Abstract
Our understanding of human germ cell development is limited in large part due to inaccessibility of early human development to molecular genetic analysis. Pluripotent human embryonic stem cells (hESCs) and induced pluripotent stem cells (iPSCs) have been shown to differentiate to cells of all three embryonic germ layers, as well as germ cells in vitro, and thus may provide a model for the study of the genetics and epigenetics of human germline. Here, we examined whether intrinsic germ cell translational, rather than transcriptional, factors might drive germline formation and/or differentiation from human pluripotent stem cells in vitro. We observed that, with overexpression of VASA (DDX4) and/or DAZL (Deleted in Azoospermia Like), both hESCs and iPSCs differentiated to primordial germ cells, and maturation and progression through meiosis was enhanced. These results demonstrate that evolutionarily unrelated and divergent RNA-binding proteins can promote meiotic progression of human-derived germ cells in vitro. These studies describe an in vitro model for exploring specifics of human meiosis, a process that is remarkably susceptible to errors that lead to different infertility-related diseases.
Keywords: DAZL, VASA, Meiosis, Induced pluripotent stem cells, Human embryonic stem cells
Introduction
Human germ cell development is poorly understood at least in part due to inaccessibility of early stages of human embryo development, in general. Thus, most of the information regarding human germ cell development has been extrapolated from studies of mouse germ cell development in vivo [1–4]. However, other studies have explored derivation of germ cells in vitro from mouse embryonic stem cells (mESCs) [5–7], while parallel studies have reported the derivation of germ cells from human embryonic stem cells (hESCs) and induced pluripotent stem cells (iPSCs) [8–12]. However, in both the mouse and human in vitro systems, progression of meiosis has limited the success of production of mature gametes, perhaps due to deficiencies in internal or external factors that regulate meiosis.
Germ cell development is inherently dependent upon transcriptional regulation; however, there is extensive transcriptional silence at several stages of development, indicative of the major role of translation regulation [13–16]. A large number of RNA-binding proteins have been shown to be critical for germ cell development, with several demonstrating conservation across species [14–16]. The DAZ (Deleted in Azoospermia) gene family contains some of the most highly conserved reproductive genes and encodes RNA-binding proteins that have been implicated in multiple stages of germ cell development, including maintenance of premeiotic germ cell populations and meiotic initiation and progression, in species as diverse as flies, worms, frogs, mice, and humans [17–20]. In fact, we previously demonstrated that overexpression of distinct members of the human DAZ gene family members may function in germ cell formation and initiation and progression through meiosis in vitro [21, 22].
Although not related to the DAZ gene family, the VASA gene (also called DDX4) encodes another RNA-binding protein that is highly conserved across species and expressed specifically in the germline in several model organisms from flies, worms, frogs, mice, and humans [23–27]. VASA encodes a member of the DEAD-box family of proteins with an ATP-dependent RNA helicase; the protein is cytoplasmic and expressed in mammals from premeiotic stages (beginning during migration or shortly thereafter at arrival of germ cells in the gonadal ridge) throughout gametogenesis [23]. Although the cellular function of VASA protein remains unclear, it has been suggested that, as a member part of the DEAD (asp-gluala-asp)-box family of proteins, VASA might serve as a chaperone to ensure proper folding of different target RNAs and translational regulation [28]. In mice, disruption of Vasa resulted in male infertility whereas homozygous mutant female mice were fertile [29]. Finally, notably, it has also been reported that ectopic expression of VASA may mediate the reprogramming of chicken embryonic stem cells into germ cells [30].
Here, we used in vitro differentiation of pluripotent stem cells into germ cells to examine the results of ectopic expression of human VASA; we sought to determine if overexpression of VASA would mimic results with the divergent RNA-binding protein, DAZL. Thus, we also examined whether the divergent RNA-binding proteins encoded by the human VASA and DAZL genes might function independently or synergistically during germ cell development.
Materials and Methods
Cell Culture
Cell culture was as previously described [21, 22]. Essentially, undifferentiated pluripotent stem cell lines, hESC lines H9 (46, XX) and HSF1 (46, XY) and human adult-derived iPSC lines iHUF3 (46, XX) and iHUF4 (46, XY), and fetal-derived iPSC line iPS(IMR90) (46, XX), were maintained on irradiated mouse embryonic fibroblasts (MEFs) in Dulbecco's modified Eagle's medium (DMEM)/F12 supplemented with 20% KnockOut serum replacer, 2 mM l-glutamine, 0.1 mM nonessential amino acids, 10 ng/ml basic fibroblast growth factor (bFGF) (Invitrogen, San Diego, CA, www.invitrogen.com), and 0.1 mM 2-mercaptoethanol (Milli-pore, Billerica, MA, www.millipore.com) at 37°C, 5% CO2 . Cells were passaged every 4–5 days using 1 mg/ml Collagenase IV (Invitrogen). The feeder-free culture was maintained on Matrigel (BD Biosciences, Sparks, MD, www.bdbiosciences.com)-coated plates with conditioned medium obtained by incubating culture medium with MEFs for 24 hours. Cells were differentiated on Matrigel-coated plates with differentiation medium composed of DMEM/F12 supplemented with 20% fetal bovine serum (Hyclone, Logan, UT, www.thermoscientific.com), 2 mM l-glutamine, 0.1 mM nonessential amino acids, and 0.1 mM 2-mercaptoethanol for 7 and 14 days. All cell lines used in this study were between passages 15 and 55. All differentiation experiments were performed in triplicate.
Construction of VASA Expression Vector (iVASA)
The construction of the vector to express VASA was made by using the same backbone as the previously reported DAZL and VASA-green fluorescent protein (GFP) vectors [21]. Briefly, 2.2 kb VASA open reading frame (ORF) sequence (GenBank accession no. NM_024415.2) was isolated from human testis cDNA obtained from reverse transcription of adult human testis total RNA (Clontech). The isolated polymerase chain reaction (PCR) product was cloned into the p2K7 lentiviral backbone under the control of the elongation factor 1α (EF1α) constitutive promoter using the Gateway system (Invitrogen). Expression of VASA cDNA resulted in a 76 kDa protein as detected by mouse anti-VASA antibodies on Western blot (Abcam, Cambridge, UK, www.abcam.com).
