Abstract
Background
Zinc (Zn) and iron (Fe) are essential micronutrients for plant growth and development, their deficiency or excess severely impaired physiological and biochemical reactions of plants. Therefore, a tightly controlled zinc and iron uptake and homeostasis network has been evolved in plants. The Zinc-regulated transporters, Iron-regulated transporter-like Proteins (ZIP) are capable of uptaking and transporting divalent metal ion and are suggested to play critical roles in balancing metal uptake and homeostasis, though a detailed analysis of ZIP gene family in maize is still lacking.
Results
Nine ZIP-coding genes were identified in maize genome. It was revealed that the ZmZIP proteins share a conserved transmembrane domain and a variable region between TM-3 and TM-4. Transiently expression in onion epidermal cells revealed that all ZmZIP proteins were localized to the endoplasmic reticulum and plasma membrane. The yeast complementation analysis was performed to test the Zn or Fe transporter activity of ZmZIP proteins. Expression analysis showed that the ZmIRT1 transcripts were dramatically induced in response to Zn- and Fe-deficiency, though the expression profiles of other ZmZIP changed variously. The expression patterns of ZmZIP genes were observed in different stages of embryo and endosperm development. The accumulations of ZmIRT1 and ZmZIP6 were increased in the late developmental stages of embryo, while ZmZIP4 was up-regulated during the early development of embryo. In addition, the expression of ZmZIP5 was dramatically induced associated with middle stage development of embryo and endosperm.
Conclusions
These results suggest that ZmZIP genes encode functional Zn or Fe transporters that may be responsible for the uptake, translocation, detoxification and storage of divalent metal ion in plant cells. The various expression patterns of ZmZIP genes in embryo and endosperm indicates that they may be essential for ion translocation and storage during differential stages of embryo and endosperm development. The present study provides new insights into the evolutionary relationship and putative functional divergence of the ZmZIP gene family during the growth and development of maize.
Keywords: Embryo, Endosperm, Expression profiling, Zinc, Iron, Zinc-regulated transporters, Iron-regulated transporter-like protein (ZIP), Subcellular localization, Yeast complementation, Maize
Background
Zinc and iron are essential for plant metabolism and development [1]. Zinc serves as a key structural motif in many proteins, including DNA-binding Zn-finger protein [2,3], RING finger proteins and LIM domain containing proteins [4], which play vital roles in controlling cellular processes such as growth, development and differentiation. An adequate Zn content enhances crop productivity [5]. Iron plays an important role in electron transfer in photosynthesis and respiration, though high concentration of intracellular iron may lead to elevated Fe3+/Fe2+ redox reactions and cause damage [6]. The deficiency of Zn and Fe decreases plant growth and affecting cereal production and grain quality [7], but excess Zn and Fe may cause significant toxicity to biological systems [8,9]. Therefore, plants have established a tightly controlled system to balance the uptake, utilization and storage of these metal ions [10,11]. Because Zn cannot diffuse across cell membrane, specific zinc transporters are required to transport Zn into cytoplasm [12,13]. In recent years, a number of metal transporters have been identified in plants, including the P1B-ATPase family, zinc-regulated transporter (ZRT), iron-regulated transporter (IRT)-like protein (ZIP), natural resistance-associated macrophage protein (NRAMP) family, and cation diffusion facilitator (CDF) family [14].
ZRT, IRT-like protein (ZIP) family has been characterized ubiquitously in organisms, including archaea, bacteria, fungi, plants and mammals, and has been demonstrated to be involved in metal uptake and transport [11]. ZIP proteins generally contribute to metal ion homeostasis by transporting cations into the cytoplasm [14]. Functional complementation in yeast indicated that ZIP proteins are able to transport various divalent cations, including Fe2+, Zn2+, Mn2+, and Cd2+[15]. The ZIP proteins consist of 309-476 amino acid residues with eight potential transmembrane domains and a similar membrane topology. There is also a variable region between TM-3 and TM-4, in which the amino- and carboxyl-terminals located on the outside surface of plasma membrane. The variable region contains a potential metal-binding domain rich in conserved histidine residues [15]. Several ZIP proteins have been identified in Arabidopsis[16]. AtIRT1 (Iron-regulate transporter 1) was the first member to be identified through functional complementation of a yeast mutant defective in iron uptake, and it encodes a major Fe transporter at the root surface in Arabidopsis[17-20]. Further analysis showed that the irt1 mutant exhibited lethal chlorotic phenotypes [18-20], and had lower Ni accumulation under Fe-deficient conditions than the wild type plants. These results indicated that AtIRT1 mediates Fe and Ni translocation in Arabidopsis[21]. Likewise, overexpressing AtIRT3 leads to increased accumulation of Zn in shoots and Fe in roots. Moreover, AtIRT3 could complement the Zn and Fe uptake double yeast mutants, indicating that AtIRT3 is involved in Zn and Fe translocation [22]. Besides, expression analysis revealed that the transcripts of AtZIP1 to AtZIP5, AtZIP9 to AtZIP12, and AtIRT3 were increased in response to Zn-deficiency, suggesting that they may enhance Zn acquisition under deficient Zn status in Arabidopsis[23]. In rice, overexpression of OsIRT1 leads to increased Fe and Zn accumulations in shoots, roots and mature seeds, suggesting OsIRT1 is a functional metal transporter for iron, and it is responsible for the absorption of iron from soil, especially under Fe-deficiency [24-26]. On the contrary, over accumulation of OsZIP4 and OsZIP5 cannot increase the Zn content in seeds, though the Zn concentration in roots were dramatically increased in transgenic plants [27,28]. These results indicated that maintaining the endogenous expression pattern of ZIP genes may be essential for Zn translocation in plants. Likewise, overexpression of TdZIP1, a Zn transporter from wild emmer wheat, causes excess accumulation of Zn in cells, thus generating a toxic cytosolic environment [29]. Therefore, increase Zn content by transgenic approaches may benefit from elucidating the expression pattern of ZIP genes.
Since ZIP is the key transporter for Zn and Fe uptake and translocation in plants, considerable progress has been achieved in cloning and characterizing its functions in crop plants, including soybean and maize [30,31]. The soybean GmZIP1 is highly selective for Zn, and it might play a role in the symbiotic relationship between soybean and Bradyrhizobium japonicum[30]. The ZmZLP1 (ZmZIP-like protein) was identified from a cDNA library of Zea mays L. (maize) pollen. It was reported that ZmZLP1 localized to the endoplasmic reticulum and may be responsible for transporting zinc from the ER to the cytoplasm, though its physiological function has not been characterized [31]. The maize genome has been thoroughly sequenced and assembled. However, systematic analysis of the maize ZIP gene family is still limited. In the present study, we provide detailed information on the gene identification, chromosomal locations, subcellular localizations and expression patterns of nine ZmZIP genes. In addition, the transporter activities of ZmZIPs were tested by yeast complementation analysis. Our results suggest that ZmZIP genes may be responsible for the uptake and translocation of Zn or Fe and involve in detoxification and storage of metals in plant cells, as well as embryo and endosperm development.
