Abstract
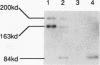
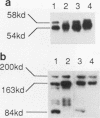
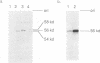
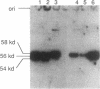
The Ty1 elements in the yeast Saccharomyces cerevisiae are a family of retrotransposons which transpose via a process similar to that of retroviral replication. We report here that the Ty1 transposition process can be blocked posttranscriptionally by treatment of cells with mating pheromones. When haploid yeast cells are treated with appropriate mating pheromones, the transposition frequency of a marked Ty1 element driven by the GAL1 promoter is greatly diminished. Ty1 viruslike particles (VLPs), the putative intermediates for transposition, can be isolated from mating pheromone-treated cells. These VLPs accumulate to normal levels but are aberrant in that they produce very few reverse transcripts of Ty1 RNA both in vivo and in vitro and contain subnormal amounts of p90-TYB and related proteins. In addition, a TYA phosphoprotein product accumulates in treated cells, and some species of TYB proteins have decreased stability. We also show that decreased transposition in mating pheromone-treated cells is not a consequence of simply blocking cell division, since Ty1 transposes at a nearly normal rate in yeast cells arrested in G2 by the drug nocodazole.
Full text
PDF







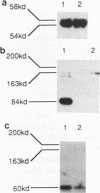
Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Adams S. E., Mellor J., Gull K., Sim R. B., Tuite M. F., Kingsman S. M., Kingsman A. J. The functions and relationships of Ty-VLP proteins in yeast reflect those of mammalian retroviral proteins. Cell. 1987 Apr 10;49(1):111–119. doi: 10.1016/0092-8674(87)90761-6. [DOI] [PubMed] [Google Scholar]
- Bender A., Sprague G. F., Jr Yeast peptide pheromones, a-factor and alpha-factor, activate a common response mechanism in their target cells. Cell. 1986 Dec 26;47(6):929–937. doi: 10.1016/0092-8674(86)90808-1. [DOI] [PubMed] [Google Scholar]
- Boeke J. D., Garfinkel D. J., Styles C. A., Fink G. R. Ty elements transpose through an RNA intermediate. Cell. 1985 Mar;40(3):491–500. doi: 10.1016/0092-8674(85)90197-7. [DOI] [PubMed] [Google Scholar]
- Boeke J. D., LaCroute F., Fink G. R. A positive selection for mutants lacking orotidine-5'-phosphate decarboxylase activity in yeast: 5-fluoro-orotic acid resistance. Mol Gen Genet. 1984;197(2):345–346. doi: 10.1007/BF00330984. [DOI] [PubMed] [Google Scholar]
- Boeke J. D., Xu H., Fink G. R. A general method for the chromosomal amplification of genes in yeast. Science. 1988 Jan 15;239(4837):280–282. doi: 10.1126/science.2827308. [DOI] [PubMed] [Google Scholar]
- Chan R. K., Otte C. A. Isolation and genetic analysis of Saccharomyces cerevisiae mutants supersensitive to G1 arrest by a factor and alpha factor pheromones. Mol Cell Biol. 1982 Jan;2(1):11–20. doi: 10.1128/mcb.2.1.11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chan R. K., Otte C. A. Physiological characterization of Saccharomyces cerevisiae mutants supersensitive to G1 arrest by a factor and alpha factor pheromones. Mol Cell Biol. 1982 Jan;2(1):21–29. doi: 10.1128/mcb.2.1.21. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cross F., Hartwell L. H., Jackson C., Konopka J. B. Conjugation in Saccharomyces cerevisiae. Annu Rev Cell Biol. 1988;4:429–457. doi: 10.1146/annurev.cb.04.110188.002241. [DOI] [PubMed] [Google Scholar]
- Dietzel C., Kurjan J. Pheromonal regulation and sequence of the Saccharomyces cerevisiae SST2 gene: a model for desensitization to pheromone. Mol Cell Biol. 1987 Dec;7(12):4169–4177. doi: 10.1128/mcb.7.12.4169. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dietzel C., Kurjan J. The yeast SCG1 gene: a G alpha-like protein implicated in the a- and alpha-factor response pathway. Cell. 1987 Sep 25;50(7):1001–1010. doi: 10.1016/0092-8674(87)90166-8. [DOI] [PubMed] [Google Scholar]
- Doolittle R. F., Feng D. F., Johnson M. S., McClure M. A. Origins and evolutionary relationships of retroviruses. Q Rev Biol. 1989 Mar;64(1):1–30. doi: 10.1086/416128. [DOI] [PubMed] [Google Scholar]
- Eichinger D. J., Boeke J. D. A specific terminal structure is required for Ty1 transposition. Genes Dev. 1990 Mar;4(3):324–330. doi: 10.1101/gad.4.3.324. [DOI] [PubMed] [Google Scholar]
- Eichinger D. J., Boeke J. D. The DNA intermediate in yeast Ty1 element transposition copurifies with virus-like particles: cell-free Ty1 transposition. Cell. 1988 Sep 23;54(7):955–966. doi: 10.1016/0092-8674(88)90110-9. [DOI] [PubMed] [Google Scholar]
