Abstract
Previous proteomics studies have partially unraveled the complexity of endothelial protein secretion but have not investigated glycosylation, a key modification of secreted and membrane proteins for cell communication. In this study, human umbilical vein endothelial cells were kept in serum-free medium before activation by phorbol-12-myristate-13 acetate, a commonly used secretagogue that induces exocytosis of endothelial vesicles. In addition to 123 secreted proteins, the secretome was particularly rich in membrane proteins. Glycopeptides were enriched by zwitterionic hydrophilic interaction liquid chromatography resins and were either treated with PNGase F and H218O or directly analyzed using a recently developed workflow combining higher-energy C-trap dissociation (HCD) with electron-transfer dissociation (ETD) for a hybrid linear ion trap–orbitrap mass spectrometer. After deglycosylation with PNGase F in the presence of H218O, 123 unique peptides displayed 18O-deamidation of asparagine, corresponding to 86 proteins with a total of 121 glycosylation sites. Direct glycopeptide analysis via HCD-ETD identified 131 glycopeptides from 59 proteins and 118 glycosylation sites, of which 41 were known, 51 were predicted, and 26 were novel. Two methods were compared: alternating HCD-ETD and HCD-product-dependent ETD. The former detected predominantly high-intensity, multiply charged glycopeptides, whereas the latter preferentially selected precursors with complex/hybrid glycans for fragmentation. Validation was performed by means of glycoprotein enrichment and analysis of the input, the flow-through, and the bound fraction. This study represents the most comprehensive characterization of endothelial protein secretion to date and demonstrates the potential of new HCD-ETD workflows for determining the glycosylation status of complex biological samples.
Cardiovascular disease manifests predominantly as myocardial ischemia, heart failure, stroke, aortic aneurysm, and peripheral vascular disease and leads to the majority of deaths and disabilities worldwide. Endothelial cells (ECs) constitute the inner lining of all blood vessels and form the interface between the circulation and the vascular wall (1). The endothelial monolayer is pivotal for maintaining vascular homeostasis through a balance of endothelium-derived factors (2, 3). ECs are preferred targets of cardiovascular risk factors such as hypercholesterolemia, diabetes, hypertension, and smoking (1, 4). Repetitive injury is associated with a varying degree of endothelial dysfunction. Alterations in its anticoagulant and anti-inflammatory properties leave the vasculature susceptible to disease (5) and play a key role in the initiation and progression of cardiovascular disease (6).
Previous proteomics studies (7–13), including one by our group (8), have investigated the secretome of unstimulated human umbilical vein ECs (HUVECs), the most widely used ECs in cardiovascular research. Only two studies have explored the secretome of HUVECs upon activation by shear stress (10) or with statin treatment (13) thus far. One study used human microvascular ECs (9), which represent a distinct population of ECs from small vessels. Yet many factors secreted by ECs were not identified, probably because of their low abundance. In this study, we used a secretagogue, phorbol ester phorbol-12-myristate-13-acetate (PMA) (14, 15), to induce maximal protein release from serum-starved HUVECs over 45 min. In addition, we applied three different proteomic strategies for the analysis of glycoproteins/glycopeptides to further enrich secreted proteins and characterize their glycosylation sites.
EXPERIMENTAL PROCEDURES
EC Culture
HUVECs (Lonza Group Ltd., Basel, Switzerland) were cultured on 0.1% gelatin-coated flasks in M199 medium supplemented with 1 ng/ml endothelial cell growth factor (Sigma), 3 μg/ml endothelial growth supplement from bovine neural tissue (Sigma), 10 U/ml heparin, 1.25 μg/ml thymidine, 10% fetal bovine serum (A15–108, PAA Laboratories, Velizy-Villacoublay, France), and 100 μg/ml penicillin and streptomycin in a humidified incubator supplemented with 5% CO2 at 37 °C. The cells were subcultured every 2 to 3 days at a ratio of 1:4 (16).
Conditioned Medium Collection
HUVECs were cultured in complete medium until confluent. Then, they were washed and incubated in M199 medium for 30 min twice before stimulation with 50 nm PMA (Sigma) in M199 medium for 45 min. The control group was incubated with M199 medium in the absence of PMA for 45 min. Conditioned media were collected and stored at −80 °C for further analysis.
Immunofluorescence Staining
HUVECs were cultured in Nunc chamber slides (Sigma-Aldrich) for 3 days. HUVECs were stimulated with 50 nm PMA in M199 medium for 45 min or incubated with M199 medium for 45 min. The cells were fixed with 4% formaldehyde in PBS for 10 min, permeabilized with 0.1% Triton X-100 in PBS for 5 min, and blocked in 5% fetal bovine serum in PBS for 30 min at 37 °C. Following 1 h of incubation with the primary antibodies, VE-cadherin (ab33168, Abcam, Cambridge, UK), and von Willebrand factor (vWF) (sc-8068, Santa Cruz Biotechnology, Santa Cruz, CA) at 37 °C, an Alexa Fluor® 594 conjugated donkey anti-rabbit IgG and an Alexa Fluor® 488 conjugated donkey anti-goat IgG, respectively, were added, and the cells were incubated at 37 °C for 30 min. Nuclei were counterstained with 4′,6-diamidino-2-phenylindole (D9542, Sigma) for 5 min. The slide was mounted in fluorescence mounting medium (DAKO, Denmark A/S, Glostrup, Denmark) and examined with an AxioPlan 2 fluorescence microscope (Carl Zeiss, Thornwood, NY) (17).
Proteomics Profiling of the Secretome
Conditioned media were concentrated with an Amicon spin column (3kD MWCO, EDO Millipore Corp., Billerica, MA) and separated via 4%–12% Bis-Tris SDS-PAGE (Invitrogen). Proteins were visualized via silver staining (PlusOne silver staining kit for proteins, GE Healthcare). Gel bands were digested with modified trypsin (Promega Corp., Madison, WI) overnight on a ProGest digestion robot (Digilab Inc., Marlborough, MA) and analyzed via reverse-phase nano-flow HPLC (PepMap C18, 3 μm, 100 Å, 25 cm × 75 μm inner diameter column, Thermo Scientific) interfaced to an LTQ Orbitrap XL MS (Thermo Scientific) (18).
Deglycosylation
Concentrated media were mixed with deglycosylation buffer (150 mm NaCl, 50 mm sodium acetate, 10 mm EDTA, proteinase inhibitors, pH 6.8) supplemented with 0.05U PNGase F (Sigma), chondroitinase ABC (C3667, Sigma), and keratanase (G6920, Sigma) and incubated at 37 °C overnight (19).
Immunoblotting
Concentrated or deglycosylated media were separated via 4%–12% Bis-Tris gel (Invitrogen). Proteins were transferred on a nitrocellulose membrane and blocked with 5% bovine serum albumin in PBS. Membranes were incubated with primary antibody overnight at 4 °C. Secondary antibodies were incubated for 1 h at room temperature. After the addition of ECL (GE Healthcare), the film was developed using a Compact X4 Automatic Processor (Xograph Healthcare Ltd., Stonehouse, UK). The following primary antibodies were used: agrin (sc-25528, Santa Cruz Biotechnology), biglycan (ab54855, Abcam), connective tissue growth factor (sc-25440, Santa Cruz Biotechnology), fibronectin (sc-56391, Santa Cruz Biotechnology), and lymphatic vessel endothelial hyaluronic acid receptor 1 (AF2089, R&D Systems).
Difference Gel Electrophoresis
Conditioned media from HUVECs treated with or without PMA were concentrated using an Amicon spin column (3kD MWCO, Millipore) and the ReadyPrep 2D clean-up kit (Bio-Rad). The pellet was resuspended in difference gel electrophoresis lysis buffer (30 mm Tris, 8 m urea, 4% w/v CHAPS, protease inhibitors, pH 8.5). For each secretome sample, 15 μg of proteins were labeled with Cy3 or Cy5. A dye swap was performed to exclude preferential labeling. Cellular extracts of HUVECs were labeled with Cy2. Cy2-, Cy3-, and Cy5-labeled samples were separated via isoelectric focusing on immobilized pH gradient dry strips (18 cm, pH 3–10 NL, GE Healthcare) with 30 KVH. The strips were equilibrated with 10 mg/ml DTT in equilibration buffer (6 m urea, 2% w/v SDS, 30% v/v glycerol, 50 mm Tris, pH 8.8) for 15 min followed by 48 mg/ml iodoacetamide in equilibration buffer for 15 min before separation via SDS-PAGE at 100 W for 4 h using an Ettan DALTsix vertical electrophoresis system (GE Healthcare) (20–22). Gels were scanned on an Ettan difference gel electrophoresis imager (GE Healthcare). Images were overlaid with ImageQuant TL software (GE Healthcare). Common spots present in both the cellular proteome and the secretome were excised, digested with trypsin, and identified using nano-flow HPLC-MS/MS. Detailed protocols are available on our research group's website.
Glycopeptide Enrichment
Conditioned media were desalted via the use of Zeba spin columns (Thermo Scientific). Proteins were then reduced by 5 mm DTT and alkylated with 25 mm iodoacetamide. After acetone precipitation overnight, the pellet was resuspended in 100 mm triethylammonium bicarbonate (pH 8.5, Sigma) and digested with modified trypsin (Promega) at 37 °C overnight. Peptides were labeled at a ratio of 100 μg peptides/0.8 mg Tandem Mass Tag Zero (TMT0) (Thermo Scientific) according to the manufacturer's instruction. Labeled peptides were further enriched for glycopeptides using zwitterionic hydrophilic interaction liquid chromatography resin (Merck) (23).
LC/MS of Intact Glycopeptides
The glycopeptide enriched fraction was separated using the EASY-nLCTM nano-HPLC system (Thermo Scientific) with a Magic C18 spray tip 15 cm × 75 μm inner diameter column (Bruker-Michrom, Auburn, CA). Gradient elution was performed with 4% to 30% acetonitrile in 0.1% formic acid over 60 min at a flow rate of 300 nl/min. The samples were analyzed with an Orbitrap Elite hybrid MS with electron-transfer dissociation (ETD) (Thermo Scientific). The following MS and MS/MS settings were used: Fourier transform: MSn automatic gain control target = 5E4; MS/MS = 1 μscans, max ion time = 200 ms; MS = 300–1800 m/z, resolution = 60,000 at m/z 400, MS target = 1E6; dynamic exclusion = repeat count 1, duration 30 s, exclusion duration 90 s; higher-energy C-trap dissociation (HCD): collision energy = 35%, resolution = 15,000; MSn target ion trap = 1E4, 2 μscans, max ion time = 150 ms; ETD anion automatic gain control target = 2E5, charge-dependent ETD reaction time enabled. For alternating HCD-ETD MS/MS, the top 10 ions were analyzed. For HCD-product-dependent ETD, the top 10 ions were analyzed via HCD, and product-dependent ETD acquisition was triggered by product (oxonium) ions (m/z 163.0812 for Hex; m/z 204.0864 for HexNAc; m/z 138.0554 for HexNAc fragment ion) (24).
Deglycosylation with PNGase F and H218O
Zwitterionic hydrophilic interaction liquid chromatography resin enriched glycopeptides were resuspended in 50 mm ammonium bicarbonate in H218O (97 atom % 18O, Sigma) and deglycosylated with PNGase F (Sigma) for 4 h at 37 °C. The samples were separated via reverse-phase nano-flow HPLC (PepMap C18, 3 μm, 100 Å, 25 cm × 75 μm inner diameter column, Thermo Scientific) before analysis on an LTQ Orbitrap XL MS (Thermo Scientific).
Glycoprotein Enrichment and LC/MS
ConA1 lectin resins (Thermo Scientific) were used to enrich glycoproteins from concentrated conditioned media according to the manufacturer's protocol. The input, glycoprotein-enriched fraction, and flow-through samples were subjected to trypsin digestion. The in-solution digests were separated on a Thermo Scientific Dionex UltiMate 3000 Rapid Separation LC (RSLC) system using a PepMap C18 column (3 μm, 100 Å, 50 cm × 75 μm inner diameter column, Thermo Scientific). The rapid separation LC system was interfaced to a Q Exactive MS (Thermo Scientific), and samples were analyzed using a top-10 HCD method.
Database Search and Data analysis
The following parameters were used for different experiments.
(i) Gel-LC-MS/MS: Peak lists were generated by Mascot daemon (version 2.3.0, Matrix Science Ltd., London, UK) using extract_msn_com.exe and searched against the UniProt/Swiss-Prot mammalian database (version 2012.03, 65,780 entries) using Mascot (version 2.3.01, Matrix Science) with peptide tolerance = 10 ppm, MS/MS tolerance = 0.8 Da, carbamidomethylation of cysteine as a fixed modification, oxidation of methionine as a variable modification, and a maximum of two missed cleavage sites. The search results were loaded into Scaffold software (version 3.6.2, Proteome Software Software, Inc., Portland, OR). A protein probability greater than 99%, a peptide probability greater than 95%, and a minimum number of two peptides per protein were applied as filters to generate the protein list. Bovine contaminant proteins are listed separately.
(ii) PNGase F + H218O experiment: Thermo Scientific Proteome Discoverer software version 1.3 was used to search against the UniProt/Swiss-Prot mammalian database (version 2012.03) using Mascot (version 2.3.01, Matrix Science) with a peptide tolerance of 10 ppm; an MS/MS tolerance of 0.8 Da; carbamidomethylation of cysteine as a fixed modification; oxidation of methionine, TMT0 label on lysine and peptide N-terminus, and deamidation (spontaneous deamidation in ordinary water) and O18-deamidation (deglycosylation by PNGase F in H218O) of asparagine as variable modifications; and a maximum of two missed cleavage sites. Proteome Discoverer produced a custom database containing 136 target proteins based on this search.
(iii) Orbitrap Elite MS: Raw files were searched against the 136-protein database (along with reversed proteins as decoys) using ByonicTM (25) with a peptide tolerance of 10 ppm; an MS/MS tolerance of 20 ppm for HCD and 0.6 Da for ETD; the carbamidomethylated cysteine, TMT0 label on lysine and peptide N-terminus as fixed modifications; and oxidation of methionine, deamidation of asparagine and glutamine, and phosphorylation of serine and threonine as variable modifications. ByonicTM allowed one N-glycan modification on the N-X(not P)-S/T consensus motif per peptide, with mass and composition chosen from its “common human” glycan database containing 350 glycan masses up to 6000 Da. Glycan modifications were verified by the presence of corresponding glycan fragment ions, such as the HexNAc oxonium ion at 204.087 Da in HCD spectra. Peptide sequences were identified by ByonicTM from the ETD spectra and verified manually.
(iv) Q Exactive MS: Raw files were searched against the UniProt/Swiss-Prot human database (version 57.13, 20,266 entries) using Proteome Discoverer (version 1.3, Thermo Scientific) with Mascot (version 2.3.0, Matrix Science) and a peptide tolerance of 10 ppm, an MS/MS tolerance of 10 mmu, carbamidomethylation of cysteine as a fixed modification, oxidation of methionine as a variable modification, and a maximum of two missed cleavage sites.
