Abstract
Nutrient starvation in the yeast Saccharomyces cerevisiae leads to a number of physiological changes that accompany entry into stationary phase. The expression of genes whose products play a role in stress adaptation is regulated in a manner that allows the cell to sense and respond to changing environmental conditions. We have identified a novel yeast gene, YGP1, that displays homology to the sporulation-specific SPS100 gene. The expression of YGP1 is regulated by nutrient availability. The gene is expressed at a basal level during "respiro-fermentative" (logarithmic) growth. When the glucose concentration in the medium falls below 1%, the YGP1 gene is derepressed and the gene product, gp37, is synthesized at levels up to 50-fold above the basal level. The glucose-sensing mechanism is independent of the SNF1 pathway and does not operate when cells are directly shifted to a low glucose concentration. The expression of YGP1 also responds to the depletion of nitrogen and phosphate, indicating a general response to nutrient deprivation. These results suggest that the YGP1 gene product may be involved in cellular adaptations prior to stationary phase and may be a useful marker protein for monitoring early events associated with the stress response.
Full text
PDF










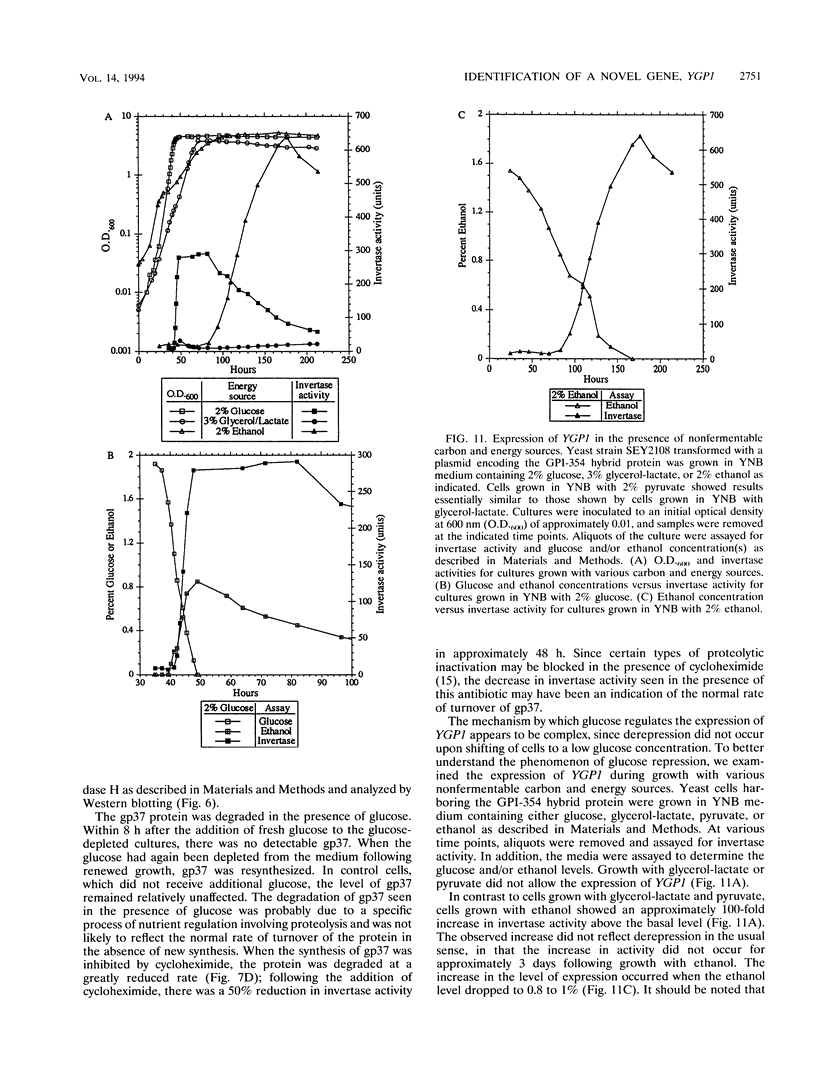



Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Bankaitis V. A., Johnson L. M., Emr S. D. Isolation of yeast mutants defective in protein targeting to the vacuole. Proc Natl Acad Sci U S A. 1986 Dec;83(23):9075–9079. doi: 10.1073/pnas.83.23.9075. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Birnboim H. C., Doly J. A rapid alkaline extraction procedure for screening recombinant plasmid DNA. Nucleic Acids Res. 1979 Nov 24;7(6):1513–1523. doi: 10.1093/nar/7.6.1513. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Boucherie H. Protein synthesis during transition and stationary phases under glucose limitation in Saccharomyces cerevisiae. J Bacteriol. 1985 Jan;161(1):385–392. doi: 10.1128/jb.161.1.385-392.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Casadaban M. J., Cohen S. N. Analysis of gene control signals by DNA fusion and cloning in Escherichia coli. J Mol Biol. 1980 Apr;138(2):179–207. doi: 10.1016/0022-2836(80)90283-1. [DOI] [PubMed] [Google Scholar]
- Chirgwin J. M., Przybyla A. E., MacDonald R. J., Rutter W. J. Isolation of biologically active ribonucleic acid from sources enriched in ribonuclease. Biochemistry. 1979 Nov 27;18(24):5294–5299. doi: 10.1021/bi00591a005. [DOI] [PubMed] [Google Scholar]
- DODYK F., ROTHSTEIN A. FACTORS INFLUENCING THE APPEARANCE OF INVERTASE IN SACCHAROMYCES CEREVISIAE. Arch Biochem Biophys. 1964 Mar;104:478–486. doi: 10.1016/0003-9861(64)90492-8. [DOI] [PubMed] [Google Scholar]
