SUMMARY
While several hundred regions of the human genome harbor signals of positive natural selection, few of the relevant adaptive traits and variants have been elucidated. Using full-genome sequence variation from the 1000 Genomes Project (1000G) and the Composite of Multiple Signals (CMS) test, we investigated 412 candidate signals and leveraged functional annotation, protein structure modeling, epigenetics, and association studies to identify and extensively annotate candidate causal variants. The resulting catalog provides a tractable list for experimental follow-up; it includes thirty-five high-scoring non-synonymous variants, fifty-nine variants associated with expression levels of a nearby coding gene or lincRNA, and numerous variants associated with susceptibility to infectious disease and other phenotypes. We experimentally characterized one candidate non-synonymous variant in TLR5, and show that it leads to altered NF-κB signaling in response to bacterial flagellin.
INTRODUCTION
Within their recent evolutionary history, humans traveled out of Africa into a wide range of new environments, endured repeated climate change, and experienced dramatic alterations in diet and disease risk, exposing them to new and powerful forces. Although many recent selective pressures have been hypothesized, only a handful of adaptive traits have ever been characterized, such as malaria resistance in HBB (Currat et al., 2002; Ohashi et al., 2004), lactose tolerance in LCT (Bersaglieri et al., 2004), skin pigmentation in SLC24A5 (Lamason et al., 2005), and high altitude tolerance in EPAS1 (Yi et al., 2010). Each of these began with knowledge of a phenotypic trait that was hypothesized to be adaptive, and subsequently genetic evidence for selection was discovered.
The advent of genomics holds great promise for the study of human evolution, making it possible to move from hypothesis-driven candidate gene studies to hypothesis-generating genome-wide scans. Over the last decade, genome-wide scans for selection have been frequently reported, finding several hundred loci that show patterns of variation characteristic of new beneficial mutations that have spread quickly through the population (Akey, 2009; Akey et al., 2002; Bustamante et al., 2005; Frazer et al., 2007; Pickrell et al., 2009; Sabeti et al., 2007; Voight et al., 2006; Williamson et al., 2007).
Moving from candidate genomic regions, however, to the underlying adaptive mutation has been difficult. The dearth of adaptive variants elucidated from genomic scans has led some to question whether many such variants remain to be identified. For example, a study of diversity and differentiation in 1000G data estimated that ~0.5% of nonsynonymous substitutions in the past 250,000 years have been subject to positive selection, and concluded that adaptive substitutions were rare in recent human history (Hernandez et al., 2011). However, this bound still allows for 340 nonsynonymous adaptive mutations in the 1000G data as well as countless regulatory mutations. The main reason so few examples have been identified is instead the difficulty of the problem. The regions detected are large, spanning 100s of kb to Mb and containing thousands of potential variants driving the signal, while the relevant phenotypic trait is unknown. Furthermore, full sequence data, necessary to compile a complete list of candidates, has not been available.
With this challenge in mind, we previously developed the method, the Composite of Multiple Signals (CMS), designed to pinpoint a small number of candidate selected variants within a large genomic region (Grossman et al., 2010). CMS combines several independent population genetic statistics that distinguish the causal variant from neighboring neutral variants, thus reducing the number of candidates by between 20- and 100-fold while maintaining high sensitivity for the causal variant. In that study, we applied it as a proof-of-concept to simulations and to published candidate genomic regions from the International Haplotype Map (HapMap) Project, but the work was fundamentally limited in its ability to find underlying causal variants by incomplete genotype data. One obtains far greater power to narrow regions with full sequence data. Furthermore, the sequence data ensures that all potential candidate variants are examined before pursuing functional validation.
Here we present the first catalog of candidate selected mutations, rather than genomic regions, using full genome sequencing data from the 1000G Project. The result is a tractable set of variants for follow-up functional characterization. Initial functional annotation reveals coding mutations as well as many variants in regulatory regions and non-coding RNAs, and associations with a variety of phenotypes. We use this database to identify a non-synonymous mutation in TLR5 with strong evidence for selection, and show it leads to altered NF-κB signaling in response to stimulation with bacterial flagellin. This example, together with another paper in this issue characterizing a selected variant in the Ectodysplasin receptor (EDAR), present a framework for moving from a genome-wide scan, to a tractable set of candidate variants, to insights from functional annotation, to characterization of a novel, population-specific functional variant and elucidation of distinct mechanisms of human evolutionary adaptation.
RESULTS
412 Fine-mapped Signals of Selection
The deep characterization of human sequence variation produced by the 1000G Project permits for the first time examination of the vast majority of nucleotides in the genome for evidence of recent evolution. The pilot phase of 1000G included complete, low-coverage (2-6x), whole-genome sequencing of 179 individuals from four populations: Yoruba individuals from Nigeria (YRI), European-ancestry individuals from Utah (CEU), Han Chinese individuals in Beijing (CHB) and Japanese individuals in Tokyo (JPT). The resulting dataset covers approximately 85% of the reference sequence and 93% of the coding sequence of the genome, with the vast majority (97%) of inaccessible sites being high-copy repeats or segmental duplications. Comparison with overlapping samples in independent studies indicates that the methodology used in the 1000G study has 90% power to detect single nucleotide polymorphisms (SNPs) present at 5% frequency in a population and 99% power for SNPs at 10% frequency or greater. The project also identified numerous insertion/deletion polymorphisms (indels). Because standard current methods primarily identify selective sweeps with causal mutations at high frequency (>20%) in at least one population, the causal variants for detectable loci under selection are almost certainly present in the 1000G sequence data.
We recently developed the CMS method to fine-map signals of natural selection within previously-identified candidate regions (Grossman et al., 2010); however we subsequently hypothesized that the method could be modified to also detect new candidate regions. We thus developed a genome-wide CMS method (CMSGW, Methods), and found that it was complementary to long-haplotype methods used to identify published candidate regions in the HapMap Project. Long-haplotype signals (generated as the selected mutation rapidly rises in frequency, bringing neighboring variants along with it) are very powerful for detecting recent sweeps (<30,000ya), but their power falls off significantly for older events due to haplotype breakdown. By incorporating signals that persist longer, such as population differentiation and high-frequency derived alleles, CMSGW is able to capture older events, for example selection at the DARC locus in West Africa, ca. 50,000 years ago.
We used CMSGW to identify 86 new regions likely to be under selection at an FDR of 19% (Figure 1, Figure S1, Table S1), and combined these with regions previously identified using long-haplotype methods (Frazer et al., 2007). Both kinds of tests have good power to detect incomplete sweeps (where the causal allele is not yet fixed in the population), compared with frequency-spectrum-based statistics such as Tajima’s D and Fay and Wu’s H. They are thus well suited to our focus on recent (< 50,000 years old) human selection events, for which the beneficial alleles are unlikely to have reached fixation.
