Feeney L, Natale V, Cleaver JE. H2AX phosphorylation within the G1 phase after UV irradiation depends on nucleotide excision repair and not DNA double-strand breaks. Proc Natl Acad Sci USA. 2006;103:9891–9896. doi: 10.1073/pnas.0603779103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 124.Hanasoge S, Ljungman M. H2AX phosphorylation after UV irradiation is triggered by DNA repair intermediates and is mediated by the ATR kinase. Carcinogenesis. 2007;28:2298–2304. doi: 10.1093/carcin/bgm157. [DOI] [PubMed] [Google Scholar]
- 125.Stokes MP, Rush J, MavNeill J, Ren JM, Sprott K, Nardone J, Yang V, Beausoleil SA, Gygi SP, Livingstone M, Zhang H, Polakiewicz RD, Comb MJ. Profiling of UV-induced ATM/ATR signaling pathways. Proc Natl Acad Sci USA. 2007;104:19855–19860. doi: 10.1073/pnas.0707579104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 126.Halicka HD, Huang X, Traganos F, King MA, Dai W, Darzynkiewicz Z. Histone H2AX phosphorylation after cell irradiation with UV-B: Relationship to cell cycle phase and induction of apoptosis. Cell Cycle. 2005;4:339–345. [PubMed] [Google Scholar]
- 127.Zhao H, Traganos F, Darzynkiewicz Z. Kinetics of the UV-induced DNA damage response in relation to cell cycle phase. Correlation with DNA replication. Cytometry A. 2010;77A:285–293. doi: 10.1002/cyto.a.20839. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 128.Elvers I, Johansson F, Groth P, Erixon K, Helleday T. UV stalled replication forks restard by repriming in human fibroblasts. Nucleic Acids Res. 2011;39:1049–1057. doi: 10.1093/nar/gkr420. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 129.Elvers I, Hagenkort A, Johansson D, Djureinovic T, Lagerqvist N, Stoimenov I, Erixon K, Helleday T. CHK1 activity is required for continuous replication fork elongation but not stabilization of post-replicative gaps after UV radiation. Nucleic Acids Res. 2012 Jun 29; doi: 10.1093/nar/gks646. eEpub. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 130.Yajima H, Lee KJ, Zhang S, Kobayashi J, Chen BP. DNA double-strand break formation upon UV-induced replication stress activates ATM and DNA-PKcs kinases. J Mol Biol. 2009;385:800–810. doi: 10.1016/j.jmb.2008.11.036. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 131.D’Arpa P, Beardmore C, Liu LF. Involvement of nucleic acid synthesis in cell killing mechanisms of topoisomerase poisons. Cancer Res. 1990;50:6916–6924. [PubMed] [Google Scholar]
- 132.Hsiang YH, Lihou MG, Liu LF. Arrest of replication forks by drug stabilized topoisomeraseI-DNA cleavable complexes as a mechanism of cell killing by camptothecin. Cancer Res. 1989;49:5077–5082. [PubMed] [Google Scholar]
- 133.Pommier Y, Leo E, Zhang H, Marchand C. DNA topoisomerases and their poisoning by anticancer and antibacterial drugs. Chem Biol. 2010;28:421–433. doi: 10.1016/j.chembiol.2010.04.012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 134.Tanaka T, Halicka HD, Traganos F, Seiter K, Darzynkiewicz Z. Induction of ATM activation, histone H2AX phosphorylation and apoptosis by etoposide: Relation to the cell cycle phase. Cell Cycle. 2007;6:371–376. doi: 10.4161/cc.6.3.3835. [DOI] [PubMed] [Google Scholar]
- 135.Huang X, Kurose A, Tanaka T, Traganos F, Dai W, Darzynkiewicz Z. Activation of ATM and histone H2AX phosphorylation induced by mitoxantrone but not by topotecan is prevented by the antioxidant N-acetyl-L-cysteine. Cancer Biol Ther. 2006;5:959–964. doi: 10.4161/cbt.5.8.2878. [DOI] [PubMed] [Google Scholar]
