Abstract
The influenza M2 protein conducts protons through a critical histidine (His) residue, His37. Whether His37 only interacts with water to relay protons into the virion, or whether a low-barrier hydrogen bond (LBHB) also exists between the histidines to stabilize charges before proton conduction, is actively debated. To address this question, we have measured the imidazole 1HN chemical shifts of His37 at different temperatures and pH using 2D 15N-1H correlation solid-state NMR. At low temperature, the HN chemical shifts are 8–15 ppm at all pH values, indicating that the His37 sidechain forms conventional hydrogen bonds (H bonds) instead of LBHBs. At ambient temperature, the dynamically averaged HN chemical shifts are 4.8 ppm, indicating that the H-bonding partner of the imidazole is water instead of another histidine in the tetrameric channel. These data show that His37 forms H-bonds only with water, with regular strength, thus supporting the His-water proton exchange model and ruling out the low-barrier H-bonded dimer model.
The influenza M2 protein forms a tetrameric proton channel important for the virus lifecycle 1-3. Activated by the low pH environment of the endosome, the channel opens to acidify the virion, which causes viral uncoating. The mechanism of proton conduction through M2 has been long debated. Early computational studies and functional data diverged on whether proton conduction occurs by Grotthuss hopping along a water wire 4,5 or requires conformational changes of the only titratable residue in the transmembrane (TM) domain, His37 6 (Fig. 1a). Recent data has ruled out the water wire model and converged on the active participation of His37 in proton relay. Evidence for proton shuttling by His37 came from magic-angle-spinning (MAS) 15N NMR spectra showing chemical exchange of the imidazole nitrogens between the protonated (NH) and unprotonated (N) states at the physiological pH of the endosome 7. This exchange is accompanied by low-pH specific imidazolium reorientation on the microsecond timescale with an energy barrier comparable to the proton conduction barrier 8.
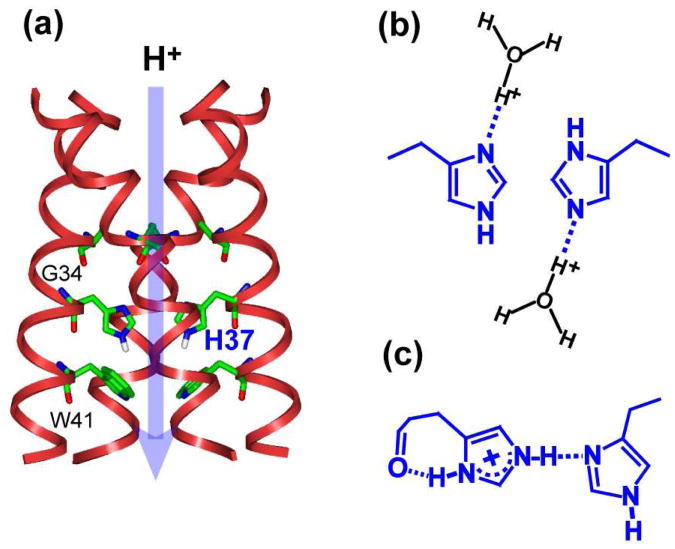
Figure 1.
Two proton conduction models for the influenza M2 channel. (a) Structure of M2TM at pH 6.5 (PDB: 3LBW), showing the location of the key His37 and Trp41. (b) His37-water proton exchange model. (b) His-His low-barrier H-bonded dimer model. The dimer of dimer state is proposed to exist stably in the +2 tetrad to stabilize charge before proton transfer to water.
Despite the general consensus that His37 shuttles protons, the mechanism by which charge is stabilized in the His37 tetrad is still actively debated. The 15N chemical exchange and imidazolium reorientation led to the proposal that His37-water H-bonding and proton exchange are sufficient for proton conduction (Fig. 1b) 7, and that excess protons are stabilized by delocalization over the His37 tetrad and the surrounding water molecules 9. In contrast, an alternative model posits a LBHB between a neutral and a cationic histidine in the +2 state of the channel (Fig. 1c) 10, which stabilizes the charges before channel activation. This model was motivated by the observation of a very high pKa of 8.2 for the first two protonation steps in DMPC/DMPG bilayer-bound M2 TM peptide (M2TM) 10, and by computational modeling of the His37 sidechain structure 11. The latter yielded His37 (χ1,χ2) torsion angles of (180°, 90°) to establish the putative Nε2-H…Nδ1 H-bond Recently reported chemical shift multiplicity of some of the TM residues 12,13 although observed at neutral pH, was also interpreted as supporting the LBHB model.