Results
Cloning of ZmZIP genes and phylogenetic analysis
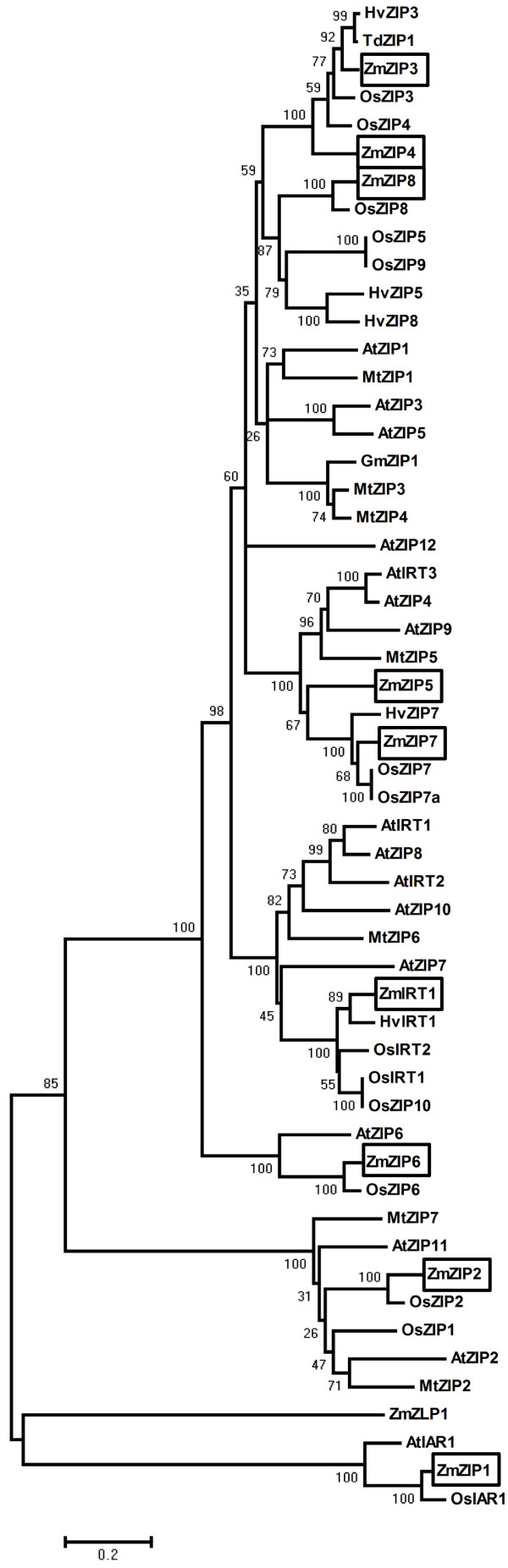
Extensive searches of public genomic databases, by using reported ZIP proteins from rice as TBLASTN queries, identified a total of nine maize ZIP genes that have complete sequences. Nine cDNA fragments containing complete opening reading frame (ORF) were cloned from leaf tissues of maize (Zea mays inbred line X178) by RT-PCR, and they were designated as ZmZIP1-8 and ZmIRT1 according to the amino acid sequence similarity with the rice and Arabidopsis (Table 1). The predicted proteins of ZmZIPs consist of 359-490 amino acids, and harbour 6-9 putative transmembrane (TM) domains. In addition, there was a variable region between TM-3 and TM-4, containing a potential metal-binding domain rich in histidine residues (Figure 1). The phylogenetic analysis showed that ZmZIP1 was closely related to AtIAR1 and OsIAR1; ZmIRT1 was related to HvIRT1 and resided in a branch comprised by iron transporter OsIRT1 and OsIRT2. It was suggested that the amino acid residue D100 in AtIRT1 is responsible for Fe and Mn transport [22], and it is conserved in ZmIRT1 (Additional file 1), which indicates that ZmIRT1 may be functional related to a Fe transporter. In addition, ZmZIP3 and ZmZIP4 formed a gene cluster with OsZIP3 and OsZIP4; ZmZIP2 was most closely related to OsZIP2 in the other branch comprised by zinc transporters OsZIP1, AtZIP2 and AtZIP11; ZmZIP5 and ZmZIP7 were observed in a branch; ZmZIP8 and ZmZIP6 were closely related to OsZIP8 and OsZIP6, respectively (Figure 2). These results revealed that ZmZIPs may be functional Zn or Fe transporters.
Table 1.
List of ZIP genes in the maize genome
| Gene name | Gene IDa | NCBI accession | Chromosome | CDS lengthb | Protein length | TM domains |
|---|---|---|---|---|---|---|
|
ZmZIP1 |
LOC100192506 |
NM_001137726 |
1 |
1473 |
490 |
7 |
|
ZmZIP2 |
LOC100286282 |
NM_001159169 |
2 |
1080 |
359 |
9 |
|
ZmZIP3 |
LOC100282628 |
NM_001155536 |
2 |
1104 |
367 |
6 |
|
ZmZIP4 |
LOC100280287 |
HM048832 |
4 |
1161 |
386 |
6 |
|
ZmZIP5 |
LOC100281339 |
NM_001154257 |
6 |
1209 |
402 |
6 |
|
ZmZIP6 |
LOC100283249 |
NM_001156151 |
8 |
1191 |
396 |
8 |
|
ZmZIP7 |
LOC100284121 |
NM_001157018 |
6 |
1164 |
387 |
6 |
|
ZmZIP8 |
LOC100281849 |
NM_001154769 |
7 |
1191 |
396 |
7 |
| ZmIRT1 | LOC100285748 | NM_001158638 | 1 | 1146 | 381 | 8 |
aThe Gene ID from MaizeGDB.
bCDS of each predicted gene sequence was obtained from Genbank by BLAST search.
Figure 1.
Amino acid alignment of predicted ZIP proteins in maize genome. Sequences were aligned using Clustal X Version 2.0 and identical or similar amino acids are shaded by BOXSHADE (http://www.ch.embnet.org/software/BOX_form.html). The membrane spanning domains are indicated as lines above the sequence, and numbered I–VIII, respectively.
Figure 2.
Phylogenetic analysis of the ZIP family members from various species. The unrooted phylogenetic tree was constructed with the deduced protein sequences of ZIP proteins from various species using the neighbor-joining method in MEGA 4.0 software. For proteins and accession numbers used for phylogenetic analysis, refer to Methods. The ZmZIP proteins are boxed, and the scale bar corresponds to a distance of 20 changes per 100 amino acid positions.
Expression patterns of ZmZIP genes in different tissues of maize
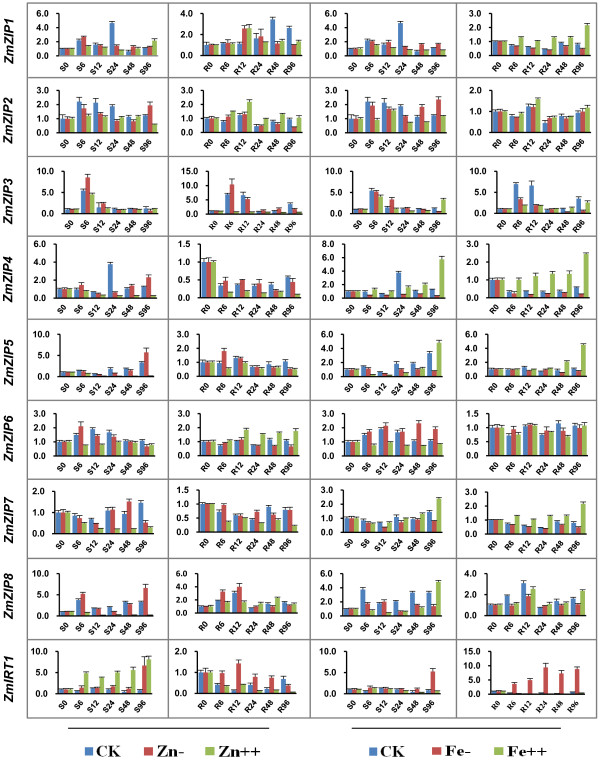
Recently, functional analysis has revealed that some ZIP genes in Arabidopsis and rice play important roles in transporting Zn or Fe [26,27,32]. To better understand the physiological functions of ZmZIP genes, their expression patterns in seedlings (under various metal conditions), embryo and endosperm were investigated by real-time RT-PCR. Under sufficient metal nutritional conditions, eight ZmZIP genes were mainly expressed in shoots, while ZmIRT1 was accumulated abundantly in both shoots and roots (Additional file 2). In addition, the expression profiles of ZmZIP genes were determined in response to fluctuating divalent metal status. Under Zn-deficient conditions, the expression levels of ZmIRT1, ZmZIP5 and ZmZIP8 were up-regulated in shoots at 96 h, whereas ZmZIP3 was induced in both shoots and roots at 6 h (Figure 3). The accumulation of ZmIRT1 was gradually increased in response to Zn-excess in shoots and reached the maximum level at 96 h after treatment, while that was suppressed in roots. Moreover, the expression of ZmZIP4, 5, 7 and 8 were suppressed and decreased gradually in shoots, while ZmZIP3 was down-regulated remarkably in roots (Figure 3). These results indicated that ZmIRT1, ZmZIP3, 4, 5, 7 and 8 are sensitive to fluctuating environmental Zn status in seedlings.