- Devereux J., Haeberli P., Smithies O. A comprehensive set of sequence analysis programs for the VAX. Nucleic Acids Res. 1984 Jan 11;12(1 Pt 1):387–395. doi: 10.1093/nar/12.1part1.387. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Drebot M. A., Barnes C. A., Singer R. A., Johnston G. C. Genetic assessment of stationary phase for cells of the yeast Saccharomyces cerevisiae. J Bacteriol. 1990 Jul;172(7):3584–3589. doi: 10.1128/jb.172.7.3584-3589.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Elliott B., Futcher B. Stress resistance of yeast cells is largely independent of cell cycle phase. Yeast. 1993 Jan;9(1):33–42. doi: 10.1002/yea.320090105. [DOI] [PubMed] [Google Scholar]
- Feinberg A. P., Vogelstein B. A technique for radiolabeling DNA restriction endonuclease fragments to high specific activity. Anal Biochem. 1983 Jul 1;132(1):6–13. doi: 10.1016/0003-2697(83)90418-9. [DOI] [PubMed] [Google Scholar]
- Ferguson J. J., Jr, Boll M., Holzer H. Yeast malate dehydrogenase: enzyme inactivation in catabolite repression. Eur J Biochem. 1967 Mar;1(1):21–25. doi: 10.1007/978-3-662-25813-2_4. [DOI] [PubMed] [Google Scholar]
- Finley D., Ozkaynak E., Varshavsky A. The yeast polyubiquitin gene is essential for resistance to high temperatures, starvation, and other stresses. Cell. 1987 Mar 27;48(6):1035–1046. doi: 10.1016/0092-8674(87)90711-2. [DOI] [PubMed] [Google Scholar]
- Goldstein A., Lampen J. O. Beta-D-fructofuranoside fructohydrolase from yeast. Methods Enzymol. 1975;42:504–511. doi: 10.1016/0076-6879(75)42159-0. [DOI] [PubMed] [Google Scholar]
- Harris S. D., Cotter D. A. Transport of yeast vacuolar trehalase to the vacuole. Can J Microbiol. 1988 Jul;34(7):835–838. doi: 10.1139/m88-143. [DOI] [PubMed] [Google Scholar]
- Holzer H. Proteolytic catabolite inactivation in Saccharomyces cerevisiae. Revis Biol Celular. 1989;21:305–319. [PubMed] [Google Scholar]
- Hubbard E. J., Yang X. L., Carlson M. Relationship of the cAMP-dependent protein kinase pathway to the SNF1 protein kinase and invertase expression in Saccharomyces cerevisiae. Genetics. 1992 Jan;130(1):71–80. doi: 10.1093/genetics/130.1.71. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ito H., Fukuda Y., Murata K., Kimura A. Transformation of intact yeast cells treated with alkali cations. J Bacteriol. 1983 Jan;153(1):163–168. doi: 10.1128/jb.153.1.163-168.1983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kaneko Y., Toh-e A., Oshima Y. Identification of the genetic locus for the structural gene and a new regulatory gene for the synthesis of repressible alkaline phosphatase in Saccharomyces cerevisiae. Mol Cell Biol. 1982 Feb;2(2):127–137. doi: 10.1128/mcb.2.2.127. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Keller F., Schellenberg M., Wiemken A. Localization of trehalase in vacuoles and of trehalose in the cytosol of yeast (Saccharomyces cerevisiae). Arch Microbiol. 1982 Jun;131(4):298–301. doi: 10.1007/BF00411175. [DOI] [PubMed] [Google Scholar]