- 136.Kurose A, Tanaka T, Huang X, Halicka HD, Traganos F, Dai W, Darzynkiewicz Z. Assessment of ATM phosphorylation on Ser-1981 induced by DNA topoisomerase I and II inhibitors in relation to Ser-139-histone H2AX phosphorylation, cell cycle phase and apoptosis. Cytometry A. 2005;68A:1–9. doi: 10.1002/cyto.a.20186. [DOI] [PubMed] [Google Scholar]
- 137.Smart DJ, Halicka HD, Schmuck G, Traganos F, Darzynkiewicz Z, Williams GM. Assessment of DNA double-strand breaks and γH2AX induced by the topoisomerase II poisons etoposide and mitoxantrone. Mutat. Res. 2008;641:43–47. doi: 10.1016/j.mrfmmm.2008.03.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 138.Tanaka T, Kurose A, Huang X, Dai W, Darzynkiewicz Z. ATM kinase activation and histone H2AX phosphorylation as indicators of DNA damage by DNA topoisomerase I inhibitor topotecan and during apoptosis. Cell Prolif. 2006;39:49–60. doi: 10.1111/j.1365-2184.2006.00364.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 139.Olive PL, Banath JP, Sinnott LT. Phosphorylated histone H2AX in spheroids, tumors, and tissues of mice exposed to etoposide and 3-amino-1,2,4-benzotriazine-1,3-dioxide. Cancer Res. 2004;64:5363–5369. doi: 10.1158/0008-5472.CAN-04-0729. [DOI] [PubMed] [Google Scholar]
- 140.Smart DJ, Lynch AM. Evaluating the genotoxicity of topoisomerase-targeted antibiotics. Mutagenesis. 2012;27:359–365. doi: 10.1093/mutage/ger089. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 141.Zhao H, Traganos F, Darzynkiewicz Z. Phosphorylation of p53 on Ser15 during cell cycle caused by Topo I and Topo II inhibitors in relation to ATM and Chk2 activation. Cell Cycle. 2008;7:3048–3055. doi: 10.4161/cc.7.19.6750. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 142.Zhao H, Traganos F, Darzynkiewicz Z. Kinetics of histone H2AX phosphorylation and Chk2 activation in A549 cells treated with topotecan and mitoxantrone in relation to the cell cycle phase. Cytometry A. 2008;73A:480–489. doi: 10.1002/cyto.a.20574. [DOI] [PubMed] [Google Scholar]
- 143.Staker BL, Hjerrild K, Feese MD, Behnke CA, Burgin AB, Stewart L. The mechanism of topoisomerase I poisoning by a camptothecin analog. Proc Natl Acad Sci USA. 2002;99:15387–15392. doi: 10.1073/pnas.242259599. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 144.Kapuscinski J, Darzynkiewicz Z. Relationship between the pharmacological activity of antitumor drugs Ametantrone and Mitoxantrone (Novantrone) and their ability to condense nucleic acids. Proc Natl Acad Sci USA. 1986;83:6302–6306. doi: 10.1073/pnas.83.17.6302. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 145.Kapuscinski J, Darzynkiewicz Z, Traganos F, Melamed MR. Interactions of a new antitumor agent 1,4-dihydroxy-5-8-bis 2-(hydroxyethyl)amino)ethyl)amino)--9,10-anthracenedione with nucleic acids. Biochem Pharmacol. 1981;30:231–240. doi: 10.1016/0006-2952(81)90083-6. [DOI] [PubMed] [Google Scholar]
- 146.Wu C-C, Li TK, Fahr L, Lin L-Y, Lin L-Y, Lin T-S, Yu Y-Y, Yen T-J, Chiang C-W, Chan N-L. Structural basis of type II topoisomerase inhibition by the anticancer drug etoposide. Science. 2011;333:459–462. doi: 10.1126/science.1204117. [DOI] [PubMed] [Google Scholar]