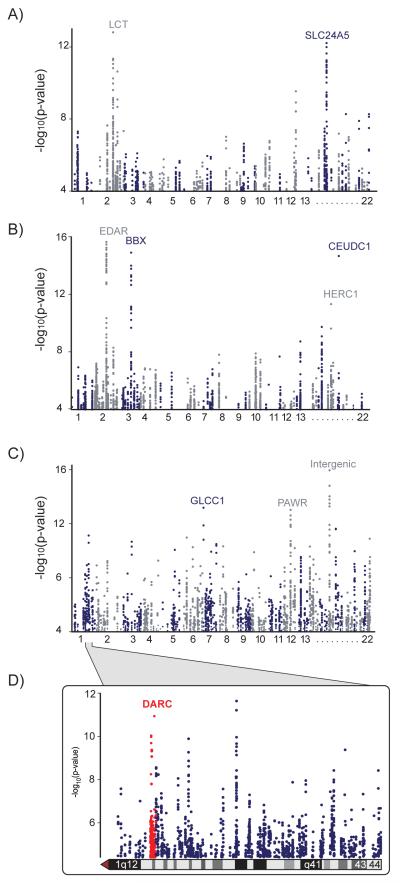
Figure 1. Genome-wide CMS.
CMSGW scores calculated from full genome sequence data from the 1000 Genomes Consortium in samples from (A) Northern Europe, (B) East Asia, and (C) West Africa. The y-axis represents the significance level (p-value on -log10 scale) for each of the variants tested across the genome, showing only variants with significance levels exceeding 10−4 (corresponding to 4 on the y-axis). See also Table S1.
With this combined set of candidate genomic regions in hand, we used our standard CMS implementation to fine-map signals and identify the candidate causal mutations within full sequence data (Grossman et al., 2010). The output was a set of 20-100 candidate variants per region (median=47), at a threshold that captured 90% of causal variants in simulations. The candidates lay within genomic regions with a median size of 27 kb (Table S2). As current functional annotation is incomplete, we included all candidate causal variants in our database, regardless of any prior knowledge of functionality.
A possible source of artifactual signals of selection is copy number variants (CNVs), which have been suggested to play a role in creating unusually long haplotypes (Gusev et al.). However, we identified only 60 instances of overlap between the localized regions and CNVs, not significantly higher than the number of overlaps expected at random (50, P = 0.23). While our analysis below focuses on SNPs, we catalogued these CNVs as they could themselves be targets of selection (Supplemental Information).
Among our fine-mapped candidates, 147 regions contain a single protein-coding gene, 88 regions contain multiple genes and 177 regions do not have any genes coding for known proteins. The regions are enriched for genes (P = 0.08) and coding variants (P < 0.01), and contain a number of genes involved in biological pathways thought to be recently targeted by selection, such as skin pigmentation and the immune system (Tables S3-S4). They also contain 48 long intergenic non-coding RNAs (lincRNAs) (Cabili et al., 2011), thirteen of which lie in regions with no protein-coding genes, suggesting another class of functional elements that may be a target of recent positive selection (Table S5).
Thirty-five Candidate Adaptive Non-synonymous Mutations
We examined the high-scoring variants and localized regions to identify both the biologically functional variants and the relevant pathways and phenotypes. We first focused on coding variation, and identified 35 high-scoring non-synonymous SNPs in 33 genes, of which seventeen are highly evolutionarily constrained (GERP score greater than 2.0) (Cooper et al., 2005) (Figure 2, Table S6). Of the 35, two have previously been characterized as adaptive mutations (in SCL24A5 and MATP, both associated with lighter skin pigmentation), and six have been associated with phenotypes in GWAS but not previously investigated as targets of selection. The latter include polymorphisms in EDAR (associated with hair thickness), ARHGEF3 (associated with greater bone mineral density), BTLA (rheumatoid arthritis), CTNS (cysteine metabolism defects), ITPR3 (type 1 diabetes and coronary aneurisms), and TLR5 (increased IFNγ secretion). We performed further structural homology modeling and conservation analysis, which pinpointed possible functional roles for SNPs in several genes including TLR5, ITGAE, and AP4B1.
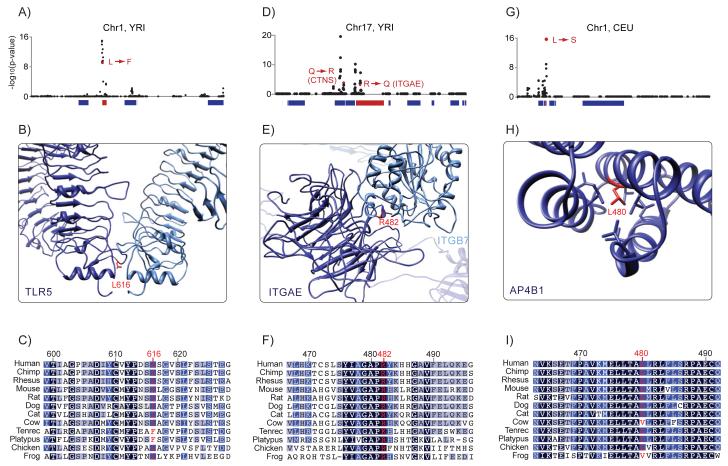
Figure 2. Candidate Non-Synonymous Mutations Identified by CMS.
CMS identified high-scoring non-synonymous mutations in the genes (A-C) TLR5, (D-F) ITGAE and (G-I) AP4B1. Panels (A), (D), and (G) show CMS scores for all variants in the regions. High-scoring non-synonymous variants and the genes in which they are located are presented in red. Panels (B), (E), and (H) show homology modeling of the genes with the residue containing the candidate variants in red. Panels (C), (F), and (I) show the amino-acid sequence in 12 vertebrate species. The color of the residue indicates the conservation score (darker color indicates greater conservation). See also Table S6.
It is striking that there are only 35 non-synonymous variants in our entire list of candidates. Based on the genomic coverage of the 1000G data, we estimate that there are no more than 38 candidate causal non-synonymous SNPs in the 412 candidate selected regions we analyzed (95% confidence interval, Methods). These data suggest that only a minority of recent adaptations are due to amino-acid changes, and that regulatory changes are likely to play a dominant role in recent human evolution.
Numerous Candidate Adaptive Regulatory Elements
To begin our investigation of potential regulatory changes in recent human evolution, we compiled several published eQTL studies in the 1000G individuals (Montgomery et al., 2010; Pickrell et al., 2010; Stranger et al., 2007), and identified candidate variants in the 412 regions that have been associated with differences in gene expression (Figure 3A, Table S7). We identified 56 regions containing SNPs associated with expression levels of nearby genes in lymphoblast cells, a twofold enrichment over the number expected by chance (P=0.02). In many of these cases, the top-scoring variants by CMS are the most strongly associated with expression levels. Fourteen high-scoring SNPs associated with expression differences lie in predicted transcription factor binding sites from the UCSC TFBS conserved track (Kent et al., 2002).
