Abstract
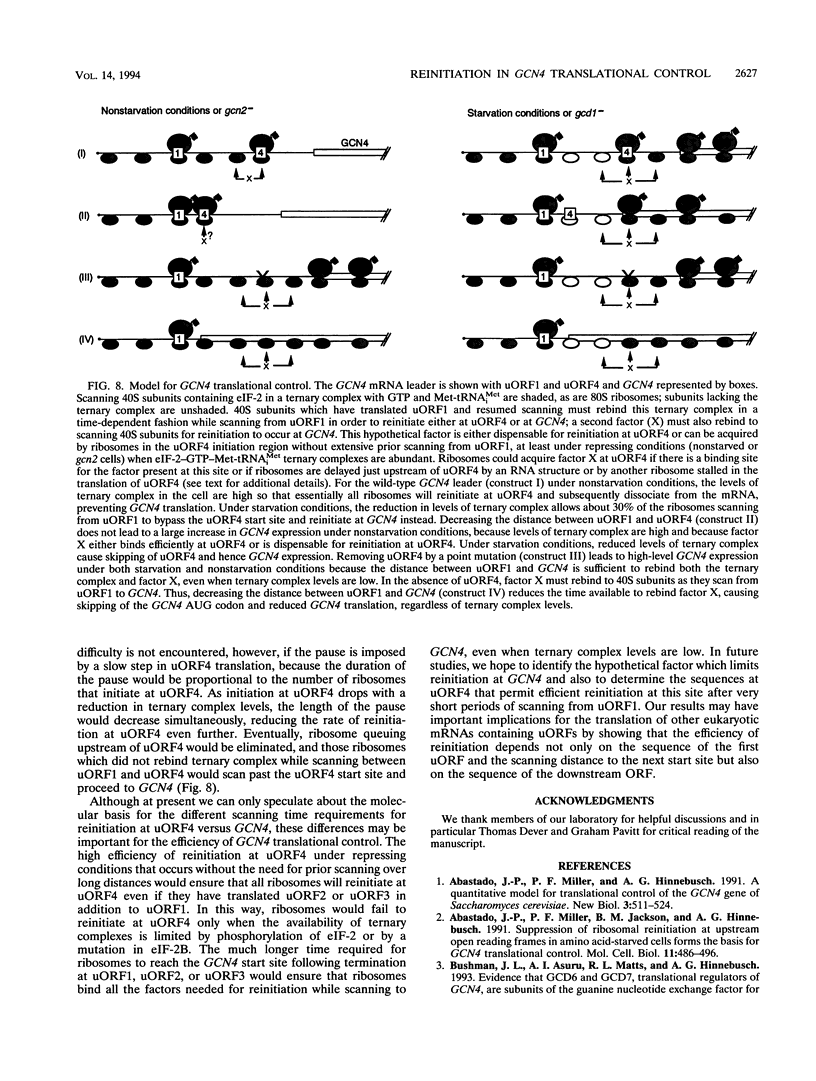
Translational control of the GCN4 gene in response to amino acid availability is mediated by four short open reading frames in the GCN4 mRNA leader (uORFs) and by phosphorylation of eukaryotic initiation factor 2 (eIF-2). We have proposed that reducing eIF-2 activity by phosphorylation of its alpha subunit or by a mutation in the eIF-2 recycling factor eIF-2B allows ribosomes which have translated the 5'-proximal uORF1 to bypass uORF2 to uORF4 and reinitiate at GCN4 instead. In this report, we present two lines of evidence that all ribosomes which synthesize GCN4 have previously translated uORF1, resumed scanning, and reinitiated at the GCN4 start site. First, GCN4 expression was abolished when uORF1 was elongated to make it overlap the beginning of the GCN4 coding region. Second, GCN4 expression was reduced as uORF1 was moved progressively closer to GCN4, decreasing to only 5% of the level seen in the absence of all uORFs when only 32 nucleotides separated uORF1 from GCN4. We additionally found that inserting small synthetic uORFs between uORF4 and GCN4 inhibited GCN4 expression under derepressing conditions, confirming the idea that reinitiation at GCN4 under conditions of diminished eIF-2 activity is proportional to the distance of the reinitiation site downstream from uORF1. While uORF4 and GCN4 appear to be equally effective at capturing ribosomes scanning downstream from the 5' cap of mRNA, these two ORFs differ greatly in their ability to capture reinitiating ribosomes scanning from uORF1. When the active form of eIF-2 is present at high levels, reinitiation appears to be much more efficient at uORF4 than at GCN4 when each is located very close to uORF1. Under conditions of reduced recycling of eIF-2, reinitiation at uORF4 is substantially suppressed, which allows ribosomes to reach the GCN4 start site; in contrast, reinitiation at GCN4 in constructs lacking uORF4 is unaffected by decreasing the level of eIF-2 activity. This last finding raises the possibility that time-dependent binding to ribosomes of a second factor besides the eIF-2-GTP-Met-tRNA(iMet) ternary complex is rate limiting for reinitiation at GCN4. Moreover, our results show that the efficiency of translational reinitiation can be strongly influenced by the nature of the downstream cistron as well as the intercistronic distance.
