Abstract
We report the implementation of an infrared laser onto our previously reported matrix-assisted laser desorption electrospray ionization (MALDESI) source with ESI post-ionization yielding multiply charged peptides and proteins. Infrared (IR)-MALDESI is demonstrated for atmospheric pressure desorption and ionization of biological molecules ranging in molecular weight from 1.2 to 17 kDa. High resolving power, high mass accuracy single-acquisition Fourier transform ion cyclotron resonance (FT-ICR) mass spectra were generated from liquid-and solid-state peptide and protein samples by desorption with an infrared laser (2.94 µm) followed by ESI post-ionization. Intact and top-down analysis of equine myoglobin (17 kDa) desorbed from the solid state with ESI post-ionization demonstrates the sequencing capabilities using IR-MALDESI coupled to FT-ICR mass spectrometry. Carbohydrates and lipids were detected through direct analysis of milk and egg yolk using both UV- and IR-MALDESI with minimal sample preparation. Three of the four classes of biological macromolecules (proteins, carbohydrates, and lipids) have been ionized and detected using MALDESI with minimal sample preparation. Sequencing of O-linked glycans, cleaved from mucin using reductive β-elimination chemistry, is also demonstrated.
The recent growth in the development of hybrid ambient direct analysis techniques such as laser desorption atmospheric pressure chemical ionization (LD-APCI) [1], direct analysis in real time (DART) [2], desorption electrospray ionization (DESI) [3], atmospheric-pressure solids analysis probe (ASAP) [4], electrospray-assisted laser desorption ionization (ELDI) [5, 6], matrix-assisted laser desorption electrospray ionization (MALDESI) [7, 8], laser ablation electrospray ionization (LAESI) [9], and infrared laser-assisted desorption electrospray ionization (IR-LADESI) [10] have rapidly expanded the analytical space accessible using mass spectrometry. LAESI and IR-LADESI were introduced around the same time and are in principle the same ionization method that may be better described as IR-MALDESI, whereas endogenous water or sacrificial analyte serves as the matrix for infrared matrix-assisted laser desorption followed by ESI post-ionization. All of these techniques achieve ionization at atmospheric pressure with minimal sample preparation, reducing sample handling and preparation times. Earlier work has reported the ionization of biomolecules from liquids held in vacuo using IR radiation [11–13]. MALDESI and other similar laser desorption with ESI post-ionization techniques generate multiply charged ions and have the added advantage of increased top-down fragmentation efficiency, thereby enhancing sequencing capabilities that are invaluable for the identification and characterization of biological molecules using a variety of fragmentation methodologies including collision-induced dissociation (CID), electron transfer dissociation (ETD), electron capture dissociation (ECD), sustained off-resonance irradiation (SORI), and infrared multiphoton dissociation (IRMPD) [14– 18]. The detection of multiply charged ions in Fourier transform ion cyclotron resonance (FT-ICR) and Orbitrap (LTQ-Orbi) yields increased resolving power and increased mass accuracy (<3 parts per million [ppm]) [19, 20], attributed to the inverse relationship between the frequency and m/z and √(m/z), respectively [21].
Poor shot-to-shot reproducibility has been an issue with UV matrix-assisted laser desorption techniques as a result of the inherent inhomogeneity of analyte/ matrix co-crystallization [22]. This inhomogeneity has been addressed by using a liquid sample, often mixed with glycerol for vacuum compatibility, thereby eliminating the co-crystallization step in sample preparation [23, 24]. In building upon this idea to improve shot-to-shot reproducibility, liquid sample droplets have been used for atmospheric pressure analysis using liquid matrix-assisted laser desorption electrospray ionization (liq-MALDESI), resulting in increased shot-to-shot reproducibility [25].
IR-MALDI has demonstrated desorption and ionization of large biomolecules using both solid and liquid matrices [26]. It is well established that infrared laser desorption generates many more neutral molecules than ions, thereby reducing mass spectrometric detection sensitivity. Although IR-MALDI has demonstrated practical utility for analysis of complex samples, the desorption and ionization mechanisms are not well understood. It has been proposed that water molecules of hydration associated with the analyte (i.e., proteins) as well as the analyte itself absorb infrared laser irradiation, resulting in O—H, C—H, and N—H bond stretching, leading to the desorption of analyte and serving as a possible source of protons for ionization [27–29]. The surrounding solvent may also act to isolate the analyte, reducing cluster formation and collisionally cool the desorbed molecules, thereby reducing fragmentation.
The advantages of IR laser desorption stem from the capability of desorption and ionization directly from samples without the need for adding exogenous matrix [27, 28, 30]. This combined with the deep penetration of infrared laser ablation results in ablation of large amounts of material, the majority of which are neutral molecules, making infrared an ideal wavelength range for laser desorption using the MALDESI platform.
Demonstrated herein we have integrated an IR laser (2.94 µm) into the existing versatile MALDESI ionization source [31]. IR laser desorption from liquid- and solid-state samples followed with ESI post-ionization at atmospheric pressure generates multiply charged peptide and protein ions. High mass accuracy for intact and top-down characterization of biomolecules (i.e., myoglobin, 17 kDa) is demonstrated. Direct analyses of commercially produced milk without sample preparation and farm-raised eggs are demonstrated using both UV (349 nm) and IR (2.94 µm) laser desorption. O-linked glycans cleaved from mucin [32–34] were sequenced from liquid-state samples using IR-MALDESI. Importantly, proteins, carbohydrates, and lipids were detected with minimal sample preparation using IR-MALDESI coupled to FT-ICR mass spectrometry.