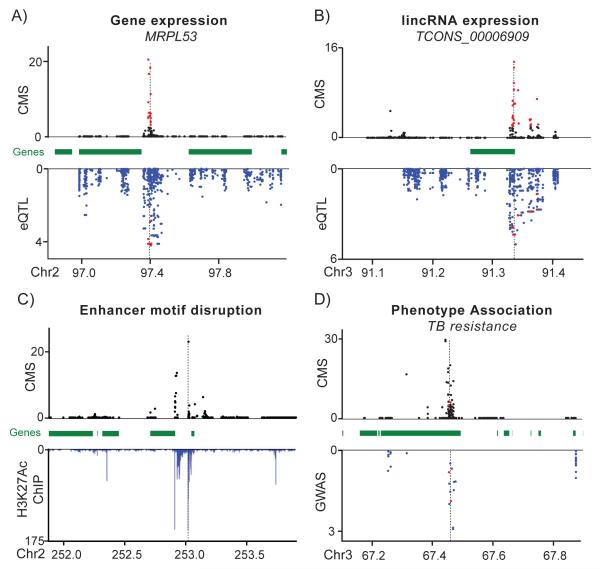
Figure 3. CMS Signals Overlapping Potential Regulatory Mutations, GWAS Signals, and Enhancers.
Example of candidate selected variants associated with gene expression (A) and lincRNA expression (B). CMS scores (top panel) and eQTL p-values (bottom panel) for all SNPs in the selected region. (C) Example of candidate selected variants that disrupt a putative enhancer element. CMS scores (top panel) and H3K27Ac ChIP-seq enrichment (bottom panel) from (Ernst et al., 2011). (D) Example of candidate selected variants associated with TB resistance. CMS scores (top panel) and association test p-values (bottom panel). Positions are given in centimorgans. High-scoring CMS variants that are with significant association scores are shown in red. See also Tables S7-S9.
We extended our study of putatively selected regulatory variants to those affecting expression levels of lincRNAs, using recently published RNA sequencing data from 1000G samples (Methods) (Montgomery et al., 2010; Pickrell et al., 2010). Three of the regions contain candidate variants under selection that are associated with differential expression of nearby lincRNAs (Figure 3B). To our knowledge, these three loci are the first candidates for recent selective pressure on mutations functionally affecting lincRNAs. LincRNAs themselves are often involved in orchestrating gene expression programs (Guttman et al., 2009; Guttman and Rinn, 2012).
Since available eQTL studies are somewhat underpowered and have only been carried out in lymphoblasts (likely missing many regulatory polymorphisms relevant in other cell types), we further used an alternative epigenetic-based strategy to identify potential regulatory variants. Using chromatin state predictions from (Ernst et al., 2011) we identified 335 SNPs (in 184 distinct candidate regions) that lie within predicted active enhancers or promoters and disrupt binding motifs of transcription factors known to be active in the cell type (Figure 3C, Table S8).
Numerous Candidate Adaptive Variations Associated with Phenotypes
In parallel to our investigation of functional variants on a molecular level, we also characterized potential phenotypes and pathways linked to the candidate variants. As natural selection can only act on mutations that drive phenotypic variation, we examined polymorphisms that have been associated with a variety of traits. Using the National Human Genome Research Institute (NHGRI) Genome-Wide Association Study (GWAS) (Hindorff et al., 2009) database, we found 165 overlaps with trait-associated SNPs, including eleven regions that contain variants associated with height and pigmentation and 79 regions associated with infectious and autoimmune disease susceptibility (Table S9) (Davila et al., 2010; Fellay et al., 2007; Ge et al., 2009; Jallow et al., 2009; Kamatani et al., 2009; Mbarek et al., 2011; Png et al., 2011; Zhang et al., 2009).
We more closely examined one example of these GWAS overlaps: susceptibility to the Mycobacterial pathogens M. tuberculosis and M. leprae, the causative agents of tuberculosis (TB) and leprosy respectively. These pathogens have been a major source of morbidity and mortality and represent a possible selective pressure in human populations. In collaboration with the Wellcome Trust Case Control Consortium, we analyzed a TB GWAS conducted in West Africa and a leprosy GWAS conducted in Northern India to find overlap between the localized selected regions and loci associated with TB and leprosy susceptibility (Thye et al., 2010; Wong et al., 2010) (see Methods). Five variants that are associated with resistance to these infectious diseases at P < 10−5 fall within the fine-mapped CMS-identified regions (Figure 3D, Table S4). The locus with the strongest association with resistance to leprosy (P = 1.25 × 10−6) contains SLC24A5, also known to influence skin pigmentation. Other loci suggested to be associated with leprosy include ALMS1, and with tuberculosis resistance include CCR9, CXCR4 and VDR, all of which play a role in immune response or pathogen binding (Liu et al., 2004).
A Candidate Adaptive Mutation in TLR5 Leads to Diminished NF-κB Signaling
We chose to functionally characterize one of the most promising non-synonymous candidates of selection, a leucine-to-phenylalanine variant (L616F) in TLR5, detected in the YRI. This is the only coding SNP in the TLR5 region with evidence of selection, and none of the high-scoring SNPs outside the ORF appear to affect TLR5 expression or disrupt regulatory regions. TLR5 is a well-described Toll-like receptor that plays a crucial role in the immunological clearance of bacterial pathogens. Its ligand, flagellin, is the principal component of the bacterial flagellum and is one of the most abundantly expressed proteins in nearly all flagellated bacteria. Receptor-ligand interaction of TLR5 activates cells of the immune system via the NF-κB pathway, leading to the production of various proinflammatory mediators. Several polymorphisms in TLR5 have been associated with differential responses to infectious diseases including Legionnaire’s disease (Hawn et al., 2003) and neonatal sepsis (Abu-Maziad et al., 2010).
Using structural modeling we found that the TLR5 L616F variant is predicted to be located in the conserved ectodomain responsible for dimerization and activation of the receptor (Figure 2B,C). To test the cellular effect of the TLR5 L616F mutation, we created stable cell lines carrying either the ancestral (L) or the derived (F) form of TLR5 in two different cell types, and measured NF-κB activation in the presence of increasing amounts of flagellin (Methods, Figure 4). In both cell types we found that the derived TLR5-616F allele produced significantly reduced NF-κB signaling in response to flagellin relative to the ancestral TLR5-616L allele (Figure 4C-D).
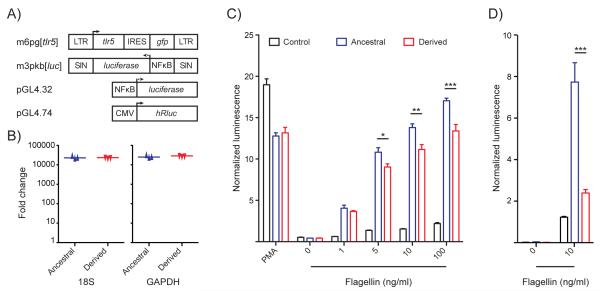
Figure 4. The Derived Form of TLR5 Leads to a Diminished NF-κB Response.
