Abstract
In an effort to understand the mechanisms by which nonsense codons affect RNA metabolism in mammalian cells, nonsense mutations were generated within the gene for the secretory major urinary protein (MUP) of mice. The translation of MUP mRNA normally begins within exon 1 and terminates within exon 6, the penultimate exon. Through the use of Northern (RNA) blot hybridization and assays that couple reverse transcription and PCR, a nonsense mutation within codon 50 of exon 2 or codon 143 of exon 5 was found to reduce the abundance of fully spliced, nuclear MUP mRNA to 10 to 20% of normal without an additional reduction in the abundance of cytoplasmic mRNA. In contrast, a nonsense mutation within codon 172 of exon 5 was found to have no effects on the abundance of MUP mRNA. These findings suggest that a boundary between nonsense mutations that do and do not reduce the abundance of nuclear mRNA exists within the exon preceding the exon that harbors the normal site of translation termination. In this way, the boundary is analogous to the boundary that exists within the penultimate exon of the human gene for the cytosolic enzyme triosephosphate isomerase. Assays for exon skipping, i.e., the removal of an exon as a part of the flanking introns during the process of splicing, reveal that 0.1, 2.0, and 0.1% of MUP mRNA normally lack exon 5, exon 6, and exons 5 plus 6, respectively. Relative to normal, the two nonsense mutations within exon 5 increase the abundance of RNA lacking exon 5 on average 20-fold and increase the abundance of RNA lacking exons 5 plus 6 on average 5-fold. Since only one of these nonsense mutations also reduces the abundance of fully spliced nuclear mRNA to 10 to 20% of normal, the two mechanisms by which a nonsense mutation can alter nuclear RNA metabolism must be distinct. The analysis of missense mutations within codons 143 and 172, some of which retain the nonsense mutation, indicates that the reduction in the abundance of fully spliced nuclear mRNA is dependent upon the premature termination of MUP mRNA translation, whereas skipping is attributable to nonsense mutation-mediated changes in exon 5 structure rather than to the premature termination of translation. The increase in exon 5 skipping by either the nonsense or missense mutations within codon 172 correlates with a decrease in the complementarity of exon 5 to U1 snRNA. This suggests that a 5' splice site may extend as far as 12 nucleotides into the upstream exon, which is, to our knowledge, the largest extension.
Full text
PDF





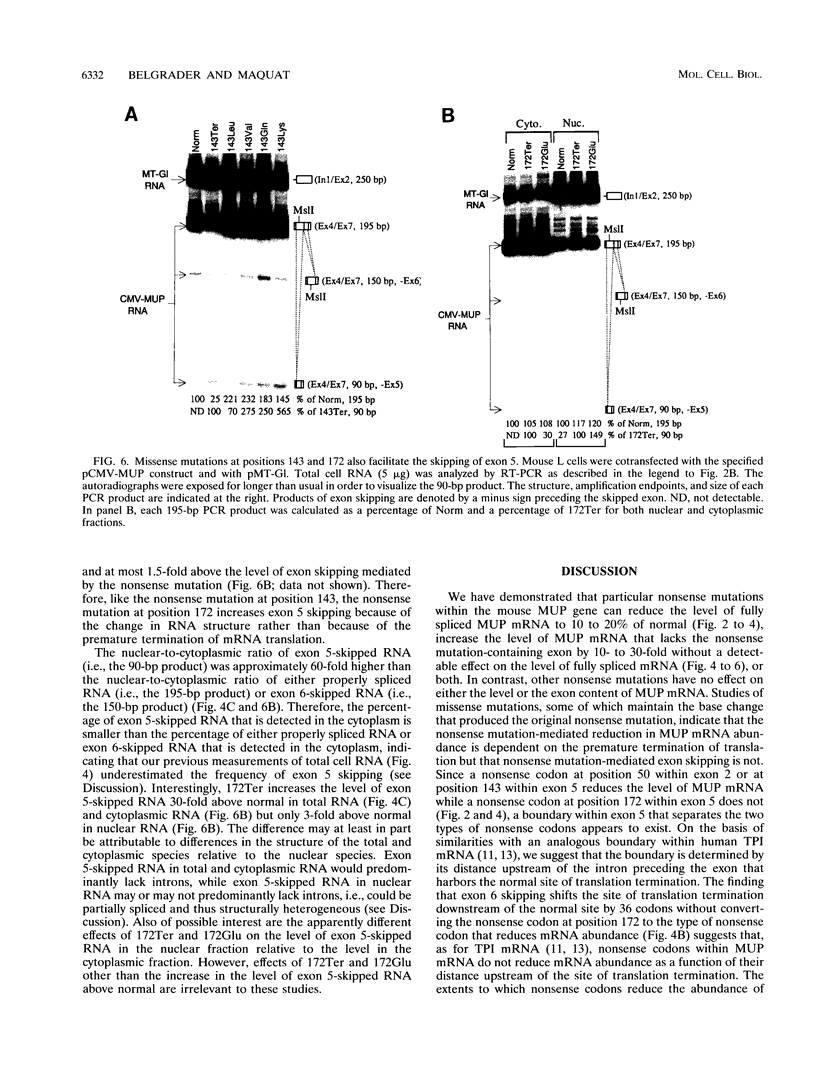




Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Bach G., Moskowitz S. M., Tieu P. T., Matynia A., Neufeld E. F. Molecular analysis of Hurler syndrome in Druze and Muslim Arab patients in Israel: multiple allelic mutations of the IDUA gene in a small geographic area. Am J Hum Genet. 1993 Aug;53(2):330–338. [PMC free article] [PubMed] [Google Scholar]
- Barker G. F., Beemon K. Nonsense codons within the Rous sarcoma virus gag gene decrease the stability of unspliced viral RNA. Mol Cell Biol. 1991 May;11(5):2760–2768. doi: 10.1128/mcb.11.5.2760. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Barker G. F., Beemon K. Rous sarcoma virus RNA stability requires an open reading frame in the gag gene and sequences downstream of the gag-pol junction. Mol Cell Biol. 1994 Mar;14(3):1986–1996. doi: 10.1128/mcb.14.3.1986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Baserga S. J., Benz E. J., Jr Beta-globin nonsense mutation: deficient accumulation of mRNA occurs despite normal cytoplasmic stability. Proc Natl Acad Sci U S A. 1992 Apr 1;89(7):2935–2939. doi: 10.1073/pnas.89.7.2935. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Baumann B., Potash M. J., Köhler G. Consequences of frameshift mutations at the immunoglobulin heavy chain locus of the mouse. EMBO J. 1985 Feb;4(2):351–359. doi: 10.1002/j.1460-2075.1985.tb03636.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Belgrader P., Cheng J., Maquat L. E. Evidence to implicate translation by ribosomes in the mechanism by which nonsense codons reduce the nuclear level of human triosephosphate isomerase mRNA. Proc Natl Acad Sci U S A. 1993 Jan 15;90(2):482–486. doi: 10.1073/pnas.90.2.482. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brown J. R., Daar I. O., Krug J. R., Maquat L. E. Characterization of the functional gene and several processed pseudogenes in the human triosephosphate isomerase gene family. Mol Cell Biol. 1985 Jul;5(7):1694–1706. doi: 10.1128/mcb.5.7.1694. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Capasso O., Bleecker G. C., Heintz N. Sequences controlling histone H4 mRNA abundance. EMBO J. 1987 Jun;6(6):1825–1831. doi: 10.1002/j.1460-2075.1987.tb02437.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Carothers A. M., Urlaub G., Grunberger D., Chasin L. A. Splicing mutants and their second-site suppressors at the dihydrofolate reductase locus in Chinese hamster ovary cells. Mol Cell Biol. 1993 Aug;13(8):5085–5098. doi: 10.1128/mcb.13.8.5085. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Carothers A. M., Urlaub G., Mucha J., Grunberger D., Chasin L. A. Point mutation analysis in a mammalian gene: rapid preparation of total RNA, PCR amplification of cDNA, and Taq sequencing by a novel method. Biotechniques. 1989 May;7(5):494-6, 498-9. [PubMed] [Google Scholar]
- Cheng J., Belgrader P., Zhou X., Maquat L. E. Introns are cis effectors of the nonsense-codon-mediated reduction in nuclear mRNA abundance. Mol Cell Biol. 1994 Sep;14(9):6317–6325. doi: 10.1128/mcb.14.9.6317. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cheng J., Fogel-Petrovic M., Maquat L. E. Translation to near the distal end of the penultimate exon is required for normal levels of spliced triosephosphate isomerase mRNA. Mol Cell Biol. 1990 Oct;10(10):5215–5225. doi: 10.1128/mcb.10.10.5215. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cheng J., Maquat L. E. Nonsense codons can reduce the abundance of nuclear mRNA without affecting the abundance of pre-mRNA or the half-life of cytoplasmic mRNA. Mol Cell Biol. 1993 Mar;13(3):1892–1902. doi: 10.1128/mcb.13.3.1892. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Clark A. J., Clissold P. M., Al Shawi R., Beattie P., Bishop J. Structure of mouse major urinary protein genes: different splicing configurations in the 3'-non-coding region. EMBO J. 1984 May;3(5):1045–1052. doi: 10.1002/j.1460-2075.1984.tb01925.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cooper T. A. In vitro splicing of cardiac troponin T precursors. Exon mutations disrupt splicing of the upstream intron. J Biol Chem. 1992 Mar 15;267(8):5330–5338. [PubMed] [Google Scholar]