- Elder R. T., St John T. P., Stinchcomb D. T., Davis R. W., Scherer S., Davis R. W. Studies on the transposable element Ty1 of yeast. I. RNA homologous to Ty1. II. Recombination and expression of Ty1 and adjacent sequences. Cold Spring Harb Symp Quant Biol. 1981;45(Pt 2):581–591. doi: 10.1101/sqb.1981.045.01.075. [DOI] [PubMed] [Google Scholar]
- Garfinkel D. J., Boeke J. D., Fink G. R. Ty element transposition: reverse transcriptase and virus-like particles. Cell. 1985 Sep;42(2):507–517. doi: 10.1016/0092-8674(85)90108-4. [DOI] [PubMed] [Google Scholar]
- Kessler S. W. Rapid isolation of antigens from cells with a staphylococcal protein A-antibody adsorbent: parameters of the interaction of antibody-antigen complexes with protein A. J Immunol. 1975 Dec;115(6):1617–1624. [PubMed] [Google Scholar]
- Mellor J., Malim M. H., Gull K., Tuite M. F., McCready S., Dibbayawan T., Kingsman S. M., Kingsman A. J. Reverse transcriptase activity and Ty RNA are associated with virus-like particles in yeast. Nature. 1985 Dec 12;318(6046):583–586. doi: 10.1038/318583a0. [DOI] [PubMed] [Google Scholar]
- Miyajima I., Nakafuku M., Nakayama N., Brenner C., Miyajima A., Kaibuchi K., Arai K., Kaziro Y., Matsumoto K. GPA1, a haploid-specific essential gene, encodes a yeast homolog of mammalian G protein which may be involved in mating factor signal transduction. Cell. 1987 Sep 25;50(7):1011–1019. doi: 10.1016/0092-8674(87)90167-x. [DOI] [PubMed] [Google Scholar]
- Müller F., Brühl K. H., Freidel K., Kowallik K. V., Ciriacy M. Processing of TY1 proteins and formation of Ty1 virus-like particles in Saccharomyces cerevisiae. Mol Gen Genet. 1987 May;207(2-3):421–429. doi: 10.1007/BF00331610. [DOI] [PubMed] [Google Scholar]
- Nakayama N., Miyajima A., Arai K. Common signal transduction system shared by STE2 and STE3 in haploid cells of Saccharomyces cerevisiae: autocrine cell-cycle arrest results from forced expression of STE2. EMBO J. 1987 Jan;6(1):249–254. doi: 10.1002/j.1460-2075.1987.tb04746.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Richardson H. E., Wittenberg C., Cross F., Reed S. I. An essential G1 function for cyclin-like proteins in yeast. Cell. 1989 Dec 22;59(6):1127–1133. doi: 10.1016/0092-8674(89)90768-x. [DOI] [PubMed] [Google Scholar]
- Teague M. A., Chaleff D. T., Errede B. Nucleotide sequence of the yeast regulatory gene STE7 predicts a protein homologous to protein kinases. Proc Natl Acad Sci U S A. 1986 Oct;83(19):7371–7375. doi: 10.1073/pnas.83.19.7371. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Towbin H., Staehelin T., Gordon J. Electrophoretic transfer of proteins from polyacrylamide gels to nitrocellulose sheets: procedure and some applications. Proc Natl Acad Sci U S A. 1979 Sep;76(9):4350–4354. doi: 10.1073/pnas.76.9.4350. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Whiteway M., Hougan L., Dignard D., Thomas D. Y., Bell L., Saari G. C., Grant F. J., O'Hara P., MacKay V. L. The STE4 and STE18 genes of yeast encode potential beta and gamma subunits of the mating factor receptor-coupled G protein. Cell. 1989 Feb 10;56(3):467–477. doi: 10.1016/0092-8674(89)90249-3. [DOI] [PubMed] [Google Scholar]
- Winston F., Durbin K. J., Fink G. R. The SPT3 gene is required for normal transcription of Ty elements in S. cerevisiae. Cell. 1984 Dec;39(3 Pt 2):675–682. doi: 10.1016/0092-8674(84)90474-4. [DOI] [PubMed] [Google Scholar]
- Xu H., Boeke J. D. High-frequency deletion between homologous sequences during retrotransposition of Ty elements in Saccharomyces cerevisiae. Proc Natl Acad Sci U S A. 1987 Dec;84(23):8553–8557. doi: 10.1073/pnas.84.23.8553. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xu H., Boeke J. D. Host genes that influence transposition in yeast: the abundance of a rare tRNA regulates Ty1 transposition frequency. Proc Natl Acad Sci U S A. 1990 Nov;87(21):8360–8364. doi: 10.1073/pnas.87.21.8360. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xu H., Boeke J. D. Localization of sequences required in cis for yeast Ty1 element transposition near the long terminal repeats: analysis of mini-Ty1 elements. Mol Cell Biol. 1990 Jun;10(6):2695–2702. doi: 10.1128/mcb.10.6.2695. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Youngren S. D., Boeke J. D., Sanders N. J., Garfinkel D. J. Functional organization of the retrotransposon Ty from Saccharomyces cerevisiae: Ty protease is required for transposition. Mol Cell Biol. 1988 Apr;8(4):1421–1431. doi: 10.1128/mcb.8.4.1421. [DOI] [PMC free article] [PubMed] [Google Scholar]