Abstract
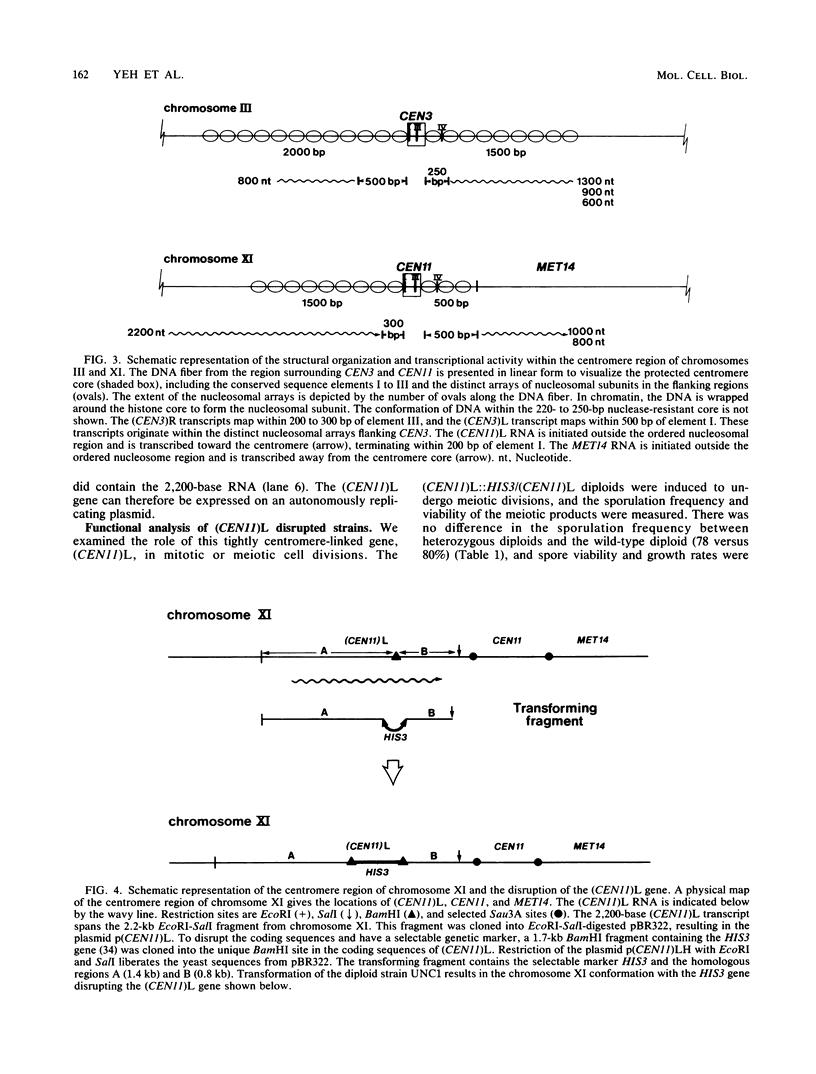
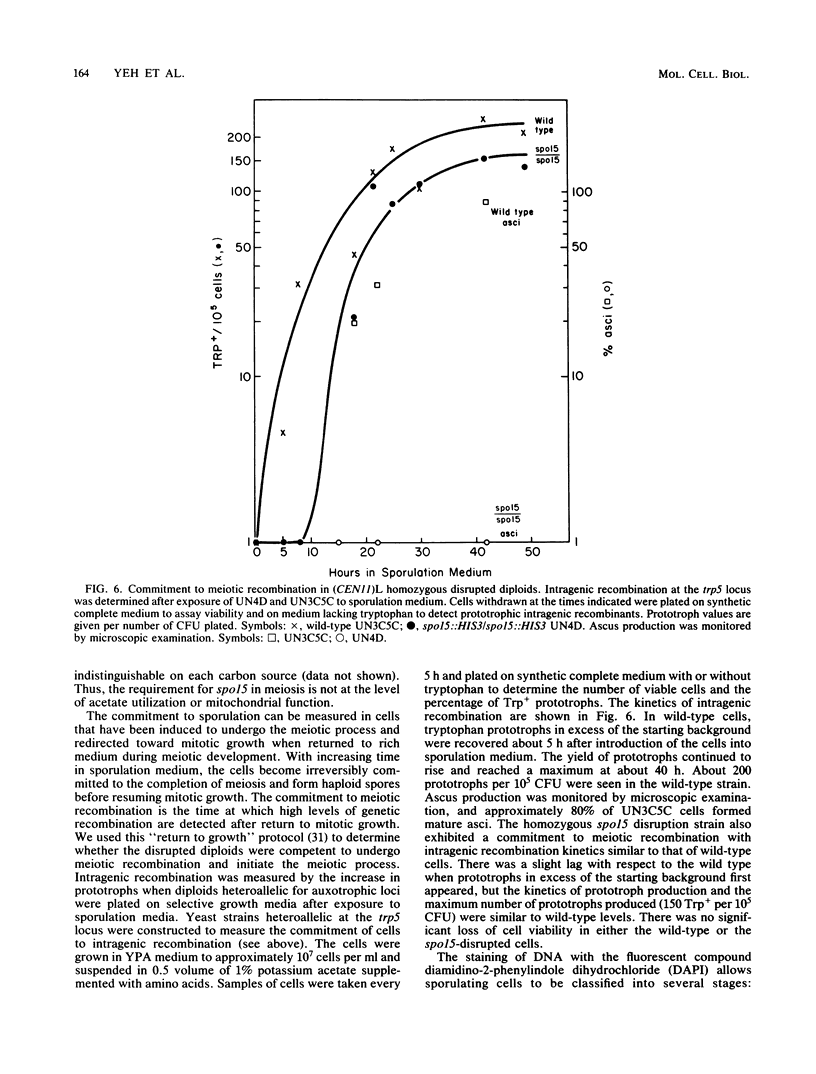
We used DNA fragments from the centromere regions of yeast (Saccharomyces cerevisiae) chromosomes III and XI to examine the transcriptional activity within this chromosomal domain. DNA transcripts were found 200 to 300 base pairs from the 250-base-pair centromere core and lie within an ordered chromatin array. No transcripts were detected from the functional centromere region. We examined the cellular function of one of these tightly centromere-linked transcripts. (CEN11)L, by disrupting the coding sequences in vivo and analyzing the phenotype of the mutant yeast cell. Diploids heterozygous for the (CEN11)L disruption sporulated at wild-type levels, and the absence of the (CEN11)L gene product had no effect on the viability or mitotic growth of haploid cells. Diploids homozygous for the (CEN11)L disruption were unable to sporulate when induced by the appropriate nutritional cues. The mutant cells were competent for intragenic recombination and appeared to be blocked at the mononucleate stage. The temporal ordering of (CEN11)L function with respect to the sporulation mutant spo13 suggests that the (CEN11)L gene product may be required at both the first and second meiotic cell divisions. This new sporulation gene has been termed SPO15.