- Cooper T. A., Ordahl C. P. Nucleotide substitutions within the cardiac troponin T alternative exon disrupt pre-mRNA alternative splicing. Nucleic Acids Res. 1989 Oct 11;17(19):7905–7921. doi: 10.1093/nar/17.19.7905. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dietz H. C., Valle D., Francomano C. A., Kendzior R. J., Jr, Pyeritz R. E., Cutting G. R. The skipping of constitutive exons in vivo induced by nonsense mutations. Science. 1993 Jan 29;259(5095):680–683. doi: 10.1126/science.8430317. [DOI] [PubMed] [Google Scholar]
- Feinberg A. P., Vogelstein B. A technique for radiolabeling DNA restriction endonuclease fragments to high specific activity. Anal Biochem. 1983 Jul 1;132(1):6–13. doi: 10.1016/0003-2697(83)90418-9. [DOI] [PubMed] [Google Scholar]
- Gaspar M. L., Meo T., Bourgarel P., Guenet J. L., Tosi M. A single base deletion in the Tfm androgen receptor gene creates a short-lived messenger RNA that directs internal translation initiation. Proc Natl Acad Sci U S A. 1991 Oct 1;88(19):8606–8610. doi: 10.1073/pnas.88.19.8606. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gibson R. A., Hajianpour A., Murer-Orlando M., Buchwald M., Mathew C. G. A nonsense mutation and exon skipping in the Fanconi anaemia group C gene. Hum Mol Genet. 1993 Jun;2(6):797–799. doi: 10.1093/hmg/2.6.797. [DOI] [PubMed] [Google Scholar]
- Graves R. A., Pandey N. B., Chodchoy N., Marzluff W. F. Translation is required for regulation of histone mRNA degradation. Cell. 1987 Feb 27;48(4):615–626. doi: 10.1016/0092-8674(87)90240-6. [DOI] [PubMed] [Google Scholar]
- Greenberg M. E., Hermanowski A. L., Ziff E. B. Effect of protein synthesis inhibitors on growth factor activation of c-fos, c-myc, and actin gene transcription. Mol Cell Biol. 1986 Apr;6(4):1050–1057. doi: 10.1128/mcb.6.4.1050. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hamada K., Kumazaki T., Mizuno K., Yokoro K. A small nuclear RNA, U5, can transform cells in vitro. Mol Cell Biol. 1989 Oct;9(10):4345–4356. doi: 10.1128/mcb.9.10.4345. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hampson R. K., La Follette L., Rottman F. M. Alternative processing of bovine growth hormone mRNA is influenced by downstream exon sequences. Mol Cell Biol. 1989 Apr;9(4):1604–1610. doi: 10.1128/mcb.9.4.1604. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Held W. A., Gallagher J. F., Hohman C. M., Kuhn N. J., Sampsell B. M., Hughes R. G., Jr Identification and characterization of functional genes encoding the mouse major urinary proteins. Mol Cell Biol. 1987 Oct;7(10):3705–3712. doi: 10.1128/mcb.7.10.3705. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Helfman D. M., Ricci W. M., Finn L. A. Alternative splicing of tropomyosin pre-mRNAs in vitro and in vivo. Genes Dev. 1988 Dec;2(12A):1627–1638. doi: 10.1101/gad.2.12a.1627. [DOI] [PubMed] [Google Scholar]
- Herrick D., Parker R., Jacobson A. Identification and comparison of stable and unstable mRNAs in Saccharomyces cerevisiae. Mol Cell Biol. 1990 May;10(5):2269–2284. doi: 10.1128/mcb.10.5.2269. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Inoue K., Hoshijima K., Higuchi I., Sakamoto H., Shimura Y. Binding of the Drosophila transformer and transformer-2 proteins to the regulatory elements of doublesex primary transcript for sex-specific RNA processing. Proc Natl Acad Sci U S A. 1992 Sep 1;89(17):8092–8096. doi: 10.1073/pnas.89.17.8092. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kessler O., Jiang Y., Chasin L. A. Order of intron removal during splicing of endogenous adenine phosphoribosyltransferase and dihydrofolate reductase pre-mRNA. Mol Cell Biol. 1993 Oct;13(10):6211–6222. doi: 10.1128/mcb.13.10.6211. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lavigueur A., La Branche H., Kornblihtt A. R., Chabot B. A splicing enhancer in the human fibronectin alternate ED1 exon interacts with SR proteins and stimulates U2 snRNP binding. Genes Dev. 1993 Dec;7(12A):2405–2417. doi: 10.1101/gad.7.12a.2405. [DOI] [PubMed] [Google Scholar]
