Abstract
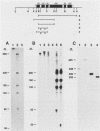
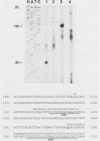
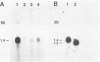
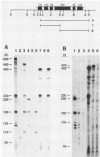
The nucleotide sequence and intron-exon structure of the Drosophila melanogaster vermilion (v) gene have been determined. In addition, the sites of several mutations and the effects of these mutations on transcription have been examined. The major v mRNA is generated upon splicing six exons of lengths (5' to 3') 83, 161, 134, 607, 94, and 227 nucleotides (nt). A minor species of v mRNA is initiated at an upstream site and has a 5' exon of at least 152 nt which overlaps the region included in the 83-nt exon of the major v RNA. The three v mutations, v1, v2, and vk, which can be suppressed by mutations at suppressor of sable, su(s), are insertions of transposon 412 at the same position in exon 1, 36 nt downstream of the major transcription initiation site. Despite the 7.5-kilobase insertion in these v alleles, a reduced level of wild-type-sized mRNA accumulates in suppressed mutant strains. The structure and transcription of several unsuppressible v alleles have also been examined. The v36f mutation is a B104/roo insertion in intron 4 near the splice donor site. A mutant carrying this alteration accumulates a very low level of mRNA that is apparently polyadenylated at a site within the B104/roo transposon. The v48a mutation, which deletes approximately 200 nt of DNA, fuses portions of exons 3 and 4 without disruption of the translational reading frame. A smaller transcript accumulates at a wild-type level, and thus an altered, nonfunctional polypeptide is likely to be synthesized in strains carrying this mutation.(ABSTRACT TRUNCATED AT 250 WORDS)
Full text
PDF








Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Baillie D. L., Chovnick A. Studies on the genetic control of tryptophan pyrrolase in Drosophila melanogaster. Mol Gen Genet. 1971;112(4):341–353. doi: 10.1007/BF00334435. [DOI] [PubMed] [Google Scholar]
- Bermingham J. R., Jr, Scott M. P. Developmentally regulated alternative splicing of transcripts from the Drosophila homeotic gene Antennapedia can produce four different proteins. EMBO J. 1988 Oct;7(10):3211–3222. doi: 10.1002/j.1460-2075.1988.tb03188.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Biggs J., Searles L. L., Greenleaf A. L. Structure of the eukaryotic transcription apparatus: features of the gene for the largest subunit of Drosophila RNA polymerase II. Cell. 1985 Sep;42(2):611–621. doi: 10.1016/0092-8674(85)90118-7. [DOI] [PubMed] [Google Scholar]
- Bingham P. M., Levis R., Rubin G. M. Cloning of DNA sequences from the white locus of D. melanogaster by a novel and general method. Cell. 1981 Sep;25(3):693–704. doi: 10.1016/0092-8674(81)90176-8. [DOI] [PubMed] [Google Scholar]
- Breathnach R., Chambon P. Organization and expression of eucaryotic split genes coding for proteins. Annu Rev Biochem. 1981;50:349–383. doi: 10.1146/annurev.bi.50.070181.002025. [DOI] [PubMed] [Google Scholar]
- Calzone F. J., Britten R. J., Davidson E. H. Mapping of gene transcripts by nuclease protection assays and cDNA primer extension. Methods Enzymol. 1987;152:611–632. doi: 10.1016/0076-6879(87)52069-9. [DOI] [PubMed] [Google Scholar]
- Chang D. Y., Wisely B., Huang S. M., Voelker R. A. Molecular cloning of suppressor of sable, a Drosophila melanogaster transposon-mediated suppressor. Mol Cell Biol. 1986 May;6(5):1520–1528. doi: 10.1128/mcb.6.5.1520. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chou T. B., Zachar Z., Bingham P. M. Developmental expression of a regulatory gene is programmed at the level of splicing. EMBO J. 1987 Dec 20;6(13):4095–4104. doi: 10.1002/j.1460-2075.1987.tb02755.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Corces V., Pellicer A., Axel R., Meselson M. Integration, transcription, and control of a Drosophila heat shock gene in mouse cells. Proc Natl Acad Sci U S A. 1981 Nov;78(11):7038–7042. doi: 10.1073/pnas.78.11.7038. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dale R. M., McClure B. A., Houchins J. P. A rapid single-stranded cloning strategy for producing a sequential series of overlapping clones for use in DNA sequencing: application to sequencing the corn mitochondrial 18 S rDNA. Plasmid. 1985 Jan;13(1):31–40. doi: 10.1016/0147-619x(85)90053-8. [DOI] [PubMed] [Google Scholar]