(A) Structure of retroviral vectors containing ancestral and derived TLR5 alleles and plasmid reporter construct transduced into 293FT and Jurkat cells. (B) Expression of TLR5 relative to non-transduced cells in transduced 293FT cells, given as normalized levels to either the 18S ribosomal subunit or GAPDH. (C) NF-κB reporter activity of 293FT cells transduced with ancestral or derived TLR5 allele or empty vector control 24 hours after stimulation with varying amounts of flagellin, normalized against the renilla luciferase signal. (D) NF-κB reporter activity of Jurkat cells transduced with ancestral or derived TLR5 allele or empty vector control 24 hours after stimulation with flagellin, normalized to non-specific activation with PMA/ionomycin. Control lane represents cells transduced with empty m6pg vector. Error-bars represent the SEM of at least three independent experiments.
DISCUSSION
The promise of the genomic age for elucidating human evolution has not yet been realized, in part due to the large size of regions identified as targets of selection, each of which can contain thousands of candidate causal variants, and in part due to the incompleteness of genotype data. Drawing on full genome sequence data from 1000G and on the CMS method, this paper presents the first comprehensive catalog of potential human adaptive mutations, instead of genomic regions. Each fine-mapped region contains 20-100 candidate variants, a small enough number to be tractable for functional characterization. As causal variants under selection typically have 10-50 perfect proxy variants, we are already near the limit of the power of population genetic tests to pinpoint the causal variant (Grossman et al., 2010). We computationally annotated all candidates and provide a proof-of-principle example of functional validation, creating a rich resource for future studies of human adaptations.
Many of the variants thus identified are associated with pathways that have emerged as targets of the strongest selective pressures on humans in recent history; the relevant traits include skin color, metabolism, and infectious disease resistance (Figure 5). In addition to the phenotypic associations and gene enrichment in these pathway discussed above, several of the eQTL SNPs regulate the expression of genes in these pathways, such as IVD, ACAS2, and CTNS (involved in metabolism) and BLK (involved in immune function). Many mutations fall in and around genes encoding the receptors or enzymes that modify the receptors for some of the most devastating pathogens in human history, including RHOA and OTUB1 (Yersinia pestis) (Edelmann et al., 2010) and DAG1 (Mycobacterium leprae), TLR5 (Salmonella typhimurium and others), LARGE (Lassa virus) (Kunz et al., 2005), DARC (Plasmodium vivax malaria) (Sabeti et al., 2006), PVRL4 (Measles virus), VDR (Mycobacterium tuberculosis), TPST1 (HIV) (Farzan et al., 1999), and CXCR4 (HIV). New pathways under selection are also coming to light: for example, in this issue of Cell Kamberov et al. elucidates selection on a nonsynonymous mutation in EDAR, which leads to a number of pleiotropic traits including altered hair and sweat gland formation.
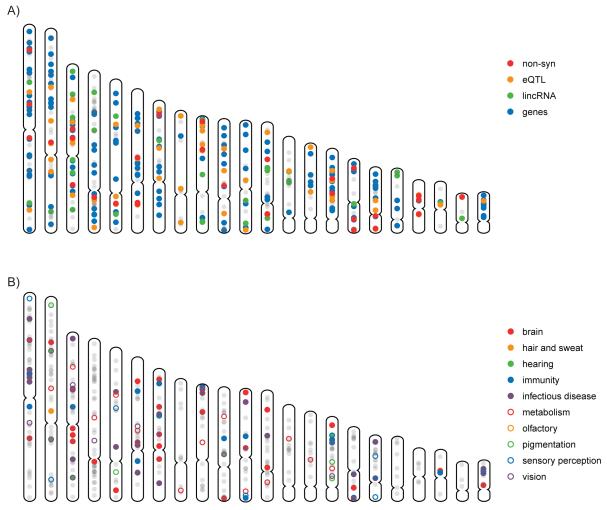
Figure 5. Characterization of Candidate Regions and Variants.
All candidate regions in the genome are shown in gray. (A) Candidate functional elements in localized regions, including regions with genes (blue), eQTLs (orange), long non-coding RNAs (green), and nonsynonymous variants (red). (B) Regions with genes relating to potential selective pressures, such as metabolism (red circle), infectious disease (purple), brain development (red), hearing (green), and hair and sweat (orange).
Our data support the mounting evidence that a great deal of recent human adaptation and phenotypic variation is based in regulatory regions (Hindorff et al., 2009; Lindblad-Toh et al., 2011; Vernot et al., 2012; Wang et al., 1995). Less than 10% of our fine-mapped regions contained high-scoring non-synonymous SNPs; candidate selected SNPs are enriched for eQTLs and include many mutations that disrupt transcription factor motifs in enhancers and promoters. Motifs for transcription factors involved in a number of different processes are disrupted, including STATs, Jun, GATAs, C/EBP, PPARγ, ETS, and IRFs. In several cases, the motif for a cell-specific transcription factor is disrupted in a cell-specific enhancer, for example an LXR:RXR motif in a hepatocyte-specific enhancer or a PU.1 motif in a monocyte-specific DNase HS site. The magnitude of the change in binding affinity varied from a minimum change in LOD score of 0.3 to a maximum of 12. More complete characterization of the regulatory variants using high-throughput cellular assays and eQTL studies in additional individuals may be illuminating.
Given the bounds on population genetic approaches to fine-map signals of selection and the limitations of current functional annotations, the true adaptive mutation must ultimately be distinguished using functional approaches. This is a challenge, especially for regions identified through genome-wide scans instead of based on a prior hypothesis of an adaptive pressure (e.g. malaria and lactose tolerance). It is impracticable to assay each variant in every possible cell type and process, and furthermore even functional variants need not be causal. While there is no way to prove what evolution did, even if we could go back in time to observe it, the standard in the evolutionary genomics field for establishing a mutation as having caused selection is strong statistical evidence of selection plus a phenotypic effect likely to enhance survival.
We chose one of the candidate variants, a nonsynonymous mutation in TLR5, to characterize experimentally. The derived allele in TLR5 with evidence of selection leads to diminished NF-κB signaling during bacterial infections. Intriguingly, another allele that decreases the function of TLR5 (a nonsense variant, TLR5-392STOP) has previously been reported to reach a frequency of 10% in European populations (Barreiro et al., 2009). The existence of common variants that decrease TLR5 signaling suggests that modulating TLR5 signaling may be advantageous in certain environments. Indeed, decreasing NF-κB signaling can have a protective effect in several bacterial infections, most significantly in bacterial sepsis (Koedel et al., 2000; Okugawa et al., 2006). Furthermore, the pathogen Salmonella typhimurium requires activated lamina propria cells (LPCs) in the intestinal epithelium to invade a host, and is consequently unable to infect mice with deficient TLR5 signaling (Uematsu et al., 2006). In a human population constantly exposed to high levels of bacterial antigens, a TLR5 variant with reduced NF-κB activation may well confer a fitness advantage.