Experimental
Materials
Substance P, somatostatin, laminin, human angiotensin I, honey bee melittin, porcine glucagon, bovine ubiquitin, bovine cytochrome c, equine myoglobin, succinic acid, and glycerol were purchased from Sigma–Aldrich (St. Louis, MO, USA) and used as received. Bovine milk (2%) was purchased from a local grocery store and used without further preparation. Farm-raised chicken eggs were prepared as indicated. Porcine mucin was purchased from Sigma–Aldrich, from which O-linked glycans were cleaved from proteins using reductive β-elimination and purification as previously reported [32–34]. The specifics for the method used are detailed in several recent publications by our group [32, 34, 35]. O-linked glycan samples were from solid-phase extraction elution 1 (10% acetonitrile) and used without further preparation. HPLC-grade acetonitrile, methanol, and high-purity water were purchased from Burdick and Jackson (Muskegon, MI, USA).
IR- and UV-MALDESI Mass Spectrometry
An erbium-doped yttrium aluminum garnet (ER-YAG) infrared laser (IR) (Bioptic Lasersysteme, Berlin, Germany) was mounted to the main working platform of the previously described MALDESI source [31]. The IR laser generated 2.8 mJ pulse energy (measured at the laser) at 10-Hz repetition rate with a pulse width of 100 ns and emission wavelength of 2.94 µm. The laser-beam path was directed by two broadband gold-coated Pyrex mirrors (Part #10D20ER.4; Newport Corporation, Irvine, CA, USA) and focused to a spot size of 200 µm using a calcium fluoride plano convex lens (Part #47174; Edmund Optics, Barrington, NJ, USA) onto the stainless steel sample target (Part #4333375; Applied Biosystems, Foster City, CA, USA). The laser fluence was calculated at 2.2 J/cm2.
The ultraviolet (UV) laser (Explorer; Newport Corporation, Irvine, CA, USA) with an emission wavelength of 349 nm, computer-controlled pulse energy (0–120 µJ) and repetition rate (0–5000 Hz) was used with the same experimental setup as previously described [31]. Briefly, two UV-enhanced aluminum mirrors (Part #10D20AL.2; Newport Corporation, Irvine, CA, USA) direct the laser beam, which is focused to a spot size of 60 µm by a fused silica lens (Part #SPX017 + AR.10, Newport Corporation, Irvine, CA, USA) onto the sample target; the laser power was set to 50 µJ, measured internally. The laser fluence was calculated at 0.4 J/cm2.
The sample target was positioned in front of and below the extended ion transfer capillary of a hybrid LTQ-FT-ICR mass spectrometer. The LTQ-FT-ICR was equipped with a 7 Tesla actively shielded superconducting magnet; the maximum ionization time was set to 1 s and resolving power set to 100,000 at 400 m/z. Electrospray ionization was generated by infusing the ESI solution (50% acetonitrile in water) through a 75-µm i.d. fused silica capillary (Polymicro Technologies, Phoenix, AZ, USA) connected with a stainless steel union (Part #ZU1XC; VICI, Houston, TX, USA) to a 30-µm fused silica tapered Picotip (New Objective Inc., Woburn, MA, USA) at a flow rate of 400–800 nL/min using a syringe pump (Harvard Apparatus, Holliston, MA, USA). The stainless steel union connecting the transfer capillary to the ESI emitter tip was biased at 3 kV. All experiments were performed at ambient temperature and pressure. Top-down data analysis was processed manually by obtaining precursor m/z values from the tandem mass spectrometry (MS/MS) header and importing the MS/MS values into ProSightPC (Thermo Electron, Waltham, MA, USA) using a THRASH [36] algorithm to determine the monoisotopic peak of the analyte signal, as detailed in a recent publication by our group [37].
Results and Discussion
Solid-State IR-MALDESI-FT-ICR
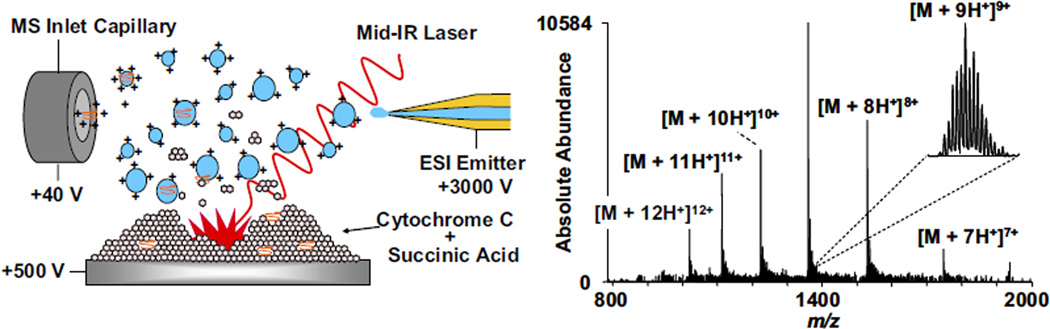
An aqueous solution of bovine cytochrome c was mixed 1:1 (v:v) with succinic acid (47 mg/mL) in 50% acetonitrile, to give a final concentration of 200 µM. The sample solution was deposited (0.8 µL) onto a stainless steel sample target and actively dried under a cool stream of air [38]. The sample target was installed onto the MALDESI platform equipped with an infrared laser for IR-MALDESI analysis. The IR laser was manually actuated while 50% acetonitrile in water was electrosprayed at 3 kV and a flow rate of 800 nL/min; the sample target was biased at 500 V. A schematic of solid-state IR-MALDESI and representative mass spectrum of cytochrome c are shown in Figure 1. Each single acquisition mass spectrum demonstrated high resolving power multiply charged ions of cytochrome c for accurate mass determination and protein identification. The average charge state (ACS) for cytochrome c was calculated using the following equation: ACS = [Σ(charge state of each peak × abundance of each peak)]/(total abundance of peaks) and found to be ACS = 9.41. Similar high resolving power multiply charged mass spectra were obtained from aqueous solutions of angiotensin I, somatostatin, laminin, melittin, glucagon, ubiquitin, and myoglobin, each mixed with succinic acid and actively dried before analysis (data not shown), demonstrating desorption and ionization over a broad peptide and protein molecular weight range (1.2–17 kDa), similar to data previously demonstrated by UV-MALDESI.