- Hultmark D., Klemenz R., Gehring W. J. Translational and transcriptional control elements in the untranslated leader of the heat-shock gene hsp22. Cell. 1986 Feb 14;44(3):429–438. doi: 10.1016/0092-8674(86)90464-2. [DOI] [PubMed] [Google Scholar]
- Kozak M. Compilation and analysis of sequences upstream from the translational start site in eukaryotic mRNAs. Nucleic Acids Res. 1984 Jan 25;12(2):857–872. doi: 10.1093/nar/12.2.857. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kuroiwa A., Hafen E., Gehring W. J. Cloning and transcriptional analysis of the segmentation gene fushi tarazu of Drosophila. Cell. 1984 Jul;37(3):825–831. doi: 10.1016/0092-8674(84)90417-3. [DOI] [PubMed] [Google Scholar]
- Levis R., O'Hare K., Rubin G. M. Effects of transposable element insertions on RNA encoded by the white gene of Drosophila. Cell. 1984 Sep;38(2):471–481. doi: 10.1016/0092-8674(84)90502-6. [DOI] [PubMed] [Google Scholar]
- Melton D. A., Krieg P. A., Rebagliati M. R., Maniatis T., Zinn K., Green M. R. Efficient in vitro synthesis of biologically active RNA and RNA hybridization probes from plasmids containing a bacteriophage SP6 promoter. Nucleic Acids Res. 1984 Sep 25;12(18):7035–7056. doi: 10.1093/nar/12.18.7035. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Modolell J., Bender W., Meselson M. Drosophila melanogaster mutations suppressible by the suppressor of Hairy-wing are insertions of a 7.3-kilobase mobile element. Proc Natl Acad Sci U S A. 1983 Mar;80(6):1678–1682. doi: 10.1073/pnas.80.6.1678. [DOI] [PMC free article] [PubMed] [Google Scholar]
- O'Hare K., Rubin G. M. Structures of P transposable elements and their sites of insertion and excision in the Drosophila melanogaster genome. Cell. 1983 Aug;34(1):25–35. doi: 10.1016/0092-8674(83)90133-2. [DOI] [PubMed] [Google Scholar]
- O'hare K., Levis R., Rubin G. M. Transcription of the white locus in Drosophila melanogaster. Proc Natl Acad Sci U S A. 1983 Nov;80(22):6917–6921. doi: 10.1073/pnas.80.22.6917. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Parkhurst S. M., Corces V. G. Developmental expression of Drosophila melanogaster retrovirus-like transposable elements. EMBO J. 1987 Feb;6(2):419–424. doi: 10.1002/j.1460-2075.1987.tb04771.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Parkhurst S. M., Corces V. G. Forked, gypsys, and suppressors in Drosophila. Cell. 1985 Jun;41(2):429–437. doi: 10.1016/s0092-8674(85)80016-7. [DOI] [PubMed] [Google Scholar]
- Parkhurst S. M., Corces V. G. Interactions among the gypsy transposable element and the yellow and the suppressor of hairy-wing loci in Drosophila melanogaster. Mol Cell Biol. 1986 Jan;6(1):47–53. doi: 10.1128/mcb.6.1.47. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Parkhurst S. M., Harrison D. A., Remington M. P., Spana C., Kelley R. L., Coyne R. S., Corces V. G. The Drosophila su(Hw) gene, which controls the phenotypic effect of the gypsy transposable element, encodes a putative DNA-binding protein. Genes Dev. 1988 Oct;2(10):1205–1215. doi: 10.1101/gad.2.10.1205. [DOI] [PubMed] [Google Scholar]
- Pirrotta V., Bröckl C. Transcription of the Drosophila white locus and some of its mutants. EMBO J. 1984 Mar;3(3):563–568. doi: 10.1002/j.1460-2075.1984.tb01847.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Poole S. J., Kauvar L. M., Drees B., Kornberg T. The engrailed locus of Drosophila: structural analysis of an embryonic transcript. Cell. 1985 Jan;40(1):37–43. doi: 10.1016/0092-8674(85)90306-x. [DOI] [PubMed] [Google Scholar]
- Proudfoot N. J., Brownlee G. G. 3' non-coding region sequences in eukaryotic messenger RNA. Nature. 1976 Sep 16;263(5574):211–214. doi: 10.1038/263211a0. [DOI] [PubMed] [Google Scholar]