RESULTS
The Secretome of Activated ECs
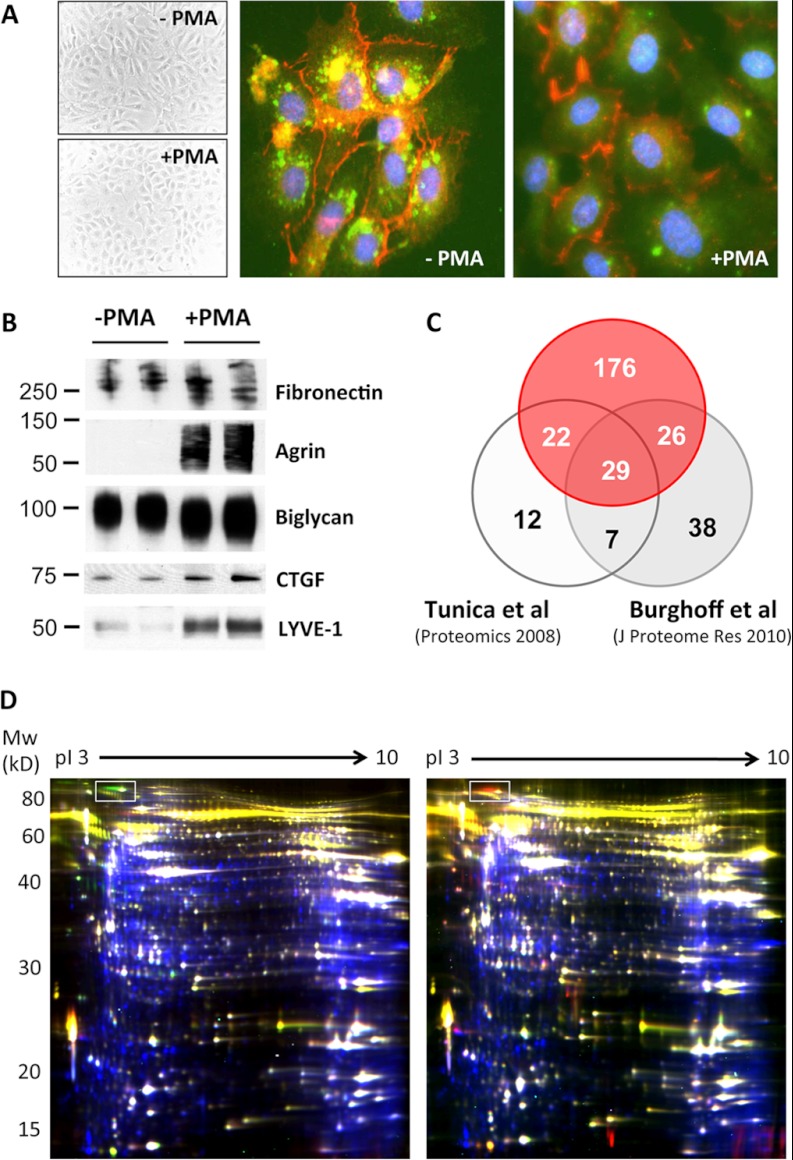
HUVECs were stimulated with PMA, a commonly used secretagogue that induces exocytosis of endothelial vesicles. As previously reported (26), the morphology of ECs changes from spindle-shaped to round upon PMA activation, and the rod-shaped Weibel-Palade bodies, unique storage vesicles within ECs containing vWF and many other secreted proteins, fuse with the cell membrane (Fig. 1A). In total, the secretomes of 17 primary ECs were analyzed via gel-LC-MS/MS, with or without deglycosylation. Apart from 123 secreted proteins, the conditioned medium of PMA-stimulated ECs was particularly rich in surface antigens and receptors, including many established endothelial markers (Table I). All identified proteins and peptides are listed in supplemental Tables S1 and S2, respectively. The distribution of the frequencies and the cumulated distribution of the number of samples in which proteins were identified are shown in supplemental Fig. S1. MS datasets of three biological replicates have been deposited in PRIDE (accession numbers 26908–27003).
Fig. 1.
PMA treatment to stimulate EC secretion. Treatment of HUVECs with PMA, a commonly used secretagogue, resulted in a characteristic morphological change indicative of activation. A, immunofluorescence staining of vWF (green) and VE-cadherin (red) shows the exocytotic effect of PMA. B, PMA increased protein secretion in the conditioned media as confirmed via immunoblotting. C, relative to previous studies, more than twice as many secreted and plasma membrane proteins were identified. D, overlay of intracellular and secreted proteins by means of difference gel electrophoresis. In the left-hand panel, proteins in conditioned media of HUVECs are stained in green (+PMA) and red (−PMA), and cellular proteins are stained in blue. Results were reproduced with different biological replicates using reverse-labeling (right-hand panel: red, +PMA; green, −PMA). The protein corresponding to von Willebrand antigen 2 is highlighted with a box. Common proteins in the secretome and the cellular proteome are numbered in supplemental Fig. S2 and listed in supplemental Table S3.
Table I. Extracellular and plasma membrane proteins identified in the HUVEC-conditioned media after PMA stimulation.
| Protein name | UniProt ID | UniProt accession number | Gene name | Cellular component | Glycoprotein | EC marker | |
|---|---|---|---|---|---|---|---|
| Calcium ion-binding proteins | |||||||
| Annexin A1 | ANXA1_HUMAN | P04083 | ANXA1 | Plasma membrane | |||
| Annexin A2a | ANXA2_HUMAN | P07355 | ANXA2 | Extracellular | Plasma membrane | ||
| Annexin A3 | ANXA3_HUMAN | P12429 | ANXA3 | Plasma membrane | |||
| Calreticulin | CALR_HUMAN | P27797 | CALR | Extracellular | Glycoprotein | ||
| Calumenin | CALU_HUMAN | O43852 | CALU | Extracellular | Glycoprotein | ||
| Calpain-1 catalytic subunit | CAN1_HUMAN | P07384 | CAPN1 | Plasma membrane | |||
| Calpain-2 catalytic subunit | CAN2_HUMAN | P17655 | CAPN2 | Plasma membrane | |||
| Calpain small subunit 1 | CPNS1_HUMAN | P04632 | CAPNS1 | Plasma membrane | |||
| Calsyntenin-1 | CSTN1_HUMAN | O94985 | CLSTN1 | Plasma membrane | Glycoprotein | ||
| Calsyntenin-3 | CSTN3_HUMAN | Q9BQT9 | CLSTN3 | Plasma membrane | Glycoprotein | ||
| Desmoglein-1 | DSG1_HUMAN | Q02413 | DSG1 | Plasma membrane | Glycoprotein | ||
| Nucleobindin-2 | NUCB2_HUMAN | P80303 | NUCB2 | Extracellular | |||
| Carbohydrate and glycan metabolism | |||||||
| Alpha-amylase 1 | AMY1_HUMAN | P04745 | AMY1C | Extracellular | Glycoprotein | ||
| Exostosin-like 2 | EXTL2_HUMAN | Q9UBQ6 | EXTL2 | Extracellular | Glycoprotein | ||
| Polypeptide N-acetylgalactosaminyltransferase 1 | GALT1_HUMAN | Q10472 | GALNT1 | Extracellular | Glycoprotein | ||
| Sialate O-acetylesterase | SIAE_HUMAN | Q9HAT2 | SIAE | Extracellular | Glycoprotein | ||
| UDP-N-acetylhexosamine pyrophosphorylase | UAP1_HUMAN | Q16222 | UAP1 | Plasma membrane | |||
| Coagulation and related proteins | |||||||
| Amyloid-like protein 2 | APLP2_HUMAN | Q06481 | APLP2 | Plasma membrane | Glycoprotein | ||
| Multimerin-1 | MMRN1_HUMAN | Q13201 | MMRN1 | Extracellular | Glycoprotein | ||
| Plasminogen activator inhibitor 1 | PAI1_HUMAN | P05121 | SERPINE1 | Extracellular | Glycoprotein | ||
| Plasminogen activator inhibitor 2 | PAI2_HUMAN | P05120 | SERPINB2 | Extracellular | Glycoprotein | ||
| Tissue factor pathway inhibitor | TFPI1_HUMAN | P10646 | TFPI | Extracellular | Glycoprotein | ||
| Tissue factor pathway inhibitor 2 | TFPI2_HUMAN | P48307 | TFPI2 | Extracellular | Glycoprotein | ||
| Tissue-type plasminogen activator | TPA_HUMAN | P00750 | PLAT | Extracellular | Glycoprotein | ||
| von Willebrand factor | VWF_HUMAN | P04275 | VWF | Extracellular | Glycoprotein | EC marker | |
| Extracellular matrix components and associated proteins | |||||||
| Agrin | AGRIN_HUMAN | O00468 | AGRN | Extracellular | Glycoprotein | ||
| Collagen alpha-2(IV) chain | CO4A2_HUMAN | P08572 | COL4A2 | Extracellular | Glycoprotein | ||
| Collagen alpha-1(VI) chain | CO6A1_HUMAN | P12109 | COL6A1 | Extracellular | Glycoprotein | ||
| Collagen alpha-1(XII) chain | COCA1_HUMAN | Q99715 | COL12A1 | Extracellular | Glycoprotein | ||
| Collagen alpha-1(XVIII) chain | COIA1_HUMAN | P39060 | COL18A1 | Extracellular | Glycoprotein | ||
| EGF-containing fibulin-like extracellular matrix protein 1 | FBLN3_HUMAN | Q12805 | EFEMP1 | Extracellular | Glycoprotein | ||
| Fibrillin-1 | FBN1_HUMAN | P35555 | FBN1 | Extracellular | Glycoprotein | ||
| Fibrillin-2 | FBN2_HUMAN | P35556 | FBN2 | Extracellular | Glycoprotein | ||
| Fibronectin | FINC_HUMAN | P02751 | FN1 | Extracellular | Glycoprotein | ||
| Hyaluronan and proteoglycan link protein 3 | HPLN3_HUMAN | Q96S86 | HAPLN3 | Extracellular | |||
| Laminin subunit alpha-4 | LAMA4_HUMAN | Q16363 | LAMA4 | Extracellular | Glycoprotein | ||
| Laminin subunit beta-1 | LAMB1_HUMAN | P07942 | LAMB1 | Extracellular | Glycoprotein | ||
| Laminin subunit gamma-1 | LAMC1_HUMAN | P11047 | LAMC1 | Extracellular | Glycoprotein | ||
| Lysyl oxidase homolog 2 | LOXL2_HUMAN | Q9Y4K0 | LOXL2 | Extracellular | Glycoprotein | ||
| Multimerin-2 | MMRN2_HUMAN | Q9H8L6 | MMRN2 | Extracellular | Glycoprotein | ||
| Nidogen-1 | NID1_HUMAN | P14543 | NID1 | Extracellular | Glycoprotein | ||
| Nidogen-2 | NID2_HUMAN | Q14112 | NID2 | Extracellular | Glycoprotein | ||
| Prolyl 3-hydroxylase 1 | P3H1_HUMAN | Q32P28 | LEPRE1 | Extracellular | Glycoprotein | ||
| Basement membrane-specific heparan sulfate proteoglycan core protein | PGBM_HUMAN | P98160 | HSPG2 | Extracellular | Glycoprotein | ||
| Biglycan | PGS1_HUMAN | P21810 | BGN | Extracellular | Glycoprotein | ||
| Peroxidasin homolog | PXDN_HUMAN | Q92626 | PXDN | Extracellular | Glycoprotein | ||
| SPARC | SPRC_HUMAN | P09486 | SPARC | Extracellular | Glycoprotein | ||
| Target of Nesh-SH3 | TARSH_HUMAN | Q7Z7G0 | ABI3BP | Extracellular | Glycoprotein | ||
| Testican-1 | TICN1_HUMAN | Q08629 | SPOCK1 | Extracellular | Glycoprotein | ||
| Thrombospondin-1 | TSP1_HUMAN | P07996 | THBS1 | Extracellular | Plasma membrane | Glycoprotein | |
| Growth factors and related proteins | |||||||
| C-type lectin domain family 11 member A | CLC11_HUMAN | Q9Y240 | CLEC11A | Extracellular | Glycoprotein | ||
| Cysteine-rich motor neuron 1 protein | CRIM1_HUMAN | Q9NZV1 | CRIM1 | Extracellular | Plasma membrane | Glycoprotein | |
| Connective tissue growth factor | CTGF_HUMAN | P29279 | CTGF | Extracellular | Glycoprotein | ||
| Protein CYR61, insulin-like growth factor-binding protein 10 | CYR61_HUMAN | O00622 | CYR61 | Extracellular | |||
| Dickkopf-related protein 3 | DKK3_HUMAN | Q9UBP4 | DKK3 | Extracellular | Glycoprotein | ||
| Follistatin-related protein 1 | FSTL1_HUMAN | Q12841 | FSTL1 | Extracellular | Glycoprotein | ||
| Hepatoma-derived growth factor | HDGF_HUMAN | P51858 | HDGF | Extracellular | |||
| Insulin-like growth factor-binding protein 2 | IBP2_HUMAN | P18065 | IGFBP2 | Extracellular | Glycoprotein | ||
| Insulin-like growth factor-binding protein 7 | IBP7_HUMAN | Q16270 | IGFBP7 | Extracellular | Glycoprotein | ||
| Latent-transforming growth factor beta-binding protein 1 | LTBP1_HUMAN | Q14766 | LTBP1 | Extracellular | Glycoprotein | ||
| Latent-transforming growth factor beta-binding protein 2 | LTBP2_HUMAN | Q14767 | LTBP2 | Extracellular | Glycoprotein | ||
| Neuronal growth regulator 1 | NEGR1_HUMAN | Q7Z3B1 | NEGR1 | Plasma membrane | Glycoprotein | ||
| Immunity- and inflammation-related proteins | |||||||
| Amyloid beta A4 protein | A4_HUMAN | P05067 | APP | Extracellular | Plasma membrane | Glycoprotein | |
| Beta-2-microglobulin | B2MG_HUMAN | P61769 | B2M | Extracellular | Glycoprotein | ||
| Complement C1q tumor necrosis factor-related protein 5 | C1QT5_HUMAN | Q9BXJ0 | C1QTNF5 | Extracellular | |||
| Complement factor H | CFAH_HUMAN | P08603 | CFH | Extracellular | Glycoprotein | ||
| Interleukin-25, UPF0556 protein C19orf10 | CS010_HUMAN | Q969H8 | C19orf10 | Extracellular | |||
| Granulins | GRN_HUMAN | P28799 | GRN | Extracellular | Glycoprotein | ||
| Interferon-induced transmembrane protein 1 | IFM1_HUMAN | P13164 | IFITM1 | Plasma membrane | |||
| Galectin-1a | LEG1_HUMAN | P09382 | LGALS1 | Extracellular | |||
| Galectin-3 | LEG3_HUMAN | P17931 | LGALS3 | Extracellular | |||
| Macrophage migration inhibitory factora | MIF_HUMAN | P14174 | MIF | Extracellular | |||
| NKG2D ligand 2 | N2DL2_HUMAN | Q9BZM5 | ULBP2 | Extracellular | Plasma membrane | Glycoprotein | |
| Pentraxin-related protein PTX3 | PTX3_HUMAN | P26022 | PTX3 | Extracellular | Glycoprotein | ||
| Protein S100-A7 | S10A7_HUMAN | P31151 | S100A7 | Extracellular | |||
| Protein S100-A8 | S10A8_HUMAN | P05109 | S100A8 | Extracellular | Plasma membrane | ||
| Tubulointerstitial nephritis antigen-like | TINAL_HUMAN | Q9GZM7 | TINAGL1 | Extracellular | Glycoprotein | ||
| Nuclease-sensitive element-binding protein 1 | YBOX1_HUMAN | P67809 | YBX1 | Extracellular | |||
| Zinc-alpha-2-glycoprotein | ZA2G_HUMAN | P25311 | AZGP1 | Extracellular | Glycoprotein | ||
| Membrane antigens and receptors | |||||||
| HLA class I histocompatibility antigen, A-24 alpha chain | 1A24_HUMAN | P05534 | HLA-A | Plasma membrane | Glycoprotein | ||
| HLA class I histocompatibility antigen, A-30 alpha chain | 1A30_HUMAN | P16188 | HLA-A | Plasma membrane | Glycoprotein | ||
| HLA class I histocompatibility antigen, Cw-12 alpha chain | 1C12_HUMAN | P30508 | HLA-C | Plasma membrane | Glycoprotein | ||
| Alpha-2-macroglobulin receptor-associated protein | AMRP_HUMAN | P30533 | LRPAP1 | Extracellular | Plasma membrane | Glycoprotein | |
| Basal cell adhesion molecule | BCAM_HUMAN | P50895 | BCAM | Plasma membrane | Glycoprotein | ||
| Complement component C1q receptor | C1QR1_HUMAN | Q9NPY3 | CD93 | Plasma membrane | Glycoprotein | EC marker | |
| Cadherin-13 | CAD13_HUMAN | P55290 | CDH13 | Plasma membrane | Glycoprotein | ||
| Cadherin-2 | CADH2_HUMAN | P19022 | CDH2 | Plasma membrane | Glycoprotein | ||
| Cadherin-5 | CADH5_HUMAN | P33151 | CDH5 | Plasma membrane | Glycoprotein | EC marker | |
| CD109 antigen | CD109_HUMAN | Q6YHK3 | CD109 | Plasma membrane | Glycoprotein | ||
| CD166 antigen | CD166_HUMAN | Q13740 | ALCAM | Plasma membrane | Glycoprotein | ||
| CD44 antigen | CD44_HUMAN | P16070 | CD44 | Plasma membrane | Glycoprotein | ||
| CD59 glycoprotein | CD59_HUMAN | P13987 | CD59 | Extracellular | Plasma membrane | Glycoprotein | |
| CD9 antigen | CD9_HUMAN | P21926 | CD9 | Plasma membrane | Glycoprotein | ||
| C-type lectin domain family 14 member A | CLC14_HUMAN | Q86T13 | CLEC14A | Plasma membrane | Glycoprotein | ||
| Dystroglycan | DAG1_HUMAN | Q14118 | DAG1 | Extracellular | Plasma membrane | Glycoprotein | |
| Endoglin | EGLN_HUMAN | P17813 | ENG | Plasma membrane | Glycoprotein | EC marker | |
| Endothelial protein C receptor | EPCR_HUMAN | Q9UNN8 | PROCR | Plasma membrane | Glycoprotein | EC marker | |
| Ephrin type-B receptor 4 | EPHB4_HUMAN | P54760 | EPHB4 | Plasma membrane | Glycoprotein | ||
| Endothelial cell-selective adhesion molecule | ESAM_HUMAN | Q96AP7 | ESAM | Plasma membrane | Glycoprotein | EC marker | |
| Leucine-rich repeat transmembrane protein FLRT2 | FLRT2_HUMAN | O43155 | FLRT2 | Plasma membrane | Glycoprotein | ||
| Guanine nucleotide-binding protein subunit beta-2-like 1a | GBLP_HUMAN | P63244 | GNB2L1 | Plasma membrane | |||
| HLA class I histocompatibility antigen, alpha chain E | HLAE_HUMAN | P13747 | HLA-E | Plasma membrane | Glycoprotein | ||
| Intercellular adhesion molecule 1 | ICAM1_HUMAN | P05362 | ICAM1 | Extracellular | Plasma membrane | Glycoprotein | EC marker |
| Intercellular adhesion molecule 2 | ICAM2_HUMAN | P13598 | ICAM2 | Plasma membrane | Glycoprotein | EC marker | |
| Integrin alpha-2 | ITA2_HUMAN | P17301 | ITGA2 | Plasma membrane | Glycoprotein | ||
| Integrin alpha-5 | ITA5_HUMAN | P08648 | ITGA5 | Plasma membrane | Glycoprotein | ||
| Integrin alpha-6 | ITA6_HUMAN | P23229 | ITGA6 | Plasma membrane | Glycoprotein | ||
| Integrin beta-1 | ITB1_HUMAN | P05556 | ITGB1 | Plasma membrane | Glycoprotein | EC marker | |
| Protein jagged-1 | JAG1_HUMAN | P78504 | JAG1 | Plasma membrane | Glycoprotein | ||
| Protein jagged-2 | JAG2_HUMAN | Q9Y219 | JAG2 | Plasma membrane | Glycoprotein | ||
| Junctional adhesion molecule A | JAM1_HUMAN | Q9Y624 | F11R | Plasma membrane | Glycoprotein | ||
| BTB/POZ domain-containing protein KCTD12 | KCD12_HUMAN | Q96CX2 | KCTD12 | Plasma membrane | |||
| Kinectin | KTN1_HUMAN | Q86UP2 | KTN1 | Plasma membrane | Glycoprotein | ||