Transduction of Cell Lines
Undifferentiated cells were transduced on Matrigel-coated plates with DAZL and VASA lentiviral overexpression vectors along with viral supernatant supplemented with 8 μg/ml polybrene (Sigma-Aldrich, Gmbh Munich, Germany, www.sigmaaldrich. com) as previously described [21, 22]. After 6 hours, conditioned medium was added in a dilution 1:3, and cells were further incubated overnight. Virus supernatants were washed off, and conditioned medium was added until the following day. Subsequently, cells were selected with 2 μg/ml blasticidin (Invitrogen) for 3 days in conditioned media after which, the differentiation was started. For the transduction of the lentiviral vector carrying GFP gene under control of the VASA promoter, cells were transduced as previously described, but the selection was started the following day with 200 μg/ml geneticin (Invitrogen) for 7 days. Cells were passaged onto MEFs and then moved back to Matrigel followed by differentiation. Untransduced cells or cells transduced with the empty vehicle were used as a control in all differentiation experiments. We note that it was necessary to adjust the amount of viral particles used in all double infections to half of that used in single infection experiments to minimize viral toxicity and maximize the effectiveness of overexpression; for example, in the case of overexpression of both DAZL and VASA.
Gene Expression Analysis
Total RNA was prepared per manufacturer's instructions via the RNeasy Mini Kit (Qiagen). Approximately, 1 μg total RNA from each sample was used for random primed reverse transcription, via the SuperScript III First-Strand Synthesis System for retrotranscription-polymerase chain reaction (RT-PCR) (Invitrogen). Endogenous or ectopic VASA expression was assayed by using two different forward primers that aligned specifically with the 5′ sequence upstream the starting codon of the ORF in the first exon of the endogenous VASA RNA or with the 5′ sequence upstream the starting codon in the p2K7 lentivector. Transcription levels were determined in triplicate reactions and normalized to the average of the housekeeping gene glyceraldehyde 3-phosphate dehydrogenase (GAPDH) with the formula 2−(ΔCt). Primer sequences are as follows: VASA-ENDO F 5′-CACGTGCAGCCGTTTAAGT-3′; VASA-ECTOPIC F 5′-CAAGTTTGTACAAAAAAGCAGGCT-3′; VASA R 5′-TGATGATGAAGCTGGAGTCCT-3′. For microfluidic PCR analysis, cDNA was preamplified using a mix of 0.2× Taqman assays (Applied Biosystems) as primers in a reaction containing 5 μl 2× buffer (from CellsDirect One-Step qRT-PCR kit, Invitrogen), 2.5 μl Taqman assay mix, 1 μl Platinum Taq (Invitrogen), and 1.5 μl cDNA. Reactions were completed using a PCR reaction of 95°C for 10 minutes and 14 cycles of 95°C for 15 seconds, and 60°C for 4 minutes. Preamplified cDNA was diluted 1:2 with TE buffer and used for gene expression analysis on the Biomark 96.96 microfluidic qPCR chip (Fluidigm). Expression was determined in triplicate reactions, and fold change for each marker was calculated based on the geometric mean of multiple control genes [31] by the Qbase Plus software (Biogazelle).
Western Blot Analysis
Cells were harvested and lysed in RIPA buffer (Sigma) plus 1 × protease inhibitors (Complete Mini, Roche Diagnostics, GmbH Mannheim, Germany, www.roche.com). Samples were rocked on ice for 20 minutes and centrifuged for 20 minutes at 16,000g at 4°C. Supernatant was measured for protein concentration, and 20 μg per sample was denatured with 6× Laemmli buffer at 95°C for 5 minutes and then loaded onto 10% sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE) gel. The gels were run at 150 V for 70 minutes and transferred to a polyvinylidene fluoride (PVDF) membrane for 1 hour at 100 V in CAPS buffer (10 mM CAPS, 10% methanol, pH 11). Membranes were blocked in a 5% solution of nonfat dry milk for 1 hour at room temperature and then incubated overnight at 4°C with primary antibodies: 1:1,000 for monoclonal anti-VASA (clone number MM0598-11G15) and 1:10,000 polyclonal anti-GAPDH (all Abcam). Subsequently, membranes were washed with TBS-T (TBS with 0.1% Tween 20) and incubated with horseradish peroxidase (HRP)-conjugated secondary antibodies (Calbiochem, Darmstadt, Germany, www.emdchemichals.com) in a 1:10,000 dilution in 5% nonfat milk for 1 hour at room temperature. HRP signal was detected by incubating washed membranes with Supersignal West Dura Extended Duration Substrate (Thermo Scientific, Barrington, IL, www.thermoscientific.com) on film.
Meiotic Spreads
Meiotic spreads were performed as previously described [21, 22]. Essentially, cells were collected and resuspended in hypoextraction buffer (30 mM Tris [Sigma-Aldrich], 50 mM sucrose [Sigma-Aldrich], 17 mM citric acid [Sigma-Aldrich], 5 mM EDTA [Invitrogen], two tablets of Complete Mini [Roche]; pH 8.2). Solution was incubated for 30 minutes on ice and cytospun onto slides. The slides were fixed with 4% paraformaldehyde for 15 minutes, washed with phosphate buffer salt (PBS), and permeabilized 5 minutes with 0.04% photoflo (KODAK, St. Rochester, NY, www.kodak.com) followed by blocking for 60 minutes with 4% goat serum in 1% bovine serum albumin (BSA), 0.1% Tween 20 in PBS. Rabbit polyclonal anti-synaptonemal complex protein 3 (SCP3) primary antibody (Novus Biologicals, Littleton, CO, www.novusbio.com) was used with 1:1,000 dilution and mouse polyclonal anti-centromeric protein A (CENPA) antibody (Abcam) was used at 1:500 dilution and incubated for 3 hours at room temperature. Slides were washed with 1% BSA + 0.1% Tween 20 in PBS, and goat anti-rabbit Alexa 594 and goat anti-mouse Alexa 488 secondary antibodies (Invitrogen) were applied in 1:1,000 dilution and incubated for 1 hour at room temperature. ProLong Gold antifade reagent with 4′,6-diamidino-2-phenylindole (DAPI) (Invitrogen) was applied. Finally, cells were counted under fluorescence microscope.
Fluorescence-Activated Cell Sorting Analysis
For the DNA content-fluorescence-activated cell sorting (FACS), cells were harvested into a single cell suspension and then fixed with ethanol 70% for 1 hour at room temperature. Subsequently, cells were washed off and incubated in staining solution (0.2 mg/ml RNase A and 0.02 mg/ml propidium iodide [Invitrogen] in BD Perm/Wash buffer [BD] containing saponin and fetal calf serum (FCS)) for 30 minutes at 37°C. Both VASA-GFP and DNA content-FACS were performed using a FACS Aria cytometer system (BD).