- 147.Zhao H, Dobrucki J, Rybak P, Traganos F, Darzynkiewicz Z. Relationship of DNA damage signaling induced by DNA topoisomerase inhibitors camptothecin/topotecan, mitoxantrone or etoposide and DNA replication. Cytometry A. 1012;81A:45–51. doi: 10.1002/cyto.a.21172. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 148.Leonhardt H, Rahn HP, Weinzierl P, Sporbert A, Cremen T, Zink D, Cardoso MC. Dynamics of DNA replication factories in living cells. J Cell Biol. 2000;149:271–279. doi: 10.1083/jcb.149.2.271. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 149.Halicka HD, Ozkaynak MF, Levendoglu-Tugal O, Sandoval C, Seiter K, Kajstura M, Traganos F, Jaybose S, Darzynkiewicz Z. DNA damage response as a biomarker in treatment of leukemias. Cell Cycle. 2009;8:1720–1724. doi: 10.4161/cc.8.11.8598. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 150.McKenna E, Traganos F, Zhao H, Darzynkiewicz Z. Persistent DNA damage caused by low levels of mitomycin C induces irreversible senescence of A549 cells. Cell Cycle. 2012;11:3132–3140. doi: 10.4161/cc.21506. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 151.Banuelos CA, Banath JP, Kim JY, Aquino-Parsons C, Olive P. gammaH2AX expression in tumors exposed to cisplatin and fractionated irradiation. Clin Canc Res. 2009;15:3344–3353. doi: 10.1158/1078-0432.CCR-08-3114. [DOI] [PubMed] [Google Scholar]
- 152.Banath JP, Klokov D, MacPhail SH, Banuelos CA, Olive PL. Residual gammaH2AX foci as an indication of lethal DNA lesions. BMC Cancer. 2010 Jan 5;10:4. doi: 10.1186/1471-2407-10-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 153.Demidenko ZN, Blagosklonny MV. Growth stimulation leads to cellular senescence when the cell cycle is blocked. Cell Cycle. 2008;7:3355–3361. doi: 10.4161/cc.7.21.6919. [DOI] [PubMed] [Google Scholar]
- 154.Albino AP, Huang X, Yang J, Gietl D, Jorgensen E, Traganos F, Darzynkiewicz Z. Induction of histone H2AX phosphorylation in A549 human pulmonary epithelial cells by tobacco smoke and in human bronchial epithelial cells by smoke condensate: A new assay to detect the presence of potential carcinogens in tobacco. Cell Cycle. 2004;3:1062–1068. [PubMed] [Google Scholar]
- 155.Albino AP, Jorgensen E, Rainey P, Gillman G, Clark TJ, Gietl D, Zhao H, Traganos F, Darzynkiewicz Z. γH2AX: A potential DNA damage response biomarker for assessing toxicological risk of tobacco products. Mutation Res. 2009;678:43–52. doi: 10.1016/j.mrgentox.2009.06.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 156.Zhao H, Albino AP, Jorgensen E, Traganos F, Darzynkiewicz Z. DNA damage response induced by tobacco smoke in normal human bronchial epithelial and A549 pulmonary adenocarcinoma cells assessed by laser scanning cytometry. Cytometry A. 2009;75A:840–847. doi: 10.1002/cyto.a.20778. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 157.Tanaka T, Huang X, Jorgensen E, Gietl E, Traganos F, Darzynkiewicz Z, Albino AP. ATM activation accompanies histone H2AX phosphorylation in A549 cells upon exposure to tobacco smoke. BMC Cell Biol. 2007 Jun 26;8:26. doi: 10.1186/1471-2121-8-26. Epub. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 158.Toyooka T, Ibuki Y. Cigarette sidestream smoke induces phosphorylated histone H2AX. Mutat Res. 2009;676:34–40. doi: 10.1016/j.mrgentox.2009.03.002. [DOI] [PubMed] [Google Scholar]
- 159.Jorgensen ED, Zhao H, Traganos F, Albino AP, Darzynkiewicz Z. DNA damage response induced by exposure of A549 human lung adenocarcinoma cells to smoke from tobacco- and nicotine-free cigarettes. Cell Cycle. 2010;9:2170–2176. doi: 10.4161/cc.9.11.11842. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 160.Garcia-Canton C, Anadon A, Meredith C. γH2AX as a novel endpoint to detect DNA Damage: Applications for the assessment of the in vitro genotoxicity of cigarette smoke. Toxicol In Vitro. 2012 Jun 23; doi: 10.1016/j.tiv.2012.06.006. Epub. [DOI] [PubMed] [Google Scholar]
