Abstract
The SLP1 gene, which is involved in the expression of vacuolar functions in the yeast Saccharomyces cerevisiae (K. Kitamoto, K. Yoshizawa, Y. Ohsumi, and Y. Anraku, J. Bacteriol. 170:2687-2691, 1988), has been cloned from a yeast genomic library by complementation of the slp1-1 mutation. The isolated plasmid has a 7.8-kilobase BamHI-BamHI fragment that is sufficient to complement several characteristic phenotypes of the slp1-1 mutation. The fragment was integrated at the chromosomal SLP1 locus, indicating that it contains an authentic SLP1 gene. By DNA sequencing of the SLP1 gene, an open reading frame of 2,073 base pairs coding for a polypeptide of 691 amino acid residues (Mr, 79,270) was found. Gene disruption of the chromosomal SLP1 did not cause a lethal event. Vacuolar proteins in the delta slp1 mutant are not processed to vacuolar forms but remain in Golgi-modified forms. Carboxypeptidase Y in the delta slp1 mutant is localized mainly to the outsides of the cells. delta slp1 mutant cells have no prominent vacuolar structures but contain numerous vesicles in the cytoplasm, as seen by electron microscopy. Genetic and molecular biological analyses revealed that SLP1 is identical to VPS33, which is required for vacuolar protein sorting as reported by Robinson et al. (J. S. Robinson, D. J. Klionsky, L. M. Banta, and S. D. Emr, Mol. Cell. Biol. 8:4936-4948, 1988). These results indicate that the SLP1 (VPS33) gene is involved in the sorting of vacuolar proteins from the Golgi apparatus and their targeting to the vacuole and that it is required for the morphogenesis of vacuoles and subsequent expression of vacuolar functions.
Full text
PDF




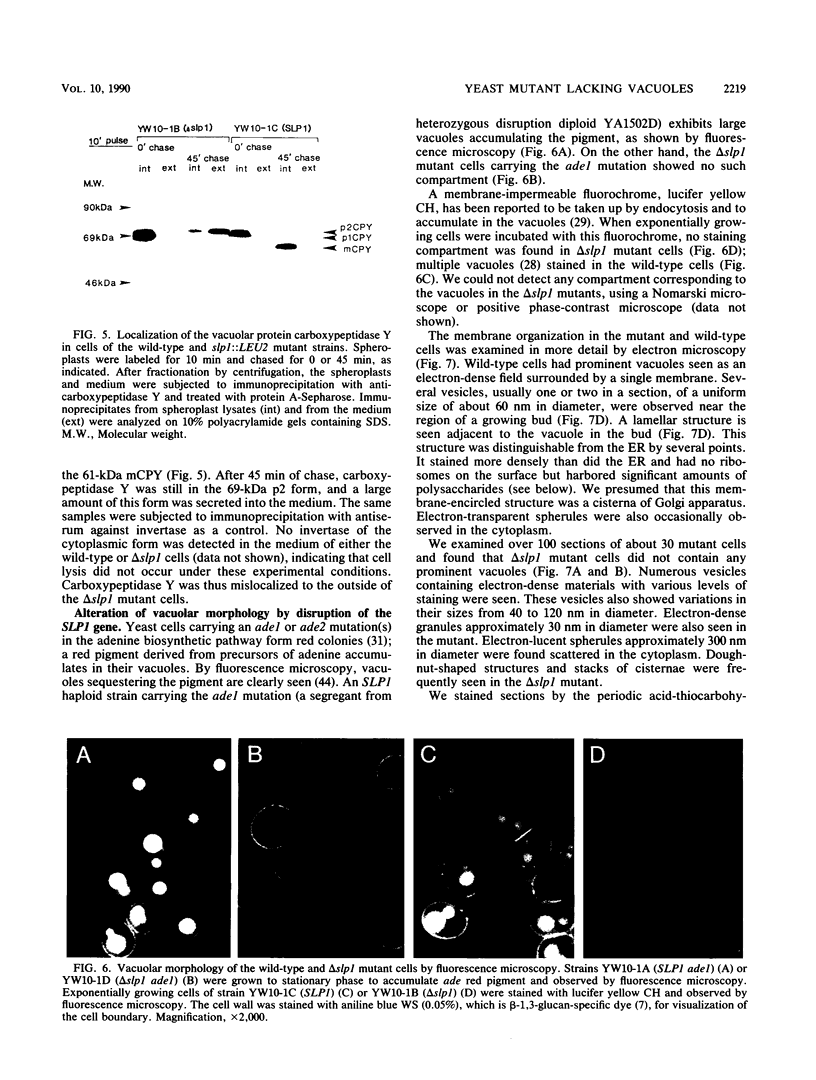




Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Ammerer G., Hunter C. P., Rothman J. H., Saari G. C., Valls L. A., Stevens T. H. PEP4 gene of Saccharomyces cerevisiae encodes proteinase A, a vacuolar enzyme required for processing of vacuolar precursors. Mol Cell Biol. 1986 Jul;6(7):2490–2499. doi: 10.1128/mcb.6.7.2490. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bankaitis V. A., Johnson L. M., Emr S. D. Isolation of yeast mutants defective in protein targeting to the vacuole. Proc Natl Acad Sci U S A. 1986 Dec;83(23):9075–9079. doi: 10.1073/pnas.83.23.9075. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Banta L. M., Robinson J. S., Klionsky D. J., Emr S. D. Organelle assembly in yeast: characterization of yeast mutants defective in vacuolar biogenesis and protein sorting. J Cell Biol. 1988 Oct;107(4):1369–1383. doi: 10.1083/jcb.107.4.1369. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Botstein D., Falco S. C., Stewart S. E., Brennan M., Scherer S., Stinchcomb D. T., Struhl K., Davis R. W. Sterile host yeasts (SHY): a eukaryotic system of biological containment for recombinant DNA experiments. Gene. 1979 Dec;8(1):17–24. doi: 10.1016/0378-1119(79)90004-0. [DOI] [PubMed] [Google Scholar]
