Abstract
The RAD51 gene of Saccharomyces cerevisiae is required both for recombination and for the repair of DNA damage caused by X rays. Here we report the sequence and transcriptional regulation of this gene. The RAD51 protein shares significant homology (approximately 50%) over a 70-amino-acid with the RAD57 protein (J.A. Kans and R.K. Mortimer, Gene 105:139-140, 1991), the product of another yeast recombinational repair gene, and also moderate (approximately 27%), but potentially significant, homology with the bacterial RecA protein. The homologies cover a region that encodes a putative nucleotide binding site of the RAD51 protein. Sequences upstream of the coding region for RAD51 protein share homology with the damage response sequence element of RAD54, an upstream activating sequence required for damage regulation of the RAD54 transcript, and also contain two sites for restriction enzyme MluI; the presence of MluI restriction sites has been associated with cell cycle regulation. A 1.6-kb transcript corresponding to RAD51 was observed, and levels of this transcript increased rapidly after exposure to relatively low doses of X-rays. Additionally, RAD51 transcript levels were found to that of a group of genes involved primarily in DNA synthesis and replication which are thought to be coordinately cell cycle regulated. Cells arrested in early G1 were still capable of increasing levels of RAD51 transcript after irradiation, indicating that increased RAD51 transcript levels after X-ray exposure are not solely due to an X-ray-induced cessation of the cell cycle at a period when the level of RAD51 expression is normally high.
Full text
PDF


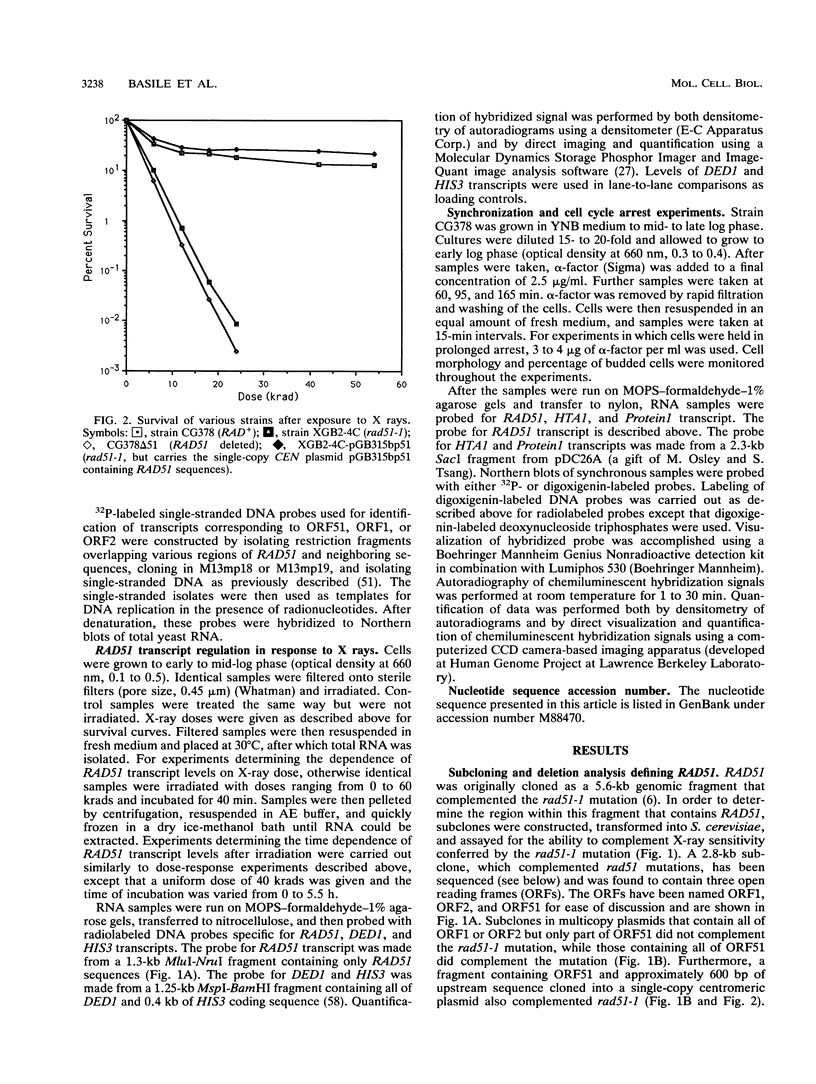


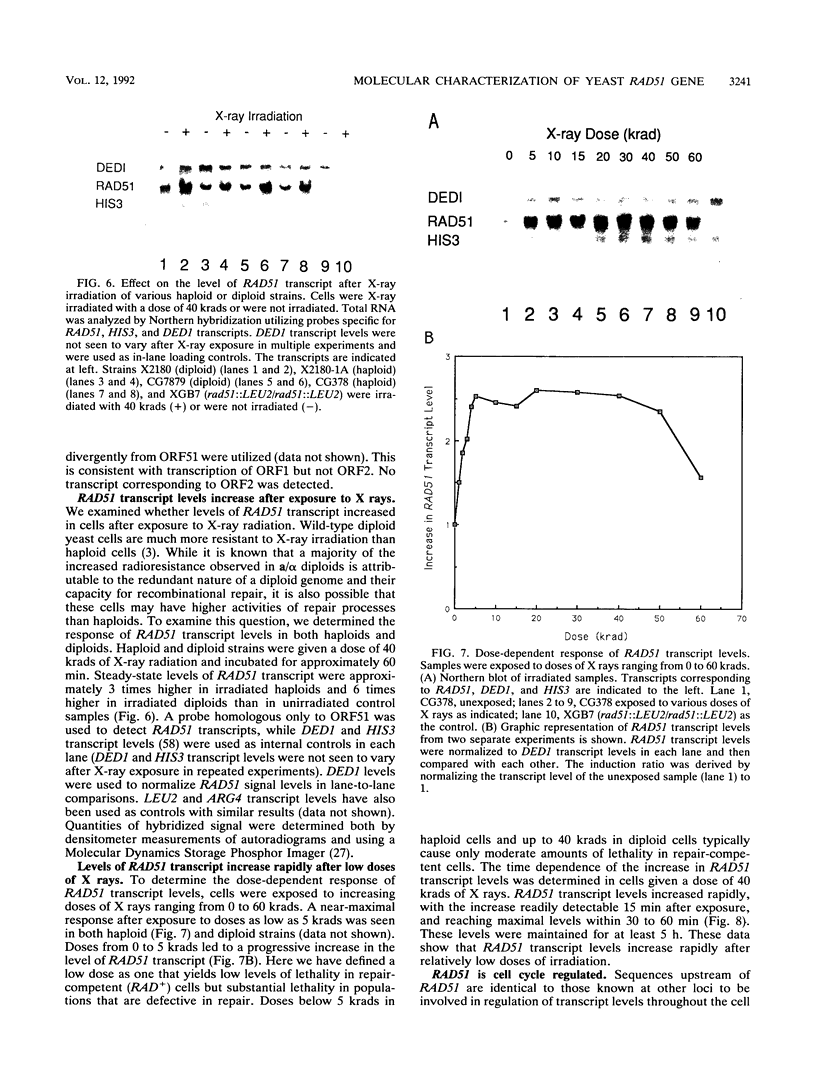
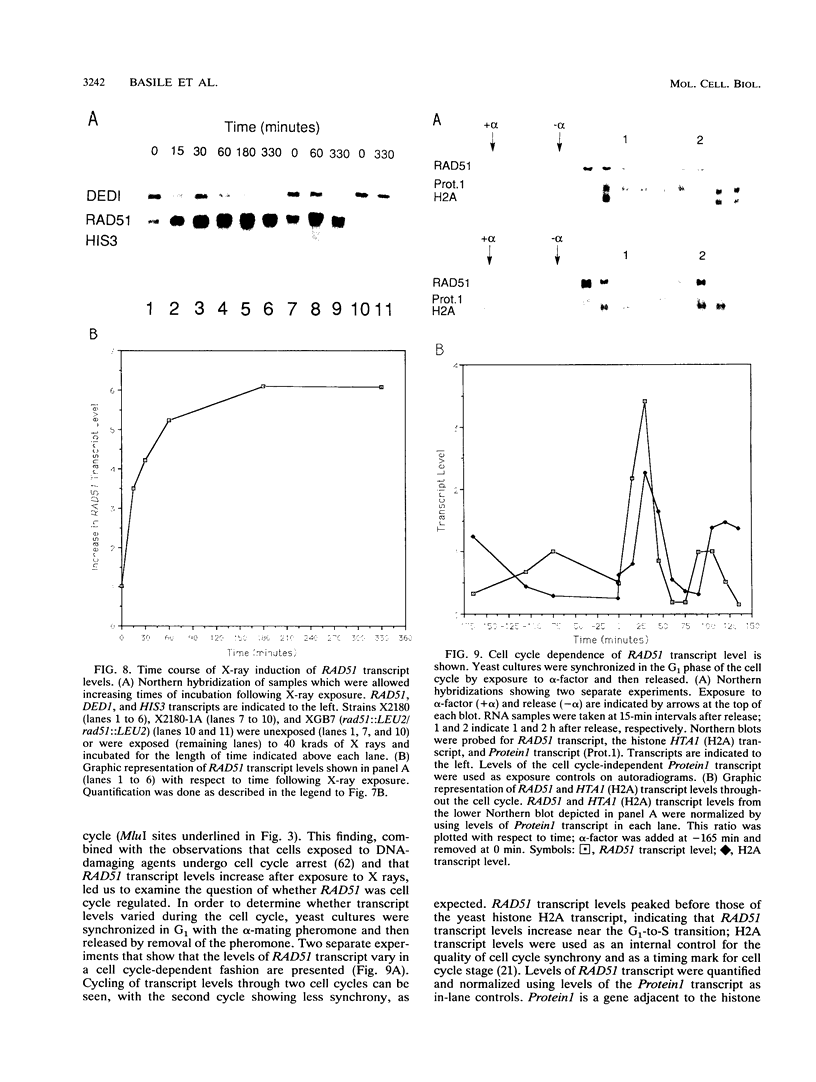




Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Adzuma K., Ogawa T., Ogawa H. Primary structure of the RAD52 gene in Saccharomyces cerevisiae. Mol Cell Biol. 1984 Dec;4(12):2735–2744. doi: 10.1128/mcb.4.12.2735. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Altschul S. F., Gish W., Miller W., Myers E. W., Lipman D. J. Basic local alignment search tool. J Mol Biol. 1990 Oct 5;215(3):403–410. doi: 10.1016/S0022-2836(05)80360-2. [DOI] [PubMed] [Google Scholar]
- BEAM C. A., MORTIMER R. K., WOLFE R. G., TOBIAS C. A. The relation of radioresistance to budding in Saccharomyces cerevisiae. Arch Biochem Biophys. 1954 Mar;49(1):110–122. doi: 10.1016/0003-9861(54)90172-1. [DOI] [PubMed] [Google Scholar]