- 161.Albino AP, Huang X, Jorgensen E, Gietl D, Traganos F, Darzynkiewicz Z. Induction of DNA double-strand breaks in A549 and normal human pulmonary epithelial cells by cigarette smoke is mediated by free radicals. Int J Oncol. 2006;28:1491–1505. [PubMed] [Google Scholar]
- 162.Maertens RM, White PA, Rickert W, Lavasseur G, Gouglas GR, Bellier PV, McNamee JP, Thuppal V, Walker M, Desjardins S. The genoptoxicity of mainstream and sidestream marijuana and tobacco smoke condensates. Chem Tes Toxicol. 2009;22:1406–1414. doi: 10.1021/tx9000286. [DOI] [PubMed] [Google Scholar]
- 163.Toduka Y, Toyooka T, Ibuki Y. Flow cytometric evaluation of nanoparticles using side-scattered light and reactive oxygen species-mediated fluorescence- correlation with genotoxicity. Environ Sci Technol. 2012;46:7629–7636. doi: 10.1021/es300433x. [DOI] [PubMed] [Google Scholar]
- 164.Toyooka T, Amano T, Ibuki Y. Titanium dioxide paricles phosphorylate histone H2AX independent of ROS production. Mutat Res. 2012;742:84–91. doi: 10.1016/j.mrgentox.2011.12.015. [DOI] [PubMed] [Google Scholar]
- 165.Ahamed M, Karns M, Goodson M, Rowe J, Hussain SM, Schlager JJ, Hong JJ. DNA damage response to different surface chemistry of silver nanoparticles in mammalian cells. Toxicol Appl Pharmacol. 2008;233:404–410. doi: 10.1016/j.taap.2008.09.015. [DOI] [PubMed] [Google Scholar]
- 166.Tsaousi A, Jones E, Case CP. The in vitro genotoxicity of orthopaedic ceramic (Al2O3) and metal CoCr alloy) particles. Mutat Res. 2010;697:1–9. doi: 10.1016/j.mrgentox.2010.01.012. [DOI] [PubMed] [Google Scholar]
- 167.Audelbert M, Dolo L, Peru E, Cravedi JP, Zalko D. Use of the γH2AX assay for assessing the genotoxicity of bisphenol A and bisphenol F in human cell lines. Arch Toxicol. 2011;85:1463–1473. doi: 10.1007/s00204-011-0721-2. [DOI] [PubMed] [Google Scholar]
- 168.Audebert M, Tiu A, Jacques C, Hillenweck A, Jamin EL, Zalko D, Cravedi JP. Use of the γH2AX assay for assessing the genotoxicity of polycyclic aromatic hydrocarbons in human cell lines. Toxicol Lett. 2010;199:82–192. doi: 10.1016/j.toxlet.2010.08.022. [DOI] [PubMed] [Google Scholar]
- 169.Toyooka T, Kubota T, Ibuki Y. Nonylphenol polyethoxylates induce phosphorylation of histone H2AX. Mutat Res. 2012;741:57–64. doi: 10.1016/j.mrgentox.2011.10.006. [DOI] [PubMed] [Google Scholar]
- 170.Carvalho CM, Menezes PF, Letenski GC, Praes CE, Feferman IH, Lorencini M. In vitro induction of apoptosis, necrosis and genotoxicity by cosmetic preservatives: application of flow cytometry as a complementary analysis by NRU. Int J Cosmet Sci. 2012;34:176–182. doi: 10.1111/j.1468-2494.2011.00698.x. [DOI] [PubMed] [Google Scholar]
- 171.Liu X, Lee J, Ji K, Takeda S, Choi K. Potentials and mechanisms of genotoxicity of xix pharmaceuticals frequently detected in freshwater environment. Toxicol Let. 2012;211:70–76. doi: 10.1016/j.toxlet.2012.03.003. [DOI] [PubMed] [Google Scholar]
- 172.Tanaka T, Kurose A, Halicka HD, Huang X, Traganos F, Darzynkiewicz Z. Nitrogen oxide-releasing aspirin induces histone H2AX phosphorylation, ATM activation, and apoptosis preferentially in S-phase cells; involvement of reactive oxygen species. Cell Cycle. 2006;5:1669–1674. doi: 10.4161/cc.5.15.3100. [DOI] [PubMed] [Google Scholar]
- 173.Buchelnikov AS, Hernandez SAA, Gonzalez FM, Vazquez RR, Davies DB, Evstigneev MP. General analysis of competitive binding in dru-interceptor-DNA systems. Eur Biophys J. 2012;41:273–283. doi: 10.1007/s00249-011-0783-3. [DOI] [PubMed] [Google Scholar]