- Sanger F., Coulson A. R., Barrell B. G., Smith A. J., Roe B. A. Cloning in single-stranded bacteriophage as an aid to rapid DNA sequencing. J Mol Biol. 1980 Oct 25;143(2):161–178. doi: 10.1016/0022-2836(80)90196-5. [DOI] [PubMed] [Google Scholar]
- Scherer G., Tschudi C., Perera J., Delius H., Pirrotta V. B104, a new dispersed repeated gene family in Drosophila melanogaster and its analogies with retroviruses. J Mol Biol. 1982 May 25;157(3):435–451. doi: 10.1016/0022-2836(82)90470-3. [DOI] [PubMed] [Google Scholar]
- Schwartz H. E., Lockett T. J., Young M. W. Analysis of transcripts from two families of nomadic DNA. J Mol Biol. 1982 May 5;157(1):49–68. doi: 10.1016/0022-2836(82)90512-5. [DOI] [PubMed] [Google Scholar]
- Searles L. L., Jokerst R. S., Bingham P. M., Voelker R. A., Greenleaf A. L. Molecular cloning of sequences from a Drosophila RNA polymerase II locus by P element transposon tagging. Cell. 1982 Dec;31(3 Pt 2):585–592. doi: 10.1016/0092-8674(82)90314-2. [DOI] [PubMed] [Google Scholar]
- Searles L. L., Voelker R. A. Molecular characterization of the Drosophila vermilion locus and its suppressible alleles. Proc Natl Acad Sci U S A. 1986 Jan;83(2):404–408. doi: 10.1073/pnas.83.2.404. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shapard P B. A Physiological Study of the Vermilion Eye Color Mutants of Drosophila Melanogaster. Genetics. 1960 Apr;45(4):359–376. doi: 10.1093/genetics/45.4.359. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shepherd B. M., Finnegan D. J. Structure of circular copies of the 412 transposable element present in Drosophila melanogaster tissue culture cells, and isolation of a free 412 long terminal repeat. J Mol Biol. 1984 Nov 25;180(1):21–40. doi: 10.1016/0022-2836(84)90428-5. [DOI] [PubMed] [Google Scholar]
- Spana C., Harrison D. A., Corces V. G. The Drosophila melanogaster suppressor of Hairy-wing protein binds to specific sequences of the gypsy retrotransposon. Genes Dev. 1988 Nov;2(11):1414–1423. doi: 10.1101/gad.2.11.1414. [DOI] [PubMed] [Google Scholar]
- Staden R. A new computer method for the storage and manipulation of DNA gel reading data. Nucleic Acids Res. 1980 Aug 25;8(16):3673–3694. doi: 10.1093/nar/8.16.3673. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tartof K. D. Interacting gene systems: I. The regulation of tryptophan pyrrolase by the vermilion-suppressor of vermilion system in Drosophila. Genetics. 1969 Jul;62(3):781–795. [PMC free article] [PubMed] [Google Scholar]
- Thomas P. S. Hybridization of denatured RNA and small DNA fragments transferred to nitrocellulose. Proc Natl Acad Sci U S A. 1980 Sep;77(9):5201–5205. doi: 10.1073/pnas.77.9.5201. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Will B. M., Bayev A. A., Finnegan D. J. Nucleotide sequence of terminal repeats of 412 transposable elements of Drosophila melanogaster. A similarity to proviral long terminal repeats and its implications for the mechanism of transposition. J Mol Biol. 1981 Dec 25;153(4):897–915. doi: 10.1016/0022-2836(81)90458-7. [DOI] [PubMed] [Google Scholar]
- Yuki S., Inouye S., Ishimaru S., Saigo K. Nucleotide sequence characterization of a Drosophila retrotransposon, 412. Eur J Biochem. 1986 Jul 15;158(2):403–410. doi: 10.1111/j.1432-1033.1986.tb09767.x. [DOI] [PubMed] [Google Scholar]
- Zachar Z., Chou T. B., Bingham P. M. Evidence that a regulatory gene autoregulates splicing of its transcript. EMBO J. 1987 Dec 20;6(13):4105–4111. doi: 10.1002/j.1460-2075.1987.tb02756.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zachar Z., Davison D., Garza D., Bingham P. M. A detailed developmental and structural study of the transcriptional effects of insertion of the Copia transposon into the white locus of Drosophila melanogaster. Genetics. 1985 Nov;111(3):495–515. doi: 10.1093/genetics/111.3.495. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zachar Z., Garza D., Chou T. B., Goland J., Bingham P. M. Molecular cloning and genetic analysis of the suppressor-of-white-apricot locus from Drosophila melanogaster. Mol Cell Biol. 1987 Jul;7(7):2498–2505. doi: 10.1128/mcb.7.7.2498. [DOI] [PMC free article] [PubMed] [Google Scholar]