Abstract
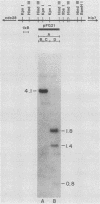
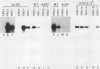
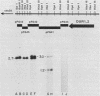
The DUR1,2 gene from Saccharomyces cerevisiae has been isolated on recombinant plasmids along with all DNA between the DUR1,2 and MET8 loci. DUR1,2 was found to encode a 5.7-kilobase transcript, which is consistent with our earlier suggestion that the DUR1 and DUR2 loci are two domains of a single multifunctional gene. Steady-state levels of the DUR1,2 transcript responded to induction and nitrogen catabolite repression in the same way as urea amidolyase activity. dal81 mutants (grown with inducer) contained barely detectable amounts of DUR1,2 RNA, whereas dal80 mutants (grown without inducer) contained the same amount as a wild-type induced culture. These observations support our earlier hypothesis that DUR1,2 is transcriptionally regulated, with control being mediated by the DAL80 and DAL81 gene products. We cloned the DUR1,2-Oh mutation and found it to be a Ty insertion near sequences required for complementation of dur1,2 mutations. The ROAM phenotype of the DUR1,2-Oh mutation is sharply different from that of cis-dominant, DUR80 mutations, which enhance DUR1,2 expression but do not affect the normal control pattern of the gene. There is evidence that DUR80 mutations may also be Ty insertions, which generate phenotypes that are different from those in DUR1,2-Oh mutations.
Full text
PDF


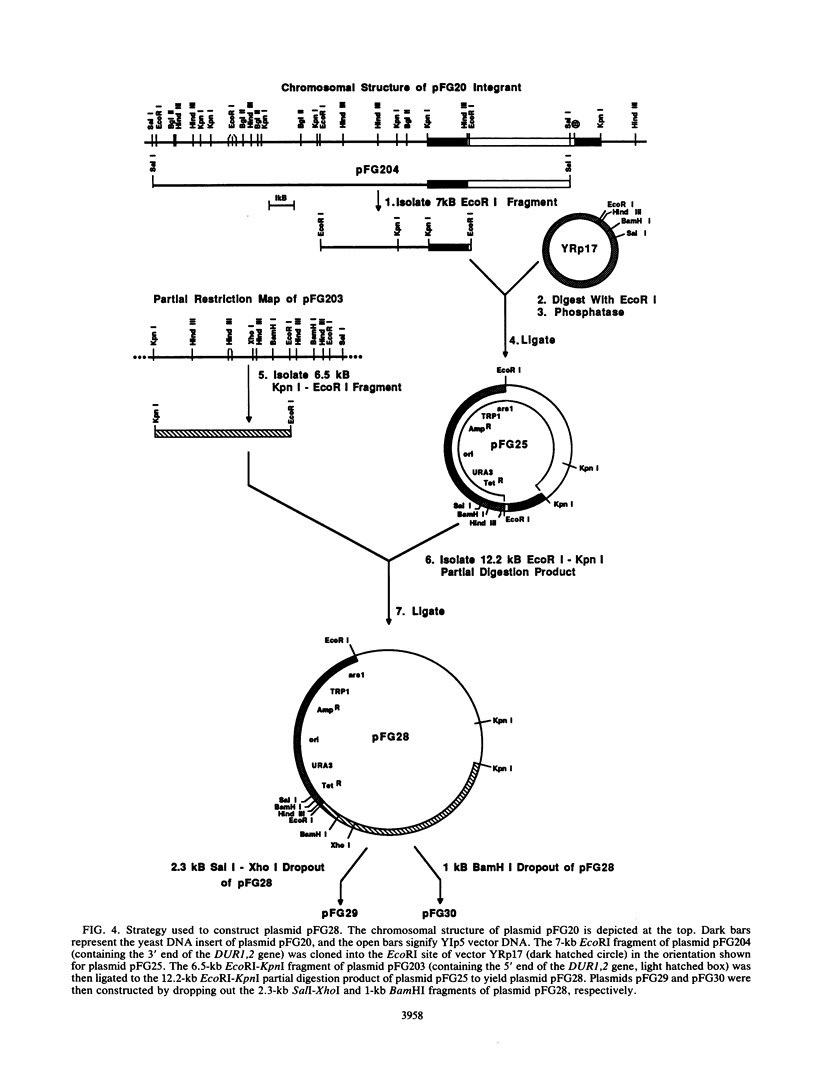
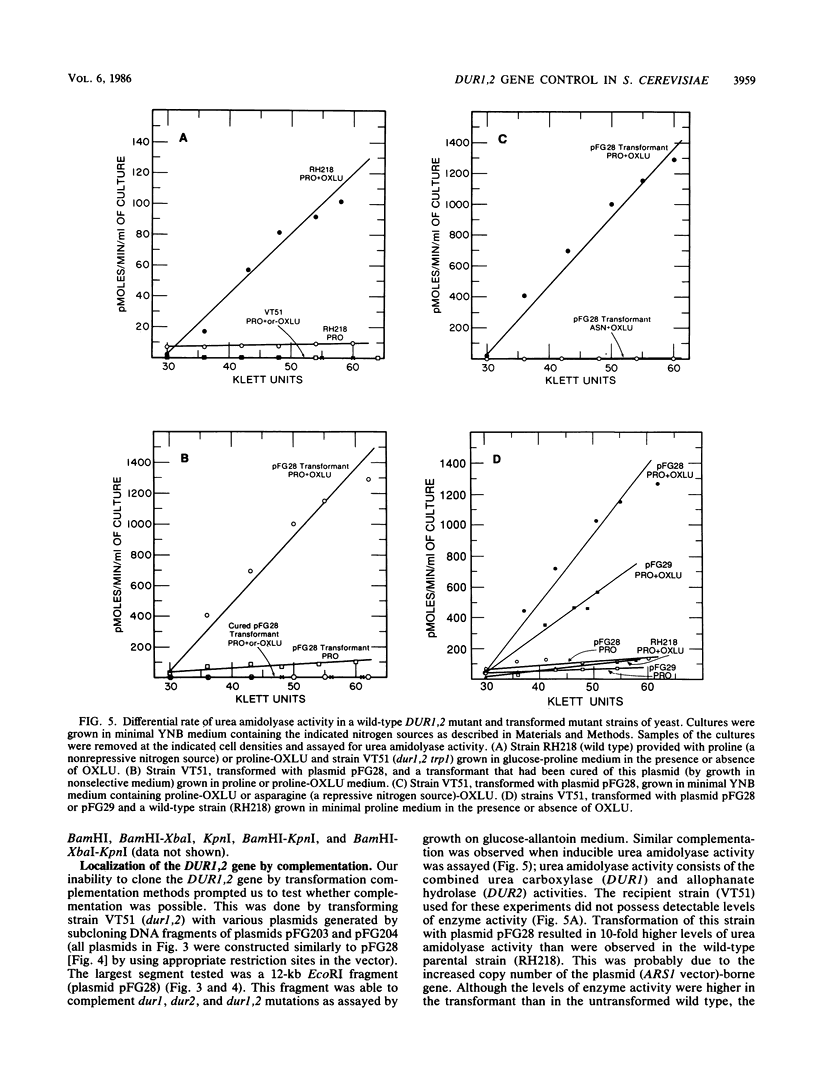





Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Bossinger J., Cooper T. G. Execution times of macromolecular synthetic processes involved in the induction of allophanate hydrolase at 15 degrees C. J Bacteriol. 1976 Oct;128(1):498–501. doi: 10.1128/jb.128.1.498-501.1976. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bossinger J., Cooper T. G. Sequence of molecular events involved in induction of allophanate hydrolase. J Bacteriol. 1976 Apr;126(1):198–204. doi: 10.1128/jb.126.1.198-204.1976. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bossinger J., Lawther R. P., Cooper T. G. Nitrogen repression of the allantoin degradative enzymes in Saccharomyces cerevisiae. J Bacteriol. 1974 Jun;118(3):821–829. doi: 10.1128/jb.118.3.821-829.1974. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bram R. J., Kornberg R. D. Specific protein binding to far upstream activating sequences in polymerase II promoters. Proc Natl Acad Sci U S A. 1985 Jan;82(1):43–47. doi: 10.1073/pnas.82.1.43. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cameron J. R., Loh E. Y., Davis R. W. Evidence for transposition of dispersed repetitive DNA families in yeast. Cell. 1979 Apr;16(4):739–751. doi: 10.1016/0092-8674(79)90090-4. [DOI] [PubMed] [Google Scholar]