| Lysosome-associated membrane glycoprotein 1 | LAMP1_HUMAN | P11279 | LAMP1 | Plasma membrane | Glycoprotein | ||
| Low-density lipoprotein receptor | LDLR_HUMAN | P01130 | LDLR | Plasma membrane | Glycoprotein | ||
| Low-density lipoprotein receptor-related protein 5 | LRP5_HUMAN | O75197 | LRP5 | Plasma membrane | Glycoprotein | ||
| Lymphatic vessel endothelial hyaluronic acid receptor 1 | LYVE1_HUMAN | Q9Y5Y7 | LYVE1 | Plasma membrane | Glycoprotein | EC marker | |
| Hepatocyte growth factor receptor | MET_HUMAN | P08581 | MET | Extracellular | Plasma membrane | Glycoprotein | |
| Cation-independent mannose-6-phosphate receptor | MPRI_HUMAN | P11717 | IGF2R | Plasma membrane | Glycoprotein | ||
| C-type mannose receptor 2 | MRC2_HUMAN | Q9UBG0 | MRC2 | Plasma membrane | Glycoprotein | ||
| Cell surface glycoprotein MUC18 | MUC18_HUMAN | P43121 | MCAM | Plasma membrane | Glycoprotein | EC marker | |
| Neuroligin-1 | NLGN1_HUMAN | Q8N2Q7 | NLGN1 | Plasma membrane | Glycoprotein | ||
| Neuronal cell adhesion molecule | NRCAM_HUMAN | Q92823 | NRCAM | Plasma membrane | Glycoprotein | ||
| Neuropilin-1 | NRP1_HUMAN | O14786 | NRP1 | Extracellular | Plasma membrane | Glycoprotein | |
| Neuropilin-2 | NRP2_HUMAN | O60462 | NRP2 | Plasma membrane | Glycoprotein | ||
| Neurotrimin | NTRI_HUMAN | Q9P121 | NTM | Plasma membrane | Glycoprotein | ||
| Protocadherin-10 | PCD10_HUMAN | Q9P2E7 | PCDH10 | Plasma membrane | Glycoprotein | ||
| Protocadherin-12 | PCD12_HUMAN | Q9NPG4 | PCDH12 | Plasma membrane | Glycoprotein | ||
| Protocadherin gamma-A11 | PCDGB_HUMAN | Q9Y5H2 | PCDHGA11 | Plasma membrane | Glycoprotein | ||
| Protocadherin gamma-A12 | PCDGC_HUMAN | O60330 | PCDHGA12 | Plasma membrane | Glycoprotein | ||
| Protocadherin gamma-B7 | PCDGJ_HUMAN | Q9Y5F8 | PCDHGB7 | Plasma membrane | Glycoprotein | ||
| Protocadherin-1 | PCDH1_HUMAN | Q08174 | PCDH1 | Plasma membrane | Glycoprotein | ||
| Protocadherin-9 | PCDH9_HUMAN | Q9HC56 | PCDH9 | Plasma membrane | Glycoprotein | ||
| Programmed cell death 1 ligand 2 | PD1L2_HUMAN | Q9BQ51 | PDCD1LG2 | Extracellular | Plasma membrane | Glycoprotein | |
| Platelet endothelial cell adhesion molecule | PECA1_HUMAN | P16284 | PECAM1 | Plasma membrane | Glycoprotein | EC marker | |
| Plexin-D1 | PLXD1_HUMAN | Q9Y4D7 | PLXND1 | Plasma membrane | Glycoprotein | ||
| Inactive tyrosine-protein kinase 7 | PTK7_HUMAN | Q13308 | PTK7 | Plasma membrane | Glycoprotein | ||
| Receptor-type tyrosine-protein phosphatase delta | PTPRD_HUMAN | P23468 | PTPRD | Plasma membrane | Glycoprotein | ||
| Receptor-type tyrosine-protein phosphatase F | PTPRF_HUMAN | P10586 | PTPRF | Plasma membrane | Glycoprotein | ||
| Receptor-type tyrosine-protein phosphatase kappa | PTPRK_HUMAN | Q15262 | PTPRK | Plasma membrane | Glycoprotein | ||
| Poliovirus receptor | PVR_HUMAN | P15151 | PVR | Extracellular | Plasma membrane | Glycoprotein | |
| Poliovirus receptor-related protein 2 | PVRL2_HUMAN | Q92692 | PVRL2 | Plasma membrane | Glycoprotein | EC marker | |
| Roundabout homolog 1 | ROBO1_HUMAN | Q9Y6N7 | ROBO1 | Plasma membrane | Glycoprotein | ||
| Roundabout homolog 4 | ROBO4_HUMAN | Q8WZ75 | ROBO4 | Plasma membrane | Glycoprotein | ||
| Syndecan-4 | SDC4_HUMAN | P31431 | SDC4 | Extracellular | Plasma membrane | Glycoprotein | |
| Semaphorin-4D | SEM4D_HUMAN | Q92854 | SEMA4D | Plasma membrane | Glycoprotein | ||
| Semaphorin-6B | SEM6B_HUMAN | Q9H3T3 | SEMA6B | Plasma membrane | Glycoprotein | ||
| Tyrosine-protein phosphatase non-receptor type substrate 1 | SHPS1_HUMAN | P78324 | SIRPA | Plasma membrane | Glycoprotein | ||
| Stabilin-1 | STAB1_HUMAN | Q9NY15 | STAB1 | Plasma membrane | Glycoprotein | EC marker | |
| Transferrin receptor protein 1 | TFR1_HUMAN | P02786 | TFRC | Extracellular | Plasma membrane | Glycoprotein | |
| Tyrosine-protein kinase receptor Tie-1 | TIE1_HUMAN | P35590 | TIE1 | Plasma membrane | Glycoprotein | ||
| Tyrosine-protein kinase receptor UFO | UFO_HUMAN | P30530 | AXL | Extracellular | Plasma membrane | Glycoprotein | |
| Vascular endothelial growth factor receptor 2 | VGFR2_HUMAN | P35968 | KDR | Extracellular | Plasma membrane | Glycoprotein | EC marker |
| Vascular endothelial growth factor receptor 3 | VGFR3_HUMAN | P35916 | FLT4 | Extracellular | Plasma membrane | Glycoprotein | EC marker |
| Very low-density lipoprotein receptor | VLDLR_HUMAN | P98155 | VLDLR | Plasma membrane | Glycoprotein | ||
| Miscellaneous membrane proteins | |||||||
| Brain acid soluble protein 1 | BASP1_HUMAN | P80723 | BASP1 | Plasma membrane | |||
| DnaJ homolog subfamily B member 4 | DNJB4_HUMAN | Q9UDY4 | DNAJB4 | Plasma membrane | |||
| RNA-binding protein EWS | EWS_HUMAN | Q01844 | EWSR1 | Plasma membrane | |||
| Nck-associated protein 1 | NCKP1_HUMAN | Q9Y2A7 | NCKAP1 | Plasma membrane | |||
| Na(+)/H(+) exchange regulatory cofactor NHE-RF2 | NHRF2_HUMAN | Q15599 | SLC9A3R2 | Plasma membrane | |||
| Polymerase I and transcript release factor | PTRF_HUMAN | Q6NZI2 | PTRF | Plasma membrane | |||
| Serum deprivation-response protein | SDPR_HUMAN | O95810 | SDPR | Plasma membrane | |||
| Sushi repeat-containing protein SRPX2 | SRPX2_HUMAN | O60687 | SRPX2 | Extracellular | |||
| Erythrocyte band 7 integral membrane protein | STOM_HUMAN | P27105 | STOM | Plasma membrane | |||
| Miscellaneous secreted proteins | |||||||
| Peptidyl-glycine alpha-amidating monooxygenase | AMD_HUMAN | P19021 | PAM | Extracellular | Glycoprotein | ||
| Angiopoietin-2 | ANGP2_HUMAN | O15123 | ANGPT2 | Extracellular | Glycoprotein | ||
| Endothelin-1 | EDN1_HUMAN | P05305 | EDN1 | Extracellular | |||
| Endothelial cell-specific molecule 1 | ESM1_HUMAN | Q9NQ30 | ESM1 | Extracellular | Glycoprotein | ||
| Protein FAM3C | FAM3C_HUMAN | Q92520 | WNT16 | Extracellular | |||
| Epididymal secretory protein E1 | NPC2_HUMAN | P61916 | NPC2 | Extracellular | Glycoprotein | ||
| Programmed cell death protein 10 | PDC10_HUMAN | Q9BUL8 | PDCD10 | Plasma membrane | |||
| Prolactin-inducible protein | PIP_HUMAN | P12273 | PIP | Extracellular | Glycoprotein | ||
| Sulfhydryl oxidase 1 | QSOX1_HUMAN | O00391 | QSOX1 | Extracellular | Glycoprotein | ||
| Secretoglobin family 1D member 2 | SG1D2_HUMAN | O95969 | SCGB1D2 | Extracellular | |||
| Thioredoxina | THIO_HUMAN | P10599 | TXN | Extracellular | |||
| Thymosin beta-4 | TYB4_HUMAN | P62328 | TMSB4X | Extracellular | |||
| Protease inhibitors | |||||||
| Cystatin-C | CYTC_HUMAN | P01034 | CST3 | Extracellular | Glycoprotein | ||
| Leukocyte elastase inhibitor | ILEU_HUMAN | P30740 | SERPINB1 | Extracellular | |||
| Inter-alpha-trypsin inhibitor heavy chain H2 | ITIH2_HUMAN | P19823 | ITIH2 | Extracellular | Glycoprotein | ||
| Serpin B9 | SPB9_HUMAN | P50453 | SERPINB9 | Extracellular | |||
| Metalloproteinase inhibitor 1 | TIMP1_HUMAN | P01033 | TIMP1 | Extracellular | Glycoprotein | ||
| Metalloproteinase inhibitor 2 | TIMP2_HUMAN | P16035 | TIMP2 | Extracellular | |||
| Proteases | |||||||
| Angiotensin-converting enzyme | ACE_HUMAN | P12821 | ACE | Extracellular | Plasma membrane | Glycoprotein | EC marker |
| Disintegrin and metalloproteinase domain-containing protein 10 | ADA10_HUMAN | O14672 | ADAM10 | Plasma membrane | Glycoprotein | ||
| Aminopeptidase B | AMPB_HUMAN | Q9H4A4 | RNPEP | Extracellular | |||
| Aminopeptidase N | AMPN_HUMAN | P15144 | ANPEP | Plasma membrane | Glycoprotein | ||
| Bone morphogenetic protein 1 | BMP1_HUMAN | P13497 | BMP1 | Extracellular | Glycoprotein | ||
| Cathepsin B | CATB_HUMAN | P07858 | CTSB | Extracellular | Glycoprotein | ||
| Cathepsin D | CATD_HUMAN | P07339 | CTSD | Extracellular | Glycoprotein | ||
| Cathepsin Z | CATZ_HUMAN | Q9UBR2 | CTSZ | Extracellular | Glycoprotein | ||
| Carboxypeptidase Q | CBPQ_HUMAN | Q9Y646 | CPQ | Extracellular | Glycoprotein | ||
| Dipeptidyl peptidase 2 | DPP2_HUMAN | Q9UHL4 | DPP7 | Extracellular | Glycoprotein | ||
| Dipeptidyl peptidase 3 | DPP3_HUMAN | Q9NY33 | DPP3 | Plasma membrane | |||
| Endoplasmic reticulum aminopeptidase 1 | ERAP1_HUMAN | Q9NZ08 | ERAP1 | Extracellular | Glycoprotein | ||
| Furin | FURIN_HUMAN | P09958 | FURIN | Plasma membrane | Glycoprotein | ||
| Gamma-glutamyl hydrolase | GGH_HUMAN | Q92820 | GGH | Extracellular | Glycoprotein | ||
| Serine protease HTRA1 | HTRA1_HUMAN | Q92743 | HTRA1 | Extracellular | |||
| Insulin-degrading enzyme | IDE_HUMAN | P14735 | IDE | Extracellular | Plasma membrane | ||
| Interstitial collagenase | MMP1_HUMAN | P03956 | MMP1 | Extracellular | Glycoprotein | ||
| Stromelysin-2 | MMP10_HUMAN | P09238 | MMP10 | Extracellular | |||
| Matrix metalloproteinase-14 | MMP14_HUMAN | P50281 | MMP14 | Plasma membrane | |||
| 72 kDa type IV collagenase | MMP2_HUMAN | P08253 | MMP2 | Extracellular | Glycoprotein | ||
| Lysosomal Pro-X carboxypeptidase | PCP_HUMAN | P42785 | PRCP | Plasma membrane | Glycoprotein | ||
| Serine protease 23 | PRS23_HUMAN | O95084 | PRSS23 | Extracellular | Glycoprotein | ||
| Ubiquitin carboxyl-terminal hydrolase 14 | UBP14_HUMAN | P54578 | USP14 | Plasma membrane | |||
| Signal transduction proteins | |||||||
| Adenylyl cyclase-associated protein 1 | CAP1_HUMAN | Q01518 | CAP1 | Plasma membrane | |||
| Cell division control protein 42 homolog | CDC42_HUMAN | P60953 | CDC42 | Plasma membrane | |||
| Contactin-associated protein-like 3 | CNTP3_HUMAN | Q9BZ76 | CNTNAP3 | Extracellular | Plasma membrane | Glycoprotein | |
| Adapter molecule crk | CRK_HUMAN | P46108 | CRK | Plasma membrane | |||
| Ras GTPase-activating protein-binding protein 1 | G3BP1_HUMAN | Q13283 | G3BP1 | Plasma membrane | |||
| Growth arrest-specific protein 6 | GAS6_HUMAN | Q14393 | GAS6 | Extracellular | Glycoprotein | ||
| Interferon-induced guanylate-binding protein 1 | GBP1_HUMAN | P32455 | GBP1 | Extracellular | |||
| Guanine nucleotide-binding protein G(i) subunit alpha-2 | GNAI2_HUMAN | P04899 | GNAI2 | Plasma membrane | |||
| Glypican-1 | GPC1_HUMAN | P35052 | GPC1 | Extracellular | Plasma membrane | Glycoprotein | |
| Hedgehog-interacting protein | HHIP_HUMAN | Q96QV1 | HHIP | Extracellular | Plasma membrane | Glycoprotein | |
| Histidine triad nucleotide-binding protein 1a | HINT1_HUMAN | P49773 | HINT1 | Plasma membrane | |||
| Integrin-linked protein kinase | ILK_HUMAN | Q13418 | ILK | Plasma membrane | |||
| Ras GTPase-activating-like protein IQGAP1 | IQGA1_HUMAN | P46940 | IQGAP1 | Plasma membrane | |||
| cAMP-dependent protein kinase type II-alpha regulatory subunit | KAP2_HUMAN | P13861 | PRKAR2A | Plasma membrane | |||
| Ras-related protein Rab-18 | RAB18_HUMAN | Q9NP72 | RAB18 | Plasma membrane | |||
| Ras-related protein Rab-5C | RAB5C_HUMAN | P51148 | RAB5C | Plasma membrane | |||
| Ras-related C3 botulinum toxin substrate 1 | RAC1_HUMAN | P63000 | RAC1 | Plasma membrane | |||
| Ras-related protein Ral-A | RALA_HUMAN | P11233 | RALA | Plasma membrane | |||
| Ras-related protein Rap-1b | RAP1B_HUMAN | P61224 | RAP1B | Plasma membrane | |||
| GTPase NRas | RASN_HUMAN | P01111 | NRAS | Plasma membrane | |||
| Ras-related protein Rab-11A | RB11A_HUMAN | P62491 | RAB11A | Plasma membrane | |||
| Rho-related GTP-binding protein RhoC | RHOC_HUMAN | P08134 | RHOC | Plasma membrane | |||
| Rho-associated protein kinase 2 | ROCK2_HUMAN | O75116 | ROCK2 | Plasma membrane | |||
| Ras-related protein R-Ras2 | RRAS2_HUMAN | P62070 | RRAS2 | Plasma membrane | |||
| Protein S100-A6a | S10A6_HUMAN | P06703 | S100A6 | Plasma membrane | |||
| Protein S100-A10a | S10AA_HUMAN | P60903 | S100A10 | Plasma membrane | |||
| Switch-associated protein 70 | SWP70_HUMAN | Q9UH65 | SWAP70 | Plasma membrane | |||
| NEDD8-activating enzyme E1 regulatory subunit | ULA1_HUMAN | Q13564 | NAE1 | Plasma membrane | |||
| Transport-related proteins | |||||||
| AP-2 complex subunit alpha-1 | AP2A1_HUMAN | O95782 | AP2A1 | Plasma membrane | |||
| AP-2 complex subunit alpha-2 | AP2A2_HUMAN | O94973 | AP2A2 | Plasma membrane | |||
| ADP-ribosylation factor 1 | ARF1_HUMAN | P84077 | ARF1 | Plasma membrane | |||
| ADP-ribosylation factor 6 | ARF6_HUMAN | P62330 | ARF6 | Plasma membrane | |||
| ADP-ribosylation factor-like protein 3 | ARL3_HUMAN | P36405 | ARL3 | Plasma membrane | |||
| Beta-arrestin-1 | ARRB1_HUMAN | P49407 | ARRB1 | Plasma membrane | |||
| Chloride intracellular channel protein 1 | CLIC1_HUMAN | O00299 | CLIC1 | Plasma membrane | |||
| Chloride intracellular channel protein 4 | CLIC4_HUMAN | Q9Y696 | CLIC4 | Plasma membrane | |||
| Clusterin | CLUS_HUMAN | P10909 | CLU | Extracellular | Glycoprotein | ||
| Coatomer subunit beta | COPB_HUMAN | P53618 | COPB1 | Plasma membrane | |||
| EH domain-containing protein 1 | EHD1_HUMAN | Q9H4M9 | EHD1 | Plasma membrane | |||
| EH domain-containing protein 2 | EHD2_HUMAN | Q9NZN4 | EHD2 | Plasma membrane | |||
| Palmitoyl-protein thioesterase 1 | PPT1_HUMAN | P50897 | PPT1 | Extracellular | Glycoprotein | ||
| Protein S100-A13 | S10AD_HUMAN | Q99584 | S100A13 | Extracellular | |||
| Solute carrier family 12 member 2 | S12A2_HUMAN | P55011 | SLC12A2 | Plasma membrane | |||
| Proactivator polypeptide | SAP_HUMAN | P07602 | PSAP | Extracellular | Glycoprotein | ||
| Syntaxin-binding protein 1 | STXB1_HUMAN | P61764 | STXBP1 | Plasma membrane | |||
| Syntaxin-binding protein 3 | STXB3_HUMAN | O00186 | STXBP3 | Plasma membrane | |||
| Transmembrane emp24 domain-containing protein 10 | TMEDA_HUMAN | P49755 | TMED10 | Plasma membrane | Glycoprotein | ||
| Vesicle-associated membrane protein-associated protein A | VAPA_HUMAN | Q9P0L0 | VAPA | Plasma membrane | |||
a These proteins were also identified in the cellular proteome according to the difference gel electrophoresis analysis presented in the supplemental data.