- Budd M., Mortimer R. K. The effect of cycloheximide on repair in a temperature conditional radiation-sensitive mutant of Saccharomyces cerevisiae. Radiat Res. 1984 Sep;99(3):582–590. [PubMed] [Google Scholar]
- Cai L., Liu S. Z. Induction of cytogenetic adaptive response of somatic and germ cells in vivo and in vitro by low-dose X-irradiation. Int J Radiat Biol. 1990 Jul;58(1):187–194. doi: 10.1080/09553009014551541. [DOI] [PubMed] [Google Scholar]
- Cole G. M., Mortimer R. K. Failure to induce a DNA repair gene, RAD54, in Saccharomyces cerevisiae does not affect DNA repair or recombination phenotypes. Mol Cell Biol. 1989 Aug;9(8):3314–3322. doi: 10.1128/mcb.9.8.3314. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Contopoulou C. R., Cook V. E., Mortimer R. K. Analysis of DNA double strand breakage and repair using orthogonal field alternation gel electrophoresis. Yeast. 1987 Jun;3(2):71–76. doi: 10.1002/yea.320030203. [DOI] [PubMed] [Google Scholar]
- Doolittle R. F. Searching through sequence databases. Methods Enzymol. 1990;183:99–110. doi: 10.1016/0076-6879(90)83008-w. [DOI] [PubMed] [Google Scholar]
- Elledge S. J., Davis R. W. Identification of the DNA damage-responsive element of RNR2 and evidence that four distinct cellular factors bind it. Mol Cell Biol. 1989 Dec;9(12):5373–5386. doi: 10.1128/mcb.9.12.5373. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Elledge S. J., Davis R. W. Two genes differentially regulated in the cell cycle and by DNA-damaging agents encode alternative regulatory subunits of ribonucleotide reductase. Genes Dev. 1990 May;4(5):740–751. doi: 10.1101/gad.4.5.740. [DOI] [PubMed] [Google Scholar]
- Emery H. S., Schild D., Kellogg D. E., Mortimer R. K. Sequence of RAD54, a Saccharomyces cerevisiae gene involved in recombination and repair. Gene. 1991 Jul 31;104(1):103–106. doi: 10.1016/0378-1119(91)90473-o. [DOI] [PubMed] [Google Scholar]
- Fornace A. J., Jr, Nebert D. W., Hollander M. C., Luethy J. D., Papathanasiou M., Fargnoli J., Holbrook N. J. Mammalian genes coordinately regulated by growth arrest signals and DNA-damaging agents. Mol Cell Biol. 1989 Oct;9(10):4196–4203. doi: 10.1128/mcb.9.10.4196. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Game J. C., Cox B. S. Synergistic interactions between rad mutations in yeast. Mutat Res. 1973 Oct;20(1):35–44. doi: 10.1016/0027-5107(73)90095-x. [DOI] [PubMed] [Google Scholar]
- Game J. C., Mortimer R. K. A genetic study of x-ray sensitive mutants in yeast. Mutat Res. 1974 Sep;24(3):281–292. doi: 10.1016/0027-5107(74)90176-6. [DOI] [PubMed] [Google Scholar]