Equilibrium conformation of His37 measured by solid-state NMR 8 and X-ray crystallography 9,14 at acidic pH have so far shown no direct His-His H-bonding: the His37 χ1 and χ2 angles were measured to be ∼180° in both lipid bilayers and detergents, which point the Nε2-H and Nδ1 toward the interior and exterior of the channel rather than toward each other. 13C-13C 2D correlation spectra of the +2 charged channel displayed no imidazole-imidazolium cross peaks 7, also challenging the LBHB model. However, the 15N NMR spectra showing N<–>NH chemical exchange, can, in principle, be due to either His-water proton transfer or His-His H-bonding. Thus, we sought more definitive evidence for the H-bonding partner of His37 as well as the strength of the His37 H-bond The strength of H-bonds can be discerned through the 1H chemical shift a proton in a low-barrier or strong H-bond should have a large chemical shift of greater than 16 ppm 15-18, whereas a proton in a regular unequal-well H-bond should have a smaller chemical shift of 8-15 ppm 19,20. The identity of the H-bonding partner for membrane proteins in hydrated lipid bilayers can be determined through the temperature dependence of the 1H chemical shift. Between -30°C and +30°C, the diffusion rates of water in the channel change significantly 21,22, thus a regular N-H…O H-bond should involve only one or few water molecules at low temperature but should undergo rapid exchange with many water molecules at physiological temperature. This should result in a 1H chemical shift close to the imidazole HN value at low temperature but a population-weighted value near the water 1H chemical shift at high temperature. In contrast, for a pKa-matched N-H…N LBHB 23,24, the central proton has a much higher activation energy for exchange with water 25; moreover proton transfer dynamics between the two nitrogens is ultrafast. Thus the 1H chemical shift will be insensitive to temperature of this range and remain large. Thus, the low-temperature 1H chemical shift reveals the H-bond strength, whereas the high-temperature shift indicates the identity of the H-bonding partner.
We measured the 1H chemical shift using the 2D 15N-1H heteronuclear correlation (HETCOR) experiment. To detect only cross peaks due to direct N-H dipolar coupling without relayed transfer, we suppressed 1H spin diffusion using 1H homonuclear decoupling during the evolution period and the 1H-15N cross polarization period 26. His37-labeled M2TM bound to a virus-mimetic lipid membrane were measured at pH 6.0, 4.5, and 8.5 8. Since all initial experiments that led to the LBHB model were conducted on M2TM, we used the same construct to avoid potential ambiguities in interpretation. Previous measurement of His37 pKa's in this virus-mimetic membrane indicated that the channel was 80% in the +2 state at pH 6, in a mixed +3 and +4 state at pH 4.5, and about 90% neutral at pH 8.5 7. Thus, the pH 6.0 sample is the closest to the putative LBHB state. The 2D HETCOR spectra were measured at 245 K to determine the H-bond strength and 296 K to determine the H-bonding partner.
Fig. 2a shows the 2D HETCOR spectra of the pH 6.0 sample. At 245 K, the imidazole Nε2 (τ tautomer) and Nδ1 (π tautomer) peaks at 160-180 ppm exhibit 1H chemical shifts of 8 - 12 ppm, similar to the backbone amide 1H chemical shift range. Thus, imidazole HN lies in a regular H-bond For comparison, histidine hydrochloride (Fig. S1) shows a large Hδ1 chemical shift of 16.8 ppm due to an intermolecular H-bond to a C=O with an N…O distance of 2.63 Å 27. Both 15N and 1H shifts reflect the strength of the H-bond: small 15N and 1H shifts indicate a stronger covalent N-H bond, while large shifts indicate a more deprotonated nitrogen or a stronger H-bond 17,18. The correlation gives a slope of ∼3 between the 15N and 1H chemical shifts (Fig. 2a). The 1H shift distribution (Fig. S2), detected for both backbone and imidazole nitrogens, indicate a distribution of H-bond strengths. The backbone distribution is likely due to varying degrees of helix ideality in an ensemble with mixed protonation states, while the imidazole HN shift distribution can be attributed to the presence of multiple N-H species, including Nε2H(τ), Nδ1H(π), and the Nε2H and Nδ1H of cationic imidazolium (Fig. S3).
Figure 2.
2D 15N-1H HETCOR spectra of membrane-bound His37-labeled M2TM at (a) pH 6.0, (b) pH 4.5, and (c) pH 8.5 at 245 K (left) and 296 K (right). The main charged states of the M2TM channel at each pH are indicated. The 1H dimension of the spectrum was measured with 1H homonuclear decoupling to eliminate spin diffusion effects. Assignments are shown in red for the neutral τ tautomer, black for the neutral π tautomer, and green for cationic His37.