- Klionsky D. J., Banta L. M., Emr S. D. Intracellular sorting and processing of a yeast vacuolar hydrolase: proteinase A propeptide contains vacuolar targeting information. Mol Cell Biol. 1988 May;8(5):2105–2116. doi: 10.1128/mcb.8.5.2105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Klionsky D. J., Emr S. D. A new class of lysosomal/vacuolar protein sorting signals. J Biol Chem. 1990 Apr 5;265(10):5349–5352. [PubMed] [Google Scholar]
- Klionsky D. J., Herman P. K., Emr S. D. The fungal vacuole: composition, function, and biogenesis. Microbiol Rev. 1990 Sep;54(3):266–292. doi: 10.1128/mr.54.3.266-292.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kozulić B., Barbarić S., Ries B., Mildner P. Study of the carbohydrate part of yeast acid phosphatase. Biochem Biophys Res Commun. 1984 Aug 16;122(3):1083–1090. doi: 10.1016/0006-291x(84)91202-6. [DOI] [PubMed] [Google Scholar]
- LOWRY O. H., ROSEBROUGH N. J., FARR A. L., RANDALL R. J. Protein measurement with the Folin phenol reagent. J Biol Chem. 1951 Nov;193(1):265–275. [PubMed] [Google Scholar]
- Law D. T., Segall J. The SPS100 gene of Saccharomyces cerevisiae is activated late in the sporulation process and contributes to spore wall maturation. Mol Cell Biol. 1988 Feb;8(2):912–922. doi: 10.1128/mcb.8.2.912. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lewis J. G., Northcott C. J., Learmonth R. P., Attfield P. V., Watson K. The need for consistent nomenclature and assessment of growth phases in diauxic cultures of Saccharomyces cerevisiae. J Gen Microbiol. 1993 Apr;139(4):835–839. doi: 10.1099/00221287-139-4-835. [DOI] [PubMed] [Google Scholar]
- Lillie S. H., Pringle J. R. Reserve carbohydrate metabolism in Saccharomyces cerevisiae: responses to nutrient limitation. J Bacteriol. 1980 Sep;143(3):1384–1394. doi: 10.1128/jb.143.3.1384-1394.1980. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Londesborough J., Varimo K. Characterization of two trehalases in baker's yeast. Biochem J. 1984 Apr 15;219(2):511–518. doi: 10.1042/bj2190511. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mittenbühler K., Holzer H. Characterization of different forms of yeast acid trehalase in the secretory pathway. Arch Microbiol. 1991;155(3):217–220. doi: 10.1007/BF00252203. [DOI] [PubMed] [Google Scholar]
- Mittenbühler K., Holzer H. Purification and characterization of acid trehalase from the yeast suc2 mutant. J Biol Chem. 1988 Jun 15;263(17):8537–8543. [PubMed] [Google Scholar]
- Moehle C. M., Tizard R., Lemmon S. K., Smart J., Jones E. W. Protease B of the lysosomelike vacuole of the yeast Saccharomyces cerevisiae is homologous to the subtilisin family of serine proteases. Mol Cell Biol. 1987 Dec;7(12):4390–4399. doi: 10.1128/mcb.7.12.4390. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Plesset J., Ludwig J. R., Cox B. S., McLaughlin C. S. Effect of cell cycle position on thermotolerance in Saccharomyces cerevisiae. J Bacteriol. 1987 Feb;169(2):779–784. doi: 10.1128/jb.169.2.779-784.