- 174.Pietrzak M, Halicka HD, Wieczorek Z, Wieczorek J, Darzynkiewicz Z. Attenuation of acridine mutagen ICR 191-DNA interactions and DNA damage by the mutagen interceptor chlorophyllin. Biophys J. 2008;135:69–75. doi: 10.1016/j.bpc.2008.03.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 175.Zhao H, Traganos F, Dobrucki J, Wlodkowic D, Darzynkiewicz Z. Induction of DNA damage response by the supravital probes of nucleic acids. Cytometry A. 2009;75A:510–519. doi: 10.1002/cyto.a.20727. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 176.Watters GP, Smart DJ, Harvey JS, Austin CA. H2AX phosphorylation as a genotoxicity endpoint. Mutat Res. 2009;679:50–58. doi: 10.1016/j.mrgentox.2009.07.007. [DOI] [PubMed] [Google Scholar]
- 177.Smart DJ, Ahmedi KP, Harvey JS, Lynch AM. Genotoxicity screening via the γH2AX by flow assay. Mutat Res. 2011;515:25–31. doi: 10.1016/j.mrfmmm.2011.07.001. [DOI] [PubMed] [Google Scholar]
- 178.Redon CE, Nakamura AJ, Martin OA, Parekh PR, Weyemi US, Bonner WM. Recent developmets in the use of γ-H2AX as a quantitative DNA double-strand break biomarker. Aging (Albany) 2011;3:168–174. doi: 10.18632/aging.100284. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 179.Redon CE, Weyemi U, Parekh PR, Huang D, Burrel AS, Bonner WM. γH2AX and other histone post-translational modifications in the clinic. Biochiom Biophys Acta. 2012;1819:743–756. doi: 10.1016/j.bbagrm.2012.02.021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 180.Skommer J, Darzynkiewicz Z, Wlodkowic D. Cell death goes LIVE: Technological advances in real-time tracking of cell death. Cell Cycle. 2010;9:2330–2341. doi: 10.4161/cc.9.12.11911. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 181.Wlodkowic D, Skommer J, Darzynkiewicz Z. Cytometry in cell necrobiology revisited. Recent advances and new vistas. Cytometry Part A. 2010;77:591–606. doi: 10.1002/cyto.a.20889. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 182.George TC, Basiji DA, Hall BE, Lynch DH, Ortyn WE, Perry DJ, Seo MJ, Zimmerman CA, Morrissey PJ. Distinguishing modes of cell death using the ImageStream multispectral imaging flow cytometer. Cytometry A. 2004;59:237–245. doi: 10.1002/cyto.a.20048. [DOI] [PubMed] [Google Scholar]
- 183.George TC, Fanning SL, Fitzgerald-Bocarsly P, Medeiros RB, Highfill S, Shimizu Y, Hall BE, Frost K, Basiji D, Ortyn WE, Morrissey PJ, Lynch DH. Quantitative measurement of nuclear translocation events using similarity analysis of multispectral cellular images obtained in flow. J Immunol Methods. 2006;311:117–129. doi: 10.1016/j.jim.2006.01.018. [DOI] [PubMed] [Google Scholar]
- 184.Bourton EC, Plowman PN, Zahir SA, Senguloglu GU, Serrai H, Bottley G, Parris CN. Multispectral imaging flow cytometry reveals distinct frequencies of γ-H2AX foci induction in DNA double strand break repair defective human cell lines. Cytometry Part A. 2012;81:130–137. doi: 10.1002/cyto.a.21171. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 185.Darzynkiewicz Z. Cycling into future: Mass cytometry for the cell-cycle analysis. Cytometry Part A. 2012;81:546–548. doi: 10.1002/cyto.a.22080. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 186.Ornatsky O, Bandura D, Baranov V, Nitz M, Winnik MA, Tanner S. Highly multiparametric analysis by mass cytometry. J Immunol Methods. 2010;361:1–20. doi: 10.1016/j.jim.2010.07.002. [DOI] [PubMed] [Google Scholar]