When the temperature increased to 296 K, the imidazole 1H chemical shifts decreased uniformly to 4.8 ppm, indicating definitively that the H-bonding partner of His37 is water instead of another His. Since 1H homonuclear decoupling was applied in the experiment, both rigid and mobile protons were equally detected, thus the observed 1H chemical shift near the unperturbed water frequency indicates a large number of water molecules in exchange with the imidazole nitrogens. For comparison, the backbone HN chemical shift is unaffected by temperature, as expected for the persistence of N-H…O=C H-bonds at these temperatures.
The 2D spectra of the pH 4.5 sample (Fig. 2b) further support the His37-water interaction model. Even at low temperature, the 178-ppm 15N peak already shows a water 1H cross peak (5.7 ppm) in addition to the Hε2/Hδ1 signal (12-15 ppm), consistent with previous data showing a more hydrated channel at this low pH 28. The Hε2/Hδ1 chemical shift is larger than at pH 6, indicating stronger H-bonds. This is consistent with the previously measured N-H bond elongation at this pH 8. The 15N/1H chemical shift slope is the same as at pH 6.0 (Fig. S4), as expected for the intrinsic correlation between 15N protonation and N-H…O H-bond strength. Again, the identity of the H-bonding partner is determined by the high-temperature spectrum, which shows a 1H chemical shift at the water position of 4.8 ppm, indicating that His37 H-bonds only with water.
At pH 8.5, the high-temperature spectrum retained the dominant water cross peak, but a weak signal at ∼8 ppm was also detected and can be assigned to Hε2. Although the channel does not conduct protons at this pH, some waters are still present, for example between His37 and Trp41 8,9,29, thus allowing polarization transfer to 15N. At low temperature, the unprotonated nitrogen, mainly Nδ1, exhibits a 1H cross peak at ∼5 ppm due to Hε1, as verified by the spectrum of amino acid histidine (Fig. S1)27.
These low-temperature 1H chemical shifts are smaller than expected for an LBHB or a strong H bond, while the high-temperature 1H chemical shifts reveal the H-bonding partner to be water. Thus the data support the direct His37-water interaction model and rule out the His-His LBHB-dimer model. The 1.65 Å crystal structure at pH 6.5 9 detected tightly-clustered water molecules near the His37 tetrad, with N…O distances as short as 2.8 Å, also supporting direct His37-water interactions. On the other hand, all experimental data so far, including the initial 15N NMR spectra from which the dimer model was proposed 10, show an absence of imidazole-imidazolium H-bond. An LBHB entails either a single 15N peak at the averaged chemical shift between N and NH for equal-well potentials or two 15N peaks centered around the averaged frequency for unequal-well potentials. Instead, the 15N spectra showed a single peak away from the averaged chemical shift, without the partner peak. Molecular modeling of the HxxxW structure 11 was questionable since it used the putative LBHB as a starting distance restraint to enforce the expected geometry during MD simulations. Finally, the LBHB model implies a hydrophobic environment for the donor and acceptor with a very small pKa difference 30, which contradict the observed different proton affinities of Nε2 and Nδ1 in His37 and the high hydration of this residue.
In conclusion, temperature-dependent 1H chemical shifts of His37 sidechain indicate that His37-water H-bonding and proton exchange dominate the equilibrium structure of the His37 tetrad throughout the whole pH range. His-His interactions are indirectly mediated by water. If direct His-His H-bond is too transient to be detectable by NMR, then it cannot be an LBHB and cannot provide stabilization for the dimer state. We propose that charges are stabilized by water-mediated interactions and by cation-π interaction between His37 and Trp41.
This temperature-dependent 1H chemical shift approach avoids the difficulty of measuring N…N and N…O distances across a H-bond by NMR; moreover it reveals the structure of the most essential player in a proton relay chain. It is applicable to both biological and synthetic proton conductors to understand the nature of the H-bond in proton transport.
Supplementary Material
Acknowledgments
We thank Professor Schmidt-Rohr for useful discussions. This work is supported by NIH grant GM088204.
Footnotes
Supporting Information: Experimental procedures and additional spectra. This information is available free of charge via the internet at http://pubs.acs.org.
