Abstract
In biological environments, nanomaterials associate with proteins forming a protein corona (PC). The PC may alter the nanomaterial’s pharmacokinetics and pharmacodynamics, thereby influencing toxicity. Using a label-free mass spectrometry-based proteomics approach, the composition of the PC is examined for a set of nanotubes (NTs) including unmodified and carboxylated single- (SWCNT) and multi-walled carbon nanotubes (MWCNT), polyvinylpyrrolidone (PVP)-coated MWCNT (MWCNT-PVP), and nanoclay. NTs are incubated for 1 h in simulated cell culture conditions, then washed, resuspended in PBS, and assessed by liquid chromatography-tandem mass spectrometry (LC-MS/MS) for their associated PC. To determine those attributes that influence PC formation, the NTs are extensively characterized. NTs had negative zeta potentials in water (SWCNT-COOH < MWCNT-COOH < unmodified NTs) while carboxylation increases their hydrodynamic sizes. All NTs are also found to associate a common subset of proteins including albumin, titin, and apolipoproteins. SWCNT-COOH and MWCNT-COOH are found to bind the greatest number of proteins (181 and 133 respectively) compared to unmodified NTs (< 100), suggesting covalent binding to protein amines. Modified NTs bind a number of unique proteins compared to unmodified NTs, implying hydrogen bonding and electrostatic interactions are involved in PC formation. PVP-coating of MWCNT did not influence PC composition, further reinforcing the possibility of hydrogen bonding and electrostatic interactions. No relationships are found between PC composition and corresponding isoelectric point, hydropathy, or aliphatic index, implying minimal roles of hydrophobic interaction and pi-stacking.
1. Introduction
The field of nanotechnology is rapidly expanding and evolving with the development of numerous engineered nanomaterials. These synthesized nanomaterials are defined by having at least one dimension less than 100 nanometers and can be utilized in various fields including multiple applications in biomedical and consumer products. Nanomaterials often possess a high degree of functionality to render a variety of physicochemical characteristics including diverse chemical composition, available surface groups, shape, electrothermal conductance capabilities, and solubility. Based upon these properties, nanomaterials may be distributed to any organ system and interact on a subcellular level making them useful for both the diagnosis and treatment of diseases.
Upon introduction into a physiological environment nano-materials are rapidly coated with a layer of proteins, known as the protein “corona” (PC).[1–3] The PC alters the size and interfacial composition of the nanomaterials, imparting a biological identity distinct from their original synthetic identity that may modify their activity, bio-distribution, clearance, and toxicity. The distinct composition of the PC, and therefore the nanomaterial’s biological activity, is influenced by the biological environment and the characteristics of each nanomaterial. Ultimately, the PC for each nanomaterial appears to be unique and is determined by each nanomaterial’s individual composition, surface charge, shape and other distinguishing characteristics.[4,5] The corona and its “epitope map” can be viewed as the bioactive entity to which the cells respond.[2] It has been hypothesized that modulation of the proteins which form the PC could be useful in targeting nanomaterials to desired tissues, cells and/or subcellular targets.[6]
Research has demonstrated that the capacity of nano-materials to bind a variety of plasma proteins including those implicated in coagulation, lipid transport, ion transport, complement activation, and pathogen recognition.[5,7] Furthermore, in vitro studies have demonstrated that the PC may influence nanomaterial uptake by cells and alter cytotoxicity.[8–12] Adsorption of a variety of proteins including IgG and fibrinogen has been shown to increase macrophage phagocytosis of nanomaterials in vitro.[13,14] The ability of the PC to enhance macrophage phagocytosis and clearance may have significant implications such as modifying subsequent immune responses and increasing systemic inflammation and oxidative stress. Polysorbate-coated nanoparticles have been shown to preferentially associate with apolipoprotein E, thereby increasing distribution across the blood brain barrier possibly through mimicking low-density lipoprotein and enhancing endothelial cell uptake.[15,16] Conversely, macrophage internalization of both positively and negatively-charged silicon microparticles is enhanced in serum-free media compared to media with serum, suggesting that addition of the PC in some cases may inhibit interactions with cell surface receptors mediating uptake.[17] Manipulation of cellular uptake via modulation of the PC could therefore be therapeutically beneficial for cell targeting of nanomaterials; however, it may also have unexpected toxicological consequences through effects on biodistribution, accumulation, and clearance. Evidently, the formation and biological effects of nanomaterials and their PCs are extremely complex and require further evaluation and study.
Because of cost, ethical, and efficiency considerations, in vitro toxicity assays are widely used for screening and assessing the toxicity of nano-particles. In vitro screening of nanoparticle safety has been ineffectual due to assay interference and contrasting findings likely resulting from differences in particle suspension, cell culture media and delivery, thereby limiting their predictive value. However, their predictive capabilities could be improved by characterizing nanoparticle interactions with fetal bovine serum proteins that are often used in cell culture media to increase our understanding of how PCs affect nanoparticle-cell interaction and biological effects.[12] Previous in vitro exposure studies of both functionalized and non-functionalized carbon NTs in barrier epithelial cells demonstrated significant NT-specific effects on relevant molecular and cellular functions and canonical pathways, with little overlap across NT type, dose, or functionalization, even in the absence of overt toxicity.[18,19] These studies suggest other physicochemical characteristics, such as the PC, may be accountable for the inconsistencies. Accordingly, in the present study, we investigated characteristics of NTs, along with that of halloysite nanoclay as a carbonaceous, high aspect ratio nanotube, which contributed to the formation of the PC in fetal bovine serum often used during the in vitro evaluation of nanomaterial toxicity. We employed a comprehensive proteomics analysis to determine the identities and individual abundance of proteins that associate a form a hard corona with NTs after incubation in bovine serum-supplemented culture media. This information is necessary in understanding properties of nanomaterials that govern their interactions with proteins in biological environments and ultimately lead to the unique biological responses.
2. Results and Discussion
2.1. Nanotube Characteristics
Electron microscopy images (Figure 1) confirmed the dimensions of the carbon nanotubes (NTs; first row in Table 1): the SWCNT (single-walled carbon nanotubes) were 0.1–1 μm, the MWCNT (multi-walled carbon nanotubes) were 10–30 μm, and the nanoclay were 0.5–2 μm in length. Results from the elemental analysis of energy dispersive spectra (Table 1 and Supporting Information EDS Studies) showed the elemental composition of the NTs and demonstrated changes in surface chemistry. The existence of nitrogen confirmed the polyvinylpyrrolidone (PVP) coating of MWCNT-PVP samples, whereas the relatively high content of oxygen indicates the existence of COOH-surface functionalization on MWCNT-COOH and SWCNT-COOH samples. The halloysite nanoclay, as expected, showed an abundance of oxygen, aluminum and silicon. The existence of functional groups on the surface of the NTs was further confirmed via forurier transform infrared spectroscopy and thermogravimetric analysis (data not shown).
Figure 1.
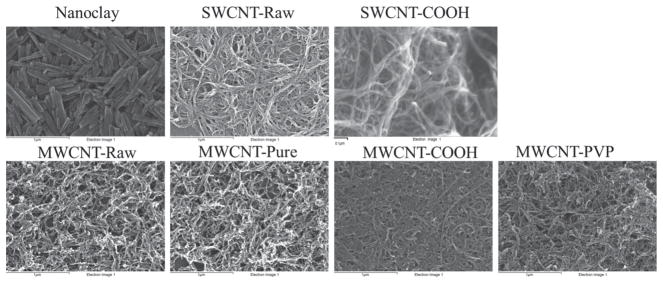
Scanning electron microscopy images of nanoclay, unmodified single walled carbon nanotubes (SWCNT-Raw), carboxylated SWCNT (SWCNT-COOH), unmodified multi-walled carbon nanotubes (MWCNT-Raw), pure MWCNT (MWCNT-Pure), carboxylated MWCNT (MWCNT-COOH), and polyvinylpyrrolidone-coated MWCNT (MWCNT-PVP) samples confirming the dimensions of all carbon NTs used in this study.
Table 1.
Length and Percent Elemental Composition of Nanoparticles.
| Nanoclay | MWCNT-Pure | MWCNT-Raw | SWCNT-Raw | MWCNT-PVP | MWCNT-COOH | SWCNT-COOH | |
|---|---|---|---|---|---|---|---|
| Length (μm) | 0.5–2 | 10–30 | 10–30 | 0.1–1 | 10–30 | 10–30 | 0.1–1 |
| Outer Diameter (nm) | 50–100 | 20–30 | 20–30 | ~0.8–1.2 | 20–30 | 20–30 | ~0.8–1.2 |
| Inner Diameter (nm) | 10–20 | 5–10 | 5–10 | – | 5–10 | 5–10 | - |
| Original Purity (%) | –a) | >95% | >95% | 85% | >95% | >95% | >95% |
| Carbon (%) | 3.81 | 98.77 | 98.65 | 94.91 | 97.59 | 97.4 | 96.92 |
| Oxygen (%) | 37.75 | 1.23 | 1.35 | 4.41 | 1.23 | 2.60 | 3.08 |
| Nitrogen (%) | 0.82 | – | – | 0.68 | 1.19 | – | – |
| Aluminum (%) | 25.80 | – | – | – | – | – | – |
| Silicon (%) | 25.42 | – | – | – | – | – | – |
| Calcium (%) | 0.20 | – | – | – | – | – | – |
| Iron (%) | 1.81 | – | – | – | – | – | – |
| Copper (%) | 4.39 | – | – | – | – | – | – |
Denotes elements that were not in sufficient quantity to be detected by energy dispersive spectroscopy.
The hydrodynamic sizes of the samples suspended in fetal bovine serum (FBS)-supplemented Dulbecco’s Modified Eagle’s medium (DMEM), in comparison to those suspended in water, revealed consistently increased size for all COOH-surface functionalized NTs, by approximately 60 to 120%, likely due to the adsorption of proteins, amino acids and lipids from the medium (Figure 2). In contrast, the hydrodynamic size of non-functionalized SWCNT (SWCNT-Raw) was decreased by approximately 25%, likely due to the debundling and dispersion as a result of PC formation. For MWCNT-PVP, however, their sizes were comparable to those suspended in water (Figure 2), suggesting exchange of PVP by the proteins in the media for coating the MWCNT “core”. For non-functionalized MWCNT(MWCNT-Raw) and purified MWCNT (MWCNT-Pure) samples, large aggregates were formed that precipitated out of the aqueous phase, suggesting the hydrophobicity of these two types of NTs was too high to be overcome by PC formation. All samples demonstrated negative zeta potentials in water (Figure 2). Zeta potential analysis of materials in DMEM was non-determinant due to the screening of the NTs by the ions and biomolecules in the medium. COOH-surface functionalization of both MWCNT and SWCNT and PVP-coating of MWCNT samples resulted in further decreased zeta potentials compared to raw NT samples, suggesting increased dispersion in water due to surface modulation.
Figure 2.
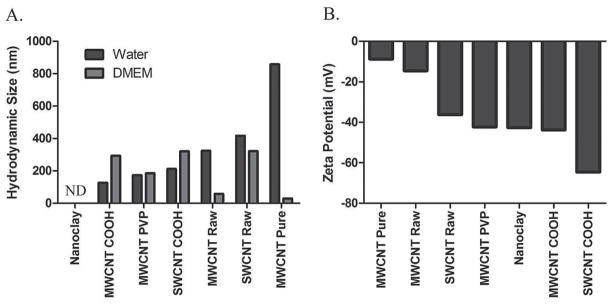
Characterization of nanoclay, unmodified single walled carbon nanotubes (SWCNT-Raw), carboxylated SWCNT (SWCNT-COOH), unmodified multi walled carbon nanotubes (MWCNT-Raw), pure MWCNT (MWCNT-Pure, carboxylated MWCNT (MWCNT-COOH), and polyvinylpyrrolidone-coated MWCNT (MWCNT-PVP) samples. 2A) Hydrodynamic size for each NT was assessed in both water and DMEM cell culture media via dynamic light scattering. 2B) Zeta potentials for each NT were determined in water via electrophroretic light scattering.
2.2. Proteomic Results
The PC that forms on nanoparticles when they are exposed to protein-containing biological fluids changes their characteristics and may be responsible for nanoparticle bioactivity in cells. Based on our previous observations that structurally similar nanoparticles can have divergent biological effects in cell culture systems, we investigated the composition of the PCs formed on different high aspect ratio nanoparticles.[18,19] The PC is a complex and dynamic entity which will evolve over time especially in its soft outer layer. The focus our current study is examining the hard corona which consists of strongly bound proteins which do not readily disassociate and will influence the dynamic interactions of the soft corona.
Proteomic analysis identified and quantified 366 different protein components of the various NT coronae. A complete list of these proteins, along with mass spectral and quantitation data, can be found in Supporting Information Tables 1 and 2, respectively. The numbers of constituent proteins detected in each NT corona are presented graphically in Figure 3A. The PC which formed on the nanoclay tubes consisted of the fewest number, 82 different proteins, whereas the SWCNT-COOH corona contained the most, at 181. For reference purposes, analysis of the 10% FBS-DMEM media alone revealed 2,507 individual proteins, polypeptides, or protein fragments/isoforms, and a list of these along with their peptide sequence and abundance data are provided in Supporting Information Table 3.
Figure 3.
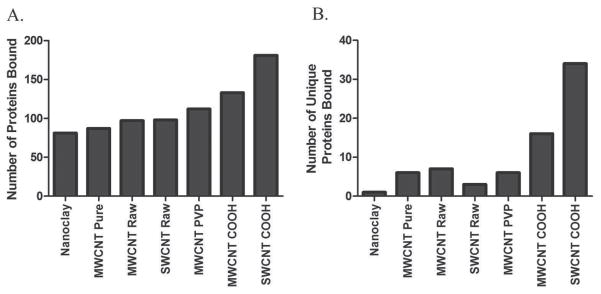
Total number and number of unique proteins found to associate with NTs after incubation in DMEM cell culture media containing 10% fetal bovine serum. Samples were analyzed via liquid chromatography-tandem mass spectrometry (LC-MS/MS) and proteins and peptides were identified using the UniProtKB Bos Taurus (Bovine) database and validated by PeptideProphet. Only proteins with a probability ≥ 0.9, or peptides with a probability ≥ 0.8, and a peptide weight ≥ 0.5 were used in the quantitation algorithm. 3A) The total number of constituent proteins detected in each NT protein corona. 3B) The number of unique proteins detected in each NT protein corona.
All NT coronae were found to consist of 14 common proteins, including alpha-1-antiproteinase, alpha-2-HS-glycoprotein, alpha-S1-casein, apolipoprotein A-I, apolipoprotein A-II, keratin, type I cytoskeletal 10, keratin, type I cytoskeletal 15, keratin, type II cytoskeletal 1, keratin, type II cytoskeletal 5, keratin, type II cytoskeletal 6A, keratin, type II cytoskeletal 6C, keratin, type II cytoskeletal 75, serum albumin, and titin listed in Table 2, in the order of decreasing abundance. The five most abundant coronal proteins (titin, serum albumin, apolipoprotein A-I, apolipoprotein A-II, and alpha-S1-casein) exhibited significant differences across the various NTs, while the relative contributions of alpha-1-antiproteinase (aka alpha-1-antitrypsin in humans), alpha-2-HS-glycoprotein, and the 7 keratins to the NT coronae were not significantly different. With the exception of titin, alpha-S1-casein and the keratins, the highly abundant serum proteins are commonly found in NP coronae formed in human plasma/serum. Titin is the 14th most abundant protein in the FBS-DMEM media whereas albumin is 1st, alpha-2-HS-glycoprotein 2nd, and alpha-1-antiproteinase 3rd and Apo-AI is 17th, while alpha-S1-casein and the keratins (other than keratin 1) are far less abundant in the culture medium (Table 2). Importantly, the presence of the latter proteins in the PC of all NPs suggests a selective enrichment that is not related to their concentrations in the media. It should also be mentioned that all of the above proteins are highly abundant in human plasma according to the most recent version of the Human Peptide Atlas database (http://www.peptideatlas.org), with the exception of alpha-S1-casein, which is not a component of human plasma.[20]
Table 2.
Abundance of the 14 proteins found in all NT protein coronae with their FBS-DMEM abundance ranking.
| Protein | FBS Rank (of 2507) | Nanoclay | MWCNT-PURE | MWCNT-RAW | SWCNT-RAW | MWCNT-PVP | MWCNT-COOH | SWCNT-COOH |
|---|---|---|---|---|---|---|---|---|
| Titin | 14 | 287,786,014c) | 249,183,558 | 235,313,725 | 221,553,081 | 222,285,701 | 204,425,865 | 220,812,507 |
| Serum albumin | 1 | 207,804,997 | 194,609,173 | 228,671,883 | 145,851,821 | 175,003,916 | 146,190,486b) | 109,549,996a) |
| Apolipoprotein A-I | 17 | 86,158,367a) | 25,189,155 | 30,866,967 | 16,884,667 | 15,747,789 | 20,862,300 | 15,827,248 |
| Apolipoprotein A-II | 85 | 47,126,637a) | 18,727,535 | 21,872,077 | 9,851,962 | 13,874,155 | 17,637,071 | 9,948,567 |
| Alpha-S1-casein | 196 | 43,612,167a) | 18,685,400 | 17,374,933 | 13,329,767 | 11,865,415 | 16,515,567 | 12,969,733 |
| Alpha-1-antiproteinase | 3 | 26,184,164 | 26,737,342 | 29,813,128 | 28,939,077 | 24,287,705 | 25,549,842 | 28,412,457 |
| Alpha-2-HS-glycoprotein | 2 | 26,276,777 | 31,171,165 | 30,278,503 | 11,817,688 | 24,758,827 | 27,448,186 | 14,549,701 |
| Keratin, type I cytoskeletal 10 | 119 | 16,497,504 | 18,964,187 | 17,308,479 | 19,955,977 | 16,343,288 | 19,080,789 | 21,111,033 |
| Keratin, type II cytoskeletal 1 | 27 | 12,015,333 | 8,305,850 | 6,632,780 | 7,534,533 | 5,764,675 | 6,095,444 | 7,003,015 |
| Keratin, type II cytoskeletal 75 | 638 | 2,059,786 | 1,568,094 | 1,923,638 | 1,674,743 | 1,814,366 | 2,075,758 | 1,837,728 |
| Keratin, type II cytoskeletal 6A | 129 | 1,961,464 | 1,711,923 | 1,862,706 | 2,198,074 | 1,603,329 | 1,753,544 | 2,445,845 |
| Keratin, type II cytoskeletal 5 | 1031 | 1,648,939 | 1,597,897 | 1,523,397 | 1,567,228 | 1,203,009 | 1,318,831 | 1,565,164 |
| Keratin, type II cytoskeletal 6C | 263 | 1,506,162 | 1,404,497 | 1,364,453 | 1,514,077 | 1,201,025 | 1,299,497 | 1,636,483 |
| Keratin, type I cytoskeletal 15 | 347 | 866,543 | 1,039,746 | 779,822 | 1,199,950 | 796,696 | 953,023 | 1,000,895 |
P < 0.05 vs. all others;
P < 0.05 vs. Nanoclay, MWCNT-PURE, MWCNT-RAW;
P < 0.05 vs. SWCNT-RAW, MWCNT-PVP, MWCNT-COOH, SWCNT-COOH.
The 25 most abundant proteins in each NT PC are listed in Table 3. Of all PC constituents, the most abundant was Xin actin-binding repeat-containing protein 2 (XIRP2) and it was found only in MWCNT-Pure, MWCNT-PVP and SWCNT-COOH coronae. XIRP2, aka mXinβ and myomaxin, is a 382 300 Da protein expressed in cardiac and skeletal muscle where it interacts with filamentous actin and α-actinin through the novel actin-binding motif, the Xin repeat.[21,22] It is also the 40th most abundant protein in the FBS-supplemented culture medium. Like titin, this largely abundant coronal protein is associated with intracellular filamentous proteins. The ample presence of XIRP2 in the media and in NT coronae may be in the form of protein fragments that are more common to fetal serum and less so in adult human or bovine sera where they are known to interact with albumin.[23] Other proteins may also be present in the PC via their association with bovine serum albumin, as part of the albuminome.[23–25] For instance, the keratins identified in the PCs may be there through their interaction with albumin directly, or indirectly via their known interaction with apolipoproteins, which also interact with albumin.[26] While it is known that both intact and fragmented proteins exist in the serum and in association with albumin and other major serum proteins, their composition is beyond the scope of this investigation.
Table 3.
25 most abundant coronal proteins associated with each nanotube.
| Nanoclay | MWCNT-PURE | MWCNT-RAW | SWCNT-RAW | MWCNT-PVP | MWCNT-COOH | SWCNT-COOH |
|---|---|---|---|---|---|---|
| TTNb) | XIRP2 | TTNb) | TTNb) | XIRP2 | TTNb) | XIRP2 |
| ALBb) | TTNb) | ALBb) | ALBb) | TTNb) | ALBb) | TTNb) |
| APOA1b) | ALBb) | Bt.105991 | TOP2Ba) | ALBb) | NCOA6 | NCOA6 |
| COL6A6 | KIF7 | APOA1b) | AARS2 | ABCA1 | Bt.105991 | ALBb) |
| APOA2b) | CCDC168 | AHSG | SERPINA1b) | ALS2 | ABCA1 | P2RX5 |
| KCTD3 | AHSG | SERPINA1b) | KRT10 | AHSG | LCT | ATAD2B |
| CSN1S1 | P2RX5 | ATAD2B | ATAT1 | SERPINA1b) | AHSG | ALS2 |
| ATAD2B | SERPINA1b) | ALS2 | APOA1b) | COL3A1 | SERPINA1b) | LCT |
| AHSG | APOA1b) | APOA2b) | CPS1 | HBA | AARS2 | KIF7 |
| SERPINA1b) | HBA | COL6A6 | CSN1S1 | KRT10b) | ATAD2B | ABCA1 |
| MTMR3 | BIRC6 | HBA | AHSG | APOA1b) | APOA1b) | CCDC168 |
| HBA | KRT10 | MTMR3 | COL4A6 | APOA2b) | KRT10b) | SERPINA1b) |
| DNAH1 | APOA2b) | CSN1S1 | APOA2b) | SLAMF9 | APOA2b) | MTMR3 |
| TTC13 | CSN1S1 | KRT10b) | TCTN1 | Bt.65579 | CSN1S1 | KRT10b) |
| FAM179B | EP400 | BSN | JAG1 | ADAMTS16 | HBA | PTPRS |
| BRPF1 | PLCE1 | CYP27B1 | RPTN | CSN1S1 | MTMR3 | EP400 |
| KRT10b) | TCTN1 | AIM1L | MAGI1 | AFAP1 | BRPF1 | ABCC1 |
| HBG | WWP1a) | CPS1 | Bt.63231 | AGL | DNAH11 | PTPRN |
| HERC5 | NT5DC3a) | PTPRN | KRT1b) | STXBP5L | Bt.65579 | APOA1b) |
| KIAA0232 | STXBP5L | SYNPO2 | DOCK4 | ABCC1 | BSN | VPS35 |
| KRT1b) | AGL | FAM179B | LOC787087 | MKI67 | ABCC1 | AHSG |
| LOC531152 | SYNPO2 | SIPA1L3 | MUC5B | GLT25D2a) | GRIN2Ba) | COL4A6 |
| PTPRS | HCFC1 | Bt.63231 | Bt.90677 | COL4A6 | HCFC1 | ATAT1 |
| MAGI1 | PDGFRL | HBG | ATP6V1B1 | BIRC6 | HBG | AFAP1 |
| NEB | KRT1b) | SYNE1 | NPTX2 | Bt.63231 | TMCO7 | CSN1S1 |
Proteins are unique to that NT corona;
Proteins are common to all 7 NT coronae (from Table 2).
Unmodified MWCNT and SWCNT were found to bind a similar number of proteins (Figure 3A). Unmodified MWCNT were found to more readily associate with α-1-antiproteinase (SERPINA1) and α-2-HS-glycoprotein (AHSG) than unmodified SWCNT (Table 3). The addition of carboxyl groups to the surface of SWCNT and MWCNT resulted in an increase in the number of types of protein which associated with the nanomaterials compared to non-functionalized SWCNT and MWCNT-Raw (Figure 3A). This increase in the number of proteins bound to carboxylated-NT is likely due to the abundance of protein amines in the medium which could readily associate with the carboxyls through electrostatic interactions. The lower zeta potential (Figure 2B) and higher protein binding capability of SWCNT COOH (Figure 3), compared with that of MWCNT COOH, can be attributed to the larger surface area and therefore higher density of COOH groups on the SWCNT surfaces. In addition, carboxylation of NTs was found to increase binding of nuclear receptor coactivator-6, lactase-phlorizin hydrolase (NCOA6) and ATP-binding cassette subfamily A member 1 (ABCA1) compared to unmodified NTs (Table 3). PVP-coated MWCNT also demonstrated a slight increase in number of proteins bound compared to raw non-functionalized MWCNT, implying the more significant roles of hydrogen bonding and/or nonspecific electrostatic interactions with protein amines than hydrophobic interaction in NT-PC formation. Furthermore, PVP coating of MWCNT was found to increase association of ATP-binding cassette subfamily A member compared to unmodified MWCNT.
To determine distinctive PC profiles, we examined proteins that were unique to each nanomaterial (Figure 3B and Table 4). With only a few exceptions (collectin-12, G-protein coupled receptor 98, basement membrane-specific heparan sulfate proteoglycan core protein, kininogen-1, receptor-type tyrosine-protein phosphatase zeta, plasma serine protease inhibitor, and vitrin), these NT-specific, low abundance coronal components are proteins of intracellular origin with few or no extracellular domains, representing virtually every subcellular compartment and organelle (via Generic Gene Ontology (GO) Term Mapper (http://go.princeton.edu/cgi-bin/GOTermMapper).[27] It is well known that the proteinaceous composition of serum/plasma includes a significant quantity of low molecular weight protein fragments derived from cell and tissue proteins, many of which are secreted and shed after degradation.[28,29] In fact, 70% of the FBS-DMEM components identified and quantified by LC-MS/MS (Supporting Information Table 1) are intracellular, as are most of the coronal components. It is likely that the cellular proteins were fragments and not whole proteins, as most were identified by 2 peptides and were in comparatively lower abundance than the conventional “serum” protein constituents. Perhaps these cellular fragments are the epitope motifs to which the cell responds upon initial interaction with the NP-corona complex, and this may account for the differential effects so often observed when cells are exposed in vitro to similar NPs with slight surface modifications.[2,30]
Table 4.
Proteins Unique to Nanotube Coronae.
| Protein ID | Gene Name | Protein Name | Quantity | |
|---|---|---|---|---|
| Reference | ||||
| Proteins* | ||||
| P15497 | APOA1 | Apolipoprotein A-I | 30,219,499 | |
| P02769 | ALB | Serum albumin | 172,526,039 | |
| F1N757 | TTN | Titin | 234,480,064 | |
| Nanoclay | ||||
| F1MPT5 | DST | Dystonin | 3,861,367 | |
| MWCNT-Pure | ||||
| G3MX12 | HERC2 | E3 ubiquitin-protein ligase HERC2 | 1,735,859 | |
| F1MLJ1 | PCDHB1 | Protocadherin beta-1 | 3,155,333 | |
| E1BEW9 | WDR87 | WD repeat-containing protein 87 | 6,299,685 | |
| E1BGJ0 | LRP1 | Prolow-density lipoprotein receptor-related protein 1 | 7,429,100 | |
| F1MYW0 | NT5DC3 | 5′-nucleotidase domain-containing protein 3 | 11,721,867 | |
| Q32PG0 | WWP1 | NEDD4-like E3 ubiquitin-protein ligase WWP1 | 12,260,233 | |
| MWCNT-Raw | ||||
| P01044 | KNG1 | Kininogen-1 | 1,012,523 | |
| F1MW73 | MGC148692 | KIAA1211 protein | 1,568,099 | |
| F1MSG6 | RAPGEF2 | Rap guanine nucleotide exchange factor 2 | 1,952,045 | |
| Q2KHZ2 | HBS1L | HBS1-like protein | 4,041,483 | |
| F1MAX6 | KATNAL1 | Katanin p60 ATPase-containing subunit A-like 1 | 4,289,983 | |
| F1N300 | Bt.76801 | Condensin complex subunit 1 | 4,804,233 | |
| F1MYB0 | STOX2 | Storkhead-box protein 2 | 7,332,400 | |
| SWCNT-Raw | ||||
| F1MER7 | HSPG2 | Basement membrane-specific heparan sulfate proteoglycan core protein | 1,610,263 | |
| Q9TTA5 | SMARCAL1 | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin | 3,302,833 | |
| F1MU48 | TOP2B | DNA topoisomerase 2-beta | 43,838,728 | |
| MWCNT-PVP | ||||
| E1BA03 | PAK6 | Serine/threonine-protein kinase PAK 6 | 1,362,630 | |
| E1BIU4 | ZC3H7B | Zinc finger CCCH domain-containing protein 7B | 3,710,533 | |
| Q0III9 | ACTN3 | Alpha-actinin-3 | 3,886,300 | |
| F1N0S1 | ASAP3 | Arf-GAP with SH3 domain, ANK repeat and PH domain-containing protein 3 | 3,987,033 | |
| E1BAU4 | PCDH17 | Protocadherin-17 | 5,714,533 | |
| F1N0V8 | GLT25D2 | Procollagen galactosyltransferase 2 | 8,720,067 | |
| MWCNT-COOH | ||||
| Q95LI2 | VIT | Vitrin | 1,199,471 | |
| E1BIN0 | FHOD1 | FH1/FH2 domain-containing protein 1 | 2,041,853 | |
| A6QLR2 | LARS | Leucine–tRNA ligase, cytoplasmic | 2,665,600 | |
| G3MXX3 | Bt.103298 | Chromodomain-helicase-DNA-binding protein 2 | 2,725,444 | |
| F1MX14 | MDGA2 | MAM domain-containing glycosylphosphatidylinositol anchor protein 2 | 2,883,907 | |
| E1BMN2 | DIAPH3 | Protein diaphanous homolog 3 | 3,074,724 | |
| Q9N2I2 | SERPINA5 | Plasma serine protease inhibitor | 3,212,431 | |
| F1MD25 | C11orf41 | UPF0606 protein C11orf41 | 3,495,796 | |
| G3MW09 | Bt.82323 | Multidrug resistance-associated protein 4 | 3,590,753 | |
| E1BA65 | PMS2 | Mismatch repair endonuclease PMS2 | 3,647,800 | |
| E1BA80 | MYO18B | Unconventional myosin-XVIIIb | 3,683,443 | |
| Q0P5J8 | FAM40A | Protein FAM40A | 4,130,100 | |
| F1MC84 | FAT3 | Protocadherin Fat 3 | 4,947,628 | |
| A6QP52 | SMTN | Smoothelin | 5,745,756 | |
| E1BG99 | EIF4ENIF1 | Eukaryotic translation initiation factor 4E transporter | 6,688,973 | |
| F1MCL3 | GRIN2B | Glutamate [NMDA] receptor subunit epsilon-2 | 9,710,800 | |
| SWCNT-COOH | ||||
| Q3B7M1 | KLHL36 | Kelch-like protein 36 | 1,188,690 | |
| F1MGS8 | NAALAD2 | N-acetylated-alpha-linked acidic dipeptidase 2 | 1,567,551 | |
| A6H709 | HSPC321 | Switch-associated protein 70 | 2,022,833 | |
| F1MLF8 | HGD | Homogentisate 1,2-dioxygenase | 2,023,368 | |
| E1AXU0 | CMYA1 | Cardiomyopathy associated protein 1 | 2,282,263 | |
| F1MUL5 | FAAH | Fatty-acid amide hydrolase 1 | 2,797,818 | |
| E1B8Z3 | DAAM2 | Disheveled-associated activator of morphogenesis 2 | 2,931,789 | |
| E1BKS0 | EPS8L1 | Epidermal growth factor receptor kinase substrate 8-like protein 1 | 3,173,085 | |
| E1BG53 | PPARGC1B | Peroxisome proliferator-activated receptor gamma coactivator 1-beta | 3,176,827 | |
| E1BCY7 | Bt.45696 | Tyrosine-protein kinase SgK223 | 3,261,906 | |
| F1MS10 | Bt.65326 | Dynein heavy chain 5, axonemal | 3,374,833 | |
| E1BGY9 | JPH3 | Junctophilin-3 | 3,405,295 | |
| G3X7Z2 | CACNA1B | Voltage-dependent N-type calcium channel subunit alpha-1B | 3,413,133 | |
| Q27991 | MYH10 | Myosin-10 | 3,774,257 | |
| E1BGB2 | PRRC2B | Protein PRRC2B | 4,048,033 | |
| E1BMG2 | DNAH5 | Dynein heavy chain 5, axonemal | 4,183,145 | |
| F1MH31 | NUP214 | Nuclear pore complex protein Nup214 | 4,619,333 | |
| F1N0J3 | ATP11A | Probable phospholipid-transporting ATPase IH | 4,793,946 | |
| G3N0C1 | ANK2 | Ankyrin-2 | 4,822,700 | |
| F1MJJ0 | DNAH9 | Dynein heavy chain 9, axonemal | 4,856,533 | |
| F1N0A6 | GPR98 | G-protein coupled receptor 98 | 5,075,200 | |
| E1BKQ9 | GALNT5 | Polypeptide N-acetylgalactosaminyltransferase 5 | 5,084,459 | |
| O02776 | PARG | Poly(ADP-ribose) glycohydrolase | 5,477,200 | |
| F1N6H4 | MACF1 | Microtubule-actin cross-linking factor 1 | 6,296,211 | |
| F1MI56 | ANKRD17 | Ankyrin repeat domain-containing protein 17 | 6,637,200 | |
| E1BC55 | LOC533883 | Uncharacterized protein KIAA1671 | 6,824,667 | |
| F1MNA8 | ZMYM2 | Zinc finger MYM-type protein 2 | 7,565,700 | |
| F1MGK5 | PTPRZ1 | Receptor-type tyrosine-protein phosphatase zeta | 7,929,133 | |
| F1MYC9 | SPTBN1 | Spectrin beta chain, brain 1 | 8,029,370 | |
| E1BM04 | TBKBP1 | TANK-binding kinase 1-binding protein 1 | 8,685,600 | |
| F1MC30 | GCNT4 | Beta-1,3-galactosyl-O-glycosyl-glycoprotein beta-1,6-N-acetylglucosaminyltransferase 4 | 9,650,800 | |
| E9QB09 | WC1 | WC1 (CD4-CD8- gamma delta T lymphocyte proteins) | 10,846,900 | |
| A6QP79 | COLEC12 | Collectin-12 | 12,443,233 | |
| F1MFX9 | PRKAG2 | 5′-AMP-activated protein kinase subunit gamma | 12,863,582 | |
found in all NT coronae, shown to emphasize protein quantity differences between these and low-abundance NT-specific proteins; mean quantity shown.
COOH-functionalization of SWCNT and MWCNT was found to increase the number of unique proteins which associated with the NTs compared to non-functionalized raw NTs (Figure 3B), pointing to the role of covalent bonding between the carboxyls of the NTs and amines of the proteins in PC formation. As illustrated in Figure 4, GRAVY scores of coronal protein constituents were similar across all NTs studied, and nearly all exhibited slight hydrophilicity as opposed to only a few with slightly hydrophobic scores. Similarly, no differences in mean GRAVY score, protein isoelectric point, or aliphatic indices of nanomaterial PCs were observed (Supporting Information Figure 2). Numerous low-abundance “cellular” proteins were found to be unique to the PCs of specific NT types. These NT-specific proteins are listed, along with their abundances, in Table 4. Despite the heterogeneity of PC composition, all but 3 of the various corona components (T-cell lymphoma invasion and metastasis 1, MW 70,704; Homeobox protein cut-like 1, MW 164,187; and receptor-transporting protein 4, MW 27,863) were also detected in FBS-DMEM media alone (see Supporting Information Table 1), indicating that the NTs did not uniquely enrich many specific groups of very low abundance media proteins that were otherwise undetectable in the FBS-DMEM analysis. As with most of the coronal proteins, these 3 proteins that were not identified in the media proteome likely are low-abundance fragments of cellular proteins.
Figure 4.
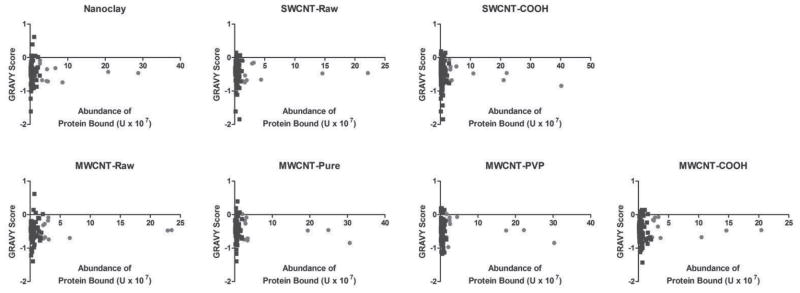
Grand average of hydophathicity (GRAVY) scores for all identified proteins found to associate with NTs after incubation in DMEM cell culture media containing 10% fetal bovine serum compared to their relative abundance. The ten most abundant peptides/proteins are denoted in red within the graph. GRAVY scores were calculated for a peptide or protein based on the sum of hydropathy values for all amino acids and divided by the number of residues in the sequence.
Surprisingly, despite the prevalence of in vitro nanotoxicology investigations, only two studies have attempted to identify and characterize fetal bovine serum proteins and their quantitative composition via SDS-PAGE separation and identification by LC-MS/MS in PCs formed during in vitro nano-particle exposures: citrate capped gold nanoparticle coronae and magnetic iron oxide nanoparticle coronae.[9,31] The electrophoretic approach used to separate and detect coronal constituents in these studies may have limited the number of proteins actually identified. All other previous studies of PC composition of nanoparticles using proteomic techniques have focused on human plasma/sera or cytosols and include: amorphous silica; polystyrene; sulfonated polystyrene and silica; atheronal-b and cholesterol coated quantum dots; lipoplexes and liposomes; carboxyl-modified polystyrene; carbon NTs and metal oxide; and surface-functionalized gold in cell lysate proteins.[6,32–42] Similar to the two studies using fetal bovine serum, these studies with human plasma/sera used SDS-PAGE followed by LC-MS/MS identification.
Similar to our current study, Zhang et al. identified and quantified 88 distinct human plasma proteins by stable isotope labeling and LC-MS/MS on polystyrene nano-particles in which PC composition was surface modification-dependent.[33] Twelve of the 88 proteins identified in the PC of these polystyrene nanoparticles were also common to our FBS-DMEM PC profile for all nanomaterials assessed (plasma serine protease inhibitor, apolipoprotein A-I, apolipoprotein A-II, fibrinogen alpha chain, alpha-2-HS-glyco-protein, serotransferrin, kininogen-1, alpha-1-antitrypsin, vitamin D-binding protein, albumin, complement C3 and complement C4). Unlike the high proportion of cellular PC constituents observed in our study, only about 34% of Zhang et al.’s coronal proteins identified were intracellular. Interestingly, when Capriotti et al. used LC-MS/MS to study the protein composition of coronae that formed on nanosized cationic liposomes (CLs), lipoplexes, and lipid/polycation/DNA (LPD) complexes exposed to human plasma roughly 70% of the 218 proteins were intracellular, similar to our results in high aspect-ratio NTs.[36] In a subsequent quantitative analysis, coronal protein variety found on lipoplexes and LPD complexes was greater than that found on cationic liposomes while individual protein abundance differed as well, again, similar to our observations in NT coronae. Compared to these studies in human plasma/sera or cytosols our current study provides information useful in interpreting and evaluating in vitro nanomaterial toxicity studies.[40] Taken together these previous studies and our current study may assist with the extrapolation of in vitro nanomaterial toxicity data to relevant in vivo interactions and human exposures.
3. Conclusion
NT PCs formed in vitro by exposure to FBS-DMEM media are extremely complex as others using comprehensive proteomics and human plasma have observed. Although typical serum proteins are abundant components of the PC, coronae also contain a large amount of proteins/protein fragments of cellular origin. This provides a diverse composition of each nanomaterial’s PC, which varies based on physicochemical differences. GRAVY, IEP, and aliphatic indices of corona constituents across the nanomaterials evaluated in this study are similar, therefore other factors such as nonspecific hydrogen bonding, electrostatic interaction and the specific covalent bonding between the carboxyls of the NTs and amines of the proteins are likely responsible for the differences in PC composition. Since functionalized NTs bound similar quantities of proteins compared to pristine NTs, hydrophobic interactions and pi-stacking between the aromatic moieties of the proteins and the aromatic groups of the NTs are deemed less significant in NT-PC formation. Although SWCNT-COOH and MWCNT-COOH were found to possess comparable hydrodynamic sizes, the conceivably more rugged surface morphology (due to bundling) and higher charge density of the former led to a slightly more robust binding of plasma proteins in both total number and structural uniqueness. These unique constituents of PC, even those in low abundance may cause unique cellular effects and bioactivity in in vitro nanotoxicology assessments.
4. Experimental Section
Reagents and Materials
DL-Dithiothreitol (DTT), urea, triethylphosphine, iodoethanol, and ammonium bicarbonate were purchased from Sigma-Aldrich (St. Louis, MO, USA). LC-MS grade 0.1% formic acid in acetonitrile and 0.1% formic acid in water were purchased from Burdick & Jackson (Muskegon, MI, USA). Modified sequencing grade porcine trypsin was obtained from Princeton Separations (Freehold, NJ, USA). Dulbecco’s Modified Eagle’s Medium (DMEM) with glutamax and 10% heat inactivated fetal bovine serum were purchased from Invitrogen (Carlsbad CA).
SWCNT were purchased from Unidym (Sunnyvale, CA) and MWCNT were purchased from Cheap Tubes Inc. (Brattleboro, VT). SWCNT-COOH and MWCNT-COOH were generated in a Microwave Accelerated Reaction System (Mode: CEM Mars) fitted with internal temperature and pressure controls as previously described.[43,44] Pre-weighed amounts of purified MWCNT were treated with a mixture of concentrated H2SO4 and HNO3 solution by subjecting them to microwave radiation at 140 °C for 20 min. The product was filtered through a 10 μm membrane filter, washed with water to a neutral pH, and dried under vacuum at 80 °C to a constant weight. SWCNT-COOH were also functionalized in the Microwave Accelerated Reaction System according a procedure developed by our laboratory.[45] Pre-weighed amounts of purified SWCNT were treated with a 1:1 mixture of concentrated H2SO4 and HNO3 solution by subjecting them to microwave radiation at 120 °C for 3 min. The mixture was then diluted with distilled water and filtered through 10 μm membrane filter paper. The filtrate was transferred to a dialysis bag and placed in a container filled with DI water, which was continually replaced until it achieved neutral pH. The filtrate was then dried overnight at 50 °C under vacuum. This led to the formation of carboxylic acid groups on the surface of the NTs resulting in high aqueous dispersibility. MWCNT-PVPs were prepared according to a procedure previously reported.[46] Purified MWCNTs were dispersed in deionized water at a concentration of 50 mg/L with the aid of 1% sodium dodecyl sulfate (SDS). 1% by weight of PVP was added to the mixture, which was then incubated at 50 °C for 12 h. The carbon NTs were then filtered through a 10 μm membrane filter, washed with deionized water followed by three cycles of ultrasonic redispersion in deionized water to remove any residual SDS. The sample was filtered and dried under vacuum at room temperature to a constant weight.
Transmission Electron Microscopy (TEM) and Energy Dispersive Spectroscopy (EDS)
All individual halloysite (nanoclay), SWCNT, and MWCNT samples were mixed with ethanol and sonicated in a water bath (Branson) for 10–15 min until well dispersed. For each sample a droplet of the suspension was placed on a copper grid and dried at room temperature. TEM imaging and EDS element analysis were performed using a Hitachi HD 2000 STEM equipped with an Oxford INCA energy 200 EDS.
Hydrodynamic Size and Zeta Potential Characterization
Approximately 0.1 mg of each sample was mixed with 1 mL 10% FBS supplemented Dulbecco’s Modified Eagle’s Medium (DMEM), and then dispersed via water bath sonication for 2 min. Samples were then incubated on a rotator for 1 h. Hydrodynamic sizes of the suspended samples were measured using a dynamic light scattering device (Malvern Instruments, Nanosizer S90). Zeta potentials of the suspended samples in water were measured using electrophroretic light scattering (Malvern Instruments, ZetaSizer Nano).
Protein Corona Generation and Proteomic Characterization
Using a modification of Tenzer’s method, 1 mg of each NT type was suspended in 10 mL of DMEM culture media supplemented with 10% fetal bovine serum (FBS), briefly sonicated in a bath sonicator, diluted 1:10 in FBS/media, and incubated for 1 h at 37 °C (to simulate in vitro exposure protocols).[32] Stable PCs are at equilibrium within 5 min.[33] The samples were centrifuged (15 min at 3000 × g/22 °C) and the pellets containing the NT-protein complexes were washed and pelleted three times with PBS. After the third and final wash, the supernatant was free of protein. PCs were solubilized in situ using a lysis buffer specific for label-free quantitative mass spectrometry (LFQMS) (8 M urea, 10 mM DTT freshly prepared). For comparative reference purposes, 100 μg of FBS supplemented culture media proteins were also solubilized for LC-MS/MS analysis. Briefly, protein samples were reduced and alkylated by triethylphosphine and iodoethanol and proteolyzed using porcine trypsin.[47] Exactly 20 μg of each tryptic digest sample was injected randomly as two technical replicates onto a C18 reversed phase column for a 3 h HPLC gradient separation, electrospray ionization, and analysis using an LTQ-PROTEOMEX ion trap mass spectrometer. A blank was injected between each sample to clean and balance the column and eliminate carryover. The acquired data were searched against the most up-to-date Uni-ProtKB Bos taurus (Bovine) database using SEQUEST (v. 28 rev. 12) algorithms in Bioworks (v. 3.3). Peptide and protein identifications were validated by PeptideProphet and ProteinProphet in the Trans-Proteomic Pipeline (TPP, v. 3.3.0).[48,49] Only proteins and peptides with (a) protein probability ≥ 0.9, (b) peptide probability ≥ 0.8, and (c) peptide weight ≥ 0.5 were used in the quantitation algorithm. Identified bovine proteins whose names appeared as ‘uncharacterized’ were annotated using homologous human proteins identified by UniProt Blast based on similarity in amino acid sequence.
Protein abundance was determined using IdentiQuantXL.[50] After chromatogram alignment and peptide retention time determination, a weighted mean m/z of each peptide is calculated and a tab delimited file was created to extract peptide intensity using MASIC.[51] Peptides were then filtered according to intensity CV across all samples and intensity correlation for those identifying a particular protein. Protein abundance (intensity) was calculated from all qualified peptides corresponding to a particular protein. Protein abundance/quantity calculated in this way has no units, and therefore are represented by unitless numerical values in Tables 2 and 4 and in Supporting Information Tables 2 and 3. Comparison of the mean abundance of individual protein in each PC, generated by LFQMS, was performed within the IdentiQuantXL platform using one-way ANOVA and Pairwise Multiple Comparisons (Holm-Sidak method). False discovery rate (FDR) was estimated using Q-value software.[52]
Protein Hydropathicity and Aliphatic Index Analysis
Grand average of hydropathicity (GRAVY) scores and aliphatic indices for all identified proteins were calculated using the Protein Identification and Analysis Tools (ProtParam) on the ExPASy Server (http://web.expasy.org/protparam/).[53] The GRAVY score for a peptide or protein is calculated as the sum of hydropathy values of all the amino acids, divided by the number of residues in the sequence. The aliphatic index of a protein is defined as the relative volume occupied by aliphatic side chains (alanine, valine, isoleucine, and leucine) and may be regarded as a positive factor for increased thermostability of globular proteins.
Supplementary Material
Acknowledgments
This study was funded by NIH R01 ES019311 (J. Brown), RC2 ES018025 and NIGMS R01GM085218 (F. Witzmann), and an NSF grant CBET-1232724 (PC. Ke). The authors would like to thank Dr. Y. Lvov of Louisiana Tech University for providing the halloysite nanotubes (nanoclay).
Footnotes
The authors declare no competing financial interests.
Supporting Information is available from the Wiley Online Library or from the author.
Contributor Information
Dr. Jonathan H. Shannahan, Department of Pharmacology and Toxicology, Brody School of Medicine, East Carolina University, Greenville, North Carolina, 27834, USA
Dr. Jared M. Brown, Department of Pharmacology and Toxicology, Brody School of Medicine, East Carolina University, Greenville, North Carolina, 27834, USA
Dr. Ran Chen, Department of Physics and Astronomy, Clemson University, Clemson, South Carolina, 29634, USA
Dr. Pu Chun Ke, Department of Physics and Astronomy, Clemson University, Clemson, South Carolina, 29634, USA
Dr. Xianyin Lai, Department of Cellular & Integrative Physiology, Indiana University School of Medicine, Indianapolis, Indiana, 46202, USA
Dr. Somenath Mitra, Department of Chemistry & Environmental Science, New Jersey Institute of Technology, Newark, New Jersey, 07102, USA
Dr. Frank A. Witzmann, Email: fwitzman@iupui.edu, Department of Cellular & Integrative Physiology, Indiana University School of Medicine, Indianapolis, Indiana, 46202, USA
References
- 1.Ehrenberg MS, Friedman AE, Finkelstein JN, Oberdorster G, McGrath JL. Biomaterials. 2009;30:603. doi: 10.1016/j.biomaterials.2008.09.050. [DOI] [PubMed] [Google Scholar]
- 2.Lynch I, Cedervall T, Lundqvist M, Cabaleiro-Lago C, Linse S, Dawson KA. Adv Colloid Interface Sci. 2007;167:134. doi: 10.1016/j.cis.2007.04.021. [DOI] [PubMed] [Google Scholar]
- 3.Blunk T, Hochstrasser DF, Sanchez JC, Muller BW, Muller RH. Electrophoresis. 1993;14:1382. doi: 10.1002/elps.11501401214. [DOI] [PubMed] [Google Scholar]
- 4.Nel AE, Madler L, Velegol D, Xia T, Hoek EM, Somasundaran P, Klaessig F, Castranova V, Thompson M. Nat Mater. 2009;8:543. doi: 10.1038/nmat2442. [DOI] [PubMed] [Google Scholar]
- 5.Walkey CD, Chan WC. Chem Soc Rev. 2012;41:2780. doi: 10.1039/c1cs15233e. [DOI] [PubMed] [Google Scholar]
- 6.Arvizo RR, Giri K, Moyano D, Miranda OR, Madden B, McCormick DJ, Bhattacharya R, Rotello VM, Kocher JP, Mukherjee P. PLoS One. 2012;7:e33650. doi: 10.1371/journal.pone.0033650. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Lundqvist M, Stigler J, Elia G, Lynch I, Cedervall T, Dawson KA. Proc Natl Acad Sci USA. 2008;105:14265. doi: 10.1073/pnas.0805135105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Lartigue L, Wilhelm C, Servais J, Factor C, Dencausse A, Bacri JC, Luciani N, Gazeau F. ACS Nano. 2012;6:2665. doi: 10.1021/nn300060u. [DOI] [PubMed] [Google Scholar]
- 9.Maiorano G, Sabella S, Sorce B, Brunetti V, Malvindi MA, Cingolani R, Pompa PP. ACS Nano. 2010;4:7481. doi: 10.1021/nn101557e. [DOI] [PubMed] [Google Scholar]
- 10.Panas A, Marquardt C, Nalcaci O, Bockhorn H, Baumann W, Paur HR, Mulhopt S, Diabate S, Weiss C. Nanotoxicology. 2012 doi: 10.3109/17435390.2011.652206. [DOI] [PubMed] [Google Scholar]
- 11.Clift MJ, Bhattacharjee S, Brown DM, Stone V. Toxicol Lett. 2010;198:358. doi: 10.1016/j.toxlet.2010.08.002. [DOI] [PubMed] [Google Scholar]
- 12.Tedja R, Lim M, Amal R, Marquis C. ACS Nano. 2012;6:4083. doi: 10.1021/nn3004845. [DOI] [PubMed] [Google Scholar]
- 13.Beduneau A, Ma Z, Grotepas CB, Kabanov A, Rabinow BE, Gong N, Mosley RL, Dou H, Boska MD, Gendelman HE. PLoS One. 2009;4:e4343. doi: 10.1371/journal.pone.0004343. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Deng ZJ, Liang M, Monteiro M, Toth I, Minchin RF. Nat Nanotechnol. 2011;6:39. doi: 10.1038/nnano.2010.250. [DOI] [PubMed] [Google Scholar]
- 15.Kreuter J. Adv Drug Deliv Rev. 2001;47:65. doi: 10.1016/s0169-409x(00)00122-8. [DOI] [PubMed] [Google Scholar]
- 16.Kreuter J, Alyautdin RN, Kharkevich DA, Ivanov AA. Brain Res. 1995;674:171. doi: 10.1016/0006-8993(95)00023-j. [DOI] [PubMed] [Google Scholar]
- 17.Serda RE, Gu J, Burks JK, Ferrari K, Ferrari C, Ferrari M. Cytometry A. 2009;75:752. doi: 10.1002/cyto.a.20769. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Blazer-Yost BL, Banga A, Amos A, Chernoff E, Lai X, Li C, Mitra S, Witzmann FA. Nanotoxicology. 2011;5:354. doi: 10.3109/17435390.2010.514076. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Lai X, Blazer-Yost BL, Clack JW, Fears SL, Mitra S, Ntim SA, Ringham HN, Witzmann FA. doi: 10.1504/IJBNN.2013.054508. unpublished. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Farrah T, Deutsch EW, Omenn GS, Campbell DS, Sun Z, Bletz JA, Mallick P, Katz JE, Malmstrom J, Ossola R, Watts JD, Lin B, Zhang H, Moritz RL, Aebersold R. Mol Cell Proteomics. 2011;10:M110 006353. doi: 10.1074/mcp.M110.006353. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Huang HT, Brand OM, Mathew M, Ignatiou C, Ewen EP, McCalmon SA, Naya FJ. J Biol Chem. 2006;281:39370. doi: 10.1074/jbc.M603244200. [DOI] [PubMed] [Google Scholar]
- 22.Pacholsky D, Vakeel P, Himmel M, Lowe T, Stradal T, Rottner K, Furst DO, van der Ven PF. J Cell Sci. 2004;117:5257. doi: 10.1242/jcs.01406. [DOI] [PubMed] [Google Scholar]
- 23.Scumaci D, Gaspari M, Saccomanno M, Argiro G, Quaresima B, Faniello CM, Ricci P, Costanzo F, Cuda G. Anal Biochem. 2011;418:161. doi: 10.1016/j.ab.2011.06.032. [DOI] [PubMed] [Google Scholar]
- 24.Gundry RL, Fu Q, Jelinek CA, Van Eyk JE, Cotter RJ. Proteomics Clin Appl. 2007;1:73. doi: 10.1002/prca.200600276. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Gundry RL, White MY, Nogee J, Tchernyshyov I, Van Eyk JE. Proteomics. 2009;9:2021. doi: 10.1002/pmic.200800686. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Zhou M, Lucas DA, Chan KC, Issaq HJ, Petricoin EF, Liotta LA, Veenstra TD, Conrads TP. Electrophoresis. 2004;25:1289. doi: 10.1002/elps.200405866. [DOI] [PubMed] [Google Scholar]
- 27.Harris MA, Clark J, Ireland A, Lomax J, Ashburner M, Foulger R, Eilbeck K, Lewis S, Marshall B, Mungall C, Richter J, Rubin GM, Blake JA, Bult C, Dolan M, Drabkin H, Eppig JT, Hill DP, Ni L, Ringwald M, Balakrishnan R, Cherry JM, Christie KR, Costanzo MC, Dwight SS, Engel S, Fisk DG, Hirschman JE, Hong EL, Nash RS, Sethuraman A, Theesfeld CL, Botstein D, Dolinski K, Feierbach B, Berardini T, Mundodi S, Rhee SY, Apweiler R, Barrell D, Camon E, Dimmer E, Lee V, Chisholm R, Gaudet P, Kibbe W, Kishore R, Schwarz EM, Sternberg P, Gwinn M, Hannick L, Wortman J, Berriman M, Wood V, de la Cruz N, Tonellato P, Jaiswal P, Seigfried T, White R. Nucleic Acids Res. 2004;32:D258. doi: 10.1093/nar/gkh036. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Liotta LA, Petricoin EF. J Clin Invest. 2006;116:26. doi: 10.1172/JCI27467. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Tirumalai RS, Chan KC, Prieto DA, Issaq HJ, Conrads TP, Veenstra TD. Mol Cell Proteomics. 2003;2:1096. doi: 10.1074/mcp.M300031-MCP200. [DOI] [PubMed] [Google Scholar]
- 30.Shemetov AA, Nabiev I, Sukhanova A. ACS Nano. 2012;6:4585. doi: 10.1021/nn300415x. [DOI] [PubMed] [Google Scholar]
- 31.Wiogo HTR, Lim M, Bulmus V, Yun J, Amal R. Langmuir. 2010;27:843. doi: 10.1021/la104278m. [DOI] [PubMed] [Google Scholar]
- 32.Tenzer S, Docter D, Rosfa S, Wlodarski A, Kuharev J, Rekik A, Knauer SK, Bantz C, Nawroth T, Bier C, Sirirattanapan J, Mann W, Treuel L, Zellner R, Maskos M, Schild H, Stauber RH. ACS Nano. 2011;5:7155. doi: 10.1021/nn201950e. [DOI] [PubMed] [Google Scholar]
- 33.Zhang H, Burnum KE, Luna ML, Petritis BO, Kim JS, Qian WJ, Moore RJ, Heredia-Langner A, Webb-Robertson BJ, Thrall BD, Camp DG, Smith RD, Pounds JG, Liu T. Proteomics. 2011;11:4569. doi: 10.1002/pmic.201100037. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Monopoli MP, Walczyk D, Campbell A, Elia G, Lynch I, Bombelli FB, Dawson KA. J Am Chem Soc. 2011;133:2525. doi: 10.1021/ja107583h. [DOI] [PubMed] [Google Scholar]
- 35.Prapainop K, Wentworth P. Eur J Pharm Biopharm. 2011;77:353. doi: 10.1016/j.ejpb.2010.12.026. [DOI] [PubMed] [Google Scholar]
- 36.Capriotti AL, Caracciolo G, Caruso G, Foglia P, Pozzi D, Samperi R, Lagana A. Anal Biochem. 2011;419:180. doi: 10.1016/j.ab.2011.08.003. [DOI] [PubMed] [Google Scholar]
- 37.Capriotti AL, Caracciolo G, Caruso G, Foglia P, Pozzi D, Samperi R, Lagana A. Proteomics. 2011;11:3349. doi: 10.1002/pmic.201000803. [DOI] [PubMed] [Google Scholar]
- 38.Caracciolo G, Callipo L, De Sanctis SC, Cavaliere C, Pozzi D, Lagana A. Biochim Biophys Acta. 2010;1798:536. doi: 10.1016/j.bbamem.2009.11.007. [DOI] [PubMed] [Google Scholar]
- 39.Capriotti AL, Caracciolo G, Cavaliere C, Crescenzi C, Pozzi D, Lagana A. Anal Bioanal Chem. 2011;401:1195. doi: 10.1007/s00216-011-5188-8. [DOI] [PubMed] [Google Scholar]
- 40.Capriotti AL, Caracciolo G, Caruso G, Cavaliere C, Pozzi D, Samperi R, Lagana A. Anal Bioanal Chem. 2012 doi: 10.1007/s00216-011-5691-y. [DOI] [PubMed] [Google Scholar]
- 41.Lundqvist M, Stigler J, Cedervall T, Berggard T, Flanagan MB, Lynch I, Elia G, Dawson K. ACS Nano. 2011;5:7503. doi: 10.1021/nn202458g. [DOI] [PubMed] [Google Scholar]
- 42.Sund J, Alenius H, Vippola M, Savolainen K, Puustinen A. ACS Nano. 2011;5:4300. doi: 10.1021/nn101492k. [DOI] [PubMed] [Google Scholar]
- 43.Chen Y, Mitra S. J Nanosci Nanotechnol. 2008;8:5770. doi: 10.1166/jnn.2008.215. [DOI] [PubMed] [Google Scholar]
- 44.Li C, Chen Y, Wang Y, Iqbal Z, Chhowalla M, Mitra S. J Mater Chem. 2007;17:2406. [Google Scholar]
- 45.Wang Y, Iqbal Z, Mitra S. Carbon. 2005;43:1015. [Google Scholar]
- 46.Ntim SA, Sae-Khow O, Witzmann FA, Mitra S. J Colloid Interface Sci. 2011;355:383. doi: 10.1016/j.jcis.2010.12.052. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Lai X, Bacallao BL, Blazer-Yost BL, Hong D, Mason SB, Witzmann FA. Proteomics Clin Appl. 2008;2:1140. doi: 10.1002/prca.200780140. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Nesvizhskii AI, Keller A, Kolker E, Aebersold R. Anal Chem. 2003;75:4646. doi: 10.1021/ac0341261. [DOI] [PubMed] [Google Scholar]
- 49.Keller A, Nesvizhskii AI, Kolker E, Aebersold R. Anal Chem. 2002;74:5383. doi: 10.1021/ac025747h. [DOI] [PubMed] [Google Scholar]
- 50.Lai X, Wang L, Tang H, Witzmann FA. J Proteome Res. 2011;10:4799. doi: 10.1021/pr2005633. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Monroe ME, Shaw JL, Daly DS, Adkins JN, Smith RD. Comput Biol Chem. 2008;32:215. doi: 10.1016/j.compbiolchem.2008.02.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Storey JD. J Roy Stat Soc B. 2002;64:479. [Google Scholar]
- 53.Gasteiger E, Hoogland C, Gattiker A, Duvaud S, Wilkins MR, Appel RD, Bairoch A. The Proteomics Protocols Handbook. Humana Press; Totowa, NJ: 2005. p. 571. [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.