Figure 1.
Schematic of solid-state IR-MALDESI with ESI post-ionization and representative mass spectrum of bovine cytochrome c mixed with succinic acid.
Control experiments were performed by laser desorption of the same solid samples with the ESI emitter removed to determine whether ionization occurs via atmospheric-pressure IR-MALDI. All control experiments resulted in no measurable analyte signal without ESI post-ionization.
In a separate experiment, the ESI emitter was removed and the recently introduced remote analyte sampling, transport, and ionization relay (RASTIR) was installed [39]. The RASTIR device serves to transport the neutral laser desorption products by induced vacuum to the ionization region positioned directly in front of the mass spectrometer inlet capillary. This device separates in space the complex electrical fields of the sample target and the ESI emitter, enabling independent optimization of each component. The electrospray emitter of the RASTIR device was biased at 3 kV, whereas the RASTIR body and the sample target were allowed to float (not biased or grounded). Cytochrome c and myoglobin, each mixed with succinic acid and actively dried, were subjected to IR laser desorption in close proximity to the collection tube of the RASTIR device. The laser desorption products were transported into the ionization region of the RASTIR device and subsequently ionized; multiply charged ions of cytochrome c and myoglobin, similar to those obtained without the RASTIR device were detected (data not shown).
The ionization mechanism we propose is similar to that proposed for solid-state UV-MALDESI [7]; laser desorbed neutral molecules or particles are absorbed into the electrospray-charged solvent droplets, which then undergo an ESI-like desorption and ionization process, generating multiply charged ions. We base our hypothesis on the similarity of the observed charge states to those obtained by ESI and the absence of ion signal with the ESI emitter removed, eliminating an AP-MALDI mechanism. The RASTIR-IR-MALDESI experiments demonstrate remote electrospray postionization of laser desorption products and can therefore rule out a DESI-like mechanism ascribed to separation of the desorption event from the ionization event in both time and space as well as eliminating the possibility of laser desorption from a charged liquid droplet from the surface (the sample target is not biased and the ESI emitter is located remote from the laser desorption region).
This initial work is the first demonstration of RASTIR combined with IR-MALDESI for the sole purpose of separating the desorption and ionization regions of MALDESI and may be invaluable in the characterization of the ionization mechanism. The complex electric fields of the desorption and ionization regions arising from the ESI emitter and sample target voltages in MALDESI can be avoided by implementing the RASTIR device, thereby allowing independent control of the ESI and sample target voltages for optimal desorption and ionization.
Liquid-State IR-MALDESI-FT-ICR
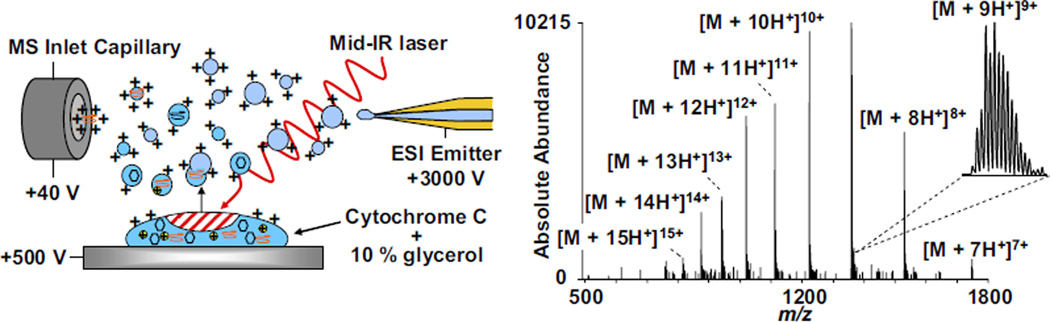
Aqueous samples of cytochrome c containing 10% glycerol were deposited (1.5 µL) and analyzed without drying by IR-MALDESI with ESI post-ionization. A schematic of liquid-state IR-MALDESI and representative mass spectrum of cytochrome c are shown in Figure 2. The sample target voltage (500 V), ESI voltage (3 kV), and ESI flow rate (800 nL/min) were the same as those used in solid-state analysis. The liquid sample drop yielded signal only while the sample remained liquid on the surface (~1 min); no signal was obtained after the droplet dried on the surface. Similar high ion abundances were observed, indicating the versatility of desorption and ionization from both solid-state and liquid-state samples using this technique. The ACS of cytochrome c desorbed from the liquid state was calculated using the same equation as for solid-state cytochrome c (see earlier text) and found to be ACS = 10.43. The average charge state for liquid-state IR-MALDESI (10.4) was found to be one whole charge higher than the average charge state for solid-state IR-MALDESI (9.4). One possible explanation for the increase in average charge state with liquid-state desorption is the absence of charge-sequestering matrix molecules that are desorbed from the solid-state sample with every laser shot. Another explanation is that in the liquid state analyte may be more efficiently desorbed (less energy is required to liberate the analyte from solution than from a solid lattice).
Figure 2.
Schematic of liquid-state IR-MALDESI with ESI post-ionization and representative mass spectrum of bovine cytochrome c mixed with 10% glycerol.