Figure 3.
Expression patterns of the nine ZmZIP genes in maize seedlings under various metal conditions. Two-week old shoots (S) and roots (R) of maize seedlings, under standard nutrient condition (CK), Zn, Fe-deficiency (Zn-), (Fe-), 200 μM ZnSO4 (Zn++) and 500 μM FeSO4 (Fe++) treated, were harvested respectively at 0 h, 6 h, 12 h, 24 h, 48 h, 96 h after treatment. Relative mRNA abundance of each gene was normalized with ZmActin1 gene. Data from real-time RT-PCR experiments were analyzed according to the 2-∆∆Ct method. The error bars indicate standard deviations.
Under Fe-deficient conditions, the expression of ZmIRT1 was dramatically up-regulated in shoots and roots. In response to Fe-excess, the transcript level of ZmIRT1 was suppressed in shoots and roots, while that of ZmZIP4, 5, 7 and 8 were increased and reached to the maximum level at 96 h in both shoots and roots (Figure 3). These results indicated that ZmIRT1 and ZmZIP4, 5, 7, 8 are sensitive to environmental Fe conditions in shoots and roots. Under Cu- and Mn-deficient conditions, the expression patterns of ZmZIP genes showed no obvious change (Additional file 3).
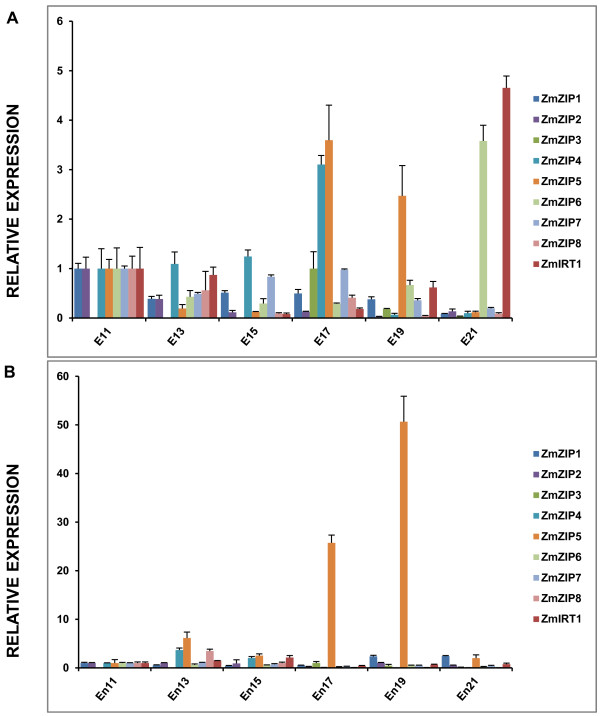
A detailed analysis of the transcriptional profiles of ZmZIP genes along embryo and endosperm development of maize were measured by real-time RT-PCR. Total RNA was isolated from embryo and endosperm on indicated days after pollination (DAP) (Figure 4A and 4B). The results revealed that the expression levels of ZmZIP4 and ZmZIP5 were up-regulated on 17 DAP, while ZmIRT1 and ZmZIP6 were mainly expressed in the late developmental stages of embryo. Moreover, the expression of ZmZIP5 was dramatically increased associated with endosperm development and reached a peak on 19 DAP (approximately 50-fold higher than that of 11 DAP) (Figure 4A and 4B). Taken together, these results suggested that ZmIRT1, ZmZIP4 and ZmZIP6 may play important roles during embryo development in maize, while ZmZIP5 may be essential for both embryo and endosperm development.
Figure 4.
Expression patterns of the nine ZmZIP genes during embryo and endosperm development. The developing embryo (E) and endosperm (En) were obtained on indicated days after pollination (11 DAP, 13 DAP, 15 DAP, 17 DAP, 19 DAP and 21 DAP). The relative mRNA abundance of each gene in embryo (A) and endosperm (B) was normalized with ZmActin1 gene. Data from real-time RT-PCR experiments were analyzed according to the 2-∆∆Ct method. The error bars indicate standard deviations.
Subcellular localization of ZmZIPs
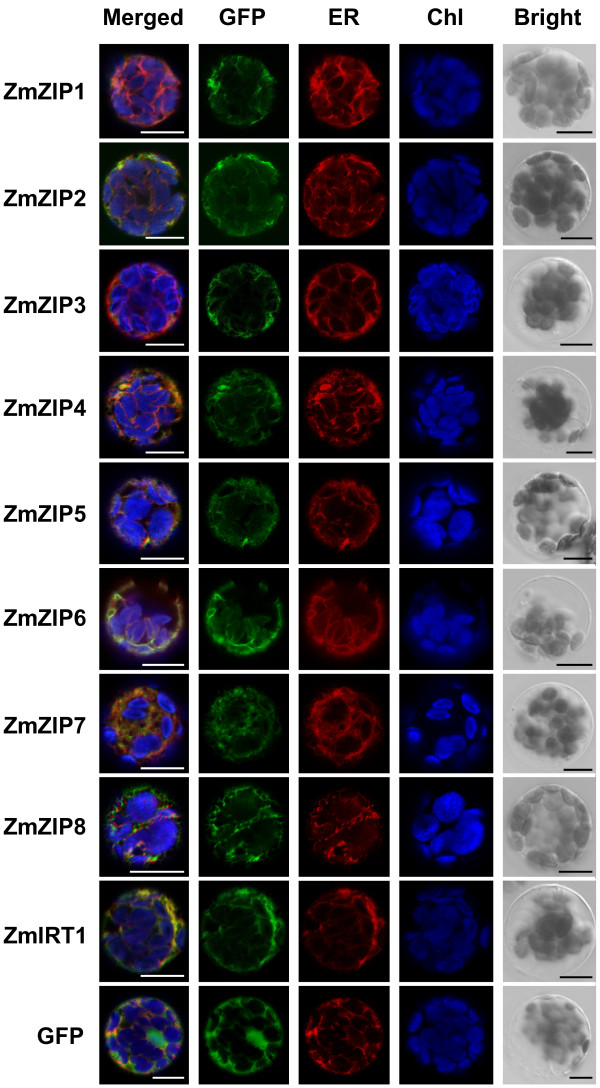
To study the localization of ZmZIP proteins, the full-length coding regions without termination codons of the ZmZIP genes were C-terminal fused with green fluorescent protein (GFP) and the fusion proteins were expressed under cauliflower mosaic virus 35S promoter. The ZmZIPs-GFP fusion proteins were transiently expressed in Arabidopsis mesophyll protoplasts and onion epidermal cells. The green fluorescence was visualized by confocal microscopy. As showed in Figure 5 and Additional file 4, all ZmZIP proteins were localized to the plasma membrane and endomembrane system, which was further identified as endoplasmic reticulum (ER) by co-localization with an ER marker in Arabidopsis mesophyll protoplasts (Figure 5). In addition, the GFP signal can be observed at the border of the cell, indicating the plasma membrane localization of ZmZIPs (Additional file 4). Moreover, plasmolysis was performed to further confirm the plasma membrane localization (Additional file 5). Cells expressing GFP were used as controls, which showed green fluorescence in nucleus and cytoplasm.