Immunoblots confirmed that proteins such as fibronectin and biglycan were constitutively secreted (Fig. 1B). Others such as agrin and lymphatic vessel endothelial hyaluronic acid receptor 1 were released upon PMA stimulation, providing an explanation for why previously unidentified proteins (8, 10) were found in the present analysis (Fig. 1C). An overlay between secreted (Cy3 and Cy 5; green and red color) and cellular (Cy 2; blue color) proteins is shown in Fig. 1D. Common spots were numbered (supplemental Fig. S2) and identified via LC-MS/MS (supplemental Table S3). Certain proteins, such as von Willebrand antigen 2 (a propeptide of vWF, AA 23–763), were clearly more abundant in the secretome of PMA-treated HUVECs.
The Endothelial Glycoproteome
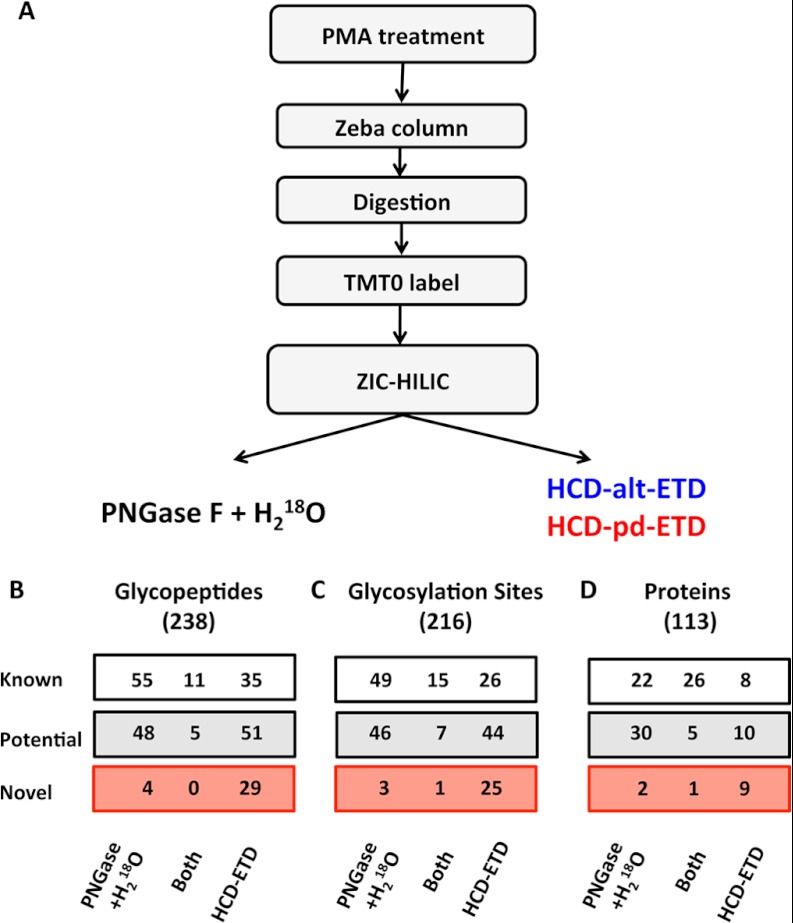
Among the 1252 identified proteins were 253 extracellular or plasma membrane proteins (approximately 20%) related to cell adhesion, blood coagulation, hemostasis, signaling transduction, and protein transportation, of which 166 were known glycoproteins (Table I). To further characterize this subproteome, we employed a glycoproteomics approach. Secreted proteins were precipitated and digested with trypsin, and tryptic peptides were labeled with TMT0 to increase their charge state prior to enrichment by means of zwitterionic hydrophilic interaction liquid chromatography purification (24). For glycosite identification, an indirect and a direct strategy were pursued (Fig. 2A): (i) digestion with PNGase F in the presence of 18O water to label the conversion of asparagine to aspartic acid upon the removal of N-glycans, and (ii) alternating HCD and ETD (HCD-alt-ETD) or HCD-product-dependent ETD (HCD-pd-ETD) fragmentation on an Orbitrap Elite MS (24).
Fig. 2.
Glycoproteomics. A, glycopeptide identification workflow. Comparison of direct and indirect glycopeptide detection using HCD-ETD and 18O-deamidation after PNGase F + H218O treatment, respectively: identified unique glycopeptides (B), unique glycosylation sites (C), and unique glycoproteins (D).
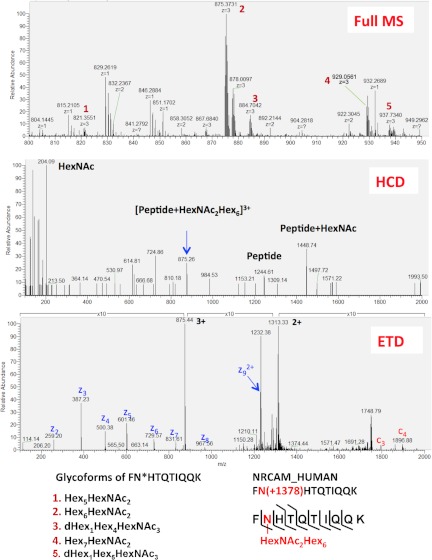
There was little overlap in the numbers of glycopeptides (Fig. 2B) and glycosylation sites (Fig. 2C) identified via the direct (HCD-ETD) and the indirect (PNGase F + H218O) methods. Better agreement was observed at the protein level (Fig. 2D). With the indirect (PNGase F + H218O) method, 27 peptides were identified with N[+2.99] modification at non-consensus sequence, out of 1139 total identified peptides with N[+2.99]. This anomaly rate of 2.4% (27/1139) combines the rate of false identifications and the rate of chance deamidations in 18O water that were not in the consensus sequence of glycosylation (i.e. N-X(not P)-S/T). All glycopeptides identified are listed in Table II and supplemental Table S4. Three spectra (full MS, HCD, and ETD) from a neuronal cell adhesion molecule (UniProt accession number Q92823) (AA - 222FNHTQTIQQK231) are presented in Fig. 3.
Table II. Glycopeptides identified via the HCD-ETD method.
| Protein name | UniProt ID | Peptide | Glycosite | Type | Glycans | Observed m/z | Z | ΔMass (ppm) |
|---|---|---|---|---|---|---|---|---|
| Afamin | AFAM_HUMAN | $DIENFN(+1702.582)^STQk | N33 | Known | Hex8HexNAc2 | 843.359 | 4 | −5.8 |
| Aminopeptidase N | AMPN_HUMAN | $kLN(+892.318)YTLSQGHRVVLR | N128 | Known | Hex3HexNAc2 | 781.916 | 4 | −2.8 |
| $NAN(+2042.720)^SSPVAsTTPSASATTNPASATTLDQSk | N42 | Novel | Hex4HexNAc4NeuAc2 | 1095.276 | 5 | −0.3 | ||
| Alpha-N-acetylglucosaminidase | ANAG_HUMAN | $SVYN(+1257.450)cSGEAcRGhNRSPLVR | N503 | Potential | Hex4HexNAc3 | 954.927 | 4 | 1.1 |
| Angiopoietin-2 | ANGP2_HUMAN | $kIVTATVN(+568.212)NSVLQk | N240 | Potential | Hex1HexNAc2 | 689.645 | 4 | −0.3 |
| $SGhTTNGIYTLTFPN(+1038.375)STEEIk | N304 | Potential | Hex3HexNAc2dHex1 | 763.565 | 5 | −1.4 | ||
| Attractin | ATRN_HUMAN | $DLDMFIN(+1241.455)ASk | N1198 | Known | Hex3HexNAc3dHex1 | 948.450 | 3 | 6.7 |
| $GcScFSDWQGPGcSVPVPAN(+1095.397)QSFWTR | N325 | Potential | Hex3HexNAc3 | 1077.708 | 4 | −2.1 | ||
| $GcScFSDWQGPGcSVPVPAN(+892.317)QSFWTREEYSnLk | N325 | Potential | Hex3HexNAc2 | 1040.061 | 5 | −2.4 | ||
| Cadherin-2 | CADH2_HUMAN | $EQIARFHLRAHAVDInGNQVENPIDIVINVIDMNDNRPEFLHQVWN(+1751.624)GTVPEGSk | N273 | Known | Hex4HexNAc4NeuAc1 | 930.011 | 9 | 2.8 |
| Cadherin-5 | CADH5_HUMAN | $EN(+1054.370)^ISEYHLTAVIVDk | N112 | Known | Hex4HexNAc2 | 815.146 | 4 | 0.7 |
| $ELDREVYPWYN(+1241.455)LTVEAk | N442 | Known | Hex3HexNAc3dHex1 | 954.972 | 4 | 10.0 | ||
| CD109 antigen | CD109_HUMAN | $LNLYLDSVN(+1038.375)^ETQFcVNIPAVR | N1355 | Novel | Hex3HexNAc2dHex1 | 938.447 | 4 | 1.5 |
| $kkN(+1540.529)ITk | N279 | Potential | Hex7HexNAc2 | 793.410 | 4 | −0.9 | ||
| $QN(+1848.640)^STMFSLTPENSWTPk | N513 | Novel | Hex8HexNAc2dHex1 | 1072.719 | 4 | 1.0 | ||
| CD59 glycoprotein | CD59_HUMAN | $TAVN(+1954.704)^csSDFDAcLITk | N43 | Known | Hex4HexNAc5NeuAc1 | 1077.708 | 4 | 9.1 |
| Complement factor I | CFAI_HUMAN | $FLNN(+1054.370)GTcTAEGk | N103 | Known | Hex4HexNAc2 | 939.094 | 3 | −4.0 |
| $LISN(+3534.244)^cSk | N494 | Potential | Hex8HexNAc6dHex1NeuAc3 | 1207.501 | 4 | 5.7 | ||
| CAP-Gly domain-containing linker protein 1 | CLIP1_HUMAN | $GEN(+1257.450)^ASAk | N1263 | Novel | Hex4HexNAc3 | 802.359 | 3 | −1.4 |
| $GEN(+1784.635)ASAk | N1263 | Novel | Hex6HexNAc4 | 970.437 | 3 | 10.7 | ||
| $EPSATPPISN(+2188.741)LTk | N187 | Novel | Hex11HexNAc2 | 999.192 | 4 | −6.9 | ||
| $ANEN(+1200.428)^ASFLQkSIEDMTVk | N971 | Novel | Hex4HexNAc2dHex1 | 980.724 | 4 | 2.6 | ||
| $ANEN(+1216.423)^ASFLQkSIEDMTVk | N971 | Novel | Hex5HexNAc2 | 985.232 | 4 | 11.0 | ||
| $ANEN(+1257.449)ASFLQkSIEDMTVk | N971 | Novel | Hex4HexNAc3 | 989.489 | 4 | 8.2 | ||
| $ANEN(+1444.534)ASFLQkSIEDMTVk | N971 | Novel | Hex3HexNAc4dHex1 | 1036.764 | 4 | 9.3 | ||
| Ephrin type-A receptor 2 | EPHA2_HUMAN | $TASVSIN(+892.317)QTEPPkVRLEGR | N435 | Known | Hex3HexNAc2 | 856.694 | 4 | −0.4 |
| Fibrous sheath-interacting protein 2 | FSIP2_HUMAN | $IGWEYESTN(+1751.624)ISR | N1423 | Novel | Hex4HexNAc4NeuAc1 | 858.373 | 4 | 0.8 |
| $TITFSAN(+1362.481)VSSHEhTYk | N1675 | Novel | Hex5HexNAc2dHex1 | 912.923 | 4 | 3.0 | ||
| $GGIN(+892.318)ISGQGSIISAQVSPTR | N215 | Novel | Hex3HexNAc2 | 1020.509 | 3 | 1.5 | ||
| $ENSN(+1200.428)FSQLALSNEILLGHkEk | N2216 | Novel | Hex4HexNAc2dHex1 | 708.363 | 6 | 6.3 | ||
| $mPIEN(+1444.534)LSSIQQk | N2824 | Novel | Hex3HexNAc4dHex1 | 825.398 | 4 | 0.8 | ||
| $YN(+2204.772)k | N427 | Novel | Hex5HexNAc4NeuAc2 | 1026.779 | 3 | 7.7 | ||
| N-acetylglucosamine-6-sulfatase | GNS_HUMAN | $LVkRLEFTGELN(+2018.708)^NTYIFYTSDnGYHTGQFSLPIDkR | N317 | Known | Hex6HexNAc3dHex1NeuAc1 | 696.027 | 10 | −9.6 |
| $GPGIkPN(+1540.529)QTSk | N362 | Potential | Hex7HexNAc2 | 835.657 | 4 | −0.8 | ||
| Golgin subfamily A member 4 | GOGA4_HUMAN | $ELEhVN(+1735.630)^LSVk | N1612 | Novel | Hex3HexNAc4dHex1NeuAc1 | 848.139 | 4 | −3.9 |
| $kELEHVN(+1038.375)LSVk | N1612 | Novel | Hex3HexNAc2dHex1 | 752.648 | 4 | 0.8 | ||
| $kELEHVN(+1524.534)LSVk | N1612 | Novel | Hex6HexNAc2dHex1 | 874.186 | 4 | −1.6 | ||
| $SLQENkN(+1257.450)QSk | N585 | Novel | Hex4HexNAc3 | 777.391 | 4 | 10.2 | ||
| $TRILELESSLEkSLQENkN(+1216.423)QSk | N585 | Novel | Hex5HexNAc2 | 938.686 | 5 | −2.3 | ||
| Intercellular adhesion molecule 2 | ICAM2_HUMAN | $GSLEVN(+2028.741)^cSTTcNQPEVGGLETSLDk | N47 | Known | Hex5HexNAc6 | 1039.861 | 5 | 5.1 |
| $HYLVSN(+568.212)ISHDTVLQcHFTcSGk | N82 | Known | Hex1HexNAc2 | 905.687 | 4 | −0.6 | ||
| ICOS ligand | ICOSL_HUMAN | $IARTPSVNIGccIENVLLQQN(+2457.877)LTVGSQTGNDIGER | N225 | Potential | Hex8HexNAc5dHex1 | 724.219 | 9 | −5.4 |
| $IARTPSVNIGccIENVLLQQN(+2594.937)^LTVGSQTGNDIGER | N225 | Potential | Hex4HexNAc6dHex1NeuAc2 | 667.906 | 10 | 0.1 | ||
| Interleukin-6 receptor subunit beta | IL6RB_HUMAN | $SHLQN(+568.212)YTVNATkLTVNLTNDRYLATLTVRNLVGk | N379 | Known | Hex1HexNAc2 | 725.691 | 7 | 6.5 |
| $SHLQN(+568.212)YTVNATkLTVNLTNDRYLATLTVRNLVGk | N379 | Known | Hex1HexNAc2 | 1015.568 | 5 | 9.4 | ||
| $LTVN(+1460.529)LTNDRYLATLTVRNLVGk | N390 | Known | Hex4HexNAc4 | 877.848 | 5 | −8.4 | ||
| $QN(+568.212)cSQHESSPDIsHFER | N818 | Novel | Hex1HexNAc2 | 733.814 | 4 | 8.8 | ||
| Integrin alpha-3 | ITA3_HUMAN | $DVRkLLLSIN(+892.317)VTNTR | N656 | Potential | Hex3HexNAc2 | 1028.554 | 3 | −2.2 |
| $AHcVWLEcPIPDAPVVTN(+1362.481)VTVk | N926 | Potential | Hex5HexNAc2dHex1 | 864.223 | 5 | 8.3 | ||
| Integrin beta-1 | ITB1_HUMAN | $kNkNVTN(+1444.534)^RSk | N97 | Potential | Hex3HexNAc4dHex1 | 888.710 | 4 | 2.8 |
| Junctional adhesion molecule C | JAM3_HUMAN | $IWN(+1581.555)VTRRDSALYRcEVVARNDR | N104 | Potential | Hex6HexNAc3 | 1139.528 | 4 | −4.8 |
| $IWN(+892.318)^VTR | N104 | Potential | Hex3HexNAc2 | 643.303 | 3 | −0.7 | ||