An accompanying article from Kamberov et al. models an adaptive human variant of EDAR in mice, and characterizes its phenotype and evolutionary origins in humans. EDARV370A, one of the 35 nonsynonymous variants detected by CMS in 1000G data, likely emerged in central China ~30,000 years ago and leads to increased sweat gland number and scalp hair thickness in mice and humans. TLR5L616F and EDARV370A demonstrate the power of our framework to move from genomic scans to the characterization of a novel adaptive mutation and elucidation of distinct mechanisms of evolution.
This paper, in conjunction with the accompanying paper on EDAR, represents a decisive shift for the field of evolutionary genomics, moving from hypothesis-driven to hypothesis-generating science. We further provide a comprehensive list of candidate adaptive mutations driving recent human selective sweeps that lay the foundation for myriad future functional studies. The data from the 1000G Project, along with functional annotations, is available on a genome-wide browser, together with software to compute CMS on any dataset (http://www.broadinstitute.org/mpg/cms). In the years ahead, unprecedented data availability and collaborations across multiple disciplines from molecular, developmental, and computational biology to history and anthropology, promise to bring key recent events that have shaped our species to light.
EXPERIMENTAL PROCEDURES
Simulations
We used the simulations described earlier in Grossman et al. (Grossman et al., 2010), with one change: a coding error was fixed in the code that simulated gene conversion during a selective sweep (neutral simulations were unaffected).
CMS
Two versions of the CMS test were used: the original (within-region) CMS test (Grossman et al., 2010) for localizing the selected variant within a candidate region, and a modified test (denoted CMSGW) for identifying candidate regions within the genome.
When using CMS to localize regions, we used the distribution of neutral SNPs within 500kb of selected SNPs as the “unselected” distribution and assumed exactly one selected SNP per region. To use CMS as a genome-wide method to detect selected regions, we made the following modifications:
SNPs in neutral regions were used as the “unselected” distribution
- We did not assume any prior hypothesis about how many SNPs are under selection. Therefore instead of calculating the posterior probability, we calculated the Bayes factor for each test
and defined the composite score as the product of the Bayes factor of each test: Scores were normalized to neutral simulations (for simulated data) or to the whole genome (for real data), rather than within each region.
Bin boundaries were adjusted as described earlier.
We identified 100kb regions in which 30% of SNPs had a normalized score above 3, a threshold which corresponded to a 0.1% FPR in simulations (i.e. in 1000 neutral simulations of a 1MB region no more than 1 contained a 100KB region meeting this criterion; the upper bound of the 95% binomial confidence interval for the FPR is 0.6%), and used this threshold to detect selected regions in the 1000 Genomes data. We note that since the 1000G data includes 2.42Gb, we expect 24 false positive regions at this threshold.
The code used for CMS analysis is available at http://www.broadinstitute.org/mpg/cms.
1000G
Quality-controlled phased SNP and indel calls for the CEU, YRI and CHB+JPT populations released by the low-coverage portion of the 1000G project were downloaded from ftp://ftp.1000genomes.ebi.ac.uk/vol1/ftp/pilot_data/release/2010_03/pilot1/, representing the March 2010 data release. All genetic variants with more than two alleles were converted to biallelic variants, by mapping all alleles to two alleles while preserving alleles’ ancestral state where known. Ancestral state was taken from the ancestral state data released by the 1000 genomes project at ftp://ftp.1000genomes.ebi.ac.uk/vol1/ftp/pilot_data/technical/reference/ancestral_alignments, constructed from a four-way alignment of human, chimp, orangutan and rhesus macaque. Monomorphic SNPs omitted from 1000G data but present in HapMap Phase II data were added back into the data.
Gene Pathway Enrichment
To test for enrichment of different classes of functional variants, we picked random sets of 412 non-overlapping regions in the genome that matched our selected regions in size, and compared the number overlaps in our selected regions to the number in the random regions.
We also manually defined functional categories that previous literature suggests may have played an important role in recent human adaptation (e.g. skin pigmentation, immune system processes). We then used INRICHv1.0 to test for enrichment of these pathways. INRICH uses a two-step permutation algorithm to test for enrichment of pathways defined by the user or derived from published databases, and corrects the number of gene sets tested, the size of each geneset, and the number of SNPs and genes within our selected regions. See http://atgu.mgh.harvard.edu/inrich/ for more information.
Protein Structure Modeling
We used Modeller9v8 to generate homology models of proteins in which we found a high-scoring non-synonymous SNP. Sequences similar to the target sequence were selected as templates for homology modeling, and the optimum model was selected as the one with the lowest energy (DOPE) score. For toll-like receptor 5 (TLR5), we used a published computationally derived model of human TLR5, provided by Wei and colleagues (Wei et al., 2011).
Cell lines
Transgenes carrying either the ancestral (tlr5a; leucine) or derived (tlr5d; phenylalanine) form of TLR5 were synthesized and cloned into the retroviral vector m6pg carrying GFP as a transgene and transduced into 293FT and Jurkat cell to create stable cell lines. TLR5 expression was measured by qPCR.
293FT and Jurkat luciferase assays
NF-κB activity in 293FT cells was measured using pGL4.32 and pGL4.74 (Promega) and in Jurkat cells using the retroviral reporter m3pkb[luc] carrying an NF-κB inducible luciferase reporter (Loizou et al., 2011).
293FT cells expressing either the ancestral or derived forms of TLR5 were transfected with pGL4.32 and pGL4.74 (Promega). 26h after transfection the cells were stimulated for an additional 24 hours with 800 ng/mL PMA or increasing levels of flagellin at 1, 5, 10 or 100 ng/mL, and luciferase and renilla luminescence was measured in a Top Count machine.
293FT cells were transfected with 6.6 μg each of m3p[luc], retroviral packaging pCL-Eco and viral envelope VsV-g DNA. Cells were incubated for 20 hours and supernatant containing viral particles was collected.
The m3pkb[luc] viral supernatant and protamine sulfate were added to Jurkat cells stably expressing either the ancestral or derived forms of TLR5 and spun at 400 x g for 2 hours at 32°C. Plates were incubated for 26h (replacing media after 20h). Cells were stimulated for 24h with 400 ng/mL PMA and 1.5 μg/mL ionomycin or 10 ng/mL flagellin and firefly luminescence was measured.
