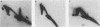Abstract
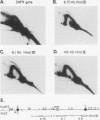
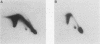
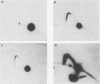
Two complementary two-dimensional gel electrophoretic techniques have recently been developed that allow initiation sites to be mapped with relative precision in eukaryotic genomes at least as complex as those of yeast and Drosophila melanogaster. We reported the first application of these mapping methods to a mammalian genome in a study on the amplified dihydrofolate reductase (DHFR) domain of the methotrexate-resistant CHO cell line CHOC 400 (J.P. Vaughn, P.A. Dijkwel, and J.L. Hamlin, Cell 61:1075-1087, 1990). Our results suggested that in this 240-kb domain, initiation of nascent DNA strands occurs at many sites within a 30- to 35-kb zone mapping immediately downstream from the DHFR gene. In the course of these studies, it was necessary to develop methods to stabilize replication intermediates against branch migration and shear. This report describes these stabilization methods in detail and presents a new enrichment protocol that extends the neutral/neutral two-dimensional gel mapping method to single-copy loci in mammalian cells. Preliminary analysis of replication intermediates purified from CHO cells by this method suggests that DNA synthesis may initiate at many sites within a broad zone in the single-copy DHFR locus as well.
Full text
PDF


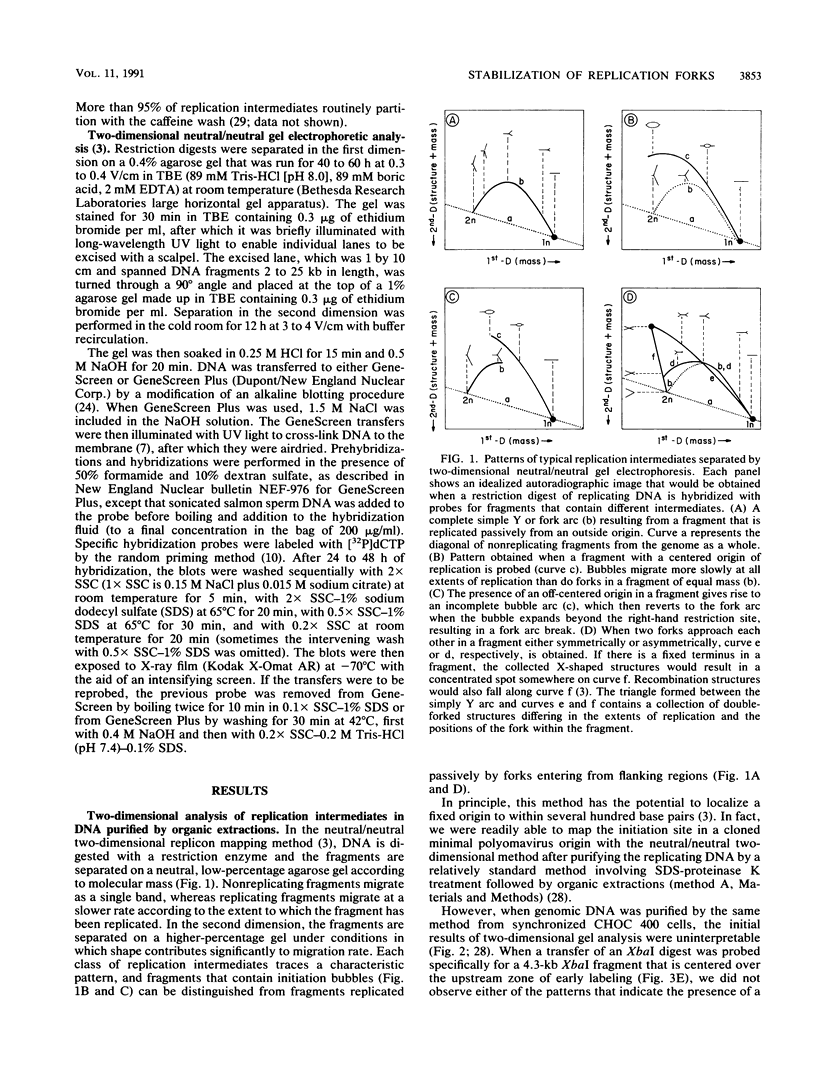



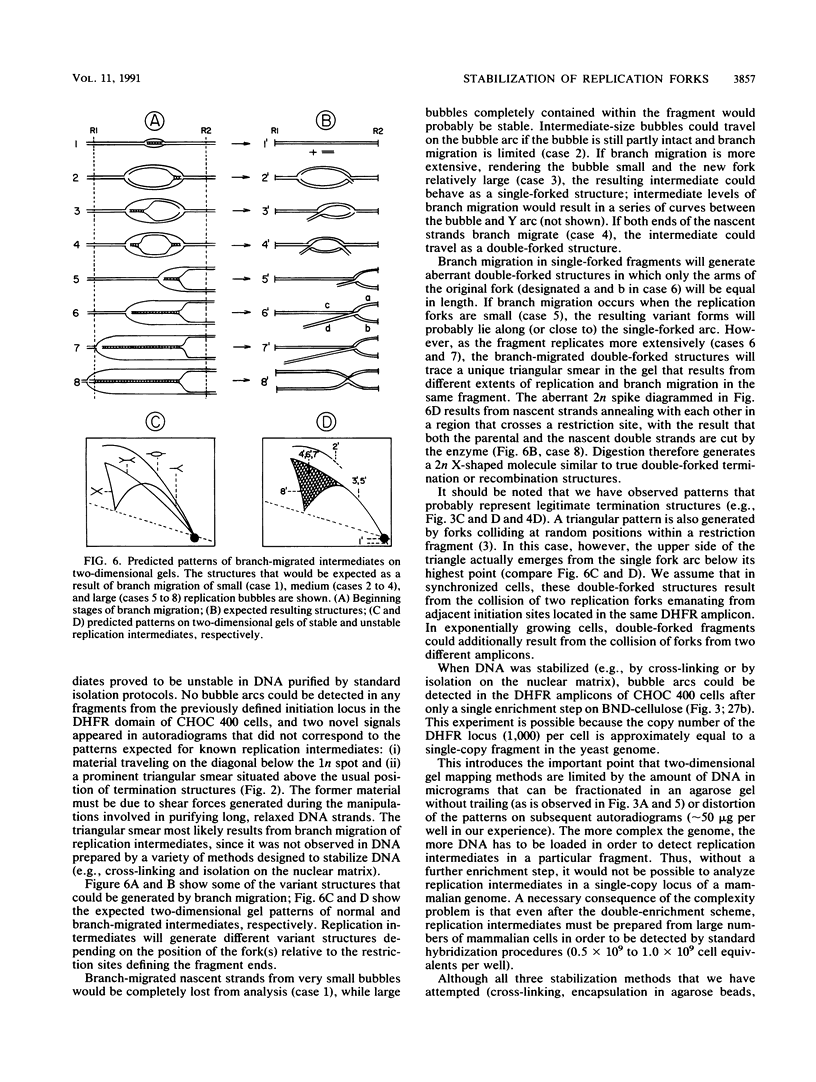


Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Anachkova B., Hamlin J. L. Replication in the amplified dihydrofolate reductase domain in CHO cells may initiate at two distinct sites, one of which is a repetitive sequence element. Mol Cell Biol. 1989 Feb;9(2):532–540. doi: 10.1128/mcb.9.2.532. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brewer B. J., Fangman W. L. A replication fork barrier at the 3' end of yeast ribosomal RNA genes. Cell. 1988 Nov 18;55(4):637–643. doi: 10.1016/0092-8674(88)90222-x. [DOI] [PubMed] [Google Scholar]
- Brewer B. J., Fangman W. L. The localization of replication origins on ARS plasmids in S. cerevisiae. Cell. 1987 Nov 6;51(3):463–471. doi: 10.1016/0092-8674(87)90642-8. [DOI] [PubMed] [Google Scholar]
- Burhans W. C., Selegue J. E., Heintz N. H. Isolation of the origin of replication associated with the amplified Chinese hamster dihydrofolate reductase domain. Proc Natl Acad Sci U S A. 1986 Oct;83(20):7790–7794. doi: 10.1073/pnas.83.20.7790. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Burhans W. C., Vassilev L. T., Caddle M. S., Heintz N. H., DePamphilis M. L. Identification of an origin of bidirectional DNA replication in mammalian chromosomes. Cell. 1990 Sep 7;62(5):955–965. doi: 10.1016/0092-8674(90)90270-o. [DOI] [PubMed] [Google Scholar]
- Church G. M., Gilbert W. Genomic sequencing. Proc Natl Acad Sci U S A. 1984 Apr;81(7):1991–1995. doi: 10.1073/pnas.81.7.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cook P. R. A general method for preparing intact nuclear DNA. EMBO J. 1984 Aug;3(8):1837–1842. doi: 10.1002/j.1460-2075.1984.tb02056.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dijkwel P. A., Mullenders L. H., Wanka F. Analysis of the attachment of replicating DNA to a nuclear matrix in mammalian interphase nuclei. Nucleic Acids Res. 1979 Jan;6(1):219–230. doi: 10.1093/nar/6.1.219. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Feinberg A. P., Vogelstein B. A technique for radiolabeling DNA restriction endonuclease fragments to high specific activity. Anal Biochem. 1983 Jul 1;132(1):6–13. doi: 10.1016/0003-2697(83)90418-9. [DOI] [PubMed] [Google Scholar]
- Handeli S., Klar A., Meuth M., Cedar H. Mapping replication units in animal cells. Cell. 1989 Jun 16;57(6):909–920. doi: 10.1016/0092-8674(89)90329-2. [DOI] [PubMed] [Google Scholar]
- Hatton K. S., Dhar V., Brown E. H., Iqbal M. A., Stuart S., Didamo V. T., Schildkraut C. L. Replication program of active and inactive multigene families in mammalian cells. Mol Cell Biol. 1988 May;8(5):2149–2158. doi: 10.1128/mcb.8.5.2149. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Heck M. M., Spradling A. C. Multiple replication origins are used during Drosophila chorion gene amplification. J Cell Biol. 1990 Apr;110(4):903–914. doi: 10.1083/jcb.110.4.903. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Heintz N. H., Hamlin J. L. An amplified chromosomal sequence that includes the gene for dihydrofolate reductase initiates replication within specific restriction fragments. Proc Natl Acad Sci U S A. 1982 Jul;79(13):4083–4087. doi: 10.1073/pnas.79.13.4083. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huberman J. A., Riggs A. D. On the mechanism of DNA replication in mammalian chromosomes. J Mol Biol. 1968 Mar 14;32(2):327–341. doi: 10.1016/0022-2836(68)90013-2. [DOI] [PubMed] [Google Scholar]
- Huberman J. A., Zhu J. G., Davis L. R., Newlon C. S. Close association of a DNA replication origin and an ARS element on chromosome III of the yeast, Saccharomyces cerevisiae. Nucleic Acids Res. 1988 Jul 25;16(14A):6373–6384. doi: 10.1093/nar/16.14.6373. [DOI] [PMC free article] [PubMed] [Google Scholar]
- James C. D., Leffak M. Polarity of DNA replication through the avian alpha-globin locus. Mol Cell Biol. 1986 Apr;6(4):976–984. doi: 10.1128/mcb.6.4.976. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Leu T. H., Hamlin J. L. High-resolution mapping of replication fork movement through the amplified dihydrofolate reductase domain in CHO cells by in-gel renaturation analysis. Mol Cell Biol. 1989 Feb;9(2):523–531. doi: 10.1128/mcb.9.2.523. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Levine A. J., Kang H. S., Billheimer F. E. DNA replication in SV40 infected cells. I. Analysis of replicating SV40 DNA. J Mol Biol. 1970 Jun 14;50(2):549–568. doi: 10.1016/0022-2836(70)90211-1. [DOI] [PubMed] [Google Scholar]
- Linskens M. H., Huberman J. A. Organization of replication of ribosomal DNA in Saccharomyces cerevisiae. Mol Cell Biol. 1988 Nov;8(11):4927–4935. doi: 10.1128/mcb.8.11.4927. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Milbrandt J. D., Heintz N. H., White W. C., Rothman S. M., Hamlin J. L. Methotrexate-resistant Chinese hamster ovary cells have amplified a 135-kilobase-pair region that includes the dihydrofolate reductase gene. Proc Natl Acad Sci U S A. 1981 Oct;78(10):6043–6047. doi: 10.1073/pnas.78.10.6043. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mirkovitch J., Mirault M. E., Laemmli U. K. Organization of the higher-order chromatin loop: specific DNA attachment sites on nuclear scaffold. Cell. 1984 Nov;39(1):223–232. doi: 10.1016/0092-8674(84)90208-3. [DOI] [PubMed] [Google Scholar]
- Nawotka K. A., Huberman J. A. Two-dimensional gel electrophoretic method for mapping DNA replicons. Mol Cell Biol. 1988 Apr;8(4):1408–1413. doi: 10.1128/mcb.8.4.1408. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Reed K. C., Mann D. A. Rapid transfer of DNA from agarose gels to nylon membranes. Nucleic Acids Res. 1985 Oct 25;13(20):7207–7221. doi: 10.1093/nar/13.20.7207. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Russev G., Vassilev L. Isolation of a DNA fraction from Ehrlich ascites tumour cells containing the putative origin of replication. J Mol Biol. 1982 Oct 15;161(1):77–87. doi: 10.1016/0022-2836(82)90279-0. [DOI] [PubMed] [Google Scholar]
- Struhl K., Stinchcomb D. T., Scherer S., Davis R. W. High-frequency transformation of yeast: autonomous replication of hybrid DNA molecules. Proc Natl Acad Sci U S A. 1979 Mar;76(3):1035–1039. doi: 10.1073/pnas.76.3.1035. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Umek R. M., Linskens M. H., Kowalski D., Huberman J. A. New beginnings in studies of eukaryotic DNA replication origins. Biochim Biophys Acta. 1989 Jan 23;1007(1):1–14. doi: 10.1016/0167-4781(89)90123-1. [DOI] [PubMed] [Google Scholar]
- Vaughn J. P., Dijkwel P. A., Hamlin J. L. Replication initiates in a broad zone in the amplified CHO dihydrofolate reductase domain. Cell. 1990 Jun 15;61(6):1075–1087. doi: 10.1016/0092-8674(90)90071-l. [DOI] [PubMed] [Google Scholar]
- Vaughn J. P., Dijkwel P. A., Mullenders L. H., Hamlin J. L. Replication forks are associated with the nuclear matrix. Nucleic Acids Res. 1990 Apr 25;18(8):1965–1969. doi: 10.1093/nar/18.8.1965. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vogelstein B., Pardoll D. M., Coffey D. S. Supercoiled loops and eucaryotic DNA replicaton. Cell. 1980 Nov;22(1 Pt 1):79–85. doi: 10.1016/0092-8674(80)90156-7. [DOI] [PubMed] [Google Scholar]
- Zannis-Hadjopoulos M., Persico M., Martin R. G. The remarkable instability of replication loops provides a general method for the isolation of origins of DNA replication. Cell. 1981 Nov;27(1 Pt 2):155–163. doi: 10.1016/0092-8674(81)90369-x. [DOI] [PubMed] [Google Scholar]