- Carlson M., Botstein D. Two differentially regulated mRNAs with different 5' ends encode secreted with intracellular forms of yeast invertase. Cell. 1982 Jan;28(1):145–154. doi: 10.1016/0092-8674(82)90384-1. [DOI] [PubMed] [Google Scholar]
- Chisholm G. E., Genbauffe F. S., Cooper T. G. tau, a repeated DNA sequence in yeast. Proc Natl Acad Sci U S A. 1984 May;81(10):2965–2969. doi: 10.1073/pnas.81.10.2965. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chisholm G., Cooper T. G. Isolation and characterization of mutants that produce the allantoin-degrading enzymes constitutively in Saccharomyces cerevisiae. Mol Cell Biol. 1982 Sep;2(9):1088–1095. doi: 10.1128/mcb.2.9.1088. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chisholm G., Cooper T. cis-Dominant mutations which dramatically enhance DUR1,2 gene expression without affecting its normal regulation. Mol Cell Biol. 1984 May;4(5):947–955. doi: 10.1128/mcb.4.5.947. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cooper T. G., Bossinger J. Selective inhibition of protein synthesis initiation in Saccharomyces cerevisiae by low concentrations of cycloheximide. J Biol Chem. 1976 Nov 25;251(22):7278–7280. [PubMed] [Google Scholar]
- Cooper T. G., Lam C., Turoscy V. Structural analysis of the dur loci in S. cerevisiae: two domains of a single multifunctional gene. Genetics. 1980 Mar;94(3):555–580. doi: 10.1093/genetics/94.3.555. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cooper T. G., Lawther R. P. Induction of the allantoin degradative enzymes in Saccharomyces cerevisiae by the last intermediate of the pathway. Proc Natl Acad Sci U S A. 1973 Aug;70(8):2340–2344. doi: 10.1073/pnas.70.8.2340. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cooper T. G., Marcelli G., Sumrada R. Factors influencing the observed half-lives of specific synthetic capacities in Saccharomyces cerevisiae. Biochim Biophys Acta. 1978 Feb 16;517(2):464–472. doi: 10.1016/0005-2787(78)90213-7. [DOI] [PubMed] [Google Scholar]
- Cooper T. G. Mutants of Saccharomyces cerevisiae possessing fully induced levels of urea amido-lyase in the absence of added inducer. Biochem Biophys Res Commun. 1978 Jun 29;82(4):1258–1263. doi: 10.1016/0006-291x(78)90323-6. [DOI] [PubMed] [Google Scholar]
- Fasiolo F., Bonnet J., Lacroute F. Cloning of the yeast methionyl-tRNA synthetase gene. J Biol Chem. 1981 Mar 10;256(5):2324–2328. [PubMed] [Google Scholar]