Full text
PDF










Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Abastado J. P., Miller P. F., Hinnebusch A. G. A quantitative model for translational control of the GCN4 gene of Saccharomyces cerevisiae. New Biol. 1991 May;3(5):511–524. [PubMed] [Google Scholar]
- Abastado J. P., Miller P. F., Jackson B. M., Hinnebusch A. G. Suppression of ribosomal reinitiation at upstream open reading frames in amino acid-starved cells forms the basis for GCN4 translational control. Mol Cell Biol. 1991 Jan;11(1):486–496. doi: 10.1128/mcb.11.1.486. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cigan A. M., Bushman J. L., Boal T. R., Hinnebusch A. G. A protein complex of translational regulators of GCN4 mRNA is the guanine nucleotide-exchange factor for translation initiation factor 2 in yeast. Proc Natl Acad Sci U S A. 1993 Jun 1;90(11):5350–5354. doi: 10.1073/pnas.90.11.5350. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cigan A. M., Donahue T. F. Sequence and structural features associated with translational initiator regions in yeast--a review. Gene. 1987;59(1):1–18. doi: 10.1016/0378-1119(87)90261-7. [DOI] [PubMed] [Google Scholar]
- Degnin C. R., Schleiss M. R., Cao J., Geballe A. P. Translational inhibition mediated by a short upstream open reading frame in the human cytomegalovirus gpUL4 (gp48) transcript. J Virol. 1993 Sep;67(9):5514–5521. doi: 10.1128/jvi.67.9.5514-5521.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dever T. E., Feng L., Wek R. C., Cigan A. M., Donahue T. F., Hinnebusch A. G. Phosphorylation of initiation factor 2 alpha by protein kinase GCN2 mediates gene-specific translational control of GCN4 in yeast. Cell. 1992 Feb 7;68(3):585–596. doi: 10.1016/0092-8674(92)90193-g. [DOI] [PubMed] [Google Scholar]
- Fütterer J., Gordon K., Sanfaçon H., Bonneville J. M., Hohn T. Positive and negative control of translation by the leader sequence of cauliflower mosaic virus pregenomic 35S RNA. EMBO J. 1990 Jun;9(6):1697–1707. doi: 10.1002/j.1460-2075.1990.tb08293.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Geballe A. P., Mocarski E. S. Translational control of cytomegalovirus gene expression is mediated by upstream AUG codons. J Virol. 1988 Sep;62(9):3334–3340. doi: 10.1128/jvi.62.9.3334-3340.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grant C. M., Hinnebusch A. G. Effect of sequence context at stop codons on efficiency of reinitiation in GCN4 translational control. Mol Cell Biol. 1994 Jan;14(1):606–618. doi: 10.1128/mcb.14.1.606. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hannig E. M., Cigan A. M., Freeman B. A., Kinzy T. G. GCD11, a negative regulator of GCN4 expression, encodes the gamma subunit of eIF-2 in Saccharomyces cerevisiae. Mol Cell Biol. 1993 Jan;13(1):506–520. doi: 10.1128/mcb.13.1.506. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hensel C. H., Petersen R. B., Hackett P. B. Effects of alterations in the leader sequence of Rous sarcoma virus RNA on initiation of translation. J Virol. 1989 Nov;63(11):4986–4990. doi: 10.1128/jvi.63.11.4986-4990.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hill J. R., Morris D. R. Cell-specific translational regulation of S-adenosylmethionine decarboxylase mRNA. Dependence on translation and coding capacity of the cis-acting upstream open reading frame. J Biol Chem. 1993 Jan 5;268(1):726–731. [PubMed] [Google Scholar]
- Hinnebusch A. G. Evidence for translational regulation of the activator of general amino acid control in yeast. Proc Natl Acad Sci U S A. 1984 Oct;81(20):6442–6446. doi: 10.1073/pnas.81.20.6442. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hinnebusch A. G. Mechanisms of gene regulation in the general control of amino acid biosynthesis in Saccharomyces cerevisiae. Microbiol Rev. 1988 Jun;52(2):248–273. doi: 10.1128/mr.52.2.248-273.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ito H., Fukuda Y., Murata K., Kimura A. Transformation of intact yeast cells treated with alkali cations. J Bacteriol. 1983 Jan;153(1):163–168. doi: 10.1128/jb.153.1.163-168.1983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Johansen H., Schümperli D., Rosenberg M. Affecting gene expression by altering the length and sequence of the 5' leader. Proc Natl Acad Sci U S A. 1984 Dec;81(24):7698–7702. doi: 10.1073/pnas.81.24.7698. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Khalili K., Brady J., Khoury G. Translational regulation of SV40 early mRNA defines a new viral protein. Cell. 1987 Feb 27;48(4):639–645. doi: 10.1016/0092-8674(87)90242-x. [DOI] [PubMed] [Google Scholar]