- Chvatchko Y., Howald I., Riezman H. Two yeast mutants defective in endocytosis are defective in pheromone response. Cell. 1986 Aug 1;46(3):355–364. doi: 10.1016/0092-8674(86)90656-2. [DOI] [PubMed] [Google Scholar]
- Dulić V., Riezman H. Characterization of the END1 gene required for vacuole biogenesis and gluconeogenic growth of budding yeast. EMBO J. 1989 May;8(5):1349–1359. doi: 10.1002/j.1460-2075.1989.tb03515.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Emr S. D., Vassarotti A., Garrett J., Geller B. L., Takeda M., Douglas M. G. The amino terminus of the yeast F1-ATPase beta-subunit precursor functions as a mitochondrial import signal. J Cell Biol. 1986 Feb;102(2):523–533. doi: 10.1083/jcb.102.2.523. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gallwitz D. Construction of a yeast actin gene intron deletion mutant that is defective in splicing and leads to the accumulation of precursor RNA in transformed yeast cells. Proc Natl Acad Sci U S A. 1982 Jun;79(11):3493–3497. doi: 10.1073/pnas.79.11.3493. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hanahan D. Studies on transformation of Escherichia coli with plasmids. J Mol Biol. 1983 Jun 5;166(4):557–580. doi: 10.1016/s0022-2836(83)80284-8. [DOI] [PubMed] [Google Scholar]
- Henikoff S. Unidirectional digestion with exonuclease III in DNA sequence analysis. Methods Enzymol. 1987;155:156–165. doi: 10.1016/0076-6879(87)55014-5. [DOI] [PubMed] [Google Scholar]
- Ito H., Fukuda Y., Murata K., Kimura A. Transformation of intact yeast cells treated with alkali cations. J Bacteriol. 1983 Jan;153(1):163–168. doi: 10.1128/jb.153.1.163-168.1983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jones E. W. Proteinase mutants of Saccharomyces cerevisiae. Genetics. 1977 Jan;85(1):23–33. doi: 10.1093/genetics/85.1.23. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kakinuma Y., Ohsumi Y., Anraku Y. Properties of H+-translocating adenosine triphosphatase in vacuolar membranes of SAccharomyces cerevisiae. J Biol Chem. 1981 Nov 10;256(21):10859–10863. [PubMed] [Google Scholar]
- Kitamoto K., Yoshizawa K., Ohsumi Y., Anraku Y. Dynamic aspects of vacuolar and cytosolic amino acid pools of Saccharomyces cerevisiae. J Bacteriol. 1988 Jun;170(6):2683–2686. doi: 10.1128/jb.170.6.2683-2686.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kitamoto K., Yoshizawa K., Ohsumi Y., Anraku Y. Mutants of Saccharomyces cerevisiae with defective vacuolar function. J Bacteriol. 1988 Jun;170(6):2687–2691. doi: 10.1128/jb.170.6.2687-2691.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kyte J., Doolittle R. F. A simple method for displaying the hydropathic character of a protein. J Mol Biol. 1982 May 5;157(1):105–132. doi: 10.1016/0022-2836(82)90515-0. [DOI] [PubMed] [Google Scholar]