- Leeds P., Peltz S. W., Jacobson A., Culbertson M. R. The product of the yeast UPF1 gene is required for rapid turnover of mRNAs containing a premature translational termination codon. Genes Dev. 1991 Dec;5(12A):2303–2314. doi: 10.1101/gad.5.12a.2303. [DOI] [PubMed] [Google Scholar]
- Leeds P., Wood J. M., Lee B. S., Culbertson M. R. Gene products that promote mRNA turnover in Saccharomyces cerevisiae. Mol Cell Biol. 1992 May;12(5):2165–2177. doi: 10.1128/mcb.12.5.2165. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Legrain P., Rosbash M. Some cis- and trans-acting mutants for splicing target pre-mRNA to the cytoplasm. Cell. 1989 May 19;57(4):573–583. doi: 10.1016/0092-8674(89)90127-x. [DOI] [PubMed] [Google Scholar]
- Levine B. J., Chodchoy N., Marzluff W. F., Skoultchi A. I. Coupling of replication type histone mRNA levels to DNA synthesis requires the stem-loop sequence at the 3' end of the mRNA. Proc Natl Acad Sci U S A. 1987 Sep;84(17):6189–6193. doi: 10.1073/pnas.84.17.6189. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Libri D., Goux-Pelletan M., Brody E., Fiszman M. Y. Exon as well as intron sequences are cis-regulating elements for the mutually exclusive alternative splicing of the beta tropomyosin gene. Mol Cell Biol. 1990 Oct;10(10):5036–5046. doi: 10.1128/mcb.10.10.5036. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ligtenberg M. J., Gennissen A. M., Vos H. L., Hilkens J. A single nucleotide polymorphism in an exon dictates allele dependent differential splicing of episialin mRNA. Nucleic Acids Res. 1991 Jan 25;19(2):297–301. doi: 10.1093/nar/19.2.297. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lim S. K., Sigmund C. D., Gross K. W., Maquat L. E. Nonsense codons in human beta-globin mRNA result in the production of mRNA degradation products. Mol Cell Biol. 1992 Mar;12(3):1149–1161. doi: 10.1128/mcb.12.3.1149. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Losson R., Lacroute F. Interference of nonsense mutations with eukaryotic messenger RNA stability. Proc Natl Acad Sci U S A. 1979 Oct;76(10):5134–5137. doi: 10.1073/pnas.76.10.5134. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mardon H. J., Sebastio G., Baralle F. E. A role for exon sequences in alternative splicing of the human fibronectin gene. Nucleic Acids Res. 1987 Oct 12;15(19):7725–7733. doi: 10.1093/nar/15.19.7725. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mayeda A., Helfman D. M., Krainer A. R. Modulation of exon skipping and inclusion by heterogeneous nuclear ribonucleoprotein A1 and pre-mRNA splicing factor SF2/ASF. Mol Cell Biol. 1993 May;13(5):2993–3001. doi: 10.1128/mcb.13.5.2993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mount S. M. A catalogue of splice junction sequences. Nucleic Acids Res. 1982 Jan 22;10(2):459–472. doi: 10.1093/nar/10.2.459. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Naeger L. K., Schoborg R. V., Zhao Q., Tullis G. E., Pintel D. J. Nonsense mutations inhibit splicing of MVM RNA in cis when they interrupt the reading frame of either exon of the final spliced product. Genes Dev. 1992 Jun;6(6):1107–1119. doi: 10.1101/gad.6.6.1107. [DOI] [PubMed] [Google Scholar]