Figure 5.
Subcellular localization of ZmZIPs in Arabidopsis mesophyll protoplasts. Full-length coding regions without stop codon of the ZmZIP genes were cloned into the pRTL2GFP vector and the resulting plasmid was transiently transformed into Arabidopsis mesophyll protoplasts by the PEG method. The fluorescence of ER marker is indicated in red, the GFP signal is shown in green and the autofluorescence of the chlorophyll (Chl) is artificially stained in blue. The images were obtained by a confocal microscope, and the cytoplasm localization of GFP is used as a control. The scale bar represents 10 μm.
Complementation in yeast cells
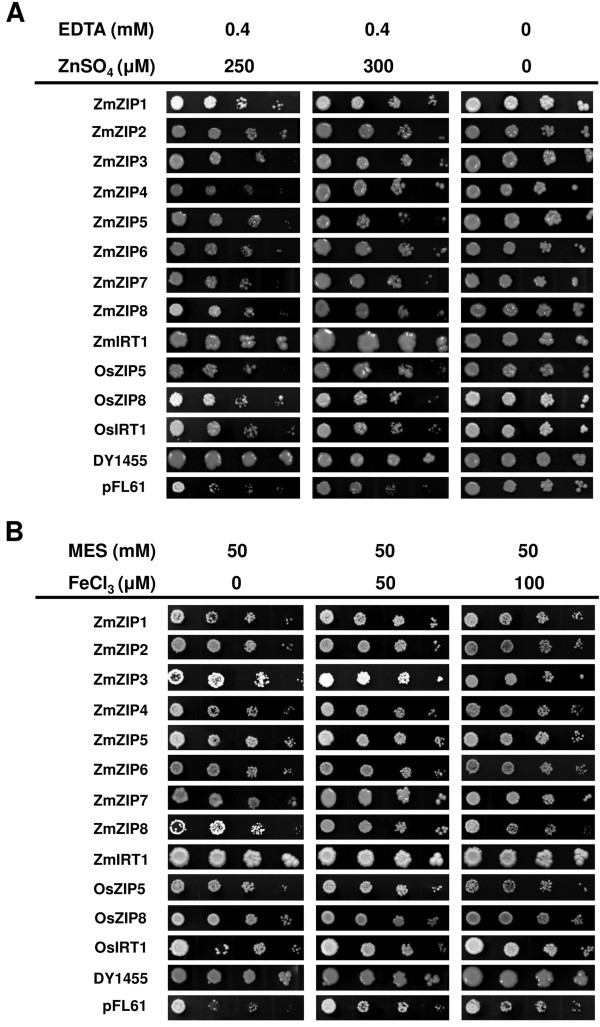
The yeast complementation analysis was performed in order to test whether ZmZIPs possess the capacity of transporting Zn or Fe in organisms. Although, it was demonstrated that Mn is a potential substrate for ZIPs [22,33,34], it has not been tested in this study. The full-length cDNAs of ZmZIP genes were expressed in the Saccharomyces cerevisiae zrt1zrt2 double mutant (ZHY3) and fet3fet4 double mutant (DEY1453). The transformed ZHY3 expressing ZmZIP genes were grew on SD media plus 0.4 mM EDTA. The result showed that the expression of ZmIRT1 protein remarkably reversed this growth defect in the yeast mutant, while other ZmZIPs reversed this growth defect as efficiently as the functional characterized ZIP proteins, OsZIP5 and OsZIP8 (Figure 6A). To test the iron transporter activity, the yeast mutants DEY1453 transformed with pFL61-ZmZIPs were grown on SD media plus 50 mM MES. The result revealed that ZmIRT1 reversed the growth defect significantly, while the effect of other proteins were relatively inferior, though that was as efficiently as a reported iron transporter OsIRT1 (Figure 6B). It was interesting to find that the yeast mutants expressing ZmIRT1 showed the strongest propagation under both Zn- and Fe-limited conditions (Figure 6A and 6B), indicating that ZmIRT1 may be essential for Zn and Fe translocation during maize growth. In addition, except for ZmIRT1, other ZmZIP proteins can complement the iron transporter mutant fet3fet4 at high pH but weakly at low pH conditions (Additional file 6). Thus, the ZmZIP genes identified in this study encoded functional Zn or Fe transporter, and ZmIRT1 may play a dominant role in ion translocation in maize.
Figure 6.
Functional complementation in yeast mutant strains. The Zn and Fe transportation mutant zrt1zrt2(A) and fet3fet4(B) were transformed with the expression vector pFL61 carrying ZmZIP1-ZmZIP8, ZmIRT1 and the functional characterized ZIP genes, OsZIP5, OsZIP8 and OsIRT1. The wild type strain DY1455 transformed with pFL6 was applied as a positive control, and the yeast mutant transformed with empty vector pFL61 was used as a negative control. The transformed yeast cells were grown under different metal conditions as indicated, and the transformed fet3fet4 was grown on medium with pH 5.8. Cell concentration was adjusted to OD600 = 1 and serial dilutions (1.0, 0.1, 0.01 and 0.001) were made. For assay, 5-μL of each dilution was spotted on plates and grown for 6 days at 30°C.
Discussion
The ZIP genes have been reported in several plants, including Arabidopsis, rice, Medicago truncatula, wild emmer wheat, Vitis vinifera L and barley [18-22,24-29,32,33,35-45]. Most of these genes are found to function as Zn or Fe transporter, though some ZIPs have major roles in Mn transport [22,33,34]. Members of the ZIP family have eight predicted transmembrane (TM) domains and a variable region between TM-3 and TM-4 where contains a potential metal-binding domain [15]. Although ZIPs have been characterized in many plants, to the best of our knowledge, there were few reports concerning the isolation and functional characterization of ZIPs in maize. Since, the maize genome sequencing project was completed, many gene families have been identified and characterized in maize [46-48]. In this study, nine cDNAs encoding ZmZIP genes were obtained from maize based on amino acids sequences similarity. In order to analyze the evolutionary relationship of the ZIP family, a phylogenetic tree consists of nine ZIPs and one ZLP1 from maize, 14 rice ZIPs (OsZIPs), 16 Arabidopsis ZIPs (AtZIPs), 5 Hordeum vulgare ZIPs (HvZIPs), 7 Medicago truncatula ZIPs (MtZIPs), Wild Emmer Wheat ZIP (TdZIP1), and Glycine max ZIP (GmZIP1) was generated (Figure 2). It was revealed that the predicted amino acid sequences of ZmZIPs were closely related to ZIPs from other plant species and they were existed as orthologs (ZmIRT1 and HvIRT1, ZmZIP1 and OsIAR1, ZmZIP2 and OsZIP2, ZmZIP3 and OsZIP3, ZmZIP4 and OsZIP4, ZmZIP6 and OsZIP6, ZmZIP7 and OsZIP7, ZmZIP8 and OsZIP8), indicating that those ZIPs from maize, rice and barley may share a common evolutionary ancestor. It has been reported that OsZIP4, OsZIP5 and OsZIP8 are functional zinc transporters and localized to the plasma membrane [27,32,36]. AtIRT2 is an iron transporter and localized to intracellular vesicles, suggesting an essential role in preventing metal toxicity through compartmentalization and remobilize iron stores from internal storage vesicles [40]. All of these proteins are able to complement the growth of the yeast strain ZHY3 or DEY1453, which is sensitive to Zn or Fe deprivation due to the mutation in both high and low Zn or Fe affinity system. In this study, similar to the Arabidopsis and rice ZIPs, the maize ZIP proteins showed different degree of zinc or iron complementary capabilities (Figure 6A and 6B). Moreover, ZmZIPs were localized to the plasma membrane and endoplasmic reticulum (Figure 5), suggesting they may functional related to excessive ion detoxification. Hence, these results demonstrated that ZmZIP genes encode Zn or Fe transporters and have various functions associated with uptake and translocation, detoxification and storage of Zn or Fe in plant cells.