- 187.Fienberg HG, Simonds EF, Fantl WJ, Nolan GP, Bodenmiller B. A platinum-based covalent viability reagent for single-cell mass cytometry. Cytometry Part A. 2012;81:467–475. doi: 10.1002/cyto.a.22067. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 188.Behbehani GK, Bendall SC, Clutter MR, Fantl WJ, Nolan GP. Single-cell mass cytometry adapted to measurements of the cell cycle. Cytometry Part A. 2012;81:552–566. doi: 10.1002/cyto.a.22075. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 189.Notingher I, Hench LL. Raman microspectroscopy: a noninvasive tool for studies of individual living cells in vitro. Expert Rev Med Devices. 2006;3:215–234. doi: 10.1586/17434440.3.2.215. [DOI] [PubMed] [Google Scholar]
- 190.Kneipp J, Kneipp H, Rice WL, Kneipp K. Optical probes for biological applications based on surface-enhanced Raman scattering from indocyanine green on gold nanoparticles. Anal Chem. 2005;77:2381–2385. doi: 10.1021/ac050109v. [DOI] [PubMed] [Google Scholar]
- 191.Jett JH. Raman spectroscopy comes to flow cytometry. Cytometry Part A. 2008;73:109–110. doi: 10.1002/cyto.a.20526. [DOI] [PubMed] [Google Scholar]
- 192.Wlodkowic D, Faley S, Zagnoni M, Wikswo JP, Cooper JM. Microfluidic single-cell array cytometry for the analysis of tumor apoptosis. Anal Chem. 2009;81:5517–5523. doi: 10.1021/ac9008463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 193.Wlodkowic D, Khoshmanesh K, Sharpe JC, Darzynkiewicz Z, Cooper JM. Apoptosis goes on a chip: advances in the microfluidic analysis of programmed cell death. Anal Chem. 2011;83:6439–6446. doi: 10.1021/ac200588g. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 194.Wlodkowic D, Cooper JM. Tumors on chips: oncology meets microfluidics. Curr Opinion Chem Biol. 2010;14:556–567. doi: 10.1016/j.cbpa.2010.08.016. [DOI] [PubMed] [Google Scholar]
- 195.Wlodkowic D, Skommer J, Faley S, Darzynkiewicz Z, Cooper JM. Dynamic analysis of apoptosis using cyanine SYTO probes: from classical to microfluidic cytometry. Exp Cell Res. 2009;315:1706–1714. doi: 10.1016/j.yexcr.2009.03.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 196.Sohn LL, Saleh OA, Facer GR, Beavis AJ, Allan RS, Notterman DA. Capacitance cytometry: measuring biological cells one by one. Proc Natl Acad Sci USA. 2000;97:10687–10690. doi: 10.1073/pnas.200361297. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 197.De Vos WH, Van Neste L, Dieriks B, Joss GH, Van Oostveldt P. High content image cytometry in the context of subnuclear organization. Cytometry Part A. 2010;77(1):64–75. doi: 10.1002/cyto.a.20807. [DOI] [PubMed] [Google Scholar]
- 198.Zhang H, Tu E, Hagen ND, Schnabel CA, Paliotti MJ, Hoo WS, Nguyen PM, Kohrumel JR, Butler WF, Chachisvillis M, Marchand PJ. Time-of-flight optophoresis analysis of live whole cells in microfluidic channels. Biomed Microdevices. 2004;6:11–21. doi: 10.1023/b:bmmd.0000013361.03291.6c. [DOI] [PubMed] [Google Scholar]
- 199.Tamaki E, Sato K, Tokeshi M, Sato K, Aihara M, Kitamori T. Single-cell analysis by a scanning thermal lens microscope with a microchip: direct monitoring of cytochrome c distribution during apoptosis process. Anal Chem. 2002;74:1560–4. doi: 10.1021/ac011092t. [DOI] [PubMed] [Google Scholar]
- 200.Wlodkowic D, Cooper JM. Microfluidic cell arrays in tumor analysis: new prospects for integrated cytomics. Expert Rev Mol Diagn. 2010;10:521–530. doi: 10.1586/erm.10.28. [DOI] [PubMed] [Google Scholar]