- Kozak M. An analysis of 5'-noncoding sequences from 699 vertebrate messenger RNAs. Nucleic Acids Res. 1987 Oct 26;15(20):8125–8148. doi: 10.1093/nar/15.20.8125. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kozak M. Effects of intercistronic length on the efficiency of reinitiation by eucaryotic ribosomes. Mol Cell Biol. 1987 Oct;7(10):3438–3445. doi: 10.1128/mcb.7.10.3438. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kozak M. Influences of mRNA secondary structure on initiation by eukaryotic ribosomes. Proc Natl Acad Sci U S A. 1986 May;83(9):2850–2854. doi: 10.1073/pnas.83.9.2850. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kozak M. Point mutations define a sequence flanking the AUG initiator codon that modulates translation by eukaryotic ribosomes. Cell. 1986 Jan 31;44(2):283–292. doi: 10.1016/0092-8674(86)90762-2. [DOI] [PubMed] [Google Scholar]
- Kozak M. Selection of initiation sites by eucaryotic ribosomes: effect of inserting AUG triplets upstream from the coding sequence for preproinsulin. Nucleic Acids Res. 1984 May 11;12(9):3873–3893. doi: 10.1093/nar/12.9.3873. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kozak M. The scanning model for translation: an update. J Cell Biol. 1989 Feb;108(2):229–241. doi: 10.1083/jcb.108.2.229. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu C. C., Simonsen C. C., Levinson A. D. Initiation of translation at internal AUG codons in mammalian cells. Nature. 1984 May 3;309(5963):82–85. doi: 10.1038/309082a0. [DOI] [PubMed] [Google Scholar]
- Matts R. L., Levin D. H., London I. M. Effect of phosphorylation of the alpha-subunit of eukaryotic initiation factor 2 on the function of reversing factor in the initiation of protein synthesis. Proc Natl Acad Sci U S A. 1983 May;80(9):2559–2563. doi: 10.1073/pnas.80.9.2559. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Merrick W. C. Mechanism and regulation of eukaryotic protein synthesis. Microbiol Rev. 1992 Jun;56(2):291–315. doi: 10.1128/mr.56.2.291-315.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Miller P. F., Hinnebusch A. G. Sequences that surround the stop codons of upstream open reading frames in GCN4 mRNA determine their distinct functions in translational control. Genes Dev. 1989 Aug;3(8):1217–1225. doi: 10.1101/gad.3.8.1217. [DOI] [PubMed] [Google Scholar]
- Mueller P. P., Hinnebusch A. G. Multiple upstream AUG codons mediate translational control of GCN4. Cell. 1986 Apr 25;45(2):201–207. doi: 10.1016/0092-8674(86)90384-3. [DOI] [PubMed] [Google Scholar]
- Petersen R. B., Moustakas A., Hackett P. B. A mutation in the short 5'-proximal open reading frame on Rous sarcoma virus RNA alters virus production. J Virol. 1989 Nov;63(11):4787–4796. doi: 10.1128/jvi.63.11.4787-4796.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rowlands A. G., Panniers R., Henshaw E. C. The catalytic mechanism of guanine nucleotide exchange factor action and competitive inhibition by phosphorylated eukaryotic initiation factor 2. J Biol Chem. 1988 Apr 25;263(12):5526–5533. [PubMed] [Google Scholar]
- Siekierka J., Mauser L., Ochoa S. Mechanism of polypeptide chain initiation in eukaryotes and its control by phosphorylation of the alpha subunit of initiation factor 2. Proc Natl Acad Sci U S A. 1982 Apr;79(8):2537–2540. doi: 10.1073/pnas.79.8.2537. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Werner M., Feller A., Messenguy F., Piérard A. The leader peptide of yeast gene CPA1 is essential for the translational repression of its expression. Cell. 1987 Jun 19;49(6):805–813. doi: 10.1016/0092-8674(87)90618-0. [DOI] [PubMed] [Google Scholar]
- Williams N. P., Hinnebusch A. G., Donahue T. F. Mutations in the structural genes for eukaryotic initiation factors 2 alpha and 2 beta of Saccharomyces cerevisiae disrupt translational control of GCN4 mRNA. Proc Natl Acad Sci U S A. 1989 Oct;86(19):7515–7519. doi: 10.1073/pnas.86.19.7515. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Williams N. P., Mueller P. P., Hinnebusch A. G. The positive regulatory function of the 5'-proximal open reading frames in GCN4 mRNA can be mimicked by heterologous, short coding sequences. Mol Cell Biol. 1988 Sep;8(9):3827–3836. doi: 10.1128/mcb.8.9.3827. [DOI] [PMC free article] [PubMed] [Google Scholar]