Gene and lincRNA eQTL analysis
We obtained expression intensities of 47293 probes representing the majority of human genes from Stranger et al (Stranger et al., 2007), and downloaded the normalized gene expression levels for 22032 genes in the YRI individuals measured by RNA seq by Pickrell et al.(Pickrell et al., 2010)
To investigate lincRNA expression, RNA reads for YRI(Pickrell et al., 2010) were obtained from http://eqtl.uchicago.edu/RNA_Seq_data/ and for CEU(Montgomery et al., 2010) from http://jungle.unige.ch/rnaseq_CEU60/ and aligned to hg19 using BWA(Li and Durbin, 2009). Aligned reads were counted across the 4,421 previously detected regions of interest. Reads were RPKM normalized against both the length of the region and the total read count in the lane (Mortazavi et al., 2008) to provide a baseline expression level for each region. LincRNAs with non-zero expression in at least half of the individuals in a population were analyzed. All non-zero expression levels were quantile-normalized within each population in order to produce a normal distribution of expression.
Normalized read counts from each gene or lincRNA were tested as quantitive traits in a standard association test with SNPs 1 MB. SNPs with significant association p-values were overlapped with the selected regions.
TB and Leprosy Association Studies
TB susceptibility data was obtained from the Wellcome Trust Case Control Consortium study in the Gambia(Thye et al., 2010) with 1,498 confirmed TB cases and 1,496 controls, genotyped on the Affymetrix GeneChip 500K Array comprising 500,568 SNPs using the CHIAMO algorithm. The primary analysis focused on single-locus tests of association using 1,320 TB cases compared to 1,384 Gambian controls for all 405,226 SNPs passing QC filters with a study-wide MAF > 1%. The trend test was performed in a logistic regression modeling framework, which was adjusted for three axes of multi-dimensional scaling, by inclusion as covariates in the logistic regression model, reducing the over-dispersion of trend tests from λ = 1.13 (no adjustment) to λ = 1.05.
Leprosy susceptibility data was obtained from the host genetics study of leprosy in Indians(Wong et al., 2010) consisting of 258 confirmed cases of leprosy and 300 controls from New Delhi. All individuals in this study were genotyped with the Illumina IBC gene-centric 50K array. Multi-dimensional scaling (MDS) and principal component analysis (PCA) were carried out with PLINK and EIGENSTRAT to remove population outliers. A total of 209 leprosy cases and 239 controls were carried forward for analysis after quality control filters. The primary test of association in the New Delhi and Kolkata cohorts was carried out with the Pearson’s χ2 allelic test, Cochran-Armitage trend test and logistic regression.
Supplementary Material
HIGHLIGHTS.
Genome-wide scan in 1000 Genomes sequence data fine-maps 412 signals of selection
Adaptive candidates include 35 non-synonymous and 59 loci with eQTLs
L→F variant in TLR5 reduces NF-κB signaling in response to bacterial flagellin
Catalog provides a tractable set of selected variants for experimental follow-up
ACKNOWLEDGEMENTS
PCS and her lab are supported by a Burroughs Wellcome Fund Career Award, a Packard Foundation Fellowship in Science and Engineering, a Broad Institute SPARC award, an NIH Innovator Award 1DP2OD006514-01, and BAA-NIAID-DAIT-NIHAI2009061. SRG is supported by NIGMS T32GM007753, KGA by a Carlsberg Foundation fellowship, DJP by NSF, and EKK by an American Cancer Society Fellowship. The mycobacterial disease studies analyzed were supported by funding from the Wellcome Trust, the UK Medical Research Council, the UK National Institute for Health Research and the European Commission; we thank the many collaborators who contributed to generating these datasets. We would like to thank S. Hart for help with figures, C. Edwards for reviews of the text, and L. Ward and C. O’Dushlaine for technical guidance.
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
REFERENCES
- A map of human genome variation from population-scale sequencing. Nature. 467:1061–1073. doi: 10.1038/nature09534. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Abu-Maziad A, Schaa K, Bell EF, Dagle JM, Cooper M, Marazita ML, Murray JC. Role of polymorphic variants as genetic modulators of infection in neonatal sepsis. Pediatr Res. 2010;68:323–329. doi: 10.1203/PDR.0b013e3181e6a068. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Akey JM. Constructing genomic maps of positive selection in humans: where do we go from here? Genome research. 2009;19:711–722. doi: 10.1101/gr.086652.108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Akey JM, Zhang G, Zhang K, Jin L, Shriver MD. Interrogating a high-density SNP map for signatures of natural selection. Genome research. 2002;12:1805–1814. doi: 10.1101/gr.631202. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Barreiro LB, Ben-Ali M, Quach H, Laval G, Patin E, Pickrell JK, Bouchier C, Tichit M, Neyrolles O, Gicquel B, et al. Evolutionary dynamics of human Toll-like receptors and their different contributions to host defense. PLoS genetics. 2009;5:e1000562. doi: 10.1371/journal.pgen.1000562. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bersaglieri T, Sabeti PC, Patterson N, Vanderploeg T, Schaffner SF, Drake JA, Rhodes M, Reich DE, Hirschhorn JN. Genetic signatures of strong recent positive selection at the lactase gene. Am J Hum Genet. 2004;74:1111–1120. doi: 10.1086/421051. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bustamante CD, Fledel-Alon A, Williamson S, Nielsen R, Hubisz MT, Glanowski S, Tanenbaum DM, White TJ, Sninsky JJ, Hernandez RD, et al. Natural selection on protein-coding genes in the human genome. Nature. 2005;437:1153–1157. doi: 10.1038/nature04240. [DOI] [PubMed] [Google Scholar]
- Cabili MN, Trapnell C, Goff L, Koziol M, Tazon-Vega B, Regev A, Rinn JL. Integrative annotation of human large intergenic noncoding RNAs reveals global properties and specific subclasses. Genes & development. 2011;25:1915–1927. doi: 10.1101/gad.17446611. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cooper GM, Stone EA, Asimenos G, Green ED, Batzoglou S, Sidow A. Distribution and intensity of constraint in mammalian genomic sequence. Genome research. 2005;15:901–913. doi: 10.1101/gr.3577405. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Currat M, Trabuchet G, Rees D, Perrin P, Harding RM, Clegg JB, Langaney A, Excoffier L. Molecular analysis of the beta-globin gene cluster in the Niokholo Mandenka population reveals a recent origin of the beta(S) Senegal mutation. Am J Hum Genet. 2002;70:207–223. doi: 10.1086/338304. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Davila S, Wright VJ, Khor CC, Sim KS, Binder A, Breunis WB, Inwald D, Nadel S, Betts H, Carrol ED, et al. Genome-wide association study identifies variants in the CFH region associated with host susceptibility to meningococcal disease. Nat Genet. 2010;42:772–776. doi: 10.1038/ng.640. [DOI] [PubMed] [Google Scholar]