- Fyrberg E. A., Kindle K. L., Davidson N., Kindle K. L. The actin genes of Drosophila: a dispersed multigene family. Cell. 1980 Feb;19(2):365–378. doi: 10.1016/0092-8674(80)90511-5. [DOI] [PubMed] [Google Scholar]
- Genbauffe F. S., Chisholm G. E., Cooper T. G. Tau, sigma, and delta. A family of repeated elements in yeast. J Biol Chem. 1984 Aug 25;259(16):10518–10525. [PubMed] [Google Scholar]
- Hinnen A., Hicks J. B., Fink G. R. Transformation of yeast. Proc Natl Acad Sci U S A. 1978 Apr;75(4):1929–1933. doi: 10.1073/pnas.75.4.1929. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hsiao C. L., Carbon J. High-frequency transformation of yeast by plasmids containing the cloned yeast ARG4 gene. Proc Natl Acad Sci U S A. 1979 Aug;76(8):3829–3833. doi: 10.1073/pnas.76.8.3829. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lawther R. P., Cooper T. G. Effects of inducer addition and removal upon the level of allophanate hydrolase in Saccharomyces cerevisiae. Biochem Biophys Res Commun. 1973 Dec 19;55(4):1100–1104. doi: 10.1016/s0006-291x(73)80008-7. [DOI] [PubMed] [Google Scholar]
- Lawther R. P., Cooper T. G. Kinetics of induced and repressed enzyme synthesis in Saccharomyces cerevisiae. J Bacteriol. 1975 Mar;121(3):1064–1073. doi: 10.1128/jb.121.3.1064-1073.1975. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lawther R. P., Phillips S. L., Cooper T. G. Lomofungin inhibition of allophanate hydrolase synthesis in Saccharomyces cerevisiae. Mol Gen Genet. 1975;137(2):89–99. doi: 10.1007/BF00341675. [DOI] [PubMed] [Google Scholar]
- Lemoine Y., Dubois E., Wiame J. M. The regulation of urea amidolyase of Saccharomyces cerevisiae: mating type influence on a constitutivity mutation acting in cis. Mol Gen Genet. 1978 Nov 9;166(3):251–258. [PubMed] [Google Scholar]
- McClure W. R. Mechanism and control of transcription initiation in prokaryotes. Annu Rev Biochem. 1985;54:171–204. doi: 10.1146/annurev.bi.54.070185.001131. [DOI] [PubMed] [Google Scholar]
- Orr-Weaver T. L., Szostak J. W., Rothstein R. J. Yeast transformation: a model system for the study of recombination. Proc Natl Acad Sci U S A. 1981 Oct;78(10):6354–6358. doi: 10.1073/pnas.78.10.6354. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Reznikoff W. S., Siegele D. A., Cowing D. W., Gross C. A. The regulation of transcription initiation in bacteria. Annu Rev Genet. 1985;19:355–387. doi: 10.1146/annurev.ge.19.120185.002035. [DOI] [PubMed] [Google Scholar]