- Nagoshi R. N., Baker B. S. Regulation of sex-specific RNA splicing at the Drosophila doublesex gene: cis-acting mutations in exon sequences alter sex-specific RNA splicing patterns. Genes Dev. 1990 Jan;4(1):89–97. doi: 10.1101/gad.4.1.89. [DOI] [PubMed] [Google Scholar]
- Naylor J. A., Green P. M., Rizza C. R., Giannelli F. Analysis of factor VIII mRNA reveals defects in everyone of 28 haemophilia A patients. Hum Mol Genet. 1993 Jan;2(1):11–17. doi: 10.1093/hmg/2.1.11. [DOI] [PubMed] [Google Scholar]
- Nesic D., Maquat L. E. Upstream introns influence the efficiency of final intron removal and RNA 3'-end formation. Genes Dev. 1994 Feb 1;8(3):363–375. doi: 10.1101/gad.8.3.363. [DOI] [PubMed] [Google Scholar]
- Newman A. J., Norman C. U5 snRNA interacts with exon sequences at 5' and 3' splice sites. Cell. 1992 Feb 21;68(4):743–754. doi: 10.1016/0092-8674(92)90149-7. [DOI] [PubMed] [Google Scholar]
- Nigro J. M., Cho K. R., Fearon E. R., Kern S. E., Ruppert J. M., Oliner J. D., Kinzler K. W., Vogelstein B. Scrambled exons. Cell. 1991 Feb 8;64(3):607–613. doi: 10.1016/0092-8674(91)90244-s. [DOI] [PubMed] [Google Scholar]
- Orkin S. H., Kazazian H. H., Jr The mutation and polymorphism of the human beta-globin gene and its surrounding DNA. Annu Rev Genet. 1984;18:131–171. doi: 10.1146/annurev.ge.18.120184.001023. [DOI] [PubMed] [Google Scholar]
- Parker R., Jacobson A. Translation and a 42-nucleotide segment within the coding region of the mRNA encoded by the MAT alpha 1 gene are involved in promoting rapid mRNA decay in yeast. Proc Natl Acad Sci U S A. 1990 Apr;87(7):2780–2784. doi: 10.1073/pnas.87.7.2780. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Peltz S. W., Brown A. H., Jacobson A. mRNA destabilization triggered by premature translational termination depends on at least three cis-acting sequence elements and one trans-acting factor. Genes Dev. 1993 Sep;7(9):1737–1754. doi: 10.1101/gad.7.9.1737. [DOI] [PubMed] [Google Scholar]
- Piechaczyk M., Lelay-Taha M. N., Sri-Widada J., Brunel C., Liautard J. P., Jeanteur P. Mouse DNA sequences complementary to small nuclear RNA U1. Nucleic Acids Res. 1982 Aug 11;10(15):4627–4640. doi: 10.1093/nar/10.15.4627. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Prowse K. R., Baumann H. Hepatocyte-stimulating factor, beta 2 interferon, and interleukin-1 enhance expression of the rat alpha 1-acid glycoprotein gene via a distal upstream regulatory region. Mol Cell Biol. 1988 Jan;8(1):42–51. doi: 10.1128/mcb.8.1.42. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Qian L., Theodor L., Carter M., Vu M. N., Sasaki A. W., Wilkinson M. F. T cell receptor-beta mRNA splicing: regulation of unusual splicing intermediates. Mol Cell Biol. 1993 Mar;13(3):1686–1696. doi: 10.1128/mcb.13.3.1686. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rapoport T. A. Transport of proteins across the endoplasmic reticulum membrane. Science. 1992 Nov 6;258(5084):931–936. doi: 10.1126/science.1332192. [DOI] [PubMed] [Google Scholar]
- Shapiro M. B., Senapathy P. RNA splice junctions of different classes of eukaryotes: sequence statistics and functional implications in gene expression. Nucleic Acids Res. 1987 Sep 11;15(17):7155–7174. doi: 10.1093/nar/15.17.7155. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shyu A. B., Belasco J. G., Greenberg M. E. Two distinct destabilizing elements in the c-fos message trigger deadenylation as a first step in rapid mRNA decay. Genes Dev. 1991 Feb;5(2):221–231. doi: 10.1101/gad.5.2.221. [DOI] [PubMed] [Google Scholar]