Full text
PDF







Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Bloom K. S., Amaya E., Carbon J., Clarke L., Hill A., Yeh E. Chromatin conformation of yeast centromeres. J Cell Biol. 1984 Nov;99(5):1559–1568. doi: 10.1083/jcb.99.5.1559. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bloom K. S., Carbon J. Yeast centromere DNA is in a unique and highly ordered structure in chromosomes and small circular minichromosomes. Cell. 1982 Jun;29(2):305–317. doi: 10.1016/0092-8674(82)90147-7. [DOI] [PubMed] [Google Scholar]
- Bloom K. S., Fitzgerald-Hayes M., Carbon J. Structural analysis and sequence organization of yeast centromeres. Cold Spring Harb Symp Quant Biol. 1983;47(Pt 2):1175–1185. doi: 10.1101/sqb.1983.047.01.133. [DOI] [PubMed] [Google Scholar]
- Cameron J. R., Loh E. Y., Davis R. W. Evidence for transposition of dispersed repetitive DNA families in yeast. Cell. 1979 Apr;16(4):739–751. doi: 10.1016/0092-8674(79)90090-4. [DOI] [PubMed] [Google Scholar]
- Carbon J., Clarke L. Structural and functional analysis of a yeast centromere (CEN3). J Cell Sci Suppl. 1984;1:43–58. doi: 10.1242/jcs.1984.supplement_1.4. [DOI] [PubMed] [Google Scholar]
- Carbon J. Yeast centromeres: structure and function. Cell. 1984 Jun;37(2):351–353. doi: 10.1016/0092-8674(84)90363-5. [DOI] [PubMed] [Google Scholar]
- Chinault A. C., Carbon J. Overlap hybridization screening: isolation and characterization of overlapping DNA fragments surrounding the leu2 gene on yeast chromosome III. Gene. 1979 Feb;5(2):111–126. doi: 10.1016/0378-1119(79)90097-0. [DOI] [PubMed] [Google Scholar]
- Clarke L., Carbon J. Genomic substitutions of centromeres in Saccharomyces cerevisiae. Nature. 1983 Sep 1;305(5929):23–28. doi: 10.1038/305023a0. [DOI] [PubMed] [Google Scholar]
- Clarke L., Carbon J. Isolation of a yeast centromere and construction of functional small circular chromosomes. Nature. 1980 Oct 9;287(5782):504–509. doi: 10.1038/287504a0. [DOI] [PubMed] [Google Scholar]
- Dawes I. W. Study of cell development using depressed mutations. Nature. 1975 Jun 26;255(5511):707–708. doi: 10.1038/255707a0. [DOI] [PubMed] [Google Scholar]
- Fitzgerald-Hayes M., Buhler J. M., Cooper T. G., Carbon J. Isolation and subcloning analysis of functional centromere DNA (CEN11) from Saccharomyces cerevisiae chromosome XI. Mol Cell Biol. 1982 Jan;2(1):82–87. doi: 10.1128/mcb.2.1.82. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fitzgerald-Hayes M., Clarke L., Carbon J. Nucleotide sequence comparisons and functional analysis of yeast centromere DNAs. Cell. 1982 May;29(1):235–244. doi: 10.1016/0092-8674(82)90108-8. [DOI] [PubMed] [Google Scholar]
- García-Bellido A. Some parameters of mitotic recombination in Drosophila melanogaster. Mol Gen Genet. 1972;115(1):54–72. doi: 10.1007/BF00272218. [DOI] [PubMed] [Google Scholar]
- Hitzeman R. A., Clarke L., Carbon J. Isolation and characterization of the yeast 3-phosphoglycerokinase gene (PGK) by an immunological screening technique. J Biol Chem. 1980 Dec 25;255(24):12073–12080. [PubMed] [Google Scholar]
- Ito H., Fukuda Y., Murata K., Kimura A. Transformation of intact yeast cells treated with alkali cations. J Bacteriol. 1983 Jan;153(1):163–168. doi: 10.1128/jb.153.1.163-168.1983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kaback D. B., Feldberg L. R. Saccharomyces cerevisiae exhibits a sporulation-specific temporal pattern of transcript accumulation. Mol Cell Biol. 1985 Apr;5(4):751–761. doi: 10.1128/mcb.5.4.751. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Klapholz S., Esposito R. E. Isolation of SPO12-1 and SPO13-1 from a natural variant of yeast that undergoes a single meiotic division. Genetics. 1980 Nov;96(3):567–588. doi: 10.1093/genetics/96.3.567. [DOI] [PMC free article] [PubMed] [Google Scholar]