| Lysosome-associated membrane glycoprotein 1 | LAMP1_HUMAN | $GHTLTLN(+1378.476)FTR | N103 | Known | Hex6HexNAc2 | 921.421 | 3 | −0.4 |
| $GHTLTLN(+1540.529)FTR | N103 | Known | Hex7HexNAc2 | 975.439 | 3 | −0.4 | ||
| $YNVSGTNGTcLLASMGLQLN(+2042.720)^LTYERkDNTTVTRLLnINPNk | N241 | Known | Hex4HexNAc4NeuAc2 | 734.661 | 10 | 6.4 | ||
| $kDN(+2059.735)TTVTR | N249 | Known | Hex5HexNAc4dHex1NeuAc1 | 861.641 | 4 | 1.0 | ||
| $kDN(+2204.772)TTVTR | N249 | Known | Hex5HexNAc4NeuAc2 | 898.156 | 4 | 5.8 | ||
| $kDN(+2350.830)TTVTR | N249 | Known | Hex5HexNAc4dHex1NeuAc2 | 934.414 | 4 | −0.2 | ||
| $kDN(+2569.905)TTVTR | N249 | Known | Hex6HexNAc5NeuAc2 | 989.440 | 4 | 6.2 | ||
| $kDN(+2570.925)TTVTR | N249 | Known | Hex6HexNAc5dHex2NeuAc1 | 989.690 | 4 | 1.0 | ||
| $kDN(+2571.946)TTVTR | N249 | Known | Hex6HexNAc5dHex4 | 989.690 | 4 | −2.4 | ||
| $kDN(+2715.963)^TTVTR | N249 | Known | Hex6HexNAc5dHex1NeuAc2 | 1031.190 | 4 | −2.0 | ||
| $kDN(+2715.963)TTVTR | N249 | Known | Hex6HexNAc5dHex1NeuAc2 | 1025.696 | 4 | −0.9 | ||
| $kDN(+2717.978)TTVTR | N249 | Known | Hex8HexNAc7 | 1025.949 | 4 | −1.2 | ||
| $kDN(+2861.000)TTVTR | N249 | Known | Hex6HexNAc5NeuAc3 | 1062.211 | 4 | 3.1 | ||
| $kDN(+2864.036)TTVTR | N249 | Known | Hex8HexNAc7dHex1 | 1062.466 | 4 | 1.2 | ||
| $kDN(+3007.058)TTVTR | N249 | Known | Hex6HexNAc5dHex1NeuAc3 | 1098.476 | 4 | 4.4 | ||
| $AAN(+1702.581)GSLR | N322 | Known | Hex8HexNAc2 | 872.711 | 3 | 2.6 | ||
| $AAN(+1864.634)GSLR | N322 | Known | Hex9HexNAc2 | 926.395 | 3 | 3.5 | ||
| Lysosome-associated membrane glycoprotein 2 | LAMP2_HUMAN | $EkPEAGTYSVNNGN(+1054.370)DTcLLATmGLQLNITQDk | N229 | Known | Hex4HexNAc2 | 1048.506 | 5 | 0.8 |
| Lysosomal alpha-glucosidase | LYAG_HUMAN | $QVVEN(+1095.397)^MTRAHFPLDVQWNDLDYMDSR | N390 | Known | Hex3HexNAc3 | 905.408 | 5 | 0.9 |
| $GAYTQVIFLARN(+1054.370)NTIVNELVR | N882 | Known | Hex4HexNAc2 | 918.718 | 4 | 1.4 | ||
| Protein-lysine 6-oxidase | LYOX_HUMAN | $AEN(+1054.370)QTAPGEVPALSNLR | N144 | Potential | Hex4HexNAc2 | 762.369 | 4 | 8.1 |
| $DPGAAVPGAANASAQQPRTPILLIRDN(+2432.884)R | N97 | Potential | Hex3HexNAc6dHex1NeuAc2 | 1106.316 | 5 | −5.0 | ||
| Lymphatic vessel endothelial hyaluronic acid receptor 1 | LYVE1_HUMAN | $AN(+1362.481)DSNPNEESkkTDk | N289 | Novel | Hex5HexNAc2dHex1 | 984.727 | 4 | 5.8 |
| Lysosomal alpha-mannosidase | MA2B1_HUMAN | $GFkDHFTFcQQLN(+892.317)ISIcPLSQTAARFQVIVYnPLGRk | N497 | Potential | Hex3HexNAc2 | 987.504 | 6 | −1.2 |
| Hepatocyte growth factor receptor | MET_HUMAN | $TLLRN(+1751.624)^SsGcEARRDEYR | N405 | Potential | Hex4HexNAc4NeuAc1 | 1040.946 | 4 | 6.4 |
| $VIVQPDQN(+2042.720)^^FTGLIAGVVSISTALLLLLGFFLWLkkRk | N930 | Potential | Hex4HexNAc4NeuAc2 | 886.225 | 8 | 3.5 | ||
| $VIVQPDQN(+2174.799)^FTGLIAGVVSISTALLLLLGFFLWLkkR | N930 | Potential | Hex5HexNAc6dHex1 | 1140.772 | 6 | 4.0 | ||
| Interstitial collagenase | MMP1_HUMAN | $AFQLWSN(+2034.703)^VTPLTFTk | N143 | Novel | Hex7HexNAc3NeuAc1 | 1065.992 | 4 | 3.1 |
| $AFQLWSN(+2059.735)VTPLTFTk | N143 | Novel | Hex5HexNAc4dHex1NeuAc1 | 1065.999 | 4 | 0.3 | ||
| $AFQLWSN(+2075.730)^VTPLTFTk | N143 | Novel | Hex6HexNAc4NeuAc1 | 1076.245 | 4 | 0.2 | ||
| $AFQLWSN(+2075.730)VTPLTFTk | N143 | Novel | Hex5HexNAc4dHex2NeuGc1 | 1070.250 | 4 | 1.4 | ||
| $AFQLWSN(+2076.750)VTPLTFTk | N143 | Novel | Hex6HexNAc4dHex2 | 1070.251 | 4 | −1.0 | ||
| $AFQLWSN(+2092.745)VTPLTFTk | N143 | Novel | Hex7HexNAc4dHex2 | 1074.251 | 4 | 0.2 | ||
| $AFQLWSN(+2350.830)^VTPLTFTk | N143 | Novel | Hex5HexNAc4dHex1NeuAc2 | 1144.769 | 4 | −0.3 | ||
| $AFQLWSN(+2350.830)VTPLTFTk | N143 | Novel | Hex5HexNAc4dHex1NeuAc2 | 1139.021 | 4 | −1.9 | ||
| $AFQLWSN(+2351.851)VTPLTFTk | N143 | Novel | Hex5HexNAc4dHex3NeuAc1 | 1139.277 | 4 | −0.9 | ||
| Multimerin-1 | MMRN1_HUMAN | $INALkkPTVN(+1200.428)LTTVLIGR | N1020 | Known | Hex4HexNAc2dHex1 | 957.031 | 4 | 1.5 |
| $GARLFVLLSSLWSGGIGLN(+1241.455)^NSk | N21 | Potential | Hex3HexNAc3dHex1 | 1001.006 | 4 | −3.0 | ||
| $LFVLLSSLWSGGIGLN(+568.212)NSk | N21 | Potential | Hex1HexNAc2 | 756.162 | 4 | −0.8 | ||
| $IDN(+1095.397)^ISLTVNDVRNTYSSLEGk | N344 | Known | Hex3HexNAc3 | 976.973 | 4 | −1.1 | ||
| $InN(+568.212)LTVSLEMEk | N576 | Potential | Hex1HexNAc2 | 803.746 | 3 | −1.8 | ||
| $kIEN(+892.317)LTSAVNSLNFIIk | N680 | Potential | Hex3HexNAc2 | 868.469 | 4 | −3.4 | ||
| $LNQSNFQkmYQMFN(+2262.815)^ETTSQVR | N828 | Potential | Hex5HexNAc5dHex1NeuAc1 | 1069.470 | 5 | 2.0 | ||
| $ALEAkSIHLSInFFSLN(+568.212)k | N921 | Potential | Hex1HexNAc2 | 819.195 | 4 | −5.2 | ||
| $SIHLSINFFSLN(+892.318)^k | N921 | Potential | Hex3HexNAc2 | 721.609 | 4 | −6.7 | ||
| C-type mannose receptor 2 | MRC2_HUMAN | $VTPAcN(+2059.735)TSLPAQR | N69 | Known | Hex5HexNAc4dHex1NeuAc1 | 925.654 | 4 | −2.1 |
| $SN(+2059.735)VTk | N954 | Potential | Hex5HexNAc4dHex1NeuAc1 | 1020.122 | 3 | 0.2 | ||
| Nck-associated protein 5 | NCKP5_HUMAN | $ERGPQGQGHGRMALNLQLSDTDDN(+2018.708)^ETFDELHIESSDEk | N585 | Novel | Hex6HexNAc3dHex1NeuAc1 | 1127.492 | 6 | −3.7 |
| Lysosomal protein NCU-G1 | NCUG1_HUMAN | $LLHTADTcQLEVALIGAsPRGN(+1241.454)R | N230 | Potential | Hex3HexNAc3dHex1 | 1010.223 | 4 | −1.4 |
| Natural cytotoxicity triggering receptor 3 ligand 1 | NR3L1_HUMAN | $LN(+2172.745)SSQEDPGTVYQcVVRHASLHTPLR | N216 | Potential | Hex10HexNAc2dHex1 | 1073.477 | 5 | −4.3 |
| Neuronal cell adhesion molecule | NRCAM_HUMAN | $IPAN(+2715.963)k | N1009 | Potential | Hex6HexNAc5dHex1NeuAc2 | 927.404 | 4 | −1.4 |
| $IPAN(+2960.068)^^k | N1009 | Potential | Hex5HexNAc7dHex1NeuAc2 | 1000.181 | 4 | 6.5 | ||
| $FN(+1378.476)HTQTIQQk | N223 | Potential | Hex6HexNAc2 | 768.611 | 4 | 0.9 | ||
| $FN(+1378.476)HTQTIQQk | N223 | Potential | Hex6HexNAc2 | 1024.480 | 3 | 2.2 | ||
| $SSRERPPTFLTPEGN(+1710.598)ASNk | N276 | Known | Hex5HexNAc3NeuAc1 | 1062.994 | 4 | −0.4 | ||
| Nuclear receptor-interacting protein 3 | NRIP3_HUMAN | $LMETN(+568.212)LSk | N72 | Novel | Hex1HexNAc2 | 651.345 | 3 | 8.7 |
| Plasminogen activator inhibitor 1 | PAI1_HUMAN | $GN(+1694.603)^MTRLPRLLVLPkFSLETEVDLRk | N288 | Known | Hex4HexNAc3dHex1NeuAc1 | 1063.953 | 5 | 3.4 |
| $GN(+2059.735)MTR | N288 | Known | Hex5HexNAc4dHex1NeuAc1 | 955.394 | 3 | 0.5 | ||
| $GN(+2350.830)^mTR | N288 | Known | Hex5HexNAc4dHex1NeuAc2 | 799.313 | 4 | −3.7 | ||
| $GN(+2351.851)^mTR | N288 | Known | Hex5HexNAc4dHex3NeuAc1 | 798.814 | 4 | −5.6 | ||
| Platelet-derived growth factor subunit B | PDGFB_HUMAN | $LLHGDPGEEDGAELDLN(+1864.634)mTRSHSGGELESLARGRR | N63 | Novel | Hex9HexNAc2 | 1176.921 | 5 | −5.5 |
| Secretory phospholipase A2 receptor | PLA2R_HUMAN | $MQDTSGHGVNTsDMYPMPNTLEYGN(+2204.772)^^RTYk | N1123 | Novel | Hex5HexNAc4NeuAc2 | 1014.913 | 6 | −1.9 |
| Procollagen-lysine,2-oxoglutarate 5-dioxygenase 2 | PLOD2_HUMAN | $YFN(+1856.656)YTVkVLGQGEEWR | N63 | Potential | Hex5HexNAc3dHex1NeuAc1 | 1074.744 | 4 | 0.2 |
| Phospholipid transfer protein | PLTP_HUMAN | $qLLYWFFYDGGYIN(+2157.783)^ASAEGVSIRTGLELSR | N117 | Potential | Hex4HexNAc6NeuAc1 | 1118.898 | 5 | 0.2 |
| $IYSN(+1403.507)HSALESLALIPLQAPLk | N398 | Known | Hex4HexNAc3dHex1 | 1033.782 | 4 | 1.5 | ||
| Plexin-C1 | PLXC1_HUMAN | $IAN(+1524.534)^FTSDVEYSDDHchLILPDSEAFQDVQGkRHR | N1308 | Novel | Hex6HexNAc2dHex1 | 1203.342 | 5 | −1.8 |
| $IAN(+2204.773)FTSDVEYSDDhcHLILPDSEAFQDVQGkR | N1308 | Novel | Hex5HexNAc4NeuAc2 | 638.689 | 10 | 5.9 | ||
| $VILGEN(+1298.476)LTSNcPEVIYEIk | N407 | Known | Hex3HexNAc4 | 985.483 | 4 | −1.8 | ||
| $ELcQN(+1378.476)k | N548 | Potential | Hex6HexNAc2 | 873.730 | 3 | 8.0 | ||
| $DVcIQFDGGNcSSVGSLSYIALPHcSLIFPATTWISGGQN(+1403.508)^ITMMGR | N771 | Potential | Hex4HexNAc3dHex1 | 607.279 | 11 | 0.5 | ||
| $ENDNFN(+1054.370)ISk | N871 | Novel | Hex4HexNAc2 | 861.725 | 3 | −4.1 | ||
| $ENDNFN(+2522.916)^ISk | N871 | Novel | Hex5HexNAc7NeuAc1 | 1019.438 | 4 | 7.5 | ||
| Tyrosine-protein kinase-like 7 | PTK7_HUMAN | $SAN(+3007.058)ASFNIk | N116 | Known | Hex6HexNAc5dHex1NeuAc3 | 1102.722 | 4 | 2.1 |
| $SAN(+3026.090)ASFNIk | N116 | Known | Hex9HexNAc7dHex1 | 1107.479 | 4 | 1.3 | ||
| $ATVFAN(+1710.598)GSLLLTQVRPR | N283 | Known | Hex5HexNAc3NeuAc1 | 945.458 | 4 | 0.0 | ||
| $RQDVN(+1686.586)ITVATVPSWLk | N405 | Potential | Hex7HexNAc2dHex1 | 991.485 | 4 | 3.2 | ||
| Pentraxin-related protein PTX3 | PTX3_HUMAN | $ATDVLN(+1419.502)kTILFsYGTk | N220 | Potential | Hex5HexNAc3 | 986.474 | 4 | −4.0 |
| Proactivator polypeptide | SAP_HUMAN | $TN(+1038.375)STFVQALVEHVk | N215 | Known | Hex3HexNAc2dHex1 | 1020.847 | 3 | 1.2 |
| $LIDNN(+1216.423)kTEk | N332 | Known | Hex5HexNAc2 | 988.825 | 3 | −0.7 | ||
| $LIDNN(+1378.476)kTEk | N332 | Known | Hex6HexNAc2 | 1042.510 | 3 | 0.7 | ||
| $LIDNN(+1694.603)kTEk | N332 | Known | Hex4HexNAc3dHex1NeuAc1 | 861.415 | 4 | −0.8 | ||