The potential for minimal sample preparation (i.e., no additional matrix) was explored by preparing analyte solutions, without the addition of the matrix glycerol. IR-MALDESI analysis of substance P, somatostatin, laminin, angiotensin I, melittin, glucagon, ubiquitin, cytochrome c, and myoglobin demonstrated desorption and ionization via IR-MALDESI without addition of matrix, generating multiply charged ions (data not shown).
Control experiments were performed using new droplets of the same analyte solutions (no matrix) with the ESI emitter removed and high potential (0–5000 V) applied directly to the sample target. Only the liquid-state IR-MALDESI mass spectrum of angiotensin I revealed analyte signal, yielding only the singly charged species; the doubly charged ion was not observed. The singly charged ion indicates an atmospheric-pressure MALDI ionization process from the liquid droplet. None of the other peptides or proteins used in these control experiments demonstrated measurable analyte signal without ESI post-ionization.
Intact and Top-Down Analysis Using IR-MALDESI-FT-ICR
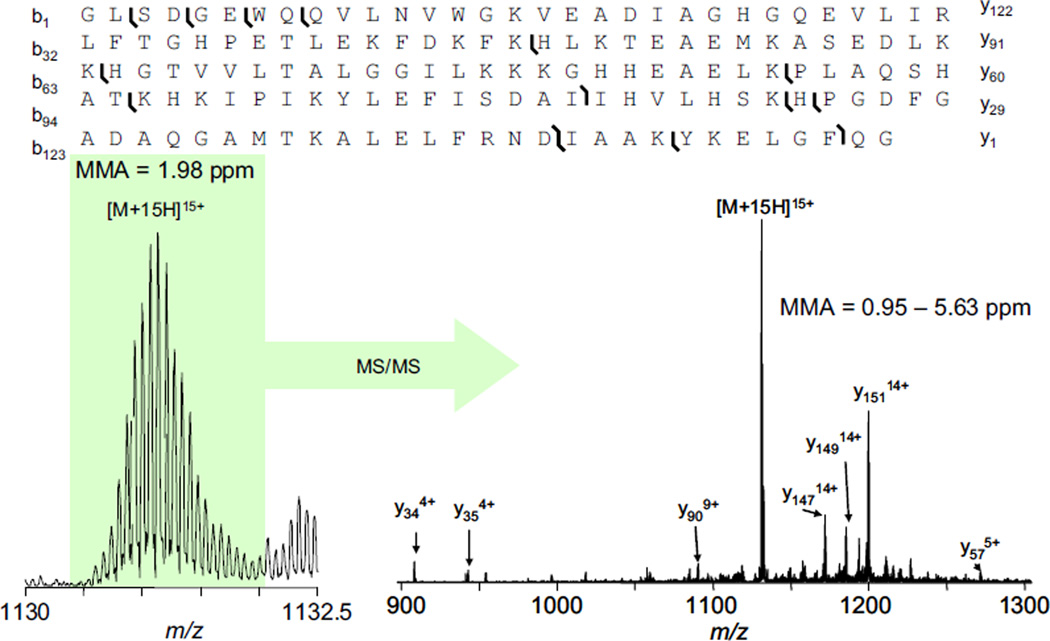
Equine myoglobin mixed with succinic acid was deposited onto the sample target and dried with a cool stream of air. The sample was desorbed and ionized by IR-MALDESI and sampled by the mass spectrometer. The most abundant charge state, [M + 15H+]15+, was isolated for CID; the precursor and fragment ions were detected in the ICR cell. The expanded region of the precursor is shown in the lower left of Figure 3, with the MS/MS spectrum with the precursor and fragment ions shown in the lower right of Figure 3. The amino acid sequence with the identified tandem MS fragments indicated is shown in the top of Figure 3. The mass measurement accuracy for the precursor (MMA = 1.98 ppm) and fragment ions (MMA = 0.95–5.63 ppm) were calculated manually.
Figure 3.
Top-down sequencing of equine myoglobin using solid-state IR-MALDESI with ESI post-ionization.
Direct Analysis Using UV- and IR-MALDESI
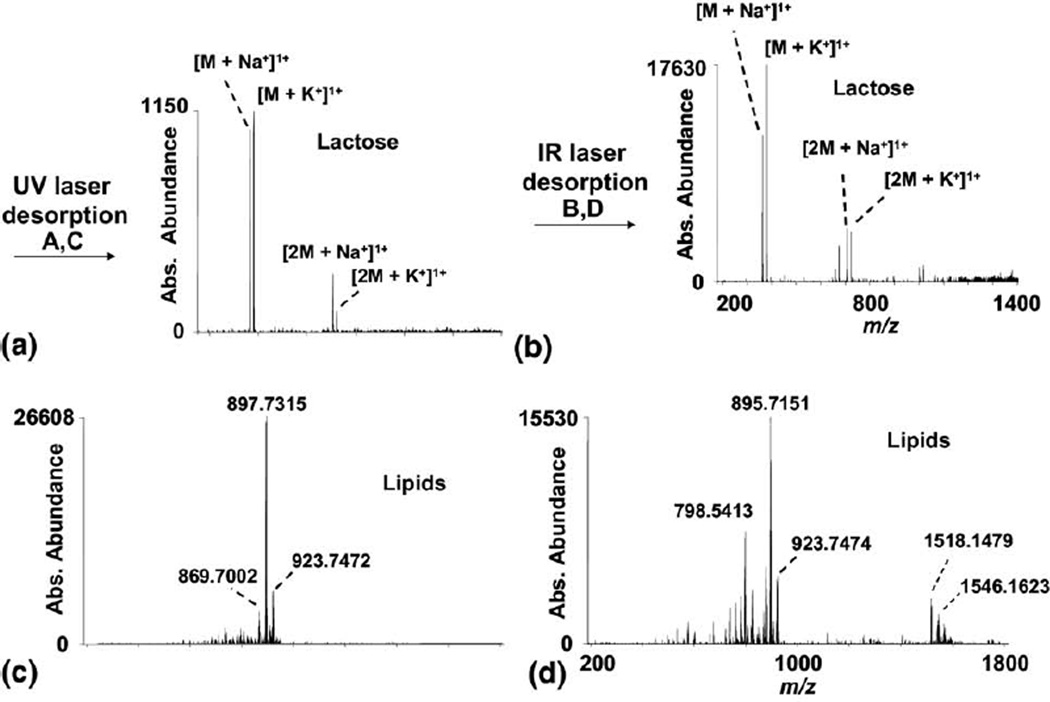
Direct analysis via UV and IR laser desorption with ESI post-ionization is demonstrated for bovine milk and chicken egg yolk as shown in Figure 4. The milk was used without preparation and analyzed in the liquid state. The UV- and IR-MALDESI mass spectra of bovine milk are shown in Figure 4a and b, respectively, both with ESI post-ionization. The main peaks observed in the mass spectra were identified as the sodium and potassium adducts of lactose as well as the sodium and potassium adducted dimers of lactose. The IR-MALDESI mass spectrum has several other unidentified peaks, indicating desorption and ionization of other components of the milk sample not observed in the UV-MALDESI mass spectrum. The absolute abundance was substantially higher when using the IR laser compared with UV laser desorption. The increased abundance may be attributed to the increased number of neutral molecules desorbed by IR laser ablation available for ESI post-ionization.