- LEWIS E. B. The phenomenon of position effect. Adv Genet. 1950;3:73–115. doi: 10.1016/s0065-2660(08)60083-8. [DOI] [PubMed] [Google Scholar]
- Maine G. T., Surosky R. T., Tye B. K. Isolation and characterization of the centromere from chromosome V (CEN5) of Saccharomyces cerevisiae. Mol Cell Biol. 1984 Jan;4(1):86–91. doi: 10.1128/mcb.4.1.86. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Moens P. B., Esposito R. E., Esposito M. S. Aberrant nuclear behavior at meiosis and anucleate spore formation by sporulation-deficient (SPO) mutants of Saccharomyces cerevisiae. Exp Cell Res. 1974 Jan;83(1):166–174. doi: 10.1016/0014-4827(74)90700-9. [DOI] [PubMed] [Google Scholar]
- Panzeri L., Philippsen P. Centromeric DNA from chromosome VI in Saccharomyces cerevisiae strains. EMBO J. 1982;1(12):1605–1611. doi: 10.1002/j.1460-2075.1982.tb01362.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pardue M. L., Gall J. G. Chromosomal localization of mouse satellite DNA. Science. 1970 Jun 12;168(3937):1356–1358. doi: 10.1126/science.168.3937.1356. [DOI] [PubMed] [Google Scholar]
- Reed S. I., Ferguson J., Groppe J. C. Preliminary characterization of the transcriptional and translational products of the Saccharomyces cerevisiae cell division cycle gene CDC28. Mol Cell Biol. 1982 Apr;2(4):412–425. doi: 10.1128/mcb.2.4.412. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Roth R. Chromosome replication during meiosis: identification of gene functions required for premeiotic DNA synthesis. Proc Natl Acad Sci U S A. 1973 Nov;70(11):3087–3091. doi: 10.1073/pnas.70.11.3087. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rothstein R. J. One-step gene disruption in yeast. Methods Enzymol. 1983;101:202–211. doi: 10.1016/0076-6879(83)01015-0. [DOI] [PubMed] [Google Scholar]
- SHERMAN F., ROMAN H. Evidence for two types of allelic recombination in yeast. Genetics. 1963 Feb;48:255–261. doi: 10.1093/genetics/48.2.255. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Southern E. M. Detection of specific sequences among DNA fragments separated by gel electrophoresis. J Mol Biol. 1975 Nov 5;98(3):503–517. doi: 10.1016/s0022-2836(75)80083-0. [DOI] [PubMed] [Google Scholar]
- Stinchcomb D. T., Mann C., Davis R. W. Centromeric DNA from Saccharomyces cerevisiae. J Mol Biol. 1982 Jun 25;158(2):157–190. doi: 10.1016/0022-2836(82)90427-2. [DOI] [PubMed] [Google Scholar]
- Struhl K., Davis R. W. A physical, genetic and transcriptional map of the cloned his3 gene region of Saccharomyces cerevisiae. J Mol Biol. 1980 Jan 25;136(3):309–332. doi: 10.1016/0022-2836(80)90376-9. [DOI] [PubMed] [Google Scholar]
- Struhl K., Stinchcomb D. T., Scherer S., Davis R. W. High-frequency transformation of yeast: autonomous replication of hybrid DNA molecules. Proc Natl Acad Sci U S A. 1979 Mar;76(3):1035–1039. doi: 10.1073/pnas.76.3.1035. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thomas P. S. Hybridization of denatured RNA and small DNA fragments transferred to nitrocellulose. Proc Natl Acad Sci U S A. 1980 Sep;77(9):5201–5205. doi: 10.1073/pnas.77.9.5201. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tschumper G., Carbon J. Copy number control by a yeast centromere. Gene. 1983 Aug;23(2):221–232. doi: 10.1016/0378-1119(83)90054-9. [DOI] [PubMed] [Google Scholar]
- Williamson D. H., Fennell D. J. The use of fluorescent DNA-binding agent for detecting and separating yeast mitochondrial DNA. Methods Cell Biol. 1975;12:335–351. doi: 10.1016/s0091-679x(08)60963-2. [DOI] [PubMed] [Google Scholar]
- Yeh E., Bloom K. Characterization of a tightly centromere-linked gene essential for meiosis in the yeast Saccharomyces cerevisiae. Basic Life Sci. 1985;36:231–242. doi: 10.1007/978-1-4613-2127-9_15. [DOI] [PubMed] [Google Scholar]