References
- 1.Pinto LH, Lamb RA. J Biol Chem. 2006;281:8997–9000. doi: 10.1074/jbc.R500020200. [DOI] [PubMed] [Google Scholar]
- 2.Cady SD, Luo WB, Hu F, Hong M. Biochemistry. 2009;48:7356–7364. doi: 10.1021/bi9008837. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Wang J, Qiu JX, Soto CS, DeGrado WF. Curr Opin Struct Biol. 2011;21:68–80. doi: 10.1016/j.sbi.2010.12.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Sansom MSP, Kerr ID, Smith GR, Son HS. Virology. 1997;233:163–173. doi: 10.1006/viro.1997.8578. [DOI] [PubMed] [Google Scholar]
- 5.Okada A, Miura T, Takeuchi H. Biochemistry. 2001;40:6053–6060. doi: 10.1021/bi0028441. [DOI] [PubMed] [Google Scholar]
- 6.Pinto LH, Dieckmann GR, Gandhi CS, Papworth CG, Braman J, Shaughnessy MA, Lear JD, Lamb RA, DeGrado WF. Proc Natl Acad Sci USA. 1997;94:11301–11306. doi: 10.1073/pnas.94.21.11301. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Hu F, Schmidt-Rohr K, Hong M. J Am Chem Soc. 2012;134:3703–3713. doi: 10.1021/ja2081185. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Hu F, Luo W, Hong M. Science. 2010;330:505–508. doi: 10.1126/science.1191714. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Acharya A, Carnevale V, Fiorin G, Levine BG, Polishchuk A, Balannick V, Samish I, Lamb RA, Pinto LH, DeGrado WF, Klein ML. Proc Natl Acad Sci U S A. 2010;107:15075–15080. doi: 10.1073/pnas.1007071107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Hu J, Fu R, Nishimura K, Zhang L, Zhou HX, Busath DD, Vijayvergiya V, Cross TA. Proc Natl Acad Sci USA. 2006;103:6865–6870. doi: 10.1073/pnas.0601944103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Sharma M, Yi M, Dong H, Qin H, Peterson E, Busath D, Zhou HX, Cross TA. Science. 2010;330:509–512. doi: 10.1126/science.1191750. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Andreas LB, Eddy MT, Chou JJ, Griffin RG. J Am Chem Soc. 2012;134:7215–7218. doi: 10.1021/ja3003606. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Can TV, Sharma M, Hung I, Gor'kov PL, Brey WW, Cross TA. J Am Chem Soc. 2012;134:9022–9025. doi: 10.1021/ja3004039. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Stouffer AL, Acharya R, Salom D, Levine AS, Di Costanzo L, Soto CS, Tereshko V, Nanda V, Stayrook S, DeGrado WF. Nature. 2008;451:596–599. doi: 10.1038/nature06528. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Brown SP. Solid State Nucl Magn Reson. 2012;41:1–27. doi: 10.1016/j.ssnmr.2011.11.006. [DOI] [PubMed] [Google Scholar]
- 16.Gilli P, Bertolasi V, Ferretti V, Gilli G. J Am Chem Soc. 2000;122:10405–10417. [Google Scholar]
- 17.Lorente P, Shenderovich IG, Golubev NS, Denisov GS, Buntkowsky G, Limbach HH. Magn Reson Chem. 2001;39:S18–S29. [Google Scholar]
- 18.Sharif S, Schagen D, Toney MD, Limbach HH. J Am Chem Soc. 2007;129:4440–4455. doi: 10.1021/ja070296+. [DOI] [PubMed] [Google Scholar]
- 19.Berglund B, Vaughan RW. J Chem Phys. 1980;73:2037–2043. [Google Scholar]
- 20.Barfield M. J Am Chem Soc. 2002;124:4158–4168. doi: 10.1021/ja012674v. [DOI] [PubMed] [Google Scholar]
- 21.Nicotera I, Coppola L, Rossi CO, Youssry M, Ranieri GA. J Phys Chem B. 2009;113:13935–13941. doi: 10.1021/jp904691g. [DOI] [PubMed] [Google Scholar]
- 22.Takahara S, Nakano M, Kittaka S, Kuroda Y, Mori T, Hamano H, Yamaguchi T. J Phys Chem B. 1999;103:5814–5819. [Google Scholar]
- 23.Song XJ, McDermott AE. Magn Reson Chem. 2001;39:S37–S43. [Google Scholar]
- 24.Wehrle B, Zimmermann H, Limbach HH. J Am Chem Soc. 1988;110:7014–7024. [Google Scholar]
- 25.Lin J, Westler WM, Cleland WW, Markley JL, Frey PA. Proc Natl Acad Sci U S A. 1998;95:14664–14668. doi: 10.1073/pnas.95.25.14664. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Hong M, Yao XL, Jakes K, Huster D. J Phys Chem B. 2002;106:7355–7364. [Google Scholar]
- 27.Li S, Hong M. J Am Chem Soc. 2011;133:1534–1544. doi: 10.1021/ja108943n. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Luo W, Hong M. J Am Chem Soc. 2010;132:2378–2384. doi: 10.1021/ja9096219. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Su Y, Hu F, Hong M. J Am Chem Soc. 2012;134:8693–8702. doi: 10.1021/ja3026328. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Schutz CN, Warshel A. Proteins. 2004;55:711–723. doi: 10.1002/prot.20096. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.