Figure 4.
Liquid-state MALDESI mass spectrum of bovine milk with no sample preparation using: (a) UV laser desorption, (b) IR laser desorption with ESI post-ionization and egg yolk using (c) UV laser desorption with DHB and (d) IR laser desorption with no sample preparation with ESI post-ionization.
The UV- and IR-MALDESI mass spectra of chicken egg yolk (mixed with 2,5-dihydroxybenzoic acid [DHB] for UV laser desorption) are shown in Figure 4c and d, respectively. The larger ion abundance observed for UV-MALDESI is likely attributable to incorporation of the UV absorbing matrix (DHB), which was required to obtain appreciable signal. The unidentified main peaks of the resulting mass spectra were determined to be lipids, based on their mass excesses. The mass excess (defined by IUPAC) is calculated by the difference between the nominal mass and the exact mass; the mass excess was determined for each of the main peaks in the UV- and IR-MALDESI mass spectra of egg yolk. The mass excesses were plotted onto a heat map of the mass excess of tryptic peptides versus the monoisotopic mass, which demonstrates the forbidden zones for tryptic peptides [40]. The mass excess of the peaks fell outside the range normally occupied by tryptic peptides, indicating that the observed ions are probably not peptides nor proteins. The mass excesses of several common lipids were plotted onto the same heat map, which fell into the same range, indicating that these are likely lipids. A small group of higher mass peaks were observed in the IR-MALDESI mass spectra of egg yolk (Figure 4d) that were not observed in the UV-MALDESI mass spectra of egg yolk (Figure 4c). These higher mass ions may indicate more efficient desorption of higher mass molecules from complex samples using IR-MALDESI compared with UV-MALDESI.
Carbohydrate Analysis Using IR-MALDESI
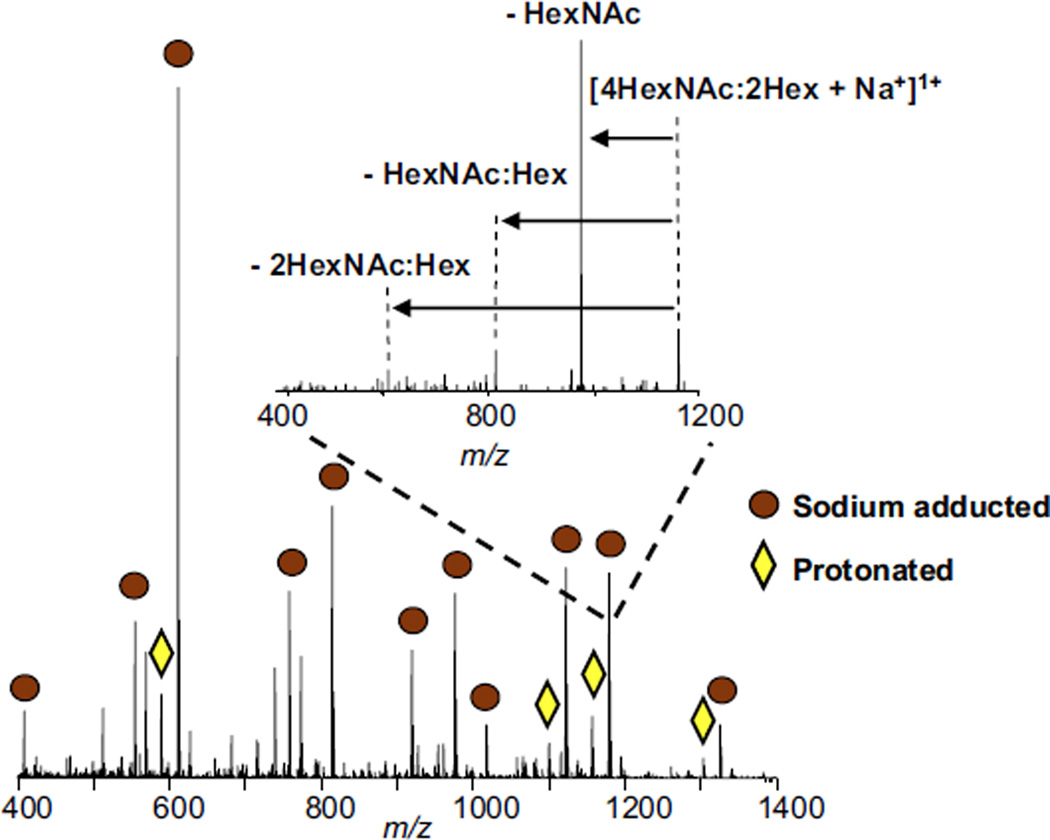
Liquid-state samples (0.8 µL) of elution 1 from the β-elimination/solid-phase extraction process were deposited onto the sample target for direct IR-MALDESI analysis with ESI post-ionization. The IR-MALDESI mass spectrum is shown in Figure 5. The circles denote identified sodium adducted O-linked glycans. The diamonds denote identified protonated O-linked glycans. The identification of the glycans was verified by exact mass and tandem MS. A representative glycan ([4HexNAc:2Hex+Na+]1, m/z = 1179.4386) was isolated and subsequently fragmented with CID and is shown in the inset of Figure 5. The fragment ions correspond to losses of HexNAc, HexNAc:Hex, and 2HexNac:Hex; the mass measurement accuracy associated with this precursor peak was −0.34 ppm. The IR-MALDESI mass spectrum was compared with MALDI-FT-ICR and DESI-FT-ICR mass spectra of the same elution, as previously reported [35]. The IR-MALDESI mass spectra yielded many of the same peaks identified using MALDI and DESI. MALDESI avoids fragmentation commonly encountered when using MALDI and, when coupled with a hybrid mass analyzer, provides top-down sequence information and high mass accuracy. MALDESI is also much more sensitive, requiring approximately one fourth the amount of material required for DESI, as demonstrated in our laboratory [35] for the analysis of carbohydrates.
Figure 5.
Liquid-state IR-MALDESI of O-linked glycans cleaved from mucin using reductive β-elimination chemistry. Circles denote sodium adducted glycans; diamonds denote protonated glycans. Inset shows MS/MS sequencing of glycan with m/z = 1179.4386.
Conclusions
IR-MALDESI is an atmospheric-pressure ionization source for analysis of biomolecules with minimal sample preparation requiring no pretreatment necessary for vacuum-sensitive samples (i.e., tissues). The liquid- or solid-state sample remains accessible during analysis, allowing flexibility in sample preparation (i.e., solid or liquid) as well as with or without added matrix and the type of matrix used, demonstrating the inherent flexibility and versatility for biomolecular analysis. IR-MALDESI intact and top-down proteomic strategies have been used for identification and characterization of biological molecules, including proteins and glycans, which are invaluable for analysis of complex samples (e.g., tissues, biofluids). IR-MALDESI direct analysis without addition of an exogenous matrix has the unique potential for direct analysis, without sample preparation, for generation of multiply charged ions from biological samples by using the endogenous matrix (solvent or sacrificial analyte). Furthermore, the inherent versatility of the MALDESI ionization source is demonstrated to include both UV and IR laser desorption with ESI post-ionization for the generation of multiply charged ions and the potential application for other desorption and ionization techniques. Future studies include using the RASTIR device to gain a clear understanding of the IR-MALDESI ionization mechanism that will not only enhance the capabilities of this technique but also potentially elucidate other laser desorption/ionization mechanisms (i.e., MALDI).
Acknowledgments
The authors thank the machine shop personnel in the College of Physical and Mathematical Sciences at North Carolina State University for their assistance with the MALDESI source construction. The authors thank R. Brent Dixon for assistance in implementing the RASTIR device, Michael S. Bereman for O-glycan mucin sample preparation, and Timothy S. Collier for database searching of the top-down data. The authors gratefully acknowledge financial support received from the National Cancer Institute, National Cancer Institute Grant R33 CA-105295, the W.M. Keck Foundation, the William R. Kenan, Jr. Fund for Engineering, Technology & Science, and North Carolina State University. One of us (KKM) acknowledges support from the National Science Foundation through Grant CHE0415360.
References
- 1.Coon JJ, Harrison WW. Laser Desorption-Atmospheric Pressure Chemical Ionization Mass Spectrometry for the Analysis of Peptides from Aqueous Solutions. Anal. Chem. 2002;74:5600–5605. doi: 10.1021/ac020402k. [DOI] [PubMed] [Google Scholar]
- 2.Cody RB, Laramee JA, Durst HD. Versatile New Ion Source for the Analysis of Materials in Open Air under Ambient Conditions. Anal. Chem. 2005;77:2297–2302. doi: 10.1021/ac050162j. [DOI] [PubMed] [Google Scholar]
- 3.Cooks RG, Ouyang Z, Takats Z, Wiseman JM. Ambient Mass Spectrometry. Science. 2006;311:1566–1570. doi: 10.1126/science.1119426. [DOI] [PubMed] [Google Scholar]
- 4.McEwen CN, McKay RG, Larsen BS. Analysis of Solids, Liquids, and Biological Tissues Using Solids Probe Introduction at Atmospheric Pressure on Commercial LC/MS Instruments. Anal. Chem. 2005;77:7826–7831. doi: 10.1021/ac051470k. [DOI] [PubMed] [Google Scholar]
- 5.Shiea J, Huang MZ, Hsu HJ, Lee CY, Yuan CH, Beech I, Sunner J. Electrospray-Assisted Laser Desorption/Ionization Mass Spectrometry for Direct Ambient Analysis of Solids. Rapid Commun. Mass Spectrom. 2005;19:3701–3704. doi: 10.1002/rcm.2243. [DOI] [PubMed] [Google Scholar]
- 6.Huang MZ, Hsu HJ, Lee JY, Jeng J, Shiea J. Direct Protein Detection from Biological Media through Electrospray-Assisted Laser Desorption Ionization/Mass Spectrometry. J. Proteome Res. 2006;5:1107–1116. doi: 10.1021/pr050442f. [DOI] [PubMed] [Google Scholar]
- 7.Sampson JS, Hawkridge AM, Muddiman DC. Generation and Detection of Multiply-Charged Peptides and Proteins by Matrix- Assisted Laser Desorption Electrospray Ionization (MALDESI) Fourier Transform Ion Cyclotron Resonance Mass Spectrometry. J. Am. Soc. Mass Spectrom. 2006;17:1712–1716. doi: 10.1016/j.jasms.2006.08.003. [DOI] [PubMed] [Google Scholar]