- Shyu A. B., Greenberg M. E., Belasco J. G. The c-fos transcript is targeted for rapid decay by two distinct mRNA degradation pathways. Genes Dev. 1989 Jan;3(1):60–72. doi: 10.1101/gad.3.1.60. [DOI] [PubMed] [Google Scholar]
- Simpson S. B., Stoltzfus C. M. Frameshift mutations in the v-src gene of avian sarcoma virus act in cis to specifically reduce v-src mRNA levels. Mol Cell Biol. 1994 Mar;14(3):1835–1844. doi: 10.1128/mcb.14.3.1835. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Steingrimsdottir H., Rowley G., Dorado G., Cole J., Lehmann A. R. Mutations which alter splicing in the human hypoxanthine-guanine phosphoribosyltransferase gene. Nucleic Acids Res. 1992 Mar 25;20(6):1201–1208. doi: 10.1093/nar/20.6.1201. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Streuli M., Saito H. Regulation of tissue-specific alternative splicing: exon-specific cis-elements govern the splicing of leukocyte common antigen pre-mRNA. EMBO J. 1989 Mar;8(3):787–796. doi: 10.1002/j.1460-2075.1989.tb03439.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sun Q., Hampson R. K., Rottman F. M. In vitro analysis of bovine growth hormone pre-mRNA alternative splicing. Involvement of exon sequences and trans-acting factor(s). J Biol Chem. 1993 Jul 25;268(21):15659–15666. [PubMed] [Google Scholar]
- Sun Q., Mayeda A., Hampson R. K., Krainer A. R., Rottman F. M. General splicing factor SF2/ASF promotes alternative splicing by binding to an exonic splicing enhancer. Genes Dev. 1993 Dec;7(12B):2598–2608. doi: 10.1101/gad.7.12b.2598. [DOI] [PubMed] [Google Scholar]
- Tian M., Maniatis T. A splicing enhancer complex controls alternative splicing of doublesex pre-mRNA. Cell. 1993 Jul 16;74(1):105–114. doi: 10.1016/0092-8674(93)90298-5. [DOI] [PubMed] [Google Scholar]
- Tsai A. Y., Streuli M., Saito H. Integrity of the exon 6 sequence is essential for tissue-specific alternative splicing of human leukocyte common antigen pre-mRNA. Mol Cell Biol. 1989 Oct;9(10):4550–4555. doi: 10.1128/mcb.9.10.4550. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Urlaub G., Mitchell P. J., Ciudad C. J., Chasin L. A. Nonsense mutations in the dihydrofolate reductase gene affect RNA processing. Mol Cell Biol. 1989 Jul;9(7):2868–2880. doi: 10.1128/mcb.9.7.2868. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wassarman D. A., Steitz J. A. Interactions of small nuclear RNA's with precursor messenger RNA during in vitro splicing. Science. 1992 Sep 25;257(5078):1918–1925. doi: 10.1126/science.1411506. [DOI] [PubMed] [Google Scholar]
- Watakabe A., Tanaka K., Shimura Y. The role of exon sequences in splice site selection. Genes Dev. 1993 Mar;7(3):407–418. doi: 10.1101/gad.7.3.407. [DOI] [PubMed] [Google Scholar]
- Wisdom R., Lee W. The protein-coding region of c-myc mRNA contains a sequence that specifies rapid mRNA turnover and induction by protein synthesis inhibitors. Genes Dev. 1991 Feb;5(2):232–243. doi: 10.1101/gad.5.2.232. [DOI] [PubMed] [Google Scholar]
- Xu R., Teng J., Cooper T. A. The cardiac troponin T alternative exon contains a novel purine-rich positive splicing element. Mol Cell Biol. 1993 Jun;13(6):3660–3674. doi: 10.1128/mcb.13.6.3660. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang L. H., Vrieling H., van Zeeland A. A., Jenssen D. Spectrum of spontaneously occurring mutations in the hprt gene of V79 Chinese hamster cells. J Mol Biol. 1992 Feb 5;223(3):627–635. doi: 10.1016/0022-2836(92)90979-t. [DOI] [PubMed] [Google Scholar]