- Edelmann MJ, Kramer HB, Altun M, Kessler BM. Post-translational modification of the deubiquitinating enzyme otubain 1 modulates active RhoA levels and susceptibility to Yersinia invasion. FEBS J. 2010;277:2515–2530. doi: 10.1111/j.1742-4658.2010.07665.x. [DOI] [PubMed] [Google Scholar]
- Ernst J, Kheradpour P, Mikkelsen TS, Shoresh N, Ward LD, Epstein CB, Zhang X, Wang L, Issner R, Coyne M, et al. Mapping and analysis of chromatin state dynamics in nine human cell types. Nature. 2011;473:43–49. doi: 10.1038/nature09906. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Farzan M, Mirzabekov T, Kolchinsky P, Wyatt R, Cayabyab M, Gerard NP, Gerard C, Sodroski J, Choe H. Tyrosine sulfation of the amino terminus of CCR5 facilitates HIV-1 entry. Cell. 1999;96:667–676. doi: 10.1016/s0092-8674(00)80577-2. [DOI] [PubMed] [Google Scholar]
- Fellay J, Shianna KV, Ge D, Colombo S, Ledergerber B, Weale M, Zhang K, Gumbs C, Castagna A, Cossarizza A, et al. A whole-genome association study of major determinants for host control of HIV-1. Science. 2007;317:944–947. doi: 10.1126/science.1143767. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Frazer KA, Ballinger DG, Cox DR, Hinds DA, Stuve LL, Gibbs RA, Belmont JW, Boudreau A, Hardenbol P, Leal SM, et al. A second generation human haplotype map of over 3.1 million SNPs. Nature. 2007;449:851–861. doi: 10.1038/nature06258. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ge D, Fellay J, Thompson AJ, Simon JS, Shianna KV, Urban TJ, Heinzen EL, Qiu P, Bertelsen AH, Muir AJ, et al. Genetic variation in IL28B predicts hepatitis C treatment-induced viral clearance. Nature. 2009;461:399–401. doi: 10.1038/nature08309. [DOI] [PubMed] [Google Scholar]
- Grossman SR, Shlyakhter I, Karlsson EK, Byrne EH, Morales S, Frieden G, Hostetter E, Angelino E, Garber M, Zuk O, et al. A composite of multiple signals distinguishes causal variants in regions of positive selection. Science. 2010;327:883–886. doi: 10.1126/science.1183863. [DOI] [PubMed] [Google Scholar]
- Gusev A, Palamara PF, Aponte G, Zhuang Z, Darvasi A, Gregersen P, Pe’er I. The Architecture of Long-Range Haplotypes Shared within and across Populations. Molecular biology and evolution. 2012;29:473–486. doi: 10.1093/molbev/msr133. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Guttman M, Amit I, Garber M, French C, Lin MF, Feldser D, Huarte M, Zuk O, Carey BW, Cassady JP, et al. Chromatin signature reveals over a thousand highly conserved large non-coding RNAs in mammals. Nature. 2009;458:223–227. doi: 10.1038/nature07672. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Guttman M, Rinn JL. Modular regulatory principles of large non-coding RNAs. Nature. 2012;482:339–346. doi: 10.1038/nature10887. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hawn TR, Verbon A, Lettinga KD, Zhao LP, Li SS, Laws RJ, Skerrett SJ, Beutler B, Schroeder L, Nachman A, et al. A common dominant TLR5 stop codon polymorphism abolishes flagellin signaling and is associated with susceptibility to legionnaires’ disease. J Exp Med. 2003;198:1563–1572. doi: 10.1084/jem.20031220. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hernandez RD, Kelley JL, Elyashiv E, Melton SC, Auton A, McVean G, Sella G, Przeworski M. Classic selective sweeps were rare in recent human evolution. Science. 2011;331:920–924. doi: 10.1126/science.1198878. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hindorff LA, Sethupathy P, Junkins HA, Ramos EM, Mehta JP, Collins FS, Manolio TA. Potential etiologic and functional implications of genome-wide association loci for human diseases and traits. Proc Natl Acad Sci U S A. 2009;106:9362–9367. doi: 10.1073/pnas.0903103106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jallow M, Teo YY, Small KS, Rockett KA, Deloukas P, Clark TG, Kivinen K, Bojang KA, Conway DJ, Pinder M, et al. Genome-wide and fine-resolution association analysis of malaria in West Africa. Nat Genet. 2009;41:657–665. doi: 10.1038/ng.388. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kamatani Y, Wattanapokayakit S, Ochi H, Kawaguchi T, Takahashi A, Hosono N, Kubo M, Tsunoda T, Kamatani N, Kumada H, et al. A genome-wide association study identifies variants in the HLA-DP locus associated with chronic hepatitis B in Asians. Nat Genet. 2009;41:591–595. doi: 10.1038/ng.348. [DOI] [PubMed] [Google Scholar]
- Kent WJ, Sugnet CW, Furey TS, Roskin KM, Pringle TH, Zahler AM, Haussler D. The human genome browser at UCSC. Genome research. 2002;12:996–1006. doi: 10.1101/gr.229102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Koedel U, Bayerlein I, Paul R, Sporer B, Pfister HW. Pharmacologic interference with NF-kappaB activation attenuates central nervous system complications in experimental Pneumococcal meningitis. The Journal of infectious diseases. 2000;182:1437–1445. doi: 10.1086/315877. [DOI] [PubMed] [Google Scholar]
- Kunz S, Rojek JM, Kanagawa M, Spiropoulou CF, Barresi R, Campbell KP, Oldstone MB. Posttranslational modification of alpha-dystroglycan, the cellular receptor for arenaviruses, by the glycosyltransferase LARGE is critical for virus binding. J Virol. 2005;79:14282–14296. doi: 10.1128/JVI.79.22.14282-14296.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lamason RL, Mohideen MA, Mest JR, Wong AC, Norton HL, Aros MC, Jurynec MJ, Mao X, Humphreville VR, Humbert JE, et al. SLC24A5, a putative cation exchanger, affects pigmentation in zebrafish and humans. Science. 2005;310:1782–1786. doi: 10.1126/science.1116238. [DOI] [PubMed] [Google Scholar]