- 8.Sampson JS, Hawkridge AM, Muddiman DC. Direct Characterization of Intact Polypeptides by Matrix-Assisted Laser Desorption Electrospray Ionization Quadrupole Fourier Transform Ion Cyclotron Resonance Mass Spectrometry. Rapid Commun. Mass Spectrom. 2007;21:1150–1154. doi: 10.1002/rcm.2947. [DOI] [PubMed] [Google Scholar]
- 9.Nemes P, Vertes A. Laser Ablation Electrospray Ionization for Atmospheric Pressure, In Vivo, and Imaging Mass Spectrometry. Anal. Chem. 2007;79:8098–8106. doi: 10.1021/ac071181r. [DOI] [PubMed] [Google Scholar]
- 10.Rezenom YH, Dong J, Murray KK. Infrared Laser-Assisted Desorption Electrospray Ionization Mass Spectrometry. Analyst. 2008;133:226–232. doi: 10.1039/b715146b. [DOI] [PubMed] [Google Scholar]
- 11.Charvat A, Abel B. How to Make Big Molecules Fly Out of Liquid Water: Applications, Features and Physics of Laser Assisted Liquid Phase Dispersion Mass Spectrometry. Phys. Chem. Chem. Phys. 2007;9:3335–3360. doi: 10.1039/b615114k. [DOI] [PubMed] [Google Scholar]
- 12.Kleinekofort W, Avdiev J, Brutschy B. A New Method of Laser Desorption Mass Spectrometry for the Study of Biological Macromolecules. Int. J. Mass Spectrom. Ion Processes. 1996;152:135–142. [Google Scholar]
- 13.Grimm RL, Beauchamp JL. Dynamics of Field-Induced Droplet Ionization: Time-Resolved Studies of Distortion, Jetting, and Progeny Formation from Charged and Neutral Methanol Droplets Exposed to Strong Electric Fields. J. Phys. Chem. B. 2005;109:8244–8250. doi: 10.1021/jp0450540. [DOI] [PubMed] [Google Scholar]
- 14.Kelleher NL, Lin HY, Valaskovic GA, Aaserud DJ, Fridriksson EK, McLafferty FW. Top Down versus Bottom Up Protein Characterization by Tandem High-Resolution Mass Spectrometry. J. Am. Chem. Soc. 1999;121:806–812. [Google Scholar]
- 15.Kelleher NL. Top-Down Proteomics. Anal. Chem. 2004;76:196A–203A. [PubMed] [Google Scholar]
- 16.Little DP, Speir JP, Senko MW, Oconnor PB, McLafferty FW. Infrared Multiphoton Dissociation of Large Multiply-Charged Ions for Biomolecule Sequencing. Anal. Chem. 1994;66:2809–2815. doi: 10.1021/ac00090a004. [DOI] [PubMed] [Google Scholar]
- 17.Zubarev RA, Kelleher NL, McLafferty FW. Electron Capture Dissociation of Multiply Charged Protein Cations. A Nonergodic Process. J. Am. Chem. Soc. 1998;120:3265–3266. [Google Scholar]
- 18.Reid GE, McLuckey SA. “Top Down” Protein Characterization via Tandem Mass Spectrometry. J. Mass Spectrom. 2002;37:663–675. doi: 10.1002/jms.346. [DOI] [PubMed] [Google Scholar]
- 19.Henry KD, Williams ER, Wang BH, McLafferty FW, Shabanowitz J, Hunt DF. Fourier-Transform Mass-Spectrometry of Large Molecules by Electrospray Ionization. Proc. Natl. Acad. Sci. U.S.A. 1989;86:9075–9078. doi: 10.1073/pnas.86.23.9075. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Smith RD, Anderson GA, Lipton MS, Pasa-Tolic L, Shen YF, Conrads TP, Veenstra TD, Udseth HR. An Accurate Mass Tag Strategy for Quantitative and High-Throughput Proteome Measurements. Proteomics. 2002;2:513–523. doi: 10.1002/1615-9861(200205)2:5<513::AID-PROT513>3.0.CO;2-W. [DOI] [PubMed] [Google Scholar]
- 21.Marshall AG, Guan SH. Advantages of High Magnetic Field for Fourier Transform Ion Cyclotron Resonance Mass Spectrometry. Rapid Commun. Mass Spectrom. 1996;10:1819–1823. doi: 10.1002/(SICI)1097-0231(199611)10:14<1829::AID-RCM697>3.0.CO;2-U. [DOI] [PubMed] [Google Scholar]
- 22.Xu YD, Bruening ML, Watson JT. Non-specific, On-probe Cleanup Methods for MALDI-MS Samples. Mass Spectrom. Rev. 2003;22:429–440. doi: 10.1002/mas.10064. [DOI] [PubMed] [Google Scholar]
- 23.Chan TWD, Colburn AW, Derrick PJ. Matrix-Assisted Laser Desorption Ionization Using a Liquid Matrix: Formation of High-Mass Cluster Ions from Proteins. Org. Mass Spectrom. 1992;27:53–56. [Google Scholar]
- 24.Kraft P, Alimpiev S, Dratz E, Sunner J. Infrared, Surface-Assisted Laser Desorption Ionization Mass Spectrometry on Frozen Aqueous Solutions of Proteins and Peptides Using Suspensions of Organic Solids. J. Am. Soc. Mass Spectrom. 1998;9:912–924. doi: 10.1016/S1044-0305(98)00063-4. [DOI] [PubMed] [Google Scholar]