The expression patterns of ZmZIP genes reflect their diverse functions during Zn or Fe translocation. It has been reported that the ZIP genes displayed various expression profiles regarding tissue specificity and response to fluctuating environmental Zn and Fe conditions. For example, OsZIP7a was induced mainly in Fe-deficient roots, while OsZIP8 was stimulated in Zn-deficient shoots and roots [41]. Histochemical localization analysis showed that the mRNA of OsZIP4 was accumulated in the vascular bundles of leaves and roots, phloem cells of the stem and the meristems [36]. In the model legume Medicago truncatula, the expression of MtZIP2 was detected in roots and stems and was induced by Zn deficiency [49], likewise MtZIP1 was expressed in Zn-deficient roots and leaves [37]. In our study, the expression of ZmIRT1 was remarkably up-regulated in roots and shoots under Fe-deficiency, and was induced in shoots at 96 h after Zn-deficiency (Figure 3). ZmZIP4, 5, 7 and 8 were induced and reached a peak in shoots and roots at 96 h after Fe-excess (Figure 3). These results suggested that ZmIRT1 may play essential roles in Fe and Zn uptake, especially under iron deficiency, while ZmZIP4, 5, 7 and 8 may associate with detoxification and storage of excessive Fe. Previous study showed that Arabidopsis thaliana transcription factors bZIP19 and bZIP23 regulate the adaptation to zinc deficiency by increase the transcription of ZIPs and other genes [50]. Therefore, maize bZIP-like transcription factors may be essential for the regulation of ZmZIP expression under Zn deficient status.
It has been demonstrated that Zn play essential roles in embryo and endosperm development [2-4]. Therefore, the ZmZIP genes that preferentially expressed in embryo and endosperm may be important for translocation of Zn2+ into sink organs. The expression analysis showed that ZmIRT1, ZmZIP4 and ZmZIP6 were mainly expressed in embryo, while ZmZIP5 was expressed in both embryo and endosperm. The accumulations of ZmZIP4 and ZmZIP5 were up-regulated in the middle stages of embryo development and then they were repressed, suggesting that they may be essential for plumule and radicle growth. In contrast, ZmIRT1 and ZmZIP6 were stimulated in the late development stages of embryo, which indicates that they may associate with the maturation of embryo. Interestingly, it was observed that the accumulation of ZmZIP5 was dramatically up-regulated during the development of endosperm and reached a peak on 19 DAP, though its expression was decreased on 21 DAP, which suggests that ZmZIP5 may involve in the accumulation of nutrient substance at early grain filling stage. By the light of well assembled genome sequence of maize, an in silico promoter analysis was performed for those ZmZIP genes preferentially expressed in embryo and endosperm. The results showed that the ZmIRT1, ZmZIP4, ZmZIP5 and ZmZIP6 contain cis-elements for seed expression (Additional file 7). Considering that there was some correlation between the VvZIP3 expression profile and the Zn accumulation pattern during the development of reproductive organs [42], we assumed that ZmIRT1, ZmZIP4, ZmZIP5 and ZmZIP6 may play various roles in both embryo and endosperm development.
Enhancing the iron and zinc content in cereal grains is important for improving human nutrition. Since the amount of metal transporter is generally rate-limiting [39], manipulating the transporters involved in translocation of micro-essential metals into sink organs could be a way to increase mineral contents. For instance, overexpressing AtZIP1 in barley resulted in a rise in the short-term zinc uptake as well as higher seed Zn and Fe contents [39]. Likewise, the iron and zinc contents were elevated in the shoots, roots and mature seeds of transgenic rice constitutively overexpressing OsIRT1[26]. However, overexpressing OsZIP4 under the control of the cauliflower mosaic virus (CaMV) 35S promoter lead to Zn accumulation in roots, while the Zn concentration in seeds were four times lower than untransgenic controls [28]. Moreover, overexpressing OsZIP5 and OsZIP8 in rice under the control of the maize ubiquitin promoter lead to increased Zn level in roots, though that in shoots and mature seeds were reduced in the transgenic plants [27,32]. These results indicate that ectopic overexpression of ZIP proteins may have little effect on the enhancement of Zn content in seeds due to overproduction of ZIP in vegetative tissues. Therefore, increasing the accumulation of ZIP proteins and maintain their expression pattern may provide an alternative way to enhance Zn or Fe contents. Since it was revealed that the expression of ZmIRT1, ZmZIP4, ZmZIP5 and ZmZIP6 are associated with seed development, it can be assumed that overexpression of these ZIP genes in a seed specific manner may provide an alternative strategy for biofortification of crops with Zn and Fe.
Conclusions
Although zinc and iron are essential micronutrients for plant growth and development, functional analysis of ZIP family in maize is still limited. The present study provides relevant information concerning the identification and functional characterization of ZmZIP family, and suggests that they may involve in metal uptake and overall cell zinc homeostasis. It is also indicated that ZmZIPs may be essential for ion translocation and storage during differential stages of embryo and endosperm development. In the present study, we provided detailed information of the evolutionary relationship and putative functional divergence of the ZmZIP gene family during the growth and development of maize.
Methods
Plant growth
Zea mays seeds were cultured in vermiculite (irrigated with Hoagland nutrient solution) in climate chambers with a light/dark cycle of 16/8 h. The 13-day-old seedlings were transferred to the Hoagland nutrient solution grown for 6 days (The standard Hoagland nutrient solution contained 2.0 μM ZnSO4, 50 μM Fe(III)-EDTA, 0.5 μM CuSO4, and 2.0 μM MnSO4), then transferred to the standard Hoagland nutrient solution and Hoagland medium lacking ZnSO4 (Zn-deficient), Fe (III)-EDTA (Fe- deficient), and 200 μM ZnSO4 (Zn-excess), 500 μM Fe(III)-EDTA (Fe-excess). Shoots and roots were selected and detached at 0 h, 6 h, 12 h, 24 h, 48 h and 96 h after the different treatment for real-time RT-PCR assays.