Fluorescent In Situ Hybridization
Sorted cell populations were collected by cytospinning as previously described [22]. Slides were fixed with Carnoy's fixative (1:3 acetic acid/methanol) for 5 minutes and air-dried. The slides were treated with 50 μg/ml pepsin in 0.01 N HCl for 5 minutes, washed twice in PBS for 5 minutes each, and then dehydrated in ice-cold 70%, 80%, and 100% ethanol, and air-dried. Fluorescence in situ hybridization (FISH) probes against chromosomes 16 and 18 (Vysis, Abbott Park, Illinois, www.abbottmolecular.com) were denatured on slides at 85°C and hybridized at 37°C overnight. Slides were then washed with 50% formamide 2× SSC at 43°C and in 2× SSC at 43°C, 5 minutes each. ProLong Gold antifade with DAPI was applied to slides. Cells were counted under fluorescence microscope.
Immunostaining of Fixed Cells
For sorted cell populations, cells were cytospun as described above; the slides were then fixed with 4% paraformaldehyde for 15 minutes and then permeabilized with 1% Triton X-100 for 15 minutes, followed by washes with 0.1% Tween 20 in PBS. Slides were blocked with 4% goat serum in PBS for 1 hour at room temperature and stained with polyclonal antibodies against ACROSIN, DAZL, or VASA (all Abcam) diluted 1:300 with 1% goat serum in 4°C overnight. Slides were washed and incubated with Alexa (Invitrogen) secondary antibodies at a dilution of 1:1,000 for 1 hour at room temperature, followed by washes and mounted as described above.
Bisulfite Sequencing
Genomic DNA was extracted via the QIAamp DNA Mini kit (Qiagen, Valencia, CA, www.qiagen.com). After determining the efficiency of bisulfite conversion of the genomic DNA, 100 ng of genomic DNA was processed using the Epitect Bisulfite Kit (Qiagen) according to manufacturer's instructions. One microliter of bisulphite-treated genomic DNA was PCR amplified following previously reported protocols 12 in the presence of 5 mM MgCl2 , 0.2 mM dNTPs, 10 pmol each primer, and 1 Unit Platinum Taq polymerase (Invitrogen). PCR amplifications for H19 were initiated at 94°C for 3 minutes followed by 45 cycles of 94°C for 30 seconds, 58°C for 30 seconds, and 72°C for 30 seconds. Primer sequences are as follows, hH19F 5′-TGTATAGTATATGGGTATTTTTG-GAGGTTT-3′, hH19R 5′-TCCTATAAATATCCTATTCCCAAA-TAACC-3′. The resultant PCR products were gel-extracted using the Qiaquick gel extraction kit (Qiagen) and cloned into the TOPO TA vector (Invitrogen). At least 20 clones were sequenced using ABI BigDye v3.1 dye terminator sequencing chemistry (Applied Biosystems, Carlsbad, CA, www.appliedbiosystems.com) and ABI PRISM 3730×l capillary DNA analyzer for sequence analysis. Sequenced clones were aligned by Geneious software (Biomatters, Auckland, New Zealand, www.biomatters.com) and CpG methylation analysis was performed by BiQ Analyzer software (Max Planck Institut Informatik, Saarbrücken, Germany, www.mpi-inf.mpg.de).
Statistical Analysis
Statistical analysis for gene expression results was performed with one-way ANOVA (analysis of variance) and Student's t pairwise comparison by SPSS software (SPSS Inc, Chicago, IL, www.ibm.com). Significance was accepted at p < .05.
Results
Germ Cell Properties of VASA-Positive Cells Differentiated In Vitro
To compare the potential of different pluripotent stem cell lines to spontaneously differentiate into primordial germ cells, we transduced two hESC lines (H9 [46, XX] and HSF1 [46, XY]) and one iPSC line (iHUF3 [46, XX]) with a VASA-GFP lentiviral reporter system, as we and our collaborators previously described [21]. Subsequently, cells were selected for antibiotic resistance, spontaneously differentiated in the absence of bFGF with fetal bovine serum (FBS) for 7 days, and isolated by FACS. Examination of FACS profiles revealed variation across lines in their potential to differentiate to VASA-GFP positive cells as shown (Fig. 1A). Immunocytochemistry of all cell lines further confirmed that endogenous VASA protein expression was observed in more than 90% of all the VASA-GFP positive sorted cells and was not observed in VASA-GFP negative cells (Fig. 1B).
Figure 1.
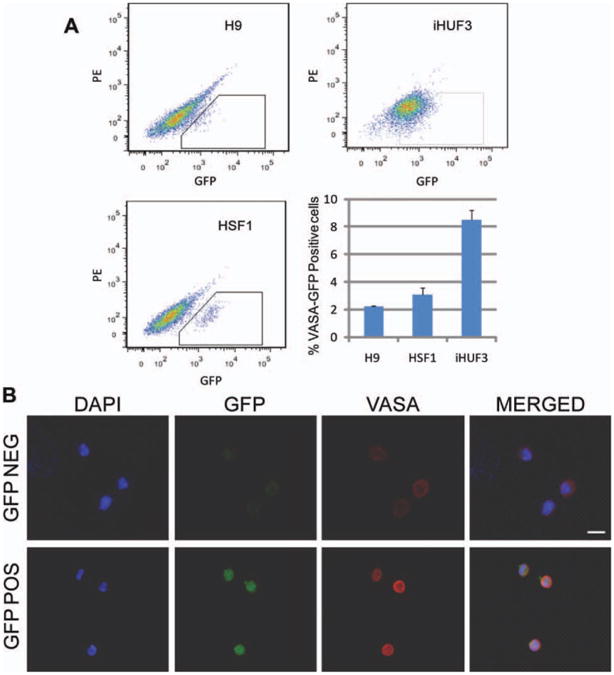
Fluorescence-activated cell sorting (FACS) of VASA-GFP positive and negative populations. (A): FACS plots and representation of the percentage of VASA-GFP positive cells we found for each cell line. Data are represented as mean ± SEM. (B): Representative VASA immunostaining of GFP-positive and -negative sorted populations. The scale bar represents a distance of 10 μm. Abbreviations: DAPI, 4′,6-diamidino-2-phenylindole; GFP, green fluorescent protein; PE, phycoerythrin.
Further examination of gene expression demonstrated that early germ cell markers were enriched in VASA-GFP positive cells relative to the GFP-negative cell populations (Fig. 2A). Comparisons of expression profiles of late and meiotic germ cell markers within the VASA-GFP positive populations also demonstrated significantly higher expression of these markers relative to the GFP-negative populations in all analyzed cell lines (Fig. 2B). These results confirmed findings from previous studies suggesting that VASA-positive sorted cell populations were enriched for germ cells that were at early to more-advanced stages of development, with some indication of meiotic initiation as evidenced by the expression of meiotic markers such as SCP3, MutL homolog 1, and dosage suppressor of MCk 1 [21, 22]. Based on these results and previous findings, we then sought to address our hypothesis that divergent intrinsic germ cell translational factors might drive germline formation and promote differentiation from human pluripotent stem cells in vitro. Thus, we examined the potential of induced expression of VASA to regulate meiotic initiation and/or progression in germ cells, alone and in combination with DAZL, a member of the DAZ gene family previously shown to enhance human germ cell specification and meiotic progression.
Figure 2.

Gene expression in VASA-GFP positive and negative sorted cell populations. (A): Comparison of early germ cell markers expression profile among GFP-positive and -negative sorted populations. (B): Comparison of late and meiotic germ cell markers expression profile among GFP-positive and their respective negative sorted populations. Asterisks represent significant differences (*, p, < .05) with controls. Data are represented as normalized fold change mean ± SEM. Abbreviations: DAZL, Deleted in Azoospermia Like; DMC1, dosage suppressor of MCk 1; GFP, green fluorescent protein; MLH1, MutL homolog 1; SCP3, synaptonemal complex protein 3.
Ectopic Expression of VASA Promotes Meiotic Progression in Spontaneously Differentiated Germ Cells
To test whether ectopic expression of VASA alters the dynamics of human germ cell formation and/or differentiation, we isolated a segment of DNA from human testis that encodes the ORF of VASA and cloned it into a lentiviral backbone with transcription under control of a strong, constitutive mammalian promoter EF1α (Supporting Information Fig. S1). We then established pluripotent cell lines by selecting them for antibiotic resistance and spontaneously differentiated them for 14 days to examine how the ectopic expression of VASA impacts germ cell development.
Ectopic expression of VASA (iVASA) was confirmed by Western blotting with VASA observed as a 76-kDa protein as expected (Supporting Information Fig. S2A). VASA expression was maintained at both RNA and protein levels in all transduced cell lines after 14 days of spontaneous differentiation (Supporting Information Fig. S2B, S2C). We also noted that the majority of VASA expression in transduced cell lines was due to the ectopic induction, as shown by RT-qPCR analysis with assays specific to endogenous and ectopic VASA transcripts using testis as a positive control for endogenous expression (Supporting Information Fig. S2D). We observed that gene expression in response to ectopic expression of VASA differed from that of controls. There was increased expression in most of the early and late germ cell markers we analyzed in both hESC lines. This result suggested that ectopic VASA expression induced spontaneously differentiated germ cells to progress in their maturation programs (Fig. 3). Surprisingly, the effect of VASA ectopic expression was lower in iHUF3 in all our experiments (n = 3), even that some of the germ cell markers we analyzed were also upregulated in transduced cells compared with their controls. However, additional preliminary experiments performed over the previously reported iPSC lines iHUF4 and iPS(IMR90) showed a response more similar to that observed in the two hESC lines, indicating intrinsic differences in the response to ectopic expression of VASA among iPSC lines (Supporting Information Fig. S3).
Figure 3.

Expression of early and late germ cell markers in cells with and without ectopic expression of VASA (iVASA) 7 days postdifferen-tiation. (A): Comparison of expression of early germ cell markers. (B): Comparison of expression of late and meiotic germ cell markers. Asterisks represent significant differences (*, p < .05) with controls. Data are presented as normalized fold change mean ± SEM. Abbreviations: DAZL, Deleted in AZoospermia Like; DMC1, dosage suppressor of MCk 1; MLH1, MutL Homolog 1; SCP3, synaptonemal complex protein 3.
We next examined the extent of differentiation via isolation of putative 1N populations from whole differentiated samples by FACS. We harvested and stained 14-day spontaneously differentiated cells with propidium iodide and then sorted for putative 1N cells. There was a significant increase from approximately 0.5% to 1.5% in the percentage of cells in the putative 1N cell population from hESC lines with ectopic expression of VASA relative to controls, and similar levels in the case of iHUF3. In addition, we observed that the putative 1N populations showed an increased percentage of cells positive for both VASA protein and the postmeiotic marker, ACROSIN protein (Fig. 4A, 4B). Ploidy analysis of the 1N, 2N, and 4N sorted cells was further examined by FISH with probes against chromosomes 16 and 18 (Fig. 4C). We found a significant increase in the percentage of cells monosomic for both chromosomes in the sorted 1N populations ranging from 4- to 10-fold depending on the cell line, when VASA was overexpressed; this suggested that overexpression of VASA likely promoted germ cell progression through meiosis, albeit at low frequency (approximately 0.15% of total cells).
Figure 4.

Analysis of the putative 1N cell populations from cell lines with ectopic expression of VASA (iVASA), 14 days post-transduction. (A): Representative DNA-fluorescence-activated cell sorting plot for the isolation of the putative 1N population and percentage of putative 1N cells we were able to isolate from each cell line. Sperm control was used to set the sorting parameters. (B): Representative double VASA and ACROSIN immunostaining of 1N sorted cells and percentage of cells positive for each marker. (C): Representative fluorescence in situ hybridization results for probes against chromosomes 16 (green) and 18 (red) over putative 1N sorted cells and percentage of haploid cells found in each cell line based on the analysis of these two probes. Scale bar = 10 μm. Asterisks represent significant differences (*, p < .05) with controls. Data are presented as mean ± SEM. Abbreviations: DAPI, 4′,6-diamidino-2-phenylindole.
Combined Overexpression of DAZL and VASA Is Not Synergistic
Next, we combined ectopic expression of VASA and DAZL and compared the efficiency of germ cell formation and maturation. Thus, we ectopically expressed DAZL (iDAZL), as previously reported [21], and VASA/DAZL (iDAZL+VASA) in all three pluripotent cell lines and compared key characteristics with VASA overexpressed cells (iVASA) and controls.
We observed significantly higher expression of some diagnostic germ cell and/or stem cell markers such as OCT4 (POUF5), LIN28, SOX2, NANOG, DNMT3B, Blimp1 (PRDM1A), PRDM14, Fragilis (IFITM1), DAZ, DAZL, VASA (DDX4), c-KIT, PELOTA, GCNF, PRM1, Acrosin, SOX9, LHR, ZP2 and ZP4 in iDAZL+iVASA overexpressed cells compared with controls. However, we observed very high variability in gene expression profiles among different cell lines at the RNA level, indicating different potential or stochastic programming of germline development (Supporting Information Fig. S4).
The localization and distribution of SCP3 along the chromosomes is one of the most significant indicators of meiosis and much more specific to meiotic progression than just RNA or protein expression [21, 22]. Subsequently, analysis of the meiotic stages of differentiated cells was accomplished via observation of the synaptonemal complex. For this purpose, we coimmunostained cells for SCP3 and CENPA, a centromeric component of chromosomes (Fig. 5A). We observed that most cells did not show any SCP3 staining, indicating either that they were not meiotic or had already completed meiosis. However, a subset of cells showed punctate SCP3 staining pattern, indicative of early meiotic stages, especially the leptotene stage of prophase I. In addition, some cells demonstrated an elongated staining pattern of SCP3, indicative of progression through zygotene, pachytene, or diplotene meiotic prophase I stages, as previously described [21, 22]. We observed that the punctate staining pattern was present in a subset of cells in all treatments and controls, ranging from 1% to 6%. In contrast, we observed a significant increase in the elongated pattern of synaptonemal complex staining indicative of meiotic progression only with ectopic expression of DAZL or VASA. Efficiency of meiotic progression was highest with overexpression of DAZL as indicated by the greatest percentage of cells with elongated SCP3 (Fig. 5B). We note here that we attempted to differentiate the SCP3-positive cells for longer time periods to assess the outcomes of extended culture. However, we found that the optimal time of differentiation remains at 14 days as previously described [21]. Further culture of the differentiating pluripotent cell lines resulted in significant reduction in the viability of differentiated cells and a reduction in germ cells suggesting a need to further optimize the culture conditions.
Figure 5.

Synaptonemal complex (SCP) formation analysis over cell lines with ectopic expression of DAZL (iDAZL), VASA (iVASA), and DAZL+VASA (iDAZL+iVASA) and differentiated for 14 days. (A): Representative SCP3 staining of meiotic spreads. Scale bar = 10 μm. (B): Percentage of cells with punctuated (leptotene) and elongated (diplotene, pachytene, or zygotene) SCP3 staining patterns. Abbreviations: CENPA, centromeric protein A; DAPI, 4′,6-diamidino-2-phenylindole; DAZL, Deleted in Azoospermia Like; SCP3, synaptonemal complex protein 3.
We next compared the putative 1N populations obtained with ectopic expression of DAZL and DAZL+VASA to those with ectopic expression of VASA alone and observed that overexpression of DAZL was also the most effective internal inducer of meiotic progression as shown by percentage of haploid cells via FISH (Fig. 6A, 6B). However, the percentage of cells positive for ACROSIN and VASA was greatest when DAZL and VASA were ectopically expressed together, suggesting potential synergism in translational regulation of late germ cell markers such as ACROSIN (Fig. 6C). Of special note, when we compare iDAZL and iDAZL+iVASA, overexpression of VASA seems to decrease the number of haploid cells when compared with DAZL overexpression. However, this result may reflect the fact that it was necessary to adjust the amount of viral particles used in the double infection to half of that used in single infection experiments to minimize the viral toxicity and maximize the effectiveness of overexpression. Thus, we cannot state whether VASA overexpression directly reduces production of haploid cells or results in arrest. This effect was also observed as a decrease of the expression levels for both markers when cells were subjected to the double transduction compared with the single transduction ones (Supporting Information Fig. S4C).
Figure 6.

Analysis of the DNA-FACS-sorted putative 1N cell populations with ectopic expression of DAZL (iDAZL), VASA (iVASA), and DAZL+VASA (iDAZL+iVASA), 14 days postdifferentiation. (A): Percentage of putative 1N cells isolated from each cell line. (B): Percentage of haploid cells confirmed by FISH in 1N sorted populations. (C): Percentage of 1N sorted cells positive for VASA and ACROSIN staining. Asterisks represent significant differences (*, p < .05) with controls. Data are presented as mean ± SEM. Abbreviations: DAZL, Deleted in AZoospermia Like.
Ectopic Expression of VASA Results in Recapitulation of Some Aspects of Germline Reprogramming at the H19 Locus
Erasure of the epigenetic marks at imprinted loci is a process that occurs specifically during mammalian germ cell development just prior to, or shortly after, germ cells arrive to the gonadal ridges. Subsequent reacquisition of a sex-specific imprinted pattern generally occurs just before or during meiosis depending of the sex [32–34]. To determine if ectopic expression of VASA alters epigenetic programs, we analyzed methylation status at the CpG islands of the imprinted, maternally expressed gene H19 [35, 36]. We observed that the H19 locus was hemimethylated in undifferentiated cells, as expected. Following spontaneous differentiation of controls, H19 methylation patterns showed a slight increase in methylation, as shown (Fig. 7A). In contrast, overexpression of VASA led to a significant decrease in the methylation ratio of H19 (Fig. 7B). This effect was specific to VASA; ectopic expression of DAZL did not lead to significant changes in methylation status at the H19 locus (Supporting Information Fig. S5).
Figure 7.

CpG methylation analysis at the DMR of H19. (A): Diagrams represent methylation status of each CpG dinucleotides on individual DNA clones. Lines represent different clones and columns are different CpG dinucleotides. Methylated CpGs are represented as filled circles and unmethylated CpGs are represented as open circles. Empty CpG sites represent the CpGs that could not be determined. (B): Percentage of methylated CpGs found in each condition at the DMR of H19. Asterisks represent significant differences (*, p < .05) with controls. Abbreviation: DMR, differentially methylated region.
Discussion
Previous studies demonstrated that both hESCs and iPSCs can form germ cells in vitro, albeit with different efficiencies observed between different cell types [21, 22]. Here, we examined the effect of ectopic expression of evolutionarily divergent RNA-binding proteins on germ cell formation, maintenance, and progression through meiosis. We confirmed key characteristics of VASA-GFP positive germline cells [21, 22]. We then extended the studies to examine effects of over-expression of VASA, DAZL, and VASA/DAZL. The human VASA gene encodes an RNA-binding protein that is highly conserved in evolution in all metazoans suggesting an essential role for germline development that may also be conserved in humans [37, 38]. Recently, one study identified 221 candidate mRNAs that may be targets of VASA in Drosophila [39]. This study suggested that VASA may act by regulating the translation of mei-P26, a repressor of micro-RNA activity and promoter of differentiation of germline stem cells; mei-P26 is a component of P-granules or germplasm, a perinuclear electron-dense ribonucleoprotein structure that specifically appears in the germline of several model organisms and also contains other highly conserved RNA-binding proteins such as Nanos and Piwi. Interestingly, disruption of both the Mvh and Piwi genes in mice indicates that they are required for meiosis and have a similar phenotype with elevated retrotransposon expression that leads to defective de novo DNA methylation due to impaired Piwi-RNA regulation; this suggests that VASA is likely implicated in their regulation [39, 40]. Based on these studies and others, we hypothesized that ectopic expression of VASA may promote maturation and progression of human germ cells through meiosis.
We observed that cells with ectopic expression of VASA expressed most of the germ cell markers analyzed, including markers characteristic of germ cells in both early and later stages of maturation, similar to results reported in different model systems [30]. However, the only iPSC line that we analyzed in any detail, iHUF3, demonstrated an apparently less robust response to overexpression of VASA compared with the hESC lines and the other two iPSC lines we tested, indicating that different cell lines may have a different response to ectopic induction of VASA. We hypothesize that these differences among the three iPSC lines we tested could be due to inherent genetic differences, differences in derivation, or alternatively linked to epigenetic memory of each cell line because of their different somatic cell origin. However, a larger comparison across many lines would allow greater understanding regarding whether this is a generalized observation of iPSC versus hESC lines.
We also detected a subset of cells with ectopic expression of VASA that appeared able to complete meiosis as judged by haploidy of two chromosome markers and expression of ACRO-SIN, a late male-specific germ cell protein widely expressed in spermatid stages [41]. Additional immunohistochemical markers are not, to our knowledge, available for robust and specific expression and localization studies of spermatids. Nonetheless, interestingly, we observed that ACROSIN expression appeared in all cell lines to different extents independently of chromosomal sex, suggesting that sex-specific expression is likely to depend on the somatic niche in addition to chromosomal sex of the germ cells, as previously described [10, 22]. Alternatively, it is possible that the ectopic expression of VASA directly or indirectly modulates the translation of proteins such as ACRO-SIN on a background of predominantly female germ cell gene expression. We note that we also found that the percentage of cells that expressed ACROSIN was greater than the percentage of haploid cells, suggesting that cells other than haploid cells may also express ACROSIN, perhaps indicating that the post-meiotic ACROSIN time-related expression found in male germ cells in vivo may be altered due to the ectopic expression of key meiotic regulators such as the RNA-binding proteins DAZL and VASA in vitro.
We also compared germ cell development in pluripotent stem cell lines with ectopic expression of VASA to those with ectopic expression of DAZL and both VASA and DAZL. We found that both DAZL and VASA promoted progression through meiosis. However, we observed that the ectopic expression of DAZL induced the formation of haploid cells most effectively, without any evidence of synergistic effect with the ectopic expression of VASA. On the other hand, we observed a clear synergistic effect of the combined overexpression of both DAZL and VASA in the percentage of ACROSIN-positive cells, indicating a possible relationship of both proteins in the upregulation of this late male germ cell marker (Fig. 6).
Moreover, we did not observe differences in the overall gene expression profiles linked to ectopic expression of either translational factor (Supporting Information Fig. S4). We suggest that this may reflect the epistatic relationship of DAZL and VASA function; previous studies have suggested that DAZL may exert translational control on VASA in the germ-line [42]. Our studies also identified clusters of VASA-positive cells with morphologies that appeared well-rounded and those that appeared elongated/migratory when subjected to ectopic expression of DAZL; both cell types frequently expressed both VASA and DAZL and were not observed in controls (Supporting Information Fig. S6). Moreover, as previously noted, ectopic expression of DAZL appears to be involved in both formation and maintenance of early populations as well as in promoting meiosis [21]. In contrast, ectopic expression of VASA resulted in a significantly higher proportion of ACROSIN-positive cells but did not appear to be involved in formation and maintenance of early germ cell populations.
Erasure of epigenetic imprints and their subsequent sex-specific re-establishment is a critical and diagnostic process in germ cell development that is necessary for embryo development, organogenesis, and viability [32–34]. Erasure and re-establishment of epigenetic signatures of imprinted loci is a germline-specific process. In this study, we observed that the maternally imprinted H19 locus showed a significant decrease in the CpG methylation pattern of its differentially methylated region when cells were subjected to ectopic expression of VASA; this effect was not observed with ectopic expression of DAZL. These results suggest that erasure of imprints may be a specialization of VASA in direct or indirect regulation of RNAs implicated in DNA methylation/demethylation. Further studies should examine the role of possible targets of VASA that may be implicated in germline epigenetic reprogramming.
Obviously, the low frequency of haploid cells produced suggests that we still have not optimized meiotic progression and that other factors must be needed to quantitatively and qualitatively regulate meiosis in vitro; as other groups have reported previously, we note that the gonadal niche is likely to be critical for proper meiotic initiation [43, 44]. Nonetheless, our results suggest that the evolutionarily divergent RNA-binding proteins, DAZL and VASA, function in promoting meiotic progression.
Conclusion
This study demonstrates that evolutionarily unrelated and divergent RNA-binding proteins can promote meiotic progression of human germ cells derived in vitro. Results provide a framework for exploring specifics of human meiosis, a program that is remarkably susceptible to errors that manifest commonly in human aneuploidy and disease. The fact that several families of RNA-binding proteins have been so highly conserved in the evolution of all metazoans suggests that the formation and maturation of germ cells is dependent on post-translational regulation by VASA and other RNA-binding proteins that may include NANOS, PUMILIO, and PIWI as well as their target RNAs and interacting proteins. Further studies of the role of these conserved factors in human germ cell development will likely assist in the correctly executed differentiation of pluripotent stem cells to fully functional germ cells in vitro.
Supplementary Material
Acknowledgments
We thank members of the Reijo Pera Laboratory and the Institute for Stem Cell Biology and Regenerative Medicine at Stanford University for helpful discussions and technical assistance. This work was supported by the California Institute of Regenerative Medicine (RL1-00670 and RC1-00137 to R.A.R.P.), the Instituto de Salud Carlos III from the Spanish Ministry of Science (FI07/00011), and the National Institutes of Health (NICHD U54 HD068158 as part of the Specialized Cooperative Centers Program in Reproduction and Infertility Research).
Footnotes
Disclosure of Potential Conflicts of Interest: The authors indicate no potential conflicts of interest.
Author contributions: J.V.M.: conception and design, collection and assembly of data, data analysis and interpretation, and manuscript writing; C.R. and H.N.N.: collection and assembly of data and data analysis and interpretation; C.S.: financial support, data analysis and interpretation, and final approval of manuscript; R.A.R.P.: conception and design, financial support, provision of study material, data analysis and interpretation, manuscript writing, and final approval of manuscript.
References
- 1.Lawson KA, Hage WJ. Clonal analysis of the origin of primordial germ cells in the mouse. Ciba Found Symp. 1994;182:68–84. doi: 10.1002/9780470514573.ch5. [DOI] [PubMed] [Google Scholar]
- 2.Ginsburg M, Snow MH, McLaren A. Primordial germ cells in the mouse embryo during gastrulation. Development. 1990;110:521–528. doi: 10.1242/dev.110.2.521. [DOI] [PubMed] [Google Scholar]
- 3.Hayashi K, de Sousa Lopes SM, Surani MA. Germ cell specification in mice. Science. 2007;316:394–396. doi: 10.1126/science.1137545. [DOI] [PubMed] [Google Scholar]
- 4.Saitou M, Barton SC, Surani MA. A molecular programme for the specification of germ cell fate in mice. Nature. 2002;418:293–300. doi: 10.1038/nature00927. [DOI] [PubMed] [Google Scholar]
- 5.Geijsen N, Horoschak M, Kim K, et al. Derivation of embryonic germ cells and male gametes from embryonic stem cells. Nature. 2004;427:148–154. doi: 10.1038/nature02247. [DOI] [PubMed] [Google Scholar]
- 6.Hubner K, Fuhrmann G, Christenson LK, et al. Derivation of oocytes from mouse embryonic stem cells. Science. 2003;300:1251–1256. doi: 10.1126/science.1083452. [DOI] [PubMed] [Google Scholar]
- 7.Nayernia K, Nolte J, Michelmann HW, et al. In vitro-differentiated embryonic stem cells give rise to male gametes that can generate offspring mice. Dev Cell. 2006;11:125–132. doi: 10.1016/j.devcel.2006.05.010. [DOI] [PubMed] [Google Scholar]
- 8.Bucay N, Yebra M, Cirulli V, et al. A novel approach for the derivation of putative primordial germ cells and sertoli cells from human embryonic stem cells. Stem Cells. 2009;27:68–77. doi: 10.1634/stemcells.2007-1018. [DOI] [PubMed] [Google Scholar]
- 9.Chen H, Kuo H, Chien C, et al. Derivation, characterization and differentiation of human embryonic stem cells: Comparing serum-containing versus serum-free media and evidence of germ cell differentiation. Hum Reprod. 2007;22:567–577. doi: 10.1093/humrep/del412. [DOI] [PubMed] [Google Scholar]
- 10.Clark AT, Bodnar MS, Fox MS, et al. Spontaneous differentiation of germ cells from human embryonic stem cells in vitro. Hum Mol Genet. 2004;13:727–739. doi: 10.1093/hmg/ddh088. [DOI] [PubMed] [Google Scholar]
- 11.Tilgner K, Atkinson S, Golebiewska A, et al. Isolation of primordial germ cells from differentiating human embryonic stem cells. Stem Cells. 2008;26:3075–3085. doi: 10.1634/stemcells.2008-0289. [DOI] [PubMed] [Google Scholar]
- 12.Park T, Galic Z, Conway A, et al. Derivation of primordial germ cells from human embryonic and induced pluripotent stem cells is significantly improved by coculture with human fetal gonadal cells. Stem Cells. 2009;27:783–795. doi: 10.1002/stem.13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.White-Cooper H, Schafer MA, Alphey LS, et al. Transcriptional and post-transcriptional control mechanisms coordinate the onset of spermatid differentiation with meiosis I in Drosophila. Development. 1998;125:125–134. doi: 10.1242/dev.125.1.125. [DOI] [PubMed] [Google Scholar]
- 14.Juliano C, Wessel G. Developmental biology. Versatile germline genes Science. 2010;329:640–641. doi: 10.1126/science.1194037. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Ruggiu M, Speed R, Taggart M, et al. The mouse Dazla gene encodes a cytoplasmic protein essential for gametogenesis. Nature. 1997;389:73–77. doi: 10.1038/37987. [DOI] [PubMed] [Google Scholar]
- 16.Xu EY, Moore FL, Reijo Pera RA. A gene family required for human germ cell development evolved from an ancient meiotic gene conserved in metazoans. Proc Natl Acad Sci USA. 2001;98:7414–7419. doi: 10.1073/pnas.131090498. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Reynolds N, Collier B, Bingham V, et al. Translation of the synaptonemal complex component Sycp3 is enhanced in vivo by the germ cell specific regulator Dazl. RNA. 2007;13:974–981. doi: 10.1261/rna.465507. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Eberhart CG, Maines JZ, Wasserman SA. Meiotic cell cycle requirement for a fly homologue of human Deleted in Azoospermia. Nature. 1996;381:783–785. doi: 10.1038/381783a0. [DOI] [PubMed] [Google Scholar]
- 19.Karashima T, Sugimoto A, Yamamoto M. Caenorhabditis elegans homologue of the human azoospermia factor DAZ is required for oogenesis but not for spermatogenesis. Development. 2000;127:1069–1079. doi: 10.1242/dev.127.5.1069. [DOI] [PubMed] [Google Scholar]
- 20.Houston DW, Zhang J, Maines JZ, et al. A Xenopus DAZ-like gene encodes an RNA component of germ plasm and is a functional homologue of Drosophila boule. Development. 1998;125:171–180. doi: 10.1242/dev.125.2.171. [DOI] [PubMed] [Google Scholar]
- 21.Kee K, Angeles V, Flores M, et al. Human DAZL, DAZ and Boule genes modulate primordial germ cell and haploid gamete formation. Nature. 2009;462:222–225. doi: 10.1038/nature08562. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Panula S, Medrano JV, Kee K, et al. Human germ cell differentiation from fetal- and adult-derived induced pluripotent stem cells. Hum Mol Genet. 2011;20:752–762. doi: 10.1093/hmg/ddq520. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Castrillon DH, Quade BJ, Wang TY, et al. The human VASA gene is specifically expressed in the germ cell lineage. Proc Natl Acad Sci USA. 2000;97:9585–9590. doi: 10.1073/pnas.160274797. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Lasko PF, Ashburner M. The product of the Drosophila gene vasa is very similar to eukaryotic initiation factor-4A. Nature. 1988;335:611–617. doi: 10.1038/335611a0. [DOI] [PubMed] [Google Scholar]
- 25.Fujiwara Y, Komiya T, Kawabata H, et al. Isolation of a Dead-family protein gene that encodes a murine homolog of Drosophila Vasa and its specific expression in germ cell lineage. Proc Natl Acad Sci USA. 1994;91:12258–12262. doi: 10.1073/pnas.91.25.12258. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Gruidl ME, Smith PA, Kuznicki KA, et al. Multiple potential germ-line helicases are components of the germ-line-specific P granules of Caenorhabditis elegans. Proc Natl Acad Sci USA. 1996;93:13837–13842. doi: 10.1073/pnas.93.24.13837. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Komiya T, Itoh K, Ikenishi K, et al. Isolation and characterization of a novel gene of the dead box protein family which is specifically expressed in germ cells of Xenopus laevis. Dev Biol. 1994;162:354–363. doi: 10.1006/dbio.1994.1093. [DOI] [PubMed] [Google Scholar]
- 28.Mohr S, Stryker JM, Lambowitz AM. A DEAD-box protein functions as an ATP-dependent RNA chaperone in group I intron splicing. Cell. 2002;109:769–779. doi: 10.1016/s0092-8674(02)00771-7. [DOI] [PubMed] [Google Scholar]
- 29.Tanaka SS, Toyooka Y, Akasu R, et al. The mouse homolog of Drosophila Vasa is required for the development of male germ cells. Gene Dev. 2000;14:841–853. [PMC free article] [PubMed] [Google Scholar]
- 30.Lavial F, Acloque H, Bachelard E, et al. Ectopic expression of Cvh (Chicken Vasa homologue) mediates the reprogramming of chicken embryonic stem cells to a germ cell fate. Dev Biol. 2009;330:73–82. doi: 10.1016/j.ydbio.2009.03.012. [DOI] [PubMed] [Google Scholar]
- 31.Vandesompele J, De Preter K, Pattyn F, et al. Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biol. 2002;3:RESEARCH0034. doi: 10.1186/gb-2002-3-7-research0034. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Monk M, Boubelik M, Lehnert S. Temporal and regional changes in DNA methylation in the embryonic, extraembryonic and germ cell lineages during mouse embryo development. Development. 1987;99:371–382. doi: 10.1242/dev.99.3.371. [DOI] [PubMed] [Google Scholar]
- 33.Sato S, Yoshimizu T, Sato E, et al. Erasure of methylation imprinting of Igf2r during mouse primordial germ-cell development. Mol Reprod Dev. 2003;65:41–50. doi: 10.1002/mrd.10264. [DOI] [PubMed] [Google Scholar]
- 34.Hajkova P, Erhardt S, Lane N, et al. Epigenetic reprogramming in mouse primordial germ cells. Mech Dev. 2002;117:15–23. doi: 10.1016/s0925-4773(02)00181-8. [DOI] [PubMed] [Google Scholar]
- 35.Zhang Y, Tycko B. Monoallelic expression of the human H19 gene. Nat Genet. 1992;1:40–44. doi: 10.1038/ng0492-40. [DOI] [PubMed] [Google Scholar]
- 36.Bartolomei MS, Zemel S, Tilghman SM. Parental imprinting of the mouse H19 gene. Nature. 1991;351:153–155. doi: 10.1038/351153a0. [DOI] [PubMed] [Google Scholar]
- 37.Gustafson EA, Yajima M, Juliano CE, et al. Post-translational regulation by gustavus contributes to selective Vasa protein accumulation in multipotent cells during embryogenesis. Dev Biol. 2011;349:440–450. doi: 10.1016/j.ydbio.2010.10.031. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Medrano JV, Marques-Mari AI, Aguilar CE, et al. Comparative analysis of the germ cell markers c-KIT, SSEA-1 And Vasa in testicular biopsies from secretory and obstructive Azoospermias. Mol Hum Reprod. 2010;16:811–817. doi: 10.1093/molehr/gaq044. [DOI] [PubMed] [Google Scholar]
- 39.Megosh HB, Cox DN, Campbell C, et al. The role of PIWI and the miRNA machinery in Drosophila germline determination. Curr Biol. 2006;16:1884–1894. doi: 10.1016/j.cub.2006.08.051. [DOI] [PubMed] [Google Scholar]
- 40.Kuramochi-Miyagawa S, Watanabe T, Gotoh K, et al. MVH in piRNA processing and gene silencing of retrotransposons. Genes Dev. 2010;24:887–892. doi: 10.1101/gad.1902110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Florke-Gerloff S, Topfer-Petersen E, Muller-Esterl W, et al. Acrosin and the acrosome in human spermatogenesis. Hum Genet. 1983;65:61–67. doi: 10.1007/BF00285030. [DOI] [PubMed] [Google Scholar]
- 42.Reynolds N, Collier B, Maratou K, et al. Dazl binds in vivo to specific transcripts and can regulate the pre-meiotic translation of Mvh in germ cells. Hum Mol Genet. 2005;14:3899–3909. doi: 10.1093/hmg/ddi414. [DOI] [PubMed] [Google Scholar]
- 43.McLaren A. Germ cells and germ cell sex. Philos Trans R Soc Lond B Biol Sci. 1995;350:229–233. doi: 10.1098/rstb.1995.0156. [DOI] [PubMed] [Google Scholar]
- 44.McLaren A. Primordial germ cells in the mouse. Dev Biol. 2003;262:1–15. doi: 10.1016/s0012-1606(03)00214-8. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.