| $N(+2245.800)^STk | N426 | Known | Hex4HexNAc5NeuAc2 | 1056.113 | 3 | −0.8 | ||
| $NLEkN(+1378.476)STk | N426 | Known | Hex6HexNAc2 | 995.485 | 3 | 2.5 | ||
| Histone deacetylase complex subunit SAP30 | SAP30_HUMAN | $N(+1095.397)kSDLk | N209 | Novel | Hex3HexNAc3 | 824.751 | 3 | −3.7 |
| $GGDAAAAVAAVVAAAAAAAsAGN(+1589.571)GTGAGTGAEVPGAGAVSAAGPPGAAGPGPGQLccLR | N34 | Novel | Hex3HexNAc4NeuAc1 | 682.815 | 10 | −0.7 | ||
| Serpin H1 | SERPH_HUMAN | $SLSN(+1378.476)^STAR | N120 | Potential | Hex6HexNAc2 | 821.017 | 3 | −1.4 |
| $SLSN(+1378.476)STAR | N120 | Potential | Hex6HexNAc2 | 813.358 | 3 | 1.9 | ||
| $SLSN(+1540.529)STAR | N120 | Potential | Hex7HexNAc2 | 867.371 | 3 | −3.6 | ||
| $SLSN(+1702.581)STAR | N120 | Potential | Hex8HexNAc2 | 922.062 | 3 | 1.6 | ||
| $SLSN(+1864.634)STAR | N120 | Potential | Hex9HexNAc2 | 975.414 | 3 | 4.5 | ||
| $N(+1540.529)VTWk | N125 | Potential | Hex7HexNAc2 | 879.399 | 3 | −0.5 | ||
| $N(+1702.581)VTWk | N125 | Potential | Hex8HexNAc2 | 933.418 | 3 | 0.6 | ||
| Tyrosine-protein phosphatase non-receptor type substrate 1 | SHPS1_HUMAN | $LQLTWLEnGN(+1200.428)VSR | N292 | Known | Hex4HexNAc2dHex1 | 739.847 | 4 | −3.5 |
| $LQLTWLENGN(+1872.651)^VSR | N292 | Known | Hex6HexNAc3NeuAc1 | 912.893 | 4 | −11.5 | ||
| SPARC | SPRC_HUMAN | $VcSNDN(+1622.582)k | N116 | Known | Hex5HexNAc4 | 970.425 | 3 | 3.5 |
| $VcSNDN(+1767.619)k | N116 | Known | Hex4HexNAc4NeuGc1 | 1019.107 | 3 | 4.9 | ||
| $VcSNDN(+1768.640)k | N116 | Known | Hex5HexNAc4dHex1 | 1018.439 | 3 | 0.0 | ||
| $VcSNDN(+1864.634)k | N116 | Known | Hex9HexNAc2 | 1051.108 | 3 | 2.2 | ||
| $VcSNDN(+1913.677)^k | N116 | Known | Hex5HexNAc4NeuAc1 | 806.336 | 4 | −1.4 | ||
| $VcSNDN(+1913.677)k | N116 | Known | Hex5HexNAc4NeuAc1 | 800.591 | 4 | −0.3 | ||
| $VcSNDN(+1913.677)k | N116 | Known | Hex5HexNAc4NeuAc1 | 1066.783 | 3 | −1.7 | ||
| $VcSNDN(+1914.697)k | N116 | Known | Hex5HexNAc4dHex2 | 800.844 | 4 | −2.5 | ||
| $VcSNDN(+1914.697)k | N116 | Known | Hex5HexNAc4dHex2 | 1067.125 | 3 | 0.6 | ||
| $VcSNDN(+2059.735)^k | N116 | Known | Hex5HexNAc4dHex1NeuAc1 | 842.598 | 4 | −3.0 | ||
| $VcSNDN(+2059.735)k | N116 | Known | Hex5HexNAc4dHex1NeuAc1 | 836.857 | 4 | 3.1 | ||
| $VcSNDN(+2059.735)k | N116 | Known | Hex5HexNAc4dHex1NeuAc1 | 1115.472 | 3 | 1.6 | ||
| $VcSNDN(+2350.830)k | N116 | Known | Hex5HexNAc4dHex1NeuAc2 | 909.629 | 4 | 0.3 | ||
| Stabilin-1 | STAB1_HUMAN | $ELLQHHGLVPQIEAATAYTIFVPTnRSLEAQGN(+2366.825)^SSHLDADTVR | N1178 | Potential | Hex6HexNAc4NeuAc2 | 732.443 | 10 | 3.2 |
| $ELkGDGPFTIFVPHADLMSN(+568.212)LSQDELARIR | N1626 | Known | Hex1HexNAc2 | 1097.310 | 4 | −5.4 | ||
| $LLPAHsGLSLIISDAGPDN(+892.317)SSWAPVAPGTVVVSR | N2424 | Known | Hex3HexNAc2 | 655.464 | 7 | −7.8 | ||
| $ILTMAnQVLAVN(+1095.397)ISEEGR | N606 | Potential | Hex3HexNAc3 | 820.390 | 4 | −9.4 | ||
| $GN(+1216.423)cSDGIQGNGAcLcFPDYk | N745 | Potential | Hex5HexNAc2 | 975.155 | 4 | −10.2 | ||
| Suppressor of G2 allele of SKP1 homolog | SUGT1_HUMAN | $RAMN(+568.212)kSFMESGGTVLSTNWsDVGk | N329 | Novel | Hex1HexNAc2 | 981.480 | 4 | 6.2 |
| Angiopoietin-1 receptor | TIE2_HUMAN | $ISN(+568.212)ITHSSAVIsWTILDGYSISSITIR | N649 | Potential | Hex1HexNAc2 | 762.382 | 5 | −3.1 |
| Tetraspanin-3 | TSN3_HUMAN | $TYN(+1825.661)GTNPDAASRAIDYVQR | N127 | Potential | Hex5HexNAc5 | 1041.454 | 4 | −7.9 |
| Thrombospondin-1 | TSP1_HUMAN | $VVN(+1913.677)STTGPGEHLR | N1067 | Known | Hex5HexNAc4NeuAc1 | 877.141 | 4 | 0.2 |
| Vascular endothelial growth factor receptor 1 | VGFR1_HUMAN | $RIIWDSRkGFIISN(+892.317)ATYk | N196 | Potential | Hex3HexNAc2 | 934.508 | 4 | 8.1 |
| $SVN(+892.318)TSVhIYDkAFITVk | N323 | Potential | Hex3HexNAc2 | 876.466 | 4 | 10.2 | ||
| $WFWHPcNHN(+1419.502)HSEARcDFcSNNEESFILDADSnMGNR | N474 | Potential | Hex5HexNAc3 | 1017.415 | 6 | 1.3 | ||
| $mAITkEhSITLNLTIMN(+1257.450)^VSLQDSGTYAcRAR | N625 | Potential | Hex4HexNAc3 | 1051.702 | 5 | −0.2 | ||
| von Willebrand factor | VWF_HUMAN | $YFN(+2059.735)^k | N156 | Potential | Hex5HexNAc4dHex1NeuAc1 | 776.330 | 4 | −4.7 |
| $YFN(+2059.735)^k | N156 | Potential | Hex5HexNAc4dHex1NeuAc1 | 1034.440 | 3 | −1.2 | ||
| $YFN(+2059.735)k | N156 | Potential | Hex5HexNAc4dHex1NeuAc1 | 1027.114 | 3 | −0.1 | ||
| $YFN(+2059.736)k | N156 | Potential | Hex5HexNAc4dHex1NeuAc1 | 770.587 | 4 | −0.6 | ||
| $YFN(+2204.772)k | N156 | Potential | Hex5HexNAc4NeuAc2 | 1075.798 | 3 | 3.0 | ||
| $YFN(+2221.789)k | N156 | Potential | Hex6HexNAc4dHex1NeuAc1 | 1081.467 | 3 | 0.8 | ||
| $YFN(+2350.830)^^k | N156 | Potential | Hex5HexNAc4dHex1NeuAc2 | 854.851 | 4 | −3.2 | ||
| $YFN(+2350.830)^k | N156 | Potential | Hex5HexNAc4dHex1NeuAc2 | 848.856 | 4 | −0.9 | ||
| $YFN(+2350.830)^k | N156 | Potential | Hex5HexNAc4dHex1NeuAc2 | 1131.473 | 3 | −0.2 | ||
| $YFN(+2350.830)k | N156 | Potential | Hex5HexNAc4dHex1NeuAc2 | 843.362 | 4 | 0.5 | ||
| $YFN(+2350.830)k | N156 | Potential | Hex5HexNAc4dHex1NeuAc2 | 1124.145 | 3 | −0.7 | ||
| $YFN(+2350.831)^k | N156 | Potential | Hex5HexNAc4dHex1NeuAc2 | 848.857 | 4 | 0.0 | ||
| $YFN(+2350.831)k | N156 | Potential | Hex5HexNAc4dHex1NeuAc2 | 843.362 | 4 | 1.2 | ||
| $YFN(+2351.851)^k | N156 | Potential | Hex5HexNAc4dHex3NeuAc1 | 849.613 | 4 | −0.2 | ||
| $YFN(+2366.825)^k | N156 | Potential | Hex5HexNAc4dHex1NeuAc1NeuGc1 | 853.098 | 4 | −9.4 | ||
| $YFN(+2366.825)k | N156 | Potential | Hex5HexNAc4dHex1NeuAc1NeuGc1 | 1129.813 | 3 | 0.5 | ||
| $YFN(+2715.963)^k | N156 | Potential | Hex6HexNAc5dHex1NeuAc2 | 940.387 | 4 | −3.7 | ||
| $YFN(+2715.963)k | N156 | Potential | Hex6HexNAc5dHex1NeuAc2 | 934.643 | 4 | −0.8 | ||
| $YFN(+2716.983)k | N156 | Potential | Hex6HexNAc5dHex3NeuAc1 | 934.898 | 4 | −1.5 | ||
| $YFN(+2861.001)k | N156 | Potential | Hex6HexNAc5NeuAc3 | 971.409 | 4 | 3.7 | ||
| $YFN(+3007.058)k | N156 | Potential | Hex6HexNAc5dHex1NeuAc3 | 1007.667 | 4 | −2.2 | ||
| $YFN(+3009.074)k | N156 | Potential | Hex8HexNAc7 | 1007.923 | 4 | 0.6 | ||
| $YFN(+3026.089)k | N156 | Potential | Hex9HexNAc7dHex1 | 1012.676 | 4 | −2.0 | ||
| $YFN(+3372.190)k | N156 | Potential | Hex7HexNAc6dHex1NeuAc3 | 1098.951 | 4 | −0.9 | ||
| $GDILQRVREIRYQGGN(+1378.476)R | N1574 | Known | Hex6HexNAc2 | 909.439 | 4 | −0.2 | ||
| $GDILQRVREIRYQGGN(+568.212)RTNTGLALR | N1574 | Known | Hex1HexNAc2 | 912.984 | 4 | −4.0 | ||
| $YQGGN(+2059.735)R | N1574 | Known | Hex5HexNAc4dHex1NeuAc1 | 993.411 | 3 | 1.2 | ||
| $ASPPSSScN(+2059.735)ISSGEmQk | N211 | Potential | Hex5HexNAc4dHex1NeuAc1 | 1073.955 | 4 | −2.4 | ||
| $ASPPSSScN(+2350.830)^ISSGEmQk | N211 | Potential | Hex5HexNAc4dHex1NeuAc2 | 1151.736 | 4 | 8.9 | ||
| $ASPPSSScN(+2350.830)ISSGEmQk | N211 | Potential | Hex5HexNAc4dHex1NeuAc2 | 1146.231 | 4 | 0.3 | ||
| $N(+2059.735)VScPQLEVPVcPSGFQLSck | N2546 | Known | Hex5HexNAc4dHex1NeuAc1 | 984.037 | 5 | −3.6 | ||
| $GQVYLQcGTPcN(+2059.735)LTcR | N666 | Potential | Hex5HexNAc4dHex1NeuAc1 | 1053.444 | 4 | 0.9 | ||
| $GQVYLQcGTPcN(+2075.730)LTcR | N666 | Potential | Hex6HexNAc4NeuAc1 | 1057.695 | 4 | 2.2 | ||
| $GQVYLQcGTPcN(+2076.750)LTcR | N666 | Potential | Hex6HexNAc4dHex2 | 1058.193 | 4 | −0.6 | ||
| $GQVYLQcGTPcN(+2350.830)LTcR | N666 | Potential | Hex5HexNAc4dHex1NeuAc2 | 1126.222 | 4 | 4.5 | ||
| Bovine proteins | ||||||||
| Alpha-1-acid glycoprotein | A1AG_BOVIN | $QN(+2861.000)^GTLSk | N104 | Potential | Hex6HexNAc5NeuAc3 | 1020.428 | 4 | 0.4 |
| $QN(+2861.000)GTLSk | N104 | Potential | Hex6HexNAc5NeuAc3 | 1014.934 | 4 | 1.8 | ||
| $QN(+2880.031)GTLSk | N104 | Potential | Hex9HexNAc7 | 1019.690 | 4 | 0.5 | ||
| $NPEYN(+2814.010)^^k | N57 | Potential | Hex5HexNAc7NeuAc2 | 1018.923 | 4 | 6.2 | ||
| Alpha-fetoprotein | FETA_BOVIN | $AEN(+1200.428)ATEcFETk | N197 | Potential | Hex4HexNAc2dHex1 | 983.428 | 3 | −5.8 |
| $AEN(+2204.772)ATEcFETk | N197 | Potential | Hex5HexNAc4NeuAc2 | 988.913 | 4 | 0.1 | ||
| $AN(+2204.772)FTEIQk | N251 | Potential | Hex5HexNAc4NeuAc2 | 901.651 | 4 | 2.5 | ||
| Alpha-2-HS-glycoprotein | FETUA_BOVIN | $kLcPDcPLLAPLN(+2204.772)DSR | N156 | Known | Hex5HexNAc4NeuAc2 | 905.409 | 5 | 0.3 |
| $kLcPDcPLLAPLN(+2861.000)DSR | N156 | Known | Hex6HexNAc5NeuAc3 | 1036.654 | 5 | −1.0 | ||
| $LcPDcPLLAPLN(+2204.772)DSR | N156 | Known | Hex5HexNAc4NeuAc2 | 1043.195 | 4 | −1.7 |
Peptide modification symbols: $, N-terminal TMT0 labelling (+224.152); ^, Na adduct on glycan (+21.982); ^^, 2 Na adduct on glycan (+43.964); c, carbamidomethylation of cysteine (+57.021); h, oxidation of histidine (+15.995); k, TMT0 labeling of lysine (+224.152); m, oxidation of methionine (+15.995); n, deamidation of asparagine (+0.984); q, Gln->pyro-Glu and loss of TMT0 (−241.179); s, phosphorylation of serine (+79.966).
Underlined glycopeptides were also detected via the PNGase F + H218O method (supplemental Table S4).
Fig. 3.