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Robinson J. S., Klionsky D. J., Banta L. M., Emr S. D. Protein sorting in Saccharomyces cerevisiae: isolation of mutants defective in the delivery and processing of multiple vacuolar hydrolases. Mol Cell Biol. 1988 Nov;8(11):4936–4948. doi: 10.1128/mcb.8.11.4936. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rose M. D., Novick P., Thomas J. H., Botstein D., Fink G. R. A Saccharomyces cerevisiae genomic plasmid bank based on a centromere-containing shuttle vector. Gene. 1987;60(2-3):237–243. doi: 10.1016/0378-1119(87)90232-0. [DOI] [PubMed] [Google Scholar]
- Sanchez Y., Taulien J., Borkovich K. A., Lindquist S. Hsp104 is required for tolerance to many forms of stress. EMBO J. 1992 Jun;11(6):2357–2364. doi: 10.1002/j.1460-2075.1992.tb05295.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sanger F., Nicklen S., Coulson A. R. DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci U S A. 1977 Dec;74(12):5463–5467. doi: 10.1073/pnas.74.12.5463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shah H. C., Carlson G. P. Alteration by phenobarbital and 3-methyl-cholanthrene of functional and structural changes in rat liver due to carbon tetrachloride inhalation. J Pharmacol Exp Ther. 1975 Apr;193(1):281–292. [PubMed] [Google Scholar]
- Shahin M. M. Relationship between yield of protoplasts and growth phase in Saccharomyces. J Bacteriol. 1972 May;110(2):769–771. doi: 10.1128/jb.110.2.769-771.1972. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Southern E. M. Detection of specific sequences among DNA fragments separated by gel electrophoresis. J Mol Biol. 1975 Nov 5;98(3):503–517. doi: 10.1016/s0022-2836(75)80083-0. [DOI] [PubMed] [Google Scholar]
- Thevelein J. M. Regulation of trehalose mobilization in fungi. Microbiol Rev. 1984 Mar;48(1):42–59. doi: 10.1128/mr.48.1.42-59.1984. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Toh-E A., Nakamura H., Oshima Y. A gene controlling the synthesis of non specific alkaline phosphatase in Saccharomyces cerevisiae. Biochim Biophys Acta. 1976 Mar 25;428(1):182–192. doi: 10.1016/0304-4165(76)90119-7. [DOI] [PubMed] [Google Scholar]
- Werner-Washburne M., Braun E., Johnston G. C., Singer R. A. Stationary phase in the yeast Saccharomyces cerevisiae. Microbiol Rev. 1993 Jun;57(2):383–401. doi: 10.1128/mr.57.2.383-401.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wickerham L. J. A Critical Evaluation of the Nitrogen Assimilation Tests Commonly Used in the Classification of Yeasts. J Bacteriol. 1946 Sep;52(3):293–301. [PMC free article] [PubMed] [Google Scholar]
- de Nobel J. G., Klis F. M., Priem J., Munnik T., van den Ende H. The glucanase-soluble mannoproteins limit cell wall porosity in Saccharomyces cerevisiae. Yeast. 1990 Nov-Dec;6(6):491–499. doi: 10.1002/yea.320060606. [DOI] [PubMed] [Google Scholar]
- von Heijne G. A new method for predicting signal sequence cleavage sites. Nucleic Acids Res. 1986 Jun 11;14(11):4683–4690. doi: 10.1093/nar/14.11.4683. [DOI] [PMC free article] [PubMed] [Google Scholar]