- Hereford L. M., Osley M. A., Ludwig T. R., 2nd, McLaughlin C. S. Cell-cycle regulation of yeast histone mRNA. Cell. 1981 May;24(2):367–375. doi: 10.1016/0092-8674(81)90326-3. [DOI] [PubMed] [Google Scholar]
- Herskowitz I. Life cycle of the budding yeast Saccharomyces cerevisiae. Microbiol Rev. 1988 Dec;52(4):536–553. doi: 10.1128/mr.52.4.536-553.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hurd H. K., Roberts J. W. Upstream regulatory sequences of the yeast RNR2 gene include a repression sequence and an activation site that binds the RAP1 protein. Mol Cell Biol. 1989 Dec;9(12):5359–5372. doi: 10.1128/mcb.9.12.5359. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Johnston L. H. Periodic events in the cell cycle. Curr Opin Cell Biol. 1990 Apr;2(2):274–279. doi: 10.1016/0955-0674(90)90019-b. [DOI] [PubMed] [Google Scholar]
- Johnston L. H., White J. H., Johnson A. L., Lucchini G., Plevani P. Expression of the yeast DNA primase gene, PRI1, is regulated within the mitotic cell cycle and in meiosis. Mol Gen Genet. 1990 Mar;221(1):44–48. doi: 10.1007/BF00280366. [DOI] [PubMed] [Google Scholar]
- Johnston L. H., White J. H., Johnson A. L., Lucchini G., Plevani P. The yeast DNA polymerase I transcript is regulated in both the mitotic cell cycle and in meiosis and is also induced after DNA damage. Nucleic Acids Res. 1987 Jul 10;15(13):5017–5030. doi: 10.1093/nar/15.13.5017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Johnston R. F., Pickett S. C., Barker D. L. Autoradiography using storage phosphor technology. Electrophoresis. 1990 May;11(5):355–360. doi: 10.1002/elps.1150110503. [DOI] [PubMed] [Google Scholar]
- Jones J. S., Prakash L., Prakash S. Regulated expression of the Saccharomyces cerevisiae DNA repair gene RAD7 in response to DNA damage and during sporulation. Nucleic Acids Res. 1990 Jun 11;18(11):3281–3285. doi: 10.1093/nar/18.11.3281. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jones J. S., Prakash L. Transcript levels of the Saccharomyces cerevisiae DNA repair gene RAD18 increase in UV irradiated cells and during meiosis but not during the mitotic cell cycle. Nucleic Acids Res. 1991 Feb 25;19(4):893–898. doi: 10.1093/nar/19.4.893. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kans J. A., Mortimer R. K. Nucleotide sequence of the RAD57 gene of Saccharomyces cerevisiae. Gene. 1991 Aug 30;105(1):139–140. doi: 10.1016/0378-1119(91)90527-i. [DOI] [PubMed] [Google Scholar]
- Kawashima H., Horii T., Ogawa T., Ogawa H. Functional domains of Escherichia coli recA protein deduced from the mutational sites in the gene. Mol Gen Genet. 1984;193(2):288–292. doi: 10.1007/BF00330682. [DOI] [PubMed] [Google Scholar]