- Struhl K., Stinchcomb D. T., Scherer S., Davis R. W. High-frequency transformation of yeast: autonomous replication of hybrid DNA molecules. Proc Natl Acad Sci U S A. 1979 Mar;76(3):1035–1039. doi: 10.1073/pnas.76.3.1035. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sumrada R. A., Cooper T. G. Isolation of the CAR1 gene from Saccharomyces cerevisiae and analysis of its expression. Mol Cell Biol. 1982 Dec;2(12):1514–1523. doi: 10.1128/mcb.2.12.1514. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sumrada R. A., Cooper T. G. Nucleotide sequence of the Saccharomyces cerevisiae arginase gene (CAR1) and its transcription under various physiological conditions. J Bacteriol. 1984 Dec;160(3):1078–1087. doi: 10.1128/jb.160.3.1078-1087.1984. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sumrada R. A., Cooper T. G. Urea carboxylase and allophanate hydrolase are components of a multifunctional protein in yeast. J Biol Chem. 1982 Aug 10;257(15):9119–9127. [PubMed] [Google Scholar]
- Sumrada R., Cooper T. G. Oxaluric acid: a non-metabolizable inducer of the allantoin degradative enzymes in Saccharomyces cerevisiae. J Bacteriol. 1974 Mar;117(3):1240–1247. doi: 10.1128/jb.117.3.1240-1247.1974. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Turoscy V., Cooper T. G. Pleiotropic control of five eucaryotic genes by multiple regulatory elements. J Bacteriol. 1982 Sep;151(3):1237–1246. doi: 10.1128/jb.151.3.1237-1246.1982. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Whitney P. A., Cooper T. G., Magasanik B. The induction of urea carboxylase and allophanate hydrolase in Saccharomyces cerevisiae. J Biol Chem. 1973 Sep 10;248(17):6203–6209. [PubMed] [Google Scholar]
- Wickerham L. J. A Critical Evaluation of the Nitrogen Assimilation Tests Commonly Used in the Classification of Yeasts. J Bacteriol. 1946 Sep;52(3):293–301. [PMC free article] [PubMed] [Google Scholar]
- Williamson V. M. Transposable elements in yeast. Int Rev Cytol. 1983;83:1–25. doi: 10.1016/s0074-7696(08)61684-8. [DOI] [PubMed] [Google Scholar]
- Winston F., Chumley F., Fink G. R. Eviction and transplacement of mutant genes in yeast. Methods Enzymol. 1983;101:211–228. doi: 10.1016/0076-6879(83)01016-2. [DOI] [PubMed] [Google Scholar]
- Yamamoto K. R. Steroid receptor regulated transcription of specific genes and gene networks. Annu Rev Genet. 1985;19:209–252. doi: 10.1146/annurev.ge.19.120185.001233. [DOI] [PubMed] [Google Scholar]
- Yoo H. S., Genbauffe F. S., Cooper T. G. Identification of the ureidoglycolate hydrolase gene in the DAL gene cluster of Saccharomyces cerevisiae. Mol Cell Biol. 1985 Sep;5(9):2279–2288. doi: 10.1128/mcb.5.9.2279. [DOI] [PMC free article] [PubMed] [Google Scholar]
- del Rey F. J., Donahue T. F., Fink G. R. sigma, a repetitive element found adjacent to tRNA genes of yeast. Proc Natl Acad Sci U S A. 1982 Jul;79(13):4138–4142. doi: 10.1073/pnas.79.13.4138. [DOI] [PMC free article] [PubMed] [Google Scholar]