- Laemmli U. K. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature. 1970 Aug 15;227(5259):680–685. doi: 10.1038/227680a0. [DOI] [PubMed] [Google Scholar]
- Miyamoto S., Ohya Y., Ohsumi Y., Anraku Y. Nucleotide sequence of the CLS4 (CDC24) gene of Saccharomyces cerevisiae. Gene. 1987;54(1):125–132. doi: 10.1016/0378-1119(87)90354-4. [DOI] [PubMed] [Google Scholar]
- Ohsumi Y., Anraku Y. Active transport of basic amino acids driven by a proton motive force in vacuolar membrane vesicles of Saccharomyces cerevisiae. J Biol Chem. 1981 Mar 10;256(5):2079–2082. [PubMed] [Google Scholar]
- Ohsumi Y., Anraku Y. Calcium transport driven by a proton motive force in vacuolar membrane vesicles of Saccharomyces cerevisiae. J Biol Chem. 1983 May 10;258(9):5614–5617. [PubMed] [Google Scholar]
- Orr-Weaver T. L., Szostak J. W., Rothstein R. J. Yeast transformation: a model system for the study of recombination. Proc Natl Acad Sci U S A. 1981 Oct;78(10):6354–6358. doi: 10.1073/pnas.78.10.6354. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Patterson T. E., Poyton R. O. COX8, the structural gene for yeast cytochrome c oxidase subunit VIII. DNA sequence and gene disruption indicate that subunit VIII is required for maximal levels of cellular respiration and is derived from a precursor which is extended at both its NH2 and COOH termini. J Biol Chem. 1986 Dec 25;261(36):17192–17197. [PubMed] [Google Scholar]
- Preston R. A., Murphy R. F., Jones E. W. Assay of vacuolar pH in yeast and identification of acidification-defective mutants. Proc Natl Acad Sci U S A. 1989 Sep;86(18):7027–7031. doi: 10.1073/pnas.86.18.7027. [DOI] [PMC free article] [PubMed] [Google Scholar]
- ROMAN H. Studies of gene mutation in Saccharomyces. Cold Spring Harb Symp Quant Biol. 1956;21:175–185. doi: 10.1101/sqb.1956.021.01.015. [DOI] [PubMed] [Google Scholar]
- Riezman H. Endocytosis in yeast: several of the yeast secretory mutants are defective in endocytosis. Cell. 1985 Apr;40(4):1001–1009. doi: 10.1016/0092-8674(85)90360-5. [DOI] [PubMed] [Google Scholar]
- Robinson J. S., Klionsky D. J., Banta L. M., Emr S. D. Protein sorting in Saccharomyces cerevisiae: isolation of mutants defective in the delivery and processing of multiple vacuolar hydrolases. Mol Cell Biol. 1988 Nov;8(11):4936–4948. doi: 10.1128/mcb.8.11.4936. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rothblatt J. A., Deshaies R. J., Sanders S. L., Daum G., Schekman R. Multiple genes are required for proper insertion of secretory proteins into the endoplasmic reticulum in yeast. J Cell Biol. 1989 Dec;109(6 Pt 1):2641–2652. doi: 10.1083/jcb.109.6.2641. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rothman J. H., Howald I., Stevens T. H. Characterization of genes required for protein sorting and vacuolar function in the yeast Saccharomyces cerevisiae. EMBO J. 1989 Jul;8(7):2057–2065. doi: 10.1002/j.1460-2075.1989.tb03614.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rothman J. H., Stevens T. H. Protein sorting in yeast: mutants defective in vacuole biogenesis mislocalize vacuolar proteins into the late secretory pathway. Cell. 1986 Dec 26;47(6):1041–1051. doi: 10.1016/0092-8674(86)90819-6. [DOI] [PubMed] [Google Scholar]
- Sanger F., Nicklen S., Coulson A. R. DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci U S A. 1977 Dec;74(12):5463–5467. doi: 10.1073/pnas.74.12.5463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sato T., Ohsumi Y., Anraku Y. Substrate specificities of active transport systems for amino acids in vacuolar-membrane vesicles of Saccharomyces cerevisiae. Evidence of seven independent proton/amino acid antiport systems. J Biol Chem. 1984 Sep 25;259(18):11505–11508. [PubMed] [Google Scholar]
- Stevens T., Esmon B., Schekman R. Early stages in the yeast secretory pathway are required for transport of carboxypeptidase Y to the vacuole. Cell. 1982 Sep;30(2):439–448. doi: 10.1016/0092-8674(82)90241-0. [DOI] [PubMed] [Google Scholar]
- Tanaka K., Kanbe T. Mitosis in the fission yeast Schizosaccharomyces pombe as revealed by freeze-substitution electron microscopy. J Cell Sci. 1986 Feb;80:253–268. doi: 10.1242/jcs.80.1.253. [DOI] [PubMed] [Google Scholar]
- Uchida E., Ohsumi Y., Anraku Y. Purification and properties of H+-translocating, Mg2+-adenosine triphosphatase from vacuolar membranes of Saccharomyces cerevisiae. J Biol Chem. 1985 Jan 25;260(2):1090–1095. [PubMed] [Google Scholar]
- Vieira J., Messing J. Production of single-stranded plasmid DNA. Methods Enzymol. 1987;153:3–11. doi: 10.1016/0076-6879(87)53044-0. [DOI] [PubMed] [Google Scholar]
- Wada Y., Ohsumi Y., Tanifuji M., Kasai M., Anraku Y. Vacuolar ion channel of the yeast, Saccharomyces cerevisiae. J Biol Chem. 1987 Dec 25;262(36):17260–17263. [PubMed] [Google Scholar]
- Weisman L. S., Bacallao R., Wickner W. Multiple methods of visualizing the yeast vacuole permit evaluation of its morphology and inheritance during the cell cycle. J Cell Biol. 1987 Oct;105(4):1539–1547. doi: 10.1083/jcb.105.4.1539. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wickerham L. J. A Critical Evaluation of the Nitrogen Assimilation Tests Commonly Used in the Classification of Yeasts. J Bacteriol. 1946 Sep;52(3):293–301. [PMC free article] [PubMed] [Google Scholar]
- Woolford C. A., Daniels L. B., Park F. J., Jones E. W., Van Arsdell J. N., Innis M. A. The PEP4 gene encodes an aspartyl protease implicated in the posttranslational regulation of Saccharomyces cerevisiae vacuolar hydrolases. Mol Cell Biol. 1986 Jul;6(7):2500–2510. doi: 10.1128/mcb.6.7.2500. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zaret K. S., Sherman F. DNA sequence required for efficient transcription termination in yeast. Cell. 1982 Mar;28(3):563–573. doi: 10.1016/0092-8674(82)90211-2. [DOI] [PubMed] [Google Scholar]