- Lipman D. J., Pearson W. R. Rapid and sensitive protein similarity searches. Science. 1985 Mar 22;227(4693):1435–1441. doi: 10.1126/science.2983426. [DOI] [PubMed] [Google Scholar]
- Lowndes N. F., Johnson A. L., Johnston L. H. Coordination of expression of DNA synthesis genes in budding yeast by a cell-cycle regulated trans factor. Nature. 1991 Mar 21;350(6315):247–250. doi: 10.1038/350247a0. [DOI] [PubMed] [Google Scholar]
- Madura K., Prakash S. Nucleotide sequence, transcript mapping, and regulation of the RAD2 gene of Saccharomyces cerevisiae. J Bacteriol. 1986 Jun;166(3):914–923. doi: 10.1128/jb.166.3.914-923.1986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Madura K., Prakash S., Prakash L. Expression of the Saccharomyces cerevisiae DNA repair gene RAD6 that encodes a ubiquitin conjugating enzyme, increases in response to DNA damage and in meiosis but remains constant during the mitotic cell cycle. Nucleic Acids Res. 1990 Feb 25;18(4):771–778. doi: 10.1093/nar/18.4.771. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Madura K., Prakash S. Transcript levels of the Saccharomyes cerevisiae DNA repair gene RAD23 increase in response to UV light and in meiosis but remain constant in the mitotic cell cycle. Nucleic Acids Res. 1990 Aug 25;18(16):4737–4742. doi: 10.1093/nar/18.16.4737. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Malone R. E., Esposito R. E. The RAD52 gene is required for homothallic interconversion of mating types and spontaneous mitotic recombination in yeast. Proc Natl Acad Sci U S A. 1980 Jan;77(1):503–507. doi: 10.1073/pnas.77.1.503. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McClanahan T., McEntee K. Specific transcripts are elevated in Saccharomyces cerevisiae in response to DNA damage. Mol Cell Biol. 1984 Nov;4(11):2356–2363. doi: 10.1128/mcb.4.11.2356. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McIntosh E. M., Atkinson T., Storms R. K., Smith M. Characterization of a short, cis-acting DNA sequence which conveys cell cycle stage-dependent transcription in Saccharomyces cerevisiae. Mol Cell Biol. 1991 Jan;11(1):329–337. doi: 10.1128/mcb.11.1.329. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Morrison D. P., Hastings P. J. Characterization of the mutator mutation mut5-1. Mol Gen Genet. 1979 Aug;175(1):57–65. doi: 10.1007/BF00267856. [DOI] [PubMed] [Google Scholar]
- Nakai S., Matsumoto S. Two types of radiation-sensitive mutant in yeast. Mutat Res. 1967 Mar-Apr;4(2):129–136. doi: 10.1016/0027-5107(67)90064-4. [DOI] [PubMed] [Google Scholar]