- Li H, Durbin R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics. 2009;25:1754–1760. doi: 10.1093/bioinformatics/btp324. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lindblad-Toh K, Garber M, Zuk O, Lin MF, Parker BJ, Washietl S, Kheradpour P, Ernst J, Jordan G, Mauceli E, et al. A high-resolution map of human evolutionary constraint using 29 mammals. Nature. 2011;478:476–482. doi: 10.1038/nature10530. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu W, Cao WC, Zhang CY, Tian L, Wu XM, Habbema JD, Zhao QM, Zhang PH, Xin ZT, Li CZ, et al. VDR and NRAMP1 gene polymorphisms in susceptibility to pulmonary tuberculosis among the Chinese Han population: a case-control study. Int J Tuberc Lung Dis. 2004;8:428–434. [PubMed] [Google Scholar]
- Loizou L, Andersen KG, Betz AG. Foxp3 interacts with c-Rel to mediate NF-kappaB repression. PloS one. 2011;6:e18670. doi: 10.1371/journal.pone.0018670. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mbarek H, Ochi H, Urabe Y, Kumar V, Kubo M, Hosono N, Takahashi A, Kamatani Y, Miki D, Abe H, et al. A genome-wide association study of chronic hepatitis B identified novel risk locus in a Japanese population. Hum Mol Genet. 2011 doi: 10.1093/hmg/ddr301. [DOI] [PubMed] [Google Scholar]
- Montgomery SB, Sammeth M, Gutierrez-Arcelus M, Lach RP, Ingle C, Nisbett J, Guigo R, Dermitzakis ET. Transcriptome genetics using second generation sequencing in a Caucasian population. Nature. 2010;464:773–777. doi: 10.1038/nature08903. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mortazavi A, Williams BA, McCue K, Schaeffer L, Wold B. Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat Methods. 2008;5:621–628. doi: 10.1038/nmeth.1226. [DOI] [PubMed] [Google Scholar]
- Ohashi J, Naka I, Patarapotikul J, Hananantachai H, Brittenham G, Looareesuwan S, Clark AG, Tokunaga K. Extended linkage disequilibrium surrounding the hemoglobin E variant due to malarial selection. Am J Hum Genet. 2004;74:1198–1208. doi: 10.1086/421330. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Okugawa S, Yanagimoto S, Tsukada K, Kitazawa T, Koike K, Kimura S, Nagase H, Hirai K, Ota Y. Bacterial flagellin inhibits T cell receptor-mediated activation of T cells by inducing suppressor of cytokine signalling-1 (SOCS-1) Cellular microbiology. 2006;8:1571–1580. doi: 10.1111/j.1462-5822.2006.00731.x. [DOI] [PubMed] [Google Scholar]
- Pickrell JK, Coop G, Novembre J, Kudaravalli S, Li JZ, Absher D, Srinivasan BS, Barsh GS, Myers RM, Feldman MW, et al. Signals of recent positive selection in a worldwide sample of human populations. Genome research. 2009;19:826–837. doi: 10.1101/gr.087577.108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pickrell JK, Marioni JC, Pai AA, Degner JF, Engelhardt BE, Nkadori E, Veyrieras JB, Stephens M, Gilad Y, Pritchard JK. Understanding mechanisms underlying human gene expression variation with RNA sequencing. Nature. 2010;464:768–772. doi: 10.1038/nature08872. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Png E, Thalamuthu A, Ong RT, Snippe H, Boland GJ, Seielstad M. A genome-wide association study of hepatitis B vaccine response in an Indonesian population reveals multiple independent risk variants in the HLA region. Hum Mol Genet. 2011 doi: 10.1093/hmg/ddr302. [DOI] [PubMed] [Google Scholar]
- Sabeti PC, Schaffner SF, Fry B, Lohmueller J, Varilly P, Shamovsky O, Palma A, Mikkelsen TS, Altshuler D, Lander ES. Positive natural selection in the human lineage. Science. 2006;312:1614–1620. doi: 10.1126/science.1124309. [DOI] [PubMed] [Google Scholar]
- Sabeti PC, Varilly P, Fry B, Lohmueller J, Hostetter E, Cotsapas C, Xie X, Byrne EH, McCarroll SA, Gaudet R, et al. Genome-wide detection and characterization of positive selection in human populations. Nature. 2007;449:913–918. doi: 10.1038/nature06250. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stranger BE, Nica AC, Forrest MS, Dimas A, Bird CP, Beazley C, Ingle CE, Dunning M, Flicek P, Koller D, et al. Population genomics of human gene expression. Nat Genet. 2007;39:1217–1224. doi: 10.1038/ng2142. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thye T, Vannberg FO, Wong SH, Owusu-Dabo E, Osei I, Gyapong J, Sirugo G, Sisay-Joof F, Enimil A, Chinbuah MA, et al. Genome-wide association analyses identifies a susceptibility locus for tuberculosis on chromosome 18q11.2. Nat Genet. 2010;42:739–741. doi: 10.1038/ng.639. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Uematsu S, Jang MH, Chevrier N, Guo Z, Kumagai Y, Yamamoto M, Kato H, Sougawa N, Matsui H, Kuwata H, et al. Detection of pathogenic intestinal bacteria by Toll-like receptor 5 on intestinal CD11c+ lamina propria cells. Nat Immunol. 2006;7:868–874. doi: 10.1038/ni1362. [DOI] [PubMed] [Google Scholar]
- Vernot B, Stergachis AB, Maurano MT, Vierstra J, Neph S, Thurman RE, Stamatoyannopoulos JA, Akey JM. Personal and population genomics of human regulatory variation. Genome research. 2012;22:1689–1697. doi: 10.1101/gr.134890.111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Voight BF, Kudaravalli S, Wen X, Pritchard JK. A map of recent positive selection in the human genome. PLoS biology. 2006;4:e72. doi: 10.1371/journal.pbio.0040072. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang Y, Harvey CB, Pratt WS, Sams VR, Sarner M, Rossi M, Auricchio S, Swallow DM. The lactase persistence/non-persistence polymorphism is controlled by a cis-acting element. Hum Mol Genet. 1995;4:657–662. doi: 10.1093/hmg/4.4.657. [DOI] [PubMed] [Google Scholar]
- Wei T, Gong J, Rossle SC, Jamitzky F, Heckl WM, Stark RW. A leucine-rich repeat assembly approach for homology modeling of the human TLR5-10 and mouse TLR11-13 ectodomains. J Mol Model. 2011;17:27–36. doi: 10.1007/s00894-010-0697-5. [DOI] [PubMed] [Google Scholar]
- Williamson SH, Hubisz MJ, Clark AG, Payseur BA, Bustamante CD, Nielsen R. Localizing recent adaptive evolution in the human genome. PLoS genetics. 2007;3:e90. doi: 10.1371/journal.pgen.0030090. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wong SH, Gochhait S, Malhotra D, Pettersson FH, Teo YY, Khor CC, Rautanen A, Chapman SJ, Mills TC, Srivastava A, et al. Leprosy and the adaptation of human toll-like receptor 1. PLoS Pathog. 2010;6:e1000979. doi: 10.1371/journal.ppat.1000979. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yi X, Liang Y, Huerta-Sanchez E, Jin X, Cuo ZX, Pool JE, Xu X, Jiang H, Vinckenbosch N, Korneliussen TS, et al. Sequencing of 50 human exomes reveals adaptation to high altitude. Science. 2010;329:75–78. doi: 10.1126/science.1190371. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang FR, Huang W, Chen SM, Sun LD, Liu H, Li Y, Cui Y, Yan XX, Yang HT, Yang RD, et al. Genomewide association study of leprosy. N Engl J Med. 2009;361:2609–2618. doi: 10.1056/NEJMoa0903753. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.