- 201.Forment JV, Walker RV, Jackson SP. A high-throughput, flow cytometry-based method to quantify DNA-end resection in mammalian cells. Cytometry Part A. 2012;81:922–928. doi: 10.1002/cyto.a.22155. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 202.Jun YW, Sheikholeslami S, Hostetter DR, Tajon C, Craik CS, Alivisatos AP. Continuous imaging of plasmon rulers in live cells reveals early-stage caspase-3 activation at the single-molecule level. Proc Natl Acad Sci USA. 2009;106:17735–17740. doi: 10.1073/pnas.0907367106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 203.Chowdhury D, Keogh MC, Ishii H, Peterson CL, Buratowski S, Lieberman J. gamma-H2AX dephosphorylation by protein phosphatase 2A facilitates DNA double-strand break repair. Mol Cell. 2005;20:801–809. doi: 10.1016/j.molcel.2005.10.003. [DOI] [PubMed] [Google Scholar]
- 204.Ichijima Y, Sakasai R, Okita N, Asahina K, Mizutani S, Teraoka H. Phosphorylation of histone H2AX at M phase in human cells without DNA damage response. Biochem Biophys Res Commun. 2005;336:807–812. doi: 10.1016/j.bbrc.2005.08.164. [DOI] [PubMed] [Google Scholar]
- 205.Ichijima Y, Yoshioka K, Yoshioka Y, Shinohe K, Fujimori H, Unno M, Goto H, Inagaki M, Mizutani S, Teraoka H. DNA lesions induced by replication stress trigger mitotic aberration and tetraploidy development. PLoS One. 2010 Jan 21;5(1):e8821. doi: 10.1371/journal.pone.0008821. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 206.Tanaka T, Huang X, Halicka HD, Zhao H, Traganos F, Albino AP, Dai W, Darzynkiewicz Z. Cytometry of ATM activation and histone H2AX phosphorylation to estimate extent of DNA damage induced by exogenous agents. Cytometry A. 2007;71A:648–661. doi: 10.1002/cyto.a.20426. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 207.Henriksen M, Miller B, Newmark J, Al-Kofahi Y, Holden E. Laser scanning cytometry and its applications: A pioneering technology in the field of quantitative imaging cytometry. Meth Cell Biol. 2011;102:161–205. doi: 10.1016/B978-0-12-374912-3.00007-9. [DOI] [PubMed] [Google Scholar]
- 208.Zuba-Surma E, Ratajczak M. Analytical capabilities of the ImageStream cytometer. Meth Cell Biol. 2011;102:207–227. doi: 10.1016/B978-0-12-374912-3.00008-0. [DOI] [PubMed] [Google Scholar]
- 209.Gonzalez JE, Lee M, Barquinero JF, Valente M, Roch-Lefebre S, Garcia O. Quantitative image analysis of gamma-H2AX foci induced by ionizing radiation applying open source programs. Anal Quant Cytol Histol. 2012;34:66–71. [PubMed] [Google Scholar]
- 210.Lobrich M, Rief N, Kuhne M, Heckman M, Fleckenstein J, Rube C, Uder M. In vivo formation and repair of DNA double-strand breaks after computed tomography examinations. Proc Natl Acad Sci USA. 2005;102:8994–8989. doi: 10.1073/pnas.0501895102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 211.Kawane K, Nagata S. Nucleases in programmed cell death. Meth Enzymol. 2008;442:271–287. doi: 10.1016/S0076-6879(08)01414-6. [DOI] [PubMed] [Google Scholar]
- 212.Jackson SP. DNA damage signaling and apoptosis. Biochem Soc Transactions. 2001;29:655–661. doi: 10.1042/0300-5127:0290655. [DOI] [PubMed] [Google Scholar]
- 213.Juan G, Gruenwald S, Darzynkiewicz Z. Phosphorylation of retinoblastoma susceptibility gene protein assayed in individual lymphocytes during their mitogenic stimulation. Exp Cell Res. 1998;239:104–110. doi: 10.1006/excr.1997.3885. [DOI] [PubMed] [Google Scholar]
- 214.Krutzik PO, Crane JM, Clutter MR, Nolan GP. High-content single-cell drug screening with phosphospecific flow cytometry. Nat Chem Biol. 2008;4:132–142. doi: 10.1038/nchembio.2007.59. [DOI] [PubMed] [Google Scholar]