- Pabo C. O., Sauer R. T. Protein-DNA recognition. Annu Rev Biochem. 1984;53:293–321. doi: 10.1146/annurev.bi.53.070184.001453. [DOI] [PubMed] [Google Scholar]
- Pearson W. R. Rapid and sensitive sequence comparison with FASTP and FASTA. Methods Enzymol. 1990;183:63–98. doi: 10.1016/0076-6879(90)83007-v. [DOI] [PubMed] [Google Scholar]
- Peterson T. A., Prakash L., Prakash S., Osley M. A., Reed S. I. Regulation of CDC9, the Saccharomyces cerevisiae gene that encodes DNA ligase. Mol Cell Biol. 1985 Jan;5(1):226–235. doi: 10.1128/mcb.5.1.226. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Resnick M. A., Martin P. The repair of double-strand breaks in the nuclear DNA of Saccharomyces cerevisiae and its genetic control. Mol Gen Genet. 1976 Jan 16;143(2):119–129. doi: 10.1007/BF00266917. [DOI] [PubMed] [Google Scholar]
- Robinson G. W., Nicolet C. M., Kalainov D., Friedberg E. C. A yeast excision-repair gene is inducible by DNA damaging agents. Proc Natl Acad Sci U S A. 1986 Mar;83(6):1842–1846. doi: 10.1073/pnas.83.6.1842. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Roca A. I., Cox M. M. The RecA protein: structure and function. Crit Rev Biochem Mol Biol. 1990;25(6):415–456. doi: 10.3109/10409239009090617. [DOI] [PubMed] [Google Scholar]
- Rothstein R. Targeting, disruption, replacement, and allele rescue: integrative DNA transformation in yeast. Methods Enzymol. 1991;194:281–301. doi: 10.1016/0076-6879(91)94022-5. [DOI] [PubMed] [Google Scholar]
- Ruby S. W., Szostak J. W. Specific Saccharomyces cerevisiae genes are expressed in response to DNA-damaging agents. Mol Cell Biol. 1985 Jan;5(1):75–84. doi: 10.1128/mcb.5.1.75. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Saeki T., Machida I., Nakai S. Genetic control of diploid recovery after gamma-irradiation in the yeast Saccharomyces cerevisiae. Mutat Res. 1980 Dec;73(2):251–265. doi: 10.1016/0027-5107(80)90192-x. [DOI] [PubMed] [Google Scholar]
- Sanger F., Nicklen S., Coulson A. R. DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci U S A. 1977 Dec;74(12):5463–5467. doi: 10.1073/pnas.74.12.5463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schmitt M. E., Brown T. A., Trumpower B. L. A rapid and simple method for preparation of RNA from Saccharomyces cerevisiae. Nucleic Acids Res. 1990 May 25;18(10):3091–3092. doi: 10.1093/nar/18.10.3091. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sebastian J., Kraus B., Sancar G. B. Expression of the yeast PHR1 gene is induced by DNA-damaging agents. Mol Cell Biol. 1990 Sep;10(9):4630–4637. doi: 10.1128/mcb.10.9.4630. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Siede W., Robinson G. W., Kalainov D., Malley T., Friedberg E. C. Regulation of the RAD2 gene of Saccharomyces cerevisiae. Mol Microbiol. 1989 Dec;3(12):1697–1707. doi: 10.1111/j.1365-2958.1989.tb00155.x. [DOI] [PubMed] [Google Scholar]
- Sikorski R. S., Hieter P. A system of shuttle vectors and yeast host strains designed for efficient manipulation of DNA in Saccharomyces cerevisiae. Genetics. 1989 May;122(1):19–27. doi: 10.1093/genetics/122.1.19. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Struhl K. Nucleotide sequence and transcriptional mapping of the yeast pet56-his3-ded1 gene region. Nucleic Acids Res. 1985 Dec 9;13(23):8587–8601. doi: 10.1093/nar/13.23.8587. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Verma R., Patapoutian A., Gordon C. B., Campbell J. L. Identification and purification of a factor that binds to the Mlu I cell cycle box of yeast DNA replication genes. Proc Natl Acad Sci U S A. 1991 Aug 15;88(16):7155–7159. doi: 10.1073/pnas.88.16.7155. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Walker G. C. Mutagenesis and inducible responses to deoxyribonucleic acid damage in Escherichia coli. Microbiol Rev. 1984 Mar;48(1):60–93. doi: 10.1128/mr.48.1.60-93.1984. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Walker J. E., Saraste M., Runswick M. J., Gay N. J. Distantly related sequences in the alpha- and beta-subunits of ATP synthase, myosin, kinases and other ATP-requiring enzymes and a common nucleotide binding fold. EMBO J. 1982;1(8):945–951. doi: 10.1002/j.1460-2075.1982.tb01276.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weinert T. A., Hartwell L. H. The RAD9 gene controls the cell cycle response to DNA damage in Saccharomyces cerevisiae. Science. 1988 Jul 15;241(4863):317–322. doi: 10.1126/science.3291120. [DOI] [PubMed] [Google Scholar]
- White J. H., Barker D. G., Nurse P., Johnston L. H. Periodic transcription as a means of regulating gene expression during the cell cycle: contrasting modes of expression of DNA ligase genes in budding and fission yeast. EMBO J. 1986 Jul;5(7):1705–1709. doi: 10.1002/j.1460-2075.1986.tb04414.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wolff S., Afzal V., Wiencke J. K., Olivieri G., Michaeli A. Human lymphocytes exposed to low doses of ionizing radiations become refractory to high doses of radiation as well as to chemical mutagens that induce double-strand breaks in DNA. Int J Radiat Biol Relat Stud Phys Chem Med. 1988 Jan;53(1):39–47. doi: 10.1080/09553008814550401. [DOI] [PubMed] [Google Scholar]