HCD-pd-ETD fragmentation. Full MS showing the different glycoforms of the same peptide sequence (A). Characteristic oxonium ion detected by HCD at m/z = 204.09 (B). This HexNAc signature triggered an ETD scan to identify the peptide sequence and confirm the glycosylation site (C).
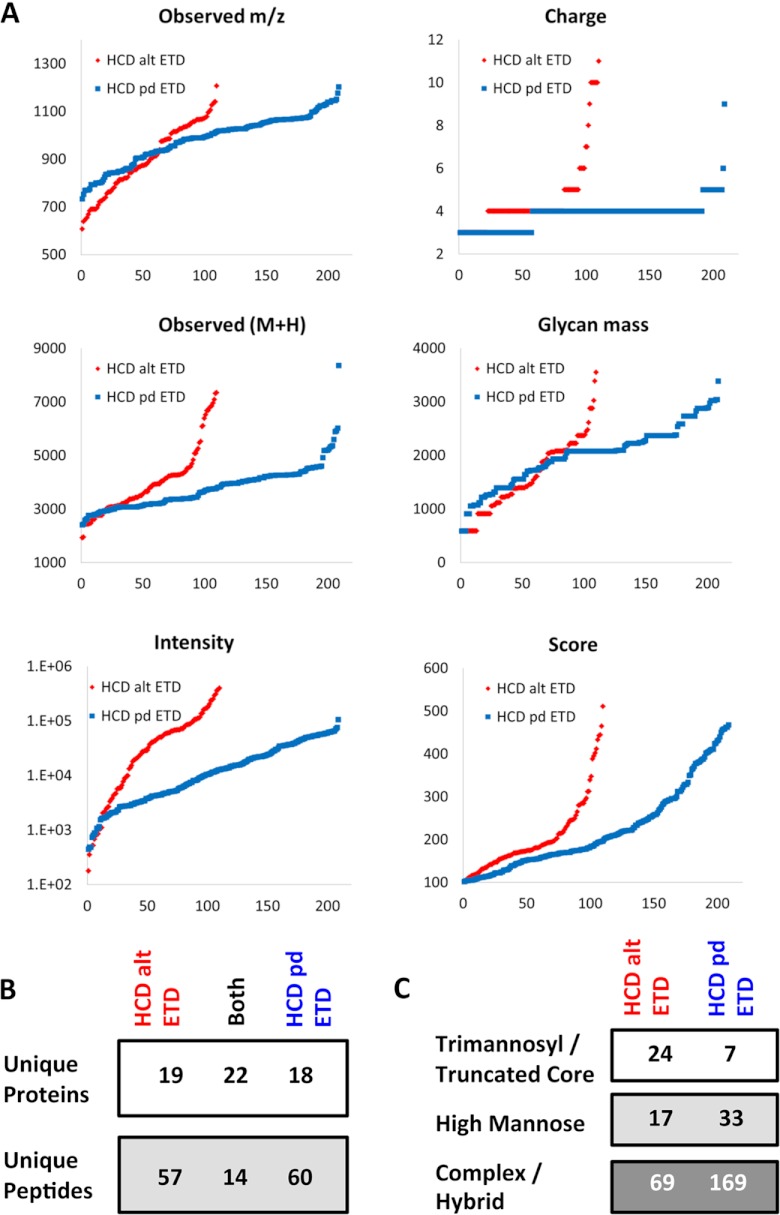
For the same samples, HCD-pd-ETD revealed 28 known, 25 potential, and 16 novel glycosylation sites based on 209 identified spectra; HCD-alt-ETD revealed 20 known, 32 potential, and 14 novel glycosylation sites from 110 identified spectra. The HCD-alt-ETD method selected mostly precursors with higher intensities, higher charge, and smaller m/z (Fig. 4A). Several large glycopeptides were detected via only HCD-alt-ETD, and more low-abundant glycopeptides were detected via HCD-pd-ETD. There was limited overlap in the identified glycopeptides but better agreement in the protein level (Fig. 4B). Among the 319 total glycopeptides identified in the conditioned media, 31 were attached with a trimannosyl core (-HexNAc2Hex3) or truncated core (-HexNAc2Hex), 50 with high mannose (-HexNAc2Hex4–9), and 238 with complex/hybrid glycans. Notably, HCD-pd-ETD detected almost twice as many complex/hybrid glycoforms as HCD-alt-ETD (Fig. 4C).
Fig. 4.
Comparison of HCD-pd-ETD and HCD-alt-ETD. The two methods, HCD-pd-ETD (blue) and HCD-alt-ETD (red), displayed distinct distributions of the observed m/z, charge state, mass of identified peptides (M+H), and glycan mass, as well as the intensity of the precursor ions and the ByonicsTM score (all y-axes). The x-axes represent index numbers after proteins were sorted by their corresponding y-axis value from lower to higher (A). There was limited overlap in the identified glycopeptides (B). C, the HCD-pd-ETD method preferentially identified complex/hybrid glycans.
Validation of Glycoproteins
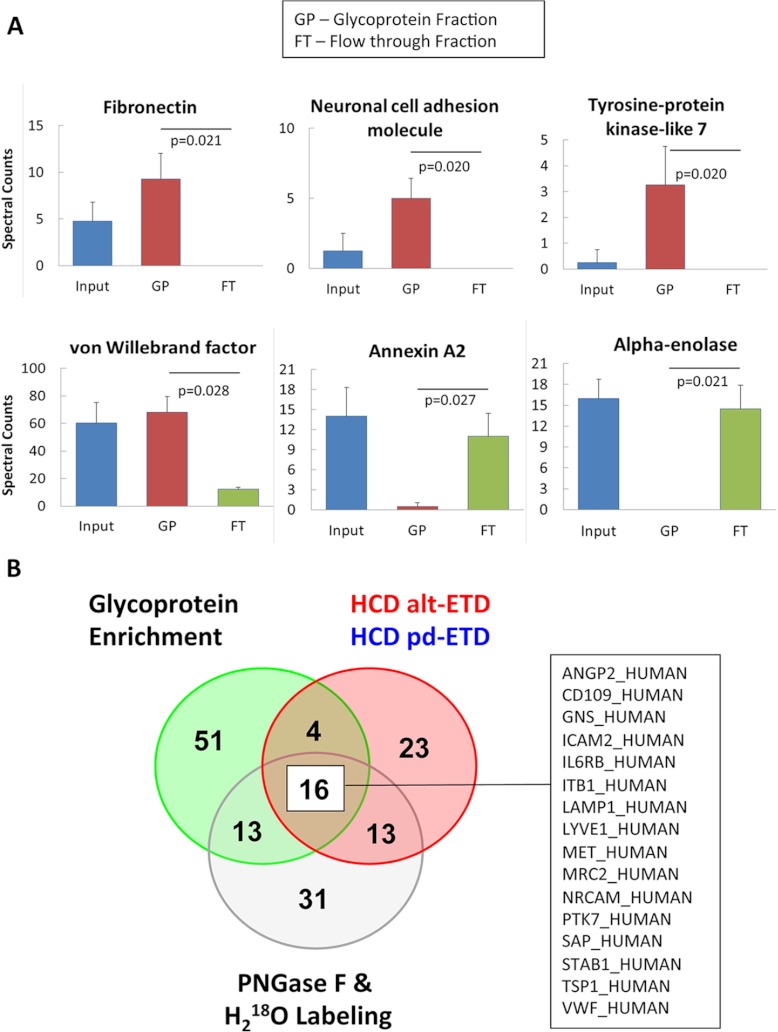
To validate the glycosylation status, we performed additional analysis before and after glycoprotein enrichment with affinity resins of ConA lectin (n = 4) using a Q Exactive MS (Thermo Scientific). We then compared the number of identified spectra in the glycoprotein-enriched fraction, the flow-through, and the input (supplemental Table S5). For most glycoproteins, a higher spectral count was observed in the glycoprotein-enriched fraction than in the original input and/or the flow-through. Representative examples (fibronectin, neuronal cell adhesion molecule, tyrosine-protein-kinase-like 7, and vWF) are shown in Fig. 5A. Non-glycosylated proteins, such as annexin A2 and alpha-enolase, were more abundant in the flow-through. Glycoproteins identified in all three methods are highlighted in Fig. 5B.
Fig. 5.
Glycoprotein enrichment for validation. A, spectral count of input, glycoprotein-enriched fraction (GP), and flow-through fraction (FT) from representative glycoproteins and non-glycoproteins. B, complementarity of the different methods (HCD-ETD, PNGase F + H218O treatment, and glycoprotein enrichment). Only 18 glycoproteins were consistently identified.
Confirmation of Predicted Glycosylation Sites
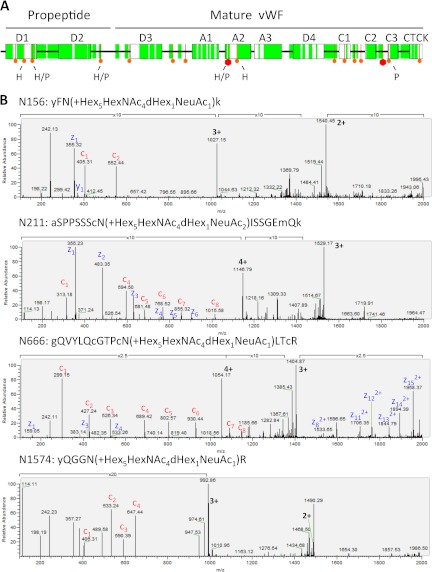
The hemostatic protein vWF is the main protein stored within Weibel-Palade bodies (27). After secretagogue stimulation, Weibel-Palade bodies undergo exocytosis, releasing vWF filaments. vWF is one of the few known proteins containing the ABO blood group signature, which is formed by different glycans. Although the released glycan composition of this protein has been investigated extensively (28, 29), experimental evidence for many putative glycosylation sites is still missing. The coverage obtained for vWF in our proteomics analysis is shown in Fig. 6A. The precursor protein consists of homologous units such as the VWF type A, C, and D domains and a C-terminal cystine know (CTCK). The vWF propeptide (D1-D2, AA 23–763) is separated from the remaining domains of mature vWF (AA 764–2813) via furin-mediated proteolytic cleavage. We confirmed 6 N-glycosylation sites. Notably, three N-glycosylation sites were located within the propeptide (AA 23–763). Examples of ETD spectra are shown in Fig. 6B.
Fig. 6.
Sequence coverage for vWF. A, schematic illustration of vWF sequence. Coverage is highlighted in green, and potential glycosylation sites are shown in red. A large hexagon indicates a glycosylation site with a reference in the Uniprot database. By using the HCD-ETD (H) or PNGase F (P) method, we confirmed six N-glycosylation sites on vWF. B, ETD spectra of glycopeptides identified via HCD-ETD (N156, N211, N666, N1574). The following abbreviations are used: a, y, g, k = TMT modified Ala, Tyr, Gly, and Lys, respectively; c = carboxyamidomethylation of Cys; m = oxidation of Met.
DISCUSSION
This study represents a significant advance over the existing proteomics literature on ECs. Unlike other cell types, ECs do not tolerate prolonged serum starvation, and their susceptibility to cell death upon serum withdrawal poses a major challenge for proteomic workflows targeting their secretome. We performed secretome analysis after 45 min of PMA stimulation combined with enrichment strategies for glycoproteins and glycopeptides. Glycopeptides were analyzed via three complementary MS techniques: the detection of 18O asparagine deamidation after digestion with PNGase F in H218O, HCD-alt-ETD, and HCD-pd-ETD using an Orbitrap Elite MS.
The Endothelial Secretome
The secretagogue PMA minimized EC death by allowing a shorter incubation period under serum-free conditions while increasing coverage in the proteomic analysis by inducing the exocytosis of intracellular storage vesicles (14) such as Weibel-Palade bodies. These unique storage vesicles in ECs play a major role in hemostasis and cell-to-cell communication. Using this approach, many more proteins were identified than in any previous proteomics study on ECs, including known endothelial surface markers such as endoglin (CD105), integrin beta-1 (CD29), tyrosine-protein kinase receptor Tie-1, and junctional adhesion molecule A; secreted growth factors (i.e. C-type lectin domain family 11 member A); co-receptors (i.e. neuropilin-1 (co-receptor for VEGF-A)); proteases(i.e. furin); and inflammatory mediators (i.e. macrophage migration inhibitory factor), to name just a few. Short-term PMA treatment does not release microparticles (30), as shedding events make it difficult to discern intracellular from secreted/membrane proteins. In a direct comparison of the cellular proteome and the secretome utilizing difference gel electrophoresis, 70 out of 96 proteins analyzed were present in both samples, representing <10% of the visible protein spots in the secretome.
Biological Importance of Glycosylation
Glycosylation is key for the stability and solubility of secreted and membrane proteins. It is the most complex post-translational modification (31) and mediates extracellular matrix network assembly, cell–cell interactions, and cell–matrix interactions. Unlike polynucleotides and polypeptides, which have a linear structure, sugars tend to be arranged in branched polymers, resulting in an exponential increase of possible polysaccharide combinations. Theoretically, just six monosaccharides can give rise to 1012 different glycan structures. This high diversity of protein-bound glycans requires a combination of different techniques. For example, new MS-based methods were developed to profile the cell surface N-glycoproteome as a differentiation marker for stem cells (32). We applied a combination of different glycoproteomics techniques to further enrich for secreted and shed membrane proteins and reveal potential glycosylation sites within the endothelial secretome. Glycoproteins play important roles in many biological processes related to ECs, such as angiogenesis, in which the structural change of the glycans will determine the attachment property of cells and influence cell-to-cell interactions (33). Interestingly, vWF is a glycoprotein produced uniquely by ECs and megakaryocytes. Previous publications investigating vWF isolated from plasma failed to identify glycosylation sites within the propeptide (29). In plasma, the concentration of the propeptide is about one-tenth of the concentration of mature vWF (34, 35). In the conditioned medium of ECs, however, we observed several glycopeptides of the propeptide. Thus, the endothelial secretome allowed us to interrogate the glycosylation sites of von Willebrand antigen 2, the N-terminal cleavage product of vWF that aids N-terminal multimerization and protein compartmentalization of mature vWF in storage granules.
Conventional Methods for Glycoproteomics
As reviewed elsewhere (36), conventional glycoproteomic methods involve the enrichment of glycoproteins (typically with lectins like ConA and wheat germ agglutinin), cleavage of the glycans, and identification of the remaining peptide sequence. The most widely used method for detecting N-glycopeptides is digestion by PNGase F. PNGase F cleaves the GlcNAc molecule closest to the peptide (37). After PNGase F treatment, formerly N-linked glycosylated peptides are identified based on the conversion of Asn to Asp (deamidation) in the consensus motif for N-linked glycosylation (sequence N-X(not P)-S/T). This method has two major caveats. The first of these is a high false positive rate due to spontaneous deamidation. Asn-Gly sites, in particular, are prone to spontaneous deamidation (38–40). To reduce false positives, PNGase F treatment is performed in 18O water, adding a larger tag of 2.99 Da. Importantly, all known glycosyltransferases that mediate N-linked glycosylation are supposed to recognize a consensus motif, and this consensus sequence for N-linked glycosylation must be taken into consideration (41). 2) The second caveat is that after PNGase F cleavage, the released sugars can be analyzed separately, but the link to the identified peptides with deamidated amino acids is lost (42, 43). Ideally, intact glycopeptides are analyzed directly via MS/MS even in complex biological samples.
Novel HCD-ETD Method
HCD fragmentation mostly breaks glycosidic bonds, whereas ETD preserves the glycan attachment and fragments the peptide backbone, providing more complete peptide sequence information. Current MS/MS acquisition strategies for glycopeptide analysis rely on the acquisition of MS/MS spectra for all precursor ions. In this study, HCD was employed to generate glycan oxonium ions and trigger an ETD spectrum in a data-dependent manner. HCD presents the sugar signatures within the low m/z range, which are otherwise lost as a result of the one-third rule of ion trap fragmentation (44). Glycopeptides with terminal HexNAc generate typically an m/z 204.0864 oxonium ion and its fragments at m/z 168.0653 and 138.0550. The oxonium ion and its fragments are measured with the high mass accuracy of the Orbitrap analyzer, and the unambiguous identification of the glycan oxonium ion generated by the HCD scan serves as a diagnostic marker for glycopeptides. This approach was compared against conventional HCD-alt-ETD scans using a complex biological sample. The HCD-alt-ETD preferentially detects higher charged and higher intensity precursor ions than HCD-pd-ETD. This might be because (i) a higher charge increases ETD fragmentation efficiency, resulting in more identified glycopeptides; (ii) high-charged precursors did not produce HCD spectra of sufficient quality to trigger ETD based on the diagnostic oxonium ions; or (iii) more abundant peptides were selected in HCD-alt-ETD because the instrument duty cycle is less efficient than in HCD-pd-ETD. Overall, the combination of multiple MS methods used in our study provides greater confidence in the identification of glycopeptides than studies relying on a single approach and offers complementary advantages in the assessment of the glycoproteome, notably, the simultaneous identification of the peptide sequence, the glycosylation site, and the glycan composition.
Study Limitations
N-linked and O-linked glycosylation are the two most common forms of glycosylation in mammals (45). Only N-linked glycosylation was analyzed in the present study. Unlike N-linked glycosylation, O-linked glycosylation has no consensus site (46). This makes the analysis of O-linked glycopeptides a more daunting task (47). Lectins are widely used for glycoprotein enrichment. There are many types of lectins binding to different sugars, such as ConA (binds to α-d-mannosyl and α-d-glucosyl residues) and wheat germ agglutinin (binds to GlcNAcβ1–4GlcNAcβ1–4GlcNAc- and N-acetylneuraminic acid). Here we used only ConA as a proof of principle to demonstrate the complementary results of multiple glycoprotein identification methods. ConA is known to display nonspecific avidity for hydrophobic ligands such as certain domains of tropomyosin (48). Furthermore, the standard protocol for the ConA glycoprotein enrichment kit is not optimized for cleanliness, and several known non-glycoproteins were also detected in the eluate samples. Sequential washes with low- and high-ionic-strength buffers before elution might have reduced this contamination (49). Also, mixing different lectins would increase the coverage of the glycoproteome in biological samples (39). Additional efforts are needed for a complete structural characterization of protein glycosylation; in particular, the quantitation of the occupancy rates and the identification of the glycan structure as complex/hybrid glycans cannot be discerned via our current MS approach.
CONCLUSIONS
Cardiovascular diseases arise from exposure to risk factors that induce complex pathophysiological perturbations of endothelial protein secretion. The recent advent of new proteomic technologies has enabled us to obtain information on the dynamic regulation of endothelial protein secretion. We present results from an extensive glycoproteomic analysis with information on glycan composition obtained via a direct MS method. Future proteomics studies linking endothelial secretory processes to cardiovascular risk factors and endothelial dysfunction will provide valuable insights about the mechanisms contributing to cardiovascular disease.
Supplementary Material
Acknowledgments
We thank Dr. Sarah Langley for assistance with the gene ontology annotation.
Footnotes
* This work was funded by the Department of Health via a National Institute for Health Research (NIHR) Biomedical Research Centre award to Guy's and St. Thomas' NHS Foundation Trust in partnership with King's College London and King's College Hospital NHS Foundation Trust. Dr. M. Mayr is supported by a Senior Research Fellowship of the British Heart Foundation.
 This article contains supplemental material.
This article contains supplemental material.
1 The abbreviations used are:
- ConA
- concanavalin A
- EC
- endothelial cell
- ETD
- electron-transfer dissociation
- GlcNAc
- N-acetylglucosamine
- HCD
- higher-energy C-trap dissociation
- Hex
- hexose
- HexNAc
- N-acetylhexosamine
- HUVEC
- human umbilical vein endothelial cell
- PMA
- phorbol-12-myristate-13-acetate
- PNGase F
- peptide: N-glycosidase F
- TMT0
- Tandem Mass Tag Zero
- vWF
- von Willebrand factor.
REFERENCES
- 1. Mayr M., Mayr U., Chung Y. L., Yin X., Griffiths J. R., Xu Q. (2004) Vascular proteomics: linking proteomic and metabolomic changes. Proteomics 4, 3751–3761 [DOI] [PubMed] [Google Scholar]
- 2. Pula G., Perera S., Prokopi M., Sidibe A., Boulanger C. M., Mayr M. (2008) Proteomic analysis of secretory proteins and vesicles in vascular research. Proteomics Clin. Appl. 2, 882–891 [DOI] [PubMed] [Google Scholar]
- 3. Pober J. S., Sessa W. C. (2007) Evolving functions of endothelial cells in inflammation. Nat. Rev. Immunol. 7, 803–815 [DOI] [PubMed] [Google Scholar]
- 4. Cai H., Harrison D. G. (2000) Endothelial dysfunction in cardiovascular diseases: the role of oxidant stress. Circ. Res. 87, 840–844 [DOI] [PubMed] [Google Scholar]
- 5. Libby P., Simon D. I. (2001) Inflammation and thrombosis: the clot thickens. Circulation 103, 1718–1720 [DOI] [PubMed] [Google Scholar]
- 6. Ross R. (1999) Atherosclerosis—an inflammatory disease. N. Engl. J. Med. 340, 115–126 [DOI] [PubMed] [Google Scholar]
- 7. Pellitteri-Hahn M. C., Warren M. C., Didier D. N., Winkler E. L., Mirza S. P., Greene A. S., Olivier M. (2006) Improved mass spectrometric proteomic profiling of the secretome of rat vascular endothelial cells. J. Proteome Res. 5, 2861–2864 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8. Tunica D. G., Yin X., Sidibe A., Stegemann C., Nissum M., Zeng L., Brunet M., Mayr M. (2009) Proteomic analysis of the secretome of human umbilical vein endothelial cells using a combination of free-flow electrophoresis and nanoflow LC-MS/MS. Proteomics 9, 4991–4996 [DOI] [PubMed] [Google Scholar]
- 9. Flora J. W., Edmiston J., Secrist R., Li G., Rana G. S., Langston T. B., McKinney W. (2008) Identification of in vitro differential cell secretions due to cigarette smoke condensate exposure using nanoflow capillary liquid chromatography and high-resolution mass spectrometry. Anal. Bioanal. Chem. 391, 2845–2856 [DOI] [PubMed] [Google Scholar]
- 10. Burghoff S., Schrader J. (2011) Secretome of human endothelial cells under shear stress. J. Proteome Res. 10, 1160–1169 [DOI] [PubMed] [Google Scholar]
- 11. Hocking S. L., Wu L. E., Guilhaus M., Chisholm D. J., James D. E. (2010) Intrinsic depot-specific differences in the secretome of adipose tissue, preadipocytes, and adipose tissue-derived microvascular endothelial cells. Diabetes 59, 3008–3016 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12. Griffoni C., Di Molfetta S., Fantozzi L., Zanetti C., Pippia P., Tomasi V., Spisni E. (2011) Modification of proteins secreted by endothelial cells during modeled low gravity exposure. J. Cell. Biochem. 112, 265–272 [DOI] [PubMed] [Google Scholar]
- 13. Brioschi M., Lento S., Tremoli E., Banfi C. (2013) Proteomic analysis of endothelial cell secretome: a means of studying the pleiotropic effects of Hmg-CoA reductase inhibitors. J. Proteomics 78, 346–361 [DOI] [PubMed] [Google Scholar]
- 14. Castagna M., Takai Y., Kaibuchi K., Sano K., Kikkawa U., Nishizuka Y. (1982) Direct activation of calcium-activated, phospholipid-dependent protein kinase by tumor-promoting phorbol esters. J. Biol. Chem. 257, 7847–7851 [PubMed] [Google Scholar]
- 15. Jacobs J. M., Waters K. M., Kathmann L. E., Camp D. G., 2nd, Wiley H. S., Smith R. D., Thrall B. D. (2008) The mammary epithelial cell secretome and its regulation by signal transduction pathways. J. Proteome Res. 7, 558–569 [DOI] [PubMed] [Google Scholar]
- 16. Zampetaki A., Kiechl S., Drozdov I., Willeit P., Mayr U., Prokopi M., Mayr A., Weger S., Oberhollenzer F., Bonora E., Shah A., Willeit J., Mayr M. (2010) Plasma microRNA profiling reveals loss of endothelial miR-126 and other microRNAs in type 2 diabetes. Circ. Res. 107, 810–817 [DOI] [PubMed] [Google Scholar]
- 17. Margariti A., Zampetaki A., Xiao Q., Zhou B., Karamariti E., Martin D., Yin X., Mayr M., Li H., Zhang Z., De Falco E., Hu Y., Cockerill G., Xu Q., Zeng L. (2010) Histone deacetylase 7 controls endothelial cell growth through modulation of beta-catenin. Circ. Res. 106, 1202–1211 [DOI] [PubMed] [Google Scholar]
- 18. Yin X., Cuello F., Mayr U., Hao Z., Hornshaw M., Ehler E., Avkiran M., Mayr M. (2010) Proteomics analysis of the cardiac myofilament subproteome reveals dynamic alterations in phosphatase subunit distribution. Mol. Cell. Proteomics 9, 497–509 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19. Didangelos A., Yin X., Mandal K., Baumert M., Jahangiri M., Mayr M. (2010) Proteomics characterization of extracellular space components in the human aorta. Mol Cell. Proteomics 9, 2048–2062 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20. Mayr M., Grainger D., Mayr U., Leroyer A. S., Leseche G., Sidibe A., Herbin O., Yin X., Gomes A., Madhu B., Griffiths J. R., Xu Q., Tedgui A., Boulanger C. M. (2009) Proteomics, metabolomics, and immunomics on microparticles derived from human atherosclerotic plaques. Circ. Cardiovasc. Genet. 2, 379–388 [DOI] [PubMed] [Google Scholar]
- 21. Pula G., Mayr U., Evans C., Prokopi M., Vara D. S., Yin X., Astroulakis Z., Xiao Q., Hill J., Xu Q., Mayr M. (2009) Proteomics identifies thymidine phosphorylase as a key regulator of the angiogenic potential of colony-forming units and endothelial progenitor cell cultures. Circ. Res. 104, 32–40 [DOI] [PubMed] [Google Scholar]
- 22. Urbich C., De Souza A. I., Rossig L., Yin X., Xing Q., Prokopi M., Drozdov I., Steiner M., Breuss J., Xu Q., Dimmeler S., Mayr M. (2011) Proteomic characterization of human early pro-angiogenic cells. J. Mol. Cell. Cardiol. 50, 333–336 [DOI] [PubMed] [Google Scholar]
- 23. Hagglund P., Bunkenborg J., Elortza F., Jensen O. N., Roepstorff P. (2004) A new strategy for identification of N-glycosylated proteins and unambiguous assignment of their glycosylation sites using HILIC enrichment and partial deglycosylation. J. Proteome Res. 3, 556–566 [DOI] [PubMed] [Google Scholar]
- 24. Saba J., Dutta S., Hemenway E., Viner R. (2012) Increasing the productivity of glycopeptides analysis by using higher-energy collision dissociation-accurate mass-product-dependent electron transfer dissociation. Int. J. Proteomics 2012, 560391. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25. Bern M., Cai Y., Goldberg D. (2007) Lookup peaks: a hybrid of de novo sequencing and database search for protein identification by tandem mass spectrometry. Anal Chem. 79, 1393–1400 [DOI] [PubMed] [Google Scholar]
- 26. Antonov A. S., Lukashev M. E., Romanov Y. A., Tkachuk V. A., Repin V. S., Smirnov V. N. (1986) Morphological alterations in endothelial cells from human aorta and umbilical vein induced by forskolin and phorbol 12-myristate 13-acetate: a synergistic action of adenylate cyclase and protein kinase C activators. Proc. Natl. Acad. Sci. U.S.A. 83, 9704–9708 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27. Sadler J. E. (1998) Biochemistry and genetics of von Willebrand factor. Annu. Rev. Biochem. 67, 395–424 [DOI] [PubMed] [Google Scholar]
- 28. Matsui T., Titani K., Mizuochi T. (1992) Structures of the asparagine-linked oligosaccharide chains of human von Willebrand factor. Occurrence of blood group A, B, and H(O) structures. J. Biol. Chem. 267, 8723–8731 [PubMed] [Google Scholar]
- 29. Canis K., McKinnon T. A., Nowak A., Haslam S. M., Panico M., Morris H. R., Laffan M. A., Dell A. (2012) Mapping the N-glycome of human von Willebrand factor. Biochem. J. 447, 217–228 [DOI] [PubMed] [Google Scholar]
- 30. Simak J., Holada K., Vostal J. G. (2002) Release of annexin V-binding membrane microparticles from cultured human umbilical vein endothelial cells after treatment with camptothecin. BMC Cell Biol. 3, 11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31. Hart G. W., Copeland R. J. (2010) Glycomics hits the big time. Cell 143, 672–676 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32. Wollscheid B., Bausch-Fluck D., Henderson C., O'Brien R., Bibel M., Schiess R., Aebersold R., Watts J. D. (2009) Mass-spectrometric identification and relative quantification of N-linked cell surface glycoproteins. Nat. Biotechnol. 27, 378–386 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33. Leth-Larsen R., Lund R. R., Ditzel H. J. (2010) Plasma membrane proteomics and its application in clinical cancer biomarker discovery. Mol. Cell. Proteomics 9, 1369–1382 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34. Madabhushi S. R., Shang C., Dayananda K. M., Rittenhouse-Olson K., Murphy M., Ryan T. E., Montgomery R. R., Neelamegham S. (2012) von Willebrand factor (VWF) propeptide binding to VWF D'D3 domain attenuates platelet activation and adhesion. Blood 119, 4769–4778 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35. van Mourik J. A., Boertjes R., Huisveld I. A., Fijnvandraat K., Pajkrt D., van Genderen P. J., Fijnheer R. (1999) von Willebrand factor propeptide in vascular disorders: a tool to distinguish between acute and chronic endothelial cell perturbation. Blood 94, 179–185 [PubMed] [Google Scholar]
- 36. Pan S., Chen R., Aebersold R., Brentnall T. A. (2011) Mass spectrometry based glycoproteomics—from a proteomics perspective. Mol. Cell. Proteomics 10, R110.003251. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37. Maley F., Trimble R. B., Tarentino A. L., Plummer T. H., Jr. (1989) Characterization of glycoproteins and their associated oligosaccharides through the use of endoglycosidases. Anal. Biochem. 180, 195–204 [DOI] [PubMed] [Google Scholar]
- 38. Flatmark T. (1967) Multiple molecular forms of bovine heart cytochrome c. V. A comparative study of their physicochemical properties and their reactions in biological systems. J. Biol. Chem. 242, 2454–2459 [PubMed] [Google Scholar]
- 39. Zielinska D. F., Gnad F., Wisniewski J. R., Mann M. (2010) Precision mapping of an in vivo N-glycoproteome reveals rigid topological and sequence constraints. Cell 141, 897–907 [DOI] [PubMed] [Google Scholar]
- 40. Angel P. M., Lim J. M., Wells L., Bergmann C., Orlando R. (2007) A potential pitfall in 18O-based N-linked glycosylation site mapping. Rapid Commun. Mass Spectrom. 21, 674–682 [DOI] [PubMed] [Google Scholar]
- 41. Palmisano G., Melo-Braga M. N., Engholm-Keller K., Parker B. L., Larsen M. R. (2012) Chemical deamidation: a common pitfall in large-scale N-linked glycoproteomic mass spectrometry-based analyses. J. Proteome Res. 11, 1949–1957 [DOI] [PubMed] [Google Scholar]
- 42. Wuhrer M., Deelder A. M., Hokke C. H. (2005) Protein glycosylation analysis by liquid chromatography-mass spectrometry. J. Chromatogr. B Analyt. Technol. Biomed. Life Sci. 825, 124–133 [DOI] [PubMed] [Google Scholar]
- 43. Morelle W., Michalski J. C. (2007) Analysis of protein glycosylation by mass spectrometry. Nat. Protoc. 2, 1585–1602 [DOI] [PubMed] [Google Scholar]
- 44. Zhang M. Y., Pace N., Kerns E. H., Kleintop T., Kagan N., Sakuma T. (2005) Hybrid triple quadrupole-linear ion trap mass spectrometry in fragmentation mechanism studies: application to structure elucidation of buspirone and one of its metabolites. J. Mass Spectrom. 40, 1017–1029 [DOI] [PubMed] [Google Scholar]
- 45. Tissot B., North S. J., Ceroni A., Pang P. C., Panico M., Rosati F., Capone A., Haslam S. M., Dell A., Morris H. R. (2009) Glycoproteomics: past, present and future. FEBS Lett. 583, 1728–1735 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46. Hang H. C., Yu C., Kato D. L., Bertozzi C. R. (2003) A metabolic labeling approach toward proteomic analysis of mucin-type O-linked glycosylation. Proc. Natl. Acad. Sci. U.S.A. 100, 14846–14851 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47. Zhao P., Viner R., Teo C. F., Boons G. J., Horn D., Wells L. (2011) Combining high-energy C-trap dissociation and electron transfer dissociation for protein O-GlcNAc modification site assignment. J. Proteome Res. 10, 4088–4104 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48. Owen J. B., Di Domenico F., Sultana R., Perluigi M., Cini C., Pierce W. M., Butterfield D. A. (2009) Proteomics-determined differences in the concanavalin-A-fractionated proteome of hippocampus and inferior parietal lobule in subjects with Alzheimer's disease and mild cognitive impairment: implications for progression of AD. J. Proteome Res. 8, 471–482 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49. Ghosh D., Krokhin O., Antonovici M., Ens W., Standing K. G., Beavis R. C., Wilkins J. A. (2004) Lectin affinity as an approach to the proteomic analysis of membrane glycoproteins. J. Proteome Res. 3, 841–850 [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.