- 25.Sampson JS, Hawkridge AM, Muddiman DC. Development and Characterization of an Ionization Technique for Analysis of Biological Macromolecules: Liquid Matrix-Assisted Laser Desorption Electrospray Ionization. Anal. Chem. 2008;80:6773–6778. doi: 10.1021/ac8001935. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Overberg A, Karas M, Bahr U, Kaufmann R, Hillenkamp F. Matrix-Assisted Infrared-Laser (2.94-Mu-M) Desorption Ionization Mass-Spectrometry of Large Biomolecules. Rapid Commun. Mass Spectrom. 1990;4:293–296. [Google Scholar]
- 27.Berkenkamp S, Karas M, Hillenkamp F. Ice as a Matrix for IR-Matrix- Assisted Laser Desorption/Ionization: Mass Spectra from a Protein Single Crystal. Proc. Natl. Acad. Sci. U.S.A. 1996;93:7003–7007. doi: 10.1073/pnas.93.14.7003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Little MW, Laboy J, Murray KK. Wavelength Dependence of Soft Infrared Laser Desorption and Ionization. J. Phys. Chem. C. 2007;111:1412–1416. [Google Scholar]
- 29.Menzel C, Dreisewerd K, Berkenkamp S, Hillenkamp F. Mechanisms of Energy Deposition in Infrared Matrix-Assisted Laser Desorption/ Ionization Mass Spectrometry. Int. J. Mass Spectrom. 2001;207:73–96. [Google Scholar]
- 30.Rousell DJ, Dutta SM, Little MW, Murray KK. Matrix-Free Infrared Soft Laser Desorption/Ionization. J. Mass Spectrom. 2004;39:1182–1189. [Google Scholar]
- 31.Sampson JS, Hawkridge AM, Muddiman DC. Construction of a Versatile High Precision Ambient Ionization Source for Direct Analysis and Imaging. J. Am. Soc. Mass Spectrom. 2008;19:1527–1534. doi: 10.1016/j.jasms.2008.06.013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Williams TI, Saggese DA, Muddiman DC. Studying O-Linked Protein Glycosylations in Human Plasma. J. Proteome Res. 2008;7:2562–2568. doi: 10.1021/pr800066e. [DOI] [PubMed] [Google Scholar]
- 33.An HJ, Miyamoto S, Lancaster KS, Kirmiz C, Li BS, Lam KS, Leiserowitz GS, Lebrilla CB. Profiling of Glycans in Serum for the Discovery of Potential Biomarkers for Ovarian Cancer. J. Proteome Res. 2006;5:1626–1635. doi: 10.1021/pr060010k. [DOI] [PubMed] [Google Scholar]
- 34.Williams TI, Saggese DA, Toups KL, Frahm JL, An HJ, Li B, Lebrilla CB, Muddiman DC. Investigations with O-Linked Protein Glycosylations by Matrix-Assisted Laser Desorption/Ionization Fourier Transform Ion Cyclotron Resonance Mass Spectrometry. J. Mass Spectrom. 2008;43:1215–1223. doi: 10.1002/jms.1398. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Bereman MS, Williams TI, Muddiman DC. Carbohydrate Analysis by Desorption Electrospray Ionization Fourier Transform Ion Cyclotron Resonance Mass Spectrometry. Anal. Chem. 2007;79:8812–8815. doi: 10.1021/ac0713858. [DOI] [PubMed] [Google Scholar]
- 36.Horn DM, Zubarev RA, McLafferty FW. Automated Reduction and Interpretation of High Resolution Electrospray Mass Spectra of Large Molecules. J. Am. Soc. Mass Spectrom. 2000;11:320–332. doi: 10.1016/s1044-0305(99)00157-9. [DOI] [PubMed] [Google Scholar]
- 37.Collier TS, Hawkridge AM, Georgianna DR, Payne GA, Muddiman DC. Top-Down Identification and Quantification of Stable Isotope Labeled Proteins from Aspergillus flavus Using Online Nano- Flow Reversed-Phase Liquid Chromatography Coupled to an LTQ-FTICR Mass Spectrometer. Anal. Chem. 2008;80:4994–5001. doi: 10.1021/ac800254z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Williams TI, Saggese DA, Wilcox RJ, Martin JD, Muddiman DC. Effect of Matrix Crystal Structure on Ion Abundance of Carbohydrates by Matrix-Assisted Laser Desorption/Ionization Fourier Transform Ion Cyclotron Resonance Mass Spectrometry. Rapid Commun. Mass Spectrom. 2007;21:807–811. doi: 10.1002/rcm.2904. [DOI] [PubMed] [Google Scholar]
- 39.Dixon RB, Sampson JS, Hawkridge AM, Muddiman DC. Ambient Aerodynamic Ionization Source for Remote Analyte Sampling and Mass Spectrometric Analysis. Anal. Chem. 2008;80:5266–5271. doi: 10.1021/ac800289f. [DOI] [PubMed] [Google Scholar]
- 40.Frahm JL, Howard BE, Heber S, Muddiman DC. Accessible Proteomics Space and Its Implications for Peak Capacity for Zero-, One- and Two-Dimensional Separations Coupled with FT-ICR and TOF Mass Spectrometry. J. Mass Spectrom. 2006;41:281–288. doi: 10.1002/jms.1024. [DOI] [PubMed] [Google Scholar]