Bioinformatics analysis
The BLAST program at GenBank (http://www.ncbi.nlm.nih.gov/blast) was used to search the maize ZIP cDNAs and the acquired cDNAs were compared to the corresponding genome database from maizesequence (http://maizesequence.org/index.html). The putative amino acid sequences were aligned with the program Clustal X Version 2.0 [51] and colored by BOXSHADE (http://www.ch.embnet.org/software/BOX_form.html). Potential transmembrane domains in protein sequences were identified using TMHMM [52]. The phylogenetic tree of 54 members of the ZIP proteins from various species was constructed using MEGA version 4.0 [53]. The accession numbers for the proteins are as follows: Arabidopsis thaliana (AtZIP1: AAC24197, AtZIP2: AAC24198, AtZIP3: AAC24199, AtZIP4: AAB65480, AtZIP5: AAL38432, AtZIP6: AAL38433, AtZIP7: AAL38434, AtZIP8: AAL83293, AtZIP9: AAL38435, AtZIP10: AAL38436, AtZIP11: AAL67953, AtZIP12: AAL38437, AtIRT1: AAB01678, AtIRT2: NP_001031670, AtIRT3: NP_564766, AtIAR1: AF216524); Oryza sativa (OsIAR1: NP_001062003, OsIRT1: AB070226, OsIRT2: AB126086, OsZIP1: AY302058, OsZIP2: AY302059, OsZIP3: AY323915, OsZIP4: AB126089, OsZIP5: AB126087, OsZIP6: AB126088, OsZIP7: AB126090, OsZIP7a: AY275180, OsZIP8: AY324148, OsZIP9: AY281300, OsZIP10: AK107681); Zea mays (ZmZIP1: NM_001137726, ZmZIP2: NM_001159169, ZmZIP3: NM_001155536, ZmZIP4: HM048832, ZmZIP5: NM_001154257, ZmZIP6: NM_001156151, ZmZIP7: NM_001157018, ZmZIP8: NM_001154769, ZmIRT1: NM_001158638, ZmZLP1: ACO50388); Hordeum vulgare (HvIRT1: EU545802, HvZIP3: FJ208991, HvZIP5: FJ208992, HvZIP7: AM182059, HvZIP8: FJ208993); Medicago truncatula (MtZIP1: AY339054, MtZIP2: AY007281, MtZIP3: AY339055, MtZIP4: AY339056, MtZIP5: AY339057, MtZIP6: AY339058, MtZIP7: AY339059); Wild Emmer Wheat (TdZIP1:AY864924); Glycine max (GmZIP1: AY029321).
RNA isolation and real-time RT-PCR
Total RNA was isolated from shoots and roots with TRIzol (Takara). For cDNA synthesis, we used 4 μg of total RNA as a template and M-MLV reverse transcriptase (Fermentas) by primering with oligo-d(T)18 in a 40-μL reaction mixture. Real-time RT-PCR was performed in a 20-μL solution containing a 5-μL aliquot of the cDNA, 0.4 μM of gene-specific primers (Additional file 8) and 10-μL SYBR Green I (Takara). The fragment was amplified by PCR in an ABI 7500 Real Time Thermal Cycler. The constitutively expressed ZmActin1 gene [GenBank: J01238.1] was amplified as the reference gene using the primers ZmActin1F and ZmActin1R (Additional file 8). Changes in expression were calculated via the ∆∆Ct method [54]. For all real-time RT-PCR analysis, two biological replicates were used and three technical replicates were performed for each biological replicate. The sizes of the amplified fragments were confirmed by gel electrophoresis.
Cloning of target genes
The cDNA sequences of nine putative maize ZIP genes with a complete ORF were obtained from the MaizeSequence database (http://www.maizesequence.org/). The primers (Additional file 8) were designed for amplifying the ORFs of these nine ZmZIP genes. Total RNA was isolated from shoots with TRIzol (Takara), and was treated with DNaseI (NEB) before being reverse transcribed. For cDNA synthesis, we used 2 μg of total RNA as a template and M-MLV reverse transcriptase (Fermentas) by primering with oligo-d(T)18 in a 20-μL reaction mixture. PCR was performed in a 20-μL solution containing a 5-μL aliquot of the cDNA, 10 μM of gene-specific primers (Additional file 8), 0.5 U Ex Taq polymerase (Takara), 8 mM dNTPs, 10-μL of 2 × GCI buffer. PCR was performed on a DNA amplification machine (BIO RAD) for a denaturation of 4 min at 94°C, followed by 33 cycles of 1 min at 94°C, 1 min at 60°C, and 1 min at 72°C and a final extension of 10 min at 72°C. The PCR products were separated on a 1% agarose gel and purified with Gel DNA Purification Kit (Shenergy Biocolor) according to the manufacturer’s instruction. The purified product was then cloned into the pGEM-T easy vector (Promega, USA) and sequenced (Openlab, China).
Subcellular localization
For subcellular localization, a C-terminal GFP fusion expression vector pRTL2GFP was used [48]. Gene-specific primers were designed and the stop codons were deleted (Additional file 8). The coding regions without the stop codon were cloned into pRTL2GFP, respectively. The plasmids were purified using the Wizard Plus Miniprep DNA Purification System (Promega). The ZmZIP-GFP fusion constructs and the mcherry labeled ER marker were co-transformed into Arabidopsis mesophyll protoplasts as described previously [55,56]. After incubation in the dark at 26°C for 14 h, the fluorescence was examined using a confocal microscope (LSM700; Carl Zeiss). The GFP signal was excitated at 488 nm, and the emission was collected at 500-530 nm; the mcherry signal was excitated at 555 nm, and the fluorescence emission was collected at 610 nm; the 630 emission filter was used to observe the autofluorescence of chlorophyll.
Yeast complementation
The ORFs of ZmZIP genes were amplified with gene specific primers (Additional file 8), and the PCR fragments were purified from an agarose gel and subsequently ligated into the NotI site of the yeast expression vector pFL61 (provided from Dr. David Eide, University of Missouri-Columbia) [57]. The resulting constructs were sequenced to ensure the correct orientations of the inserts and sequences. The yeast strains used in this experiment are zrt1zrt2 ZHY3 (MATα ade6 can1 his3 leu2 trp1 ura3 zrt1::LEU2 zrt2::HIS3), fet3fet4 DEY1453 (MATa/MATa ade2/+ can1/can1 his3/his3 leu2/leu2 trp1/trp1 ura3/ura3 fet3-2::HIS3/fet3-2::HIS3 fet4-1::LEU2/fet4-1::LEU2), and DY1455 (MATa ade6 can1 his3 leu2 trp1 ura3) (provided by Dr. David Eide) [58,59]. The pFL61-ZmZIPs were transformed into the yeast strains ZHY3 and DEY1453 using the electroporation method. The empty vector pFL61was applied as a negative control, OsIRT1 was used as the positive control for iron transporter, OsZIP5 and OsZIP8 were applied as the positive control for zinc transporter, and the wild type strain DY1455 harbouring pFL61 was used as another positive control. The yeast complementation was performed as described previously [22] with slight modification. Transformed cells were selected on synthetic agar medium (SD) containing amino acid supplements without Uracil and 2% glucose. In growth-test experiments, 5-μL drops of yeast culture at an optical density of 1.0, 0.1, 0.01 and 0.001 were spotted onto medium. The yeast strain of zrt1zrt2 ZHY3 were grown on SD/–ura medium (pH 4.4) supplemented with 0.4 mM EDTA and 250 μM or 300 μM ZnSO4. The yeast strain of fet3fet4 were grown on SD/–ura medium (pH 5.5, 5.8 containing 50 mM 2-(N-morpholino) ethanesulfonic acid (MES) supplemented with 0, 50 or 100 μM FeCl3.
Abbreviations
CaMV: Cauliflower mosaic virus; GFP: Green fluorescent protein; ER: Endoplasmic reticulum; ORF: Opening reading frame; RT: Reverse transcripts; TM: Transmembrane; ZIP: Zinc-regulated transporters, iron-regulated transporter-like protein.
Competing interests
The authors declare that they have no competing interests.
Authors’ contributions
SZL and JTC participated in the design of the study. SZL performed the bioinformatics analysis, gene cloning, real-time RT-PCR and yeast complementation. XJZ analyzed the data and drafted the manuscript. YQH assisted in gene cloning and plasmid construction. LYZ and SJZ collected the tissues for temporal and spatial expression analysis. YFZ and JJG helped in bioinformatics analysis. JTC and RMC contributed to revisions of the manuscript. All authors read and approved the final manuscript.
Supplementary Material
Amino acid alignment of ZmIRT1 with other IRT proteins. Sequences were aligned using Clustal X Version 2.0 and the identical or similar amino acids are shaded by BOXSHADE (http://www.ch.embnet.org/software/BOX_form.html). The residue D100 in ZmIRT1 is marked with an asterisk.
Expression patterns of the nine ZmZIP genes in maize seedlings. The shoots (S) and roots (R) of two weeks old hydroponically cultured maize seedlings in Hoagland nutrient solution were harvested, respectively. Relative mRNA abundance of each gene was normalized with ZmActin1 gene. Data from real-time RT-PCR experiments were analyzed according to the 2-∆∆Ct method. The error bars indicate standard deviations.
Expression patterns of the nine ZmZIP genes in maize under Cu-, Mn- deficiency. Two-week old shoots (S) and roots (R) of maize seedlings, under Cu-, Mn-deficiency treated (Cu-), (Mn-) were harvested respectively at 0 h, 6 h, 12 h, 24 h, 48 h and 96 h. Relative mRNA abundance of each gene was normalized with ZmActin1 gene. Data from real-time RT-PCR experiments were analyzed according to the 2-∆∆Ct method. The error bars indicate standard deviations.
Subcellular localization of ZmZIPs in onion epidermal cells. Full-length coding regions without stop codon of the ZmZIP genes were cloned into the pRTL2GFP vector and the resulting plasmid was transiently transformed into onion epidermal cells by bombardment. The fluorescence was observed using a confocal laser scanning microscopy. GFP was imaged using 488 nm excitation and a 500-530 nm bandpass emission filter. The scale bar represents 100 μm.
Subcellular localization of ZmZIP-GFP fusion proteins in plasmolyzed onion epidermal cells. The ZmZIP-GFP fusion proteins were transiently expressed in onion epidermal cells by bombardment, and a set of representative images are shown. The plasmolysis was performed for 15 min in 30% sucrose. The Z-stack of optical sections and single optical slice of GFP fluorescence are shown. The cytoplasm localization of GFP is used as a control. GFP was imaged using 488 nm excitation and a 500-530 nm bandpass emission filter. The scale bar represents 100 μm.
Functional complementation of the Fe transportation yeast mutant by ZmZIPs under different pH values. The Fe transportation mutant fet3fet4 were transformed with the expression vector pFL61 carrying ZmZIP1-ZmZIP8, ZmIRT1 and the functional characterized ZIP genes, OsZIP5, OsZIP8, and OsIRT1. The transformed yeast cells were grown under different pH conditions (A) pH 5.5 and (B) pH 5.8. Cell concentration was adjusted to OD600 = 1 and serial dilutions (1.0, 0.1, 0.01 and 0.001) were made. For assay, 5-μL of each dilution was spotted on plates and grown for 6 days at 30°C.
Putative cis-elements in the 2-Kb upstream promoter region of translation start site in ZmZIP genes.
Primers used in gene cloning, vector construction and real-time RT-PCR analysis.
Contributor Information
Suzhen Li, Email: lisz-1208@163.com.
Xiaojin Zhou, Email: zhouxiaojin@caas.cn.
Yaqun Huang, Email: hyqun@hebau.edu.cn.
Liying Zhu, Email: zhuliyiny@126.com.
Shaojun Zhang, Email: dashao_2011@hotmail.com.
Yongfeng Zhao, Email: zhaoyongfeng@hebau.edu.cn.
Jinjie Guo, Email: guojinjie512@163.com.
Jingtang Chen, Email: chenjingtang@126.com.
Rumei Chen, Email: chenrumei@caas.cn.
Acknowledgements
We thank Dr. David Eide (Nutritional Science Program, University of Missouri, Columbia, USA) for providing the yeast strains DEY1453, ZHY3 and yeast expression vector pFL61; Dr. Dongtao Ren (College of Biological Sciences, China Agricultural University) for providing the ER-marker. This work was supported by the National Special Program for GMO Development of China (grant number 2008ZX08003-002).
References
- Haydon MJ, Cobbett CS. Transporters of ligands for essential metal ions in plants. New Phytol. 2007;174(3):499–506. doi: 10.1111/j.1469-8137.2007.02051.x. [DOI] [PubMed] [Google Scholar]
- Rhodes D, Klug A. Zinc fingers. Sci Am. 1993;268(2):56–59. doi: 10.1038/scientificamerican0293-56. 62-55. [DOI] [PubMed] [Google Scholar]
- Vallee BL, Auld DS. Zinc coordination, function, and structure of zinc enzymes and other proteins. Biochemistry. 1990;29(24):5647–5659. doi: 10.1021/bi00476a001. [DOI] [PubMed] [Google Scholar]
- Vallee BL, Falchuk KH. The biochemical basis of zinc physiology. Physiol Rev. 1993;73(1):79–118. doi: 10.2466/pr0.1993.73.1.79. [DOI] [PubMed] [Google Scholar]
- Cakmak I. Enrichment of cereal grains with zinc: agronomic or genetic biofortification? Plant Soil. 2008;302(1–2):1–17. [Google Scholar]
- Briat JF, Lebrun M. Plant responses to metal toxicity. C R Acad Sci III. 1999;322(1):43–54. doi: 10.1016/S0764-4469(99)80016-X. [DOI] [PubMed] [Google Scholar]
- Casterline JE, Allen LH, Ruel MT. Vitamin B-12 deficiency is very prevalent in lactating Guatemalan women and their infants at three months postpartum. J Nutr. 1997;127(10):1966–1972. doi: 10.1093/jn/127.10.1966. [DOI] [PubMed] [Google Scholar]
- Påhlsson A-M. Toxicity of heavy metals (Zn, Cu, Cd, Pb) to vascular plants. Water, Air, and Soil Pollution. 1989;47(3–4):287–319. [Google Scholar]
- Price AH, Hendry GAF. Iron-catalysed oxygen radical formation and its possible contribution to drought damage in nine native grasses and three cereals. Plant Cell Environ. 1991;14(5):477–484. doi: 10.1111/j.1365-3040.1991.tb01517.x. [DOI] [Google Scholar]
- Palmgren MG, Clemens S, Williams LE, Kramer U, Borg S, Schjorring JK, Sanders D. Zinc biofortification of cereals: problems and solutions. Trends Plant Sci. 2008;13(9):464–473. doi: 10.1016/j.tplants.2008.06.005. [DOI] [PubMed] [Google Scholar]
- Grotz N, Guerinot ML. Molecular aspects of Cu, Fe and Zn homeostasis in plants. Biochim Biophys Acta. 2006;1763(7):595–608. doi: 10.1016/j.bbamcr.2006.05.014. [DOI] [PubMed] [Google Scholar]
- Kambe T, Yamaguchi-Iwai Y, Sasaki R, Nagao M. Overview of mammalian zinc transporters. Cell Mol Life Sci. 2004;61(1):49–68. doi: 10.1007/s00018-003-3148-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Taylor KM, Morgan HE, Johnson A, Nicholson RI. Structure-function analysis of HKE4, a member of the new LIV-1 subfamily of zinc transporters. Biochem J. 2004;377(Pt 1):131–139. doi: 10.1042/BJ20031183. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Colangelo EP, Guerinot ML. Put the metal to the petal: metal uptake and transport throughout plants. Curr Opin Plant Biol. 2006;9(3):322–330. doi: 10.1016/j.pbi.2006.03.015. [DOI] [PubMed] [Google Scholar]
- Guerinot ML. The ZIP family of metal transporters. Biochim Biophys Acta. 2000;1465(1–2):190–198. doi: 10.1016/s0005-2736(00)00138-3. [DOI] [PubMed] [Google Scholar]
- Maser P, Thomine S, Schroeder JI, Ward JM, Hirschi K, Sze H, Talke IN, Amtmann A, Maathuis FJ, Sanders D. et al. Phylogenetic relationships within cation transporter families of Arabidopsis. Plant Physiol. 2001;126(4):1646–1667. doi: 10.1104/pp.126.4.1646. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Eide D, Broderius M, Fett J, Guerinot ML. A novel iron-regulated metal transporter from plants identified by functional expression in yeast. Proc Natl Acad Sci U S A. 1996;93(11):5624–5628. doi: 10.1073/pnas.93.11.5624. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Henriques R, Jasik J, Klein M, Martinoia E, Feller U, Schell J, Pais MS, Koncz C. Knock-out of Arabidopsis metal transporter gene IRT1 results in iron deficiency accompanied by cell differentiation defects. Plant Mol Biol. 2002;50(4–5):587–597. doi: 10.1023/a:1019942200164. [DOI] [PubMed] [Google Scholar]
- Varotto C, Maiwald D, Pesaresi P, Jahns P, Salamini F, Leister D. The metal ion transporter IRT1 is necessary for iron homeostasis and efficient photosynthesis in Arabidopsis thaliana. Plant J. 2002;31(5):589–599. doi: 10.1046/j.1365-313X.2002.01381.x. [DOI] [PubMed] [Google Scholar]
- Vert G, Grotz N, Dedaldechamp F, Gaymard F, Guerinot ML, Briat JF, Curie C. IRT1, an Arabidopsis transporter essential for iron uptake from the soil and for plant growth. Plant Cell. 2002;14(6):1223–1233. doi: 10.1105/tpc.001388. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nishida S, Tsuzuki C, Kato A, Aisu A, Yoshida J, Mizuno T. AtIRT1, the primary iron uptake transporter in the root, mediates excess nickel accumulation in Arabidopsis thaliana. Plant Cell Physiol. 2011;52(8):1433–1442. doi: 10.1093/pcp/pcr089. [DOI] [PubMed] [Google Scholar]
- Lin YF, Liang HM, Yang SY, Boch A, Clemens S, Chen CC, Wu JF, Huang JL, Yeh KC. Arabidopsis IRT3 is a zinc-regulated and plasma membrane localized zinc/iron transporter. New Phytol. 2009;182(2):392–404. doi: 10.1111/j.1469-8137.2009.02766.x. [DOI] [PubMed] [Google Scholar]
- Kramer U, Talke IN, Hanikenne M. Transition metal transport. FEBS Lett. 2007;581(12):2263–2272. doi: 10.1016/j.febslet.2007.04.010. [DOI] [PubMed] [Google Scholar]
- Bughio N, Yamaguchi H, Nishizawa NK, Nakanishi H, Mori S. Cloning an iron-regulated metal transporter from rice. J Exp Bot. 2002;53(374):1677–1682. doi: 10.1093/jxb/erf004. [DOI] [PubMed] [Google Scholar]
- Ishimaru Y, Suzuki M, Tsukamoto T, Suzuki K, Nakazono M, Kobayashi T, Wada Y, Watanabe S, Matsuhashi S, Takahashi M. et al. Rice plants take up iron as an Fe3+-phytosiderophore and as Fe2+ Plant J. 2006;45(3):335–346. doi: 10.1111/j.1365-313X.2005.02624.x. [DOI] [PubMed] [Google Scholar]
- Lee S, An G. Over-expression of OsIRT1 leads to increased iron and zinc accumulations in rice. Plant Cell Environ. 2009;32(4):408–416. doi: 10.1111/j.1365-3040.2009.01935.x. [DOI] [PubMed] [Google Scholar]
- Lee S, Jeong HJ, Kim SA, Lee J, Guerinot ML, An G. OsZIP5 is a plasma membrane zinc transporter in rice. Plant Mol Biol. 2010;73(4–5):507–517. doi: 10.1007/s11103-010-9637-0. [DOI] [PubMed] [Google Scholar]
- Ishimaru Y, Masuda H, Suzuki M, Bashir K, Takahashi M, Nakanishi H, Mori S, Nishizawa NK. Overexpression of the OsZIP4 zinc transporter confers disarrangement of zinc distribution in rice plants. J Exp Bot. 2007;58(11):2909–2915. doi: 10.1093/jxb/erm147. [DOI] [PubMed] [Google Scholar]
- Durmaz E, Coruh C, Dinler G, Grusak M, Peleg Z, Saranga Y, Fahima T, Yazici A, Ozturk L, Cakmak I. et al. Expression and cellular localization of ZIP1 transporter under zinc deficiency in wild emmer wheat. Plant Molecular Biology Reporter. 2011;29(3):582–596. doi: 10.1007/s11105-010-0264-3. [DOI] [Google Scholar]
- Moreau S, Thomson RM, Kaiser BN, Trevaskis B, Guerinot ML, Udvardi MK, Puppo A, Day DA. GmZIP1 encodes a symbiosis-specific zinc transporter in soybean. J Biol Chem. 2002;277(7):4738–4746. doi: 10.1074/jbc.M106754200. [DOI] [PubMed] [Google Scholar]
- Xu Y, Wang B, Yu J, Ao G, Zhao Q. Cloning and characterisation of ZmZLP1, a gene encoding an endoplasmic reticulum-localised zinc transporter in Zea mays. Functional Plant Biology. 2010;37(3):194–205. doi: 10.1071/FP09045. [DOI] [Google Scholar]
- Lee S, Kim SA, Lee J, Guerinot ML, An G. Zinc deficiency-inducible OsZIP8 encodes a plasma membrane-localized zinc transporter in rice. Mol Cells. 2010;29(6):551–558. doi: 10.1007/s10059-010-0069-0. [DOI] [PubMed] [Google Scholar]
- Vert G, Briat JF, Curie C. Arabidopsis IRT2 gene encodes a root-periphery iron transporter. Plant J. 2001;26(2):181–189. doi: 10.1046/j.1365-313x.2001.01018.x. [DOI] [PubMed] [Google Scholar]
- Milner MJ, Seamon J, Craft E, Kochian LV. Transport properties of members of the ZIP family in plants and their role in Zn and Mn homeostasis. J Exp Bot. 2013;64(1):369–381. doi: 10.1093/jxb/ers315. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ramesh SA, Shin R, Eide DJ, Schachtman DP. Differential metal selectivity and gene expression of two zinc transporters from rice. Plant Physiol. 2003;133(1):126–134. doi: 10.1104/pp.103.026815. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ishimaru Y, Suzuki M, Kobayashi T, Takahashi M, Nakanishi H, Mori S, Nishizawa NK. OsZIP4, a novel zinc-regulated zinc transporter in rice. J Exp Bot. 2005;56(422):3207–3214. doi: 10.1093/jxb/eri317. [DOI] [PubMed] [Google Scholar]
- Lopez-Millan AF, Ellