Abstract
The GCD2 protein is a translational repressor of GCN4, the transcriptional activator of multiple amino acid biosynthetic genes in Saccharomyces cerevisiae. We present evidence that GCD2 has a general function in the initiation of protein synthesis in addition to its gene-specific role in translational control of GCN4 expression. Two temperature-sensitive lethal gcd2 mutations result in sensitivity to inhibitors of protein synthesis at the permissive temperature, and the gcd2-503 mutation leads to reduced incorporation of labeled leucine into total protein following a shift to the restrictive temperature of 36 degrees C. The gcd2-503 mutation also results in polysome runoff, accumulation of inactive 80S ribosomal couples, and accumulation of at least one of the subunits of the general translation initiation factor 2 (eIF-2 alpha) in 43S-48S particles following a shift to the restrictive temperature. The gcd2-502 mutation causes accumulation of 40S subunits in polysomes, known as halfmers, that are indicative of reduced 40S-60S subunit joining at the initiation codon. These phenotypes suggest that GCD2 functions in the translation initiation pathway at a step following the binding of eIF-2.GTP.Met-tRNA(iMet) to 40S ribosomal subunits. consistent with this hypothesis, we found that inhibiting 40S-60S subunit joining by deleting one copy (RPL16B) of the duplicated gene encoding the 60S ribosomal protein L16 qualitatively mimics the phenotype of gcd2 mutations in causing derepression of GCN4 expression under nonstarvation conditions. However, deletion of RPL16B also prevents efficient derepression of GCN4 under starvation conditions, indicating that lowering the concentration of 60S subunits and reducing GCD2 function affect translation initiation at GCN4 in different ways. This distinction is in accord with a recently proposed model for GCN4 translational control in which ribosomal reinitiation at short upstream open reading frames in the leader of GCN4 mRNA is suppressed under amino acid starvation conditions to allow for increased reinitiation at the GCN4 start codon.
Full text
PDF



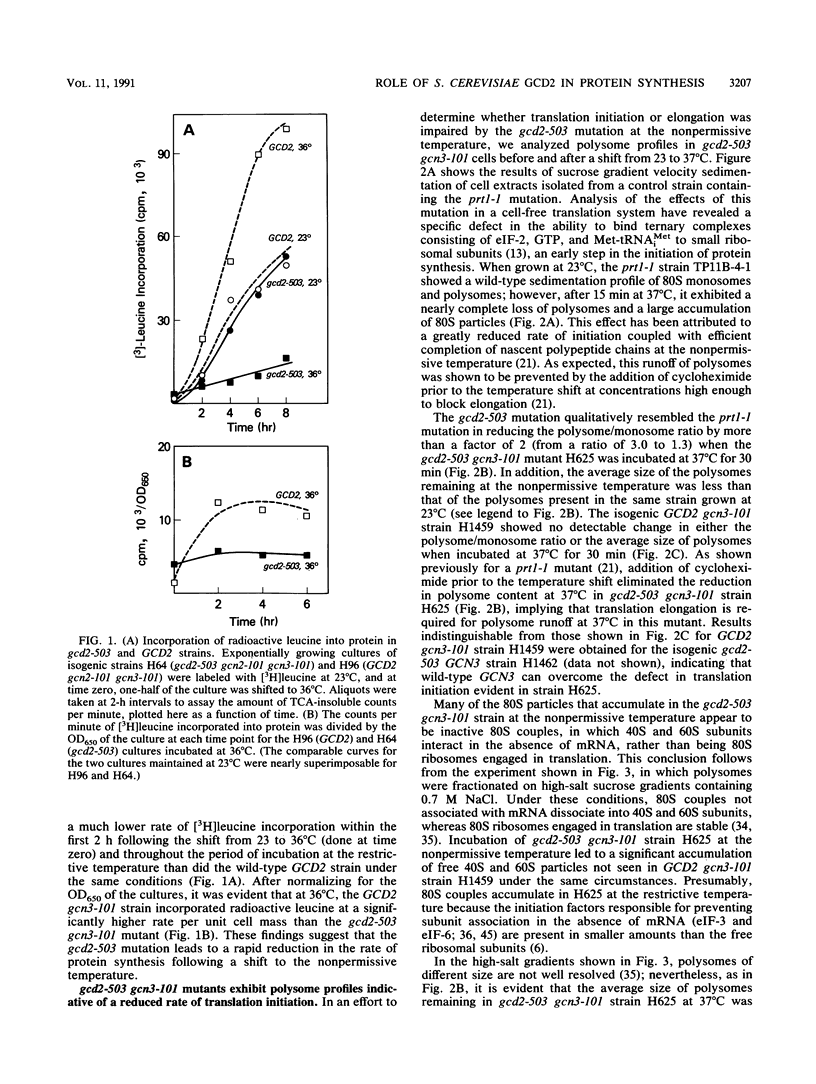









Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Abastado J. P., Miller P. F., Jackson B. M., Hinnebusch A. G. Suppression of ribosomal reinitiation at upstream open reading frames in amino acid-starved cells forms the basis for GCN4 translational control. Mol Cell Biol. 1991 Jan;11(1):486–496. doi: 10.1128/mcb.11.1.486. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Abovich N., Gritz L., Tung L., Rosbash M. Effect of RP51 gene dosage alterations on ribosome synthesis in Saccharomyces cerevisiae. Mol Cell Biol. 1985 Dec;5(12):3429–3435. doi: 10.1128/mcb.5.12.3429. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Baliga B. S., Pronczuk A. W., Munro H. N. Mechanism of cycloheximide inhibition of protein synthesis in a cell-free system prepared from rat liver. J Biol Chem. 1969 Aug 25;244(16):4480–4489. [PubMed] [Google Scholar]
- Boeke J. D., LaCroute F., Fink G. R. A positive selection for mutants lacking orotidine-5'-phosphate decarboxylase activity in yeast: 5-fluoro-orotic acid resistance. Mol Gen Genet. 1984;197(2):345–346. doi: 10.1007/BF00330984. [DOI] [PubMed] [Google Scholar]
- Botstein D., Falco S. C., Stewart S. E., Brennan M., Scherer S., Stinchcomb D. T., Struhl K., Davis R. W. Sterile host yeasts (SHY): a eukaryotic system of biological containment for recombinant DNA experiments. Gene. 1979 Dec;8(1):17–24. doi: 10.1016/0378-1119(79)90004-0. [DOI] [PubMed] [Google Scholar]
- Browning K. S., Humphreys J., Hobbs W., Smith G. B., Ravel J. M. Determination of the amounts of the protein synthesis initiation and elongation factors in wheat germ. J Biol Chem. 1990 Oct 15;265(29):17967–17973. [PubMed] [Google Scholar]
- Cigan A. M., Foiani M., Hannig E. M., Hinnebusch A. G. Complex formation by positive and negative translational regulators of GCN4. Mol Cell Biol. 1991 Jun;11(6):3217–3228. doi: 10.1128/mcb.11.6.3217. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cigan A. M., Pabich E. K., Feng L., Donahue T. F. Yeast translation initiation suppressor sui2 encodes the alpha subunit of eukaryotic initiation factor 2 and shares sequence identity with the human alpha subunit. Proc Natl Acad Sci U S A. 1989 Apr;86(8):2784–2788. doi: 10.1073/pnas.86.8.2784. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cooper T. G., Bossinger J. Selective inhibition of protein synthesis initiation in Saccharomyces cerevisiae by low concentrations of cycloheximide. J Biol Chem. 1976 Nov 25;251(22):7278–7280. [PubMed] [Google Scholar]
- De Benedetti A., Baglioni C. Inhibition of mRNA binding to ribosomes by localized activation of dsRNA-dependent protein kinase. Nature. 1984 Sep 6;311(5981):79–81. doi: 10.1038/311079a0. [DOI] [PubMed] [Google Scholar]
- De Benedetti A., Baglioni C. Phosphorylation of initiation factor eIF-2 alpha, binding of mRNA to 48 S complexes, and its reutilization in initiation of protein synthesis. J Biol Chem. 1983 Dec 10;258(23):14556–14562. [PubMed] [Google Scholar]
- Donahue T. F., Cigan A. M., Pabich E. K., Valavicius B. C. Mutations at a Zn(II) finger motif in the yeast eIF-2 beta gene alter ribosomal start-site selection during the scanning process. Cell. 1988 Aug 26;54(5):621–632. doi: 10.1016/s0092-8674(88)80006-0. [DOI] [PubMed] [Google Scholar]
- Feinberg A. P., Vogelstein B. "A technique for radiolabeling DNA restriction endonuclease fragments to high specific activity". Addendum. Anal Biochem. 1984 Feb;137(1):266–267. doi: 10.1016/0003-2697(84)90381-6. [DOI] [PubMed] [Google Scholar]
- Feinberg B., McLaughlin C. S., Moldave K. Analysis of temperature-sensitive mutant ts 187 of Saccharomyces cerevisiae altered in a component required for the initiation of protein synthesis. J Biol Chem. 1982 Sep 25;257(18):10846–10851. [PubMed] [Google Scholar]
- Gross M., Redman R., Kaplansky D. A. Evidence that the primary effect of phosphorylation of eukaryotic initiation factor 2(alpha) in rabbit reticulocyte lysate is inhibition of the release of eukaryotic initiation factor-2.GDP from 60 S ribosomal subunits. J Biol Chem. 1985 Aug 5;260(16):9491–9500. [PubMed] [Google Scholar]
- Gross M., Wing M., Rundquist C., Rubino M. S. Evidence that phosphorylation of eIF-2(alpha) prevents the eIF-2B-mediated dissociation of eIF-2 X GDP from the 60 S subunit of complete initiation complexes. J Biol Chem. 1987 May 15;262(14):6899–6907. [PubMed] [Google Scholar]
- Hannig E. M., Hinnebusch A. G. Molecular analysis of GCN3, a translational activator of GCN4: evidence for posttranslational control of GCN3 regulatory function. Mol Cell Biol. 1988 Nov;8(11):4808–4820. doi: 10.1128/mcb.8.11.4808. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hannig E. M., Williams N. P., Wek R. C., Hinnebusch A. G. The translational activator GCN3 functions downstream from GCN1 and GCN2 in the regulatory pathway that couples GCN4 expression to amino acid availability in Saccharomyces cerevisiae. Genetics. 1990 Nov;126(3):549–562. doi: 10.1093/genetics/126.3.549. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hansen L. J., Huang W. I., Jagus R. Inhibitor of translational initiation in sea urchin eggs prevents mRNA utilization. J Biol Chem. 1987 May 5;262(13):6114–6120. [PubMed] [Google Scholar]
- Harashima S., Hannig E. M., Hinnebusch A. G. Interactions between positive and negative regulators of GCN4 controlling gene expression and entry into the yeast cell cycle. Genetics. 1987 Nov;117(3):409–419. doi: 10.1093/genetics/117.3.409. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Harashima S., Hinnebusch A. G. Multiple GCD genes required for repression of GCN4, a transcriptional activator of amino acid biosynthetic genes in Saccharomyces cerevisiae. Mol Cell Biol. 1986 Nov;6(11):3990–3998. doi: 10.1128/mcb.6.11.3990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hartwell L. H., McLaughlin C. S. A mutant of yeast apparently defective in the initiation of protein synthesis. Proc Natl Acad Sci U S A. 1969 Feb;62(2):468–474. doi: 10.1073/pnas.62.2.468. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Helser T. L., Baan R. A., Dahlberg A. E. Characterization of a 40S ribosomal subunit complex in polyribosomes of Saccharomyces cerevisiae treated with cycloheximide. Mol Cell Biol. 1981 Jan;1(1):51–57. doi: 10.1128/mcb.1.1.51. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hershey J. W. Protein phosphorylation controls translation rates. J Biol Chem. 1989 Dec 15;264(35):20823–20826. [PubMed] [Google Scholar]
- Hinnebusch A. G. Evidence for translational regulation of the activator of general amino acid control in yeast. Proc Natl Acad Sci U S A. 1984 Oct;81(20):6442–6446. doi: 10.1073/pnas.81.20.6442. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hinnebusch A. G. Mechanisms of gene regulation in the general control of amino acid biosynthesis in Saccharomyces cerevisiae. Microbiol Rev. 1988 Jun;52(2):248–273. doi: 10.1128/mr.52.2.248-273.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hoerz W., McCarty K. S. Initiation of protein synthesis in a rabbit reticulocyte lysate system. Biochim Biophys Acta. 1971 Jan 28;228(2):526–535. doi: 10.1016/0005-2787(71)90058-x. [DOI] [PubMed] [Google Scholar]
- Ito H., Fukuda Y., Murata K., Kimura A. Transformation of intact yeast cells treated with alkali cations. J Bacteriol. 1983 Jan;153(1):163–168. doi: 10.1128/jb.153.1.163-168.1983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kappen L. S., Goldberg I. H. Analysis of the two steps in polypeptide chain initiation inhibited by pactamycin. Biochemistry. 1976 Feb 24;15(4):811–818. doi: 10.1021/bi00649a013. [DOI] [PubMed] [Google Scholar]
- Kappen L. S., Suzuki H., Goldberg I. H. Inhibition of reticulocyte peptide-chain initiation by pactamycin: accumulation of inactive ribosomal initiation complexes. Proc Natl Acad Sci U S A. 1973 Jan;70(1):22–26. doi: 10.1073/pnas.70.1.22. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kleid D. G., Yansura D., Small B., Dowbenko D., Moore D. M., Grubman M. J., McKercher P. D., Morgan D. O., Robertson B. H., Bachrach H. L. Cloned viral protein vaccine for foot-and-mouth disease: responses in cattle and swine. Science. 1981 Dec 4;214(4525):1125–1129. doi: 10.1126/science.6272395. [DOI] [PubMed] [Google Scholar]
- Kozak M., Shatkin A. J. Migration of 40 S ribosomal subunits on messenger RNA in the presence of edeine. J Biol Chem. 1978 Sep 25;253(18):6568–6577. [PubMed] [Google Scholar]
- Laemmli U. K. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature. 1970 Aug 15;227(5259):680–685. doi: 10.1038/227680a0. [DOI] [PubMed] [Google Scholar]
- Lucchini G., Hinnebusch A. G., Chen C., Fink G. R. Positive regulatory interactions of the HIS4 gene of Saccharomyces cerevisiae. Mol Cell Biol. 1984 Jul;4(7):1326–1333. doi: 10.1128/mcb.4.7.1326. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Martin T. E. A simple general method to determine the proportion of active ribosomes in eukaryotic cells. Exp Cell Res. 1973 Aug;80(2):496–498. doi: 10.1016/0014-4827(73)90333-9. [DOI] [PubMed] [Google Scholar]
- Martin T. E., Hartwell L. H. Resistance of active yeast ribosomes to dissociation by KCl. J Biol Chem. 1970 Mar 25;245(6):1504–1506. [PubMed] [Google Scholar]
- Moldave K. Eukaryotic protein synthesis. Annu Rev Biochem. 1985;54:1109–1149. doi: 10.1146/annurev.bi.54.070185.005333. [DOI] [PubMed] [Google Scholar]
- Mueller P. P., Harashima S., Hinnebusch A. G. A segment of GCN4 mRNA containing the upstream AUG codons confers translational control upon a heterologous yeast transcript. Proc Natl Acad Sci U S A. 1987 May;84(9):2863–2867. doi: 10.1073/pnas.84.9.2863. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mueller P. P., Hinnebusch A. G. Multiple upstream AUG codons mediate translational control of GCN4. Cell. 1986 Apr 25;45(2):201–207. doi: 10.1016/0092-8674(86)90384-3. [DOI] [PubMed] [Google Scholar]
- Niederberger P., Aebi M., Hütter R. Identification and characterization of four new GCD genes in Saccharomyces cerevisiae. Curr Genet. 1986;10(9):657–664. doi: 10.1007/BF00410913. [DOI] [PubMed] [Google Scholar]
- Paddon C. J., Hannig E. M., Hinnebusch A. G. Amino acid sequence similarity between GCN3 and GCD2, positive and negative translational regulators of GCN4: evidence for antagonism by competition. Genetics. 1989 Jul;122(3):551–559. doi: 10.1093/genetics/122.3.551. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Paddon C. J., Hinnebusch A. G. gcd12 mutations are gcn3-dependent alleles of GCD2, a negative regulator of GCN4 in the general amino acid control of Saccharomyces cerevisiae. Genetics. 1989 Jul;122(3):543–550. doi: 10.1093/genetics/122.3.543. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Peterson D. T., Safer B., Merrick W. C. Role of eukaryotic initiation factor 5 in the formation of 80 S initiation complexes. J Biol Chem. 1979 Aug 25;254(16):7730–7735. [PubMed] [Google Scholar]
- Petes T. D., Hereford L. M., Skryabin K. G. Characterization of two types of yeast ribosomal DNA genes. J Bacteriol. 1978 Apr;134(1):295–305. doi: 10.1128/jb.134.1.295-305.1978. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Raychaudhuri P., Stringer E. A., Valenzuela D. M., Maitra U. Ribosomal subunit antiassociation activity in rabbit reticulocyte lysates. Evidence for a low molecular weight ribosomal subunit antiassociation protein factor (Mr = 25,000). J Biol Chem. 1984 Oct 10;259(19):11930–11935. [PubMed] [Google Scholar]
- Rhoads R. E. Cap recognition and the entry of mRNA into the protein synthesis initiation cycle. Trends Biochem Sci. 1988 Feb;13(2):52–56. doi: 10.1016/0968-0004(88)90028-x. [DOI] [PubMed] [Google Scholar]
- Rotenberg M. O., Moritz M., Woolford J. L., Jr Depletion of Saccharomyces cerevisiae ribosomal protein L16 causes a decrease in 60S ribosomal subunits and formation of half-mer polyribosomes. Genes Dev. 1988 Feb;2(2):160–172. doi: 10.1101/gad.2.2.160. [DOI] [PubMed] [Google Scholar]
- Sonenberg N. Measures and countermeasures in the modulation of initiation factor activities by viruses. New Biol. 1990 May;2(5):402–409. [PubMed] [Google Scholar]
- Struhl K., Stinchcomb D. T., Scherer S., Davis R. W. High-frequency transformation of yeast: autonomous replication of hybrid DNA molecules. Proc Natl Acad Sci U S A. 1979 Mar;76(3):1035–1039. doi: 10.1073/pnas.76.3.1035. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Szczesna E., Filipowicz W. Faithful and efficient translation of viral and cellular eukaryotic mRNAs in a cell-free S-27 extract of Saccharomyces cerevisiae. Biochem Biophys Res Commun. 1980 Jan 29;92(2):563–569. doi: 10.1016/0006-291x(80)90370-8. [DOI] [PubMed] [Google Scholar]
- Tzamarias D., Roussou I., Thireos G. Coupling of GCN4 mRNA translational activation with decreased rates of polypeptide chain initiation. Cell. 1989 Jun 16;57(6):947–954. doi: 10.1016/0092-8674(89)90333-4. [DOI] [PubMed] [Google Scholar]
- Williams N. P., Hinnebusch A. G., Donahue T. F. Mutations in the structural genes for eukaryotic initiation factors 2 alpha and 2 beta of Saccharomyces cerevisiae disrupt translational control of GCN4 mRNA. Proc Natl Acad Sci U S A. 1989 Oct;86(19):7515–7519. doi: 10.1073/pnas.86.19.7515. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wolfner M., Yep D., Messenguy F., Fink G. R. Integration of amino acid biosynthesis into the cell cycle of Saccharomyces cerevisiae. J Mol Biol. 1975 Aug 5;96(2):273–290. doi: 10.1016/0022-2836(75)90348-4. [DOI] [PubMed] [Google Scholar]
- van Venrooij W. J., van Eenbergen J., Janssen A. P. Effect of anisomycin on the cellular level of native ribosomal subunits. Biochemistry. 1977 May 31;16(11):2343–2348. doi: 10.1021/bi00630a006. [DOI] [PubMed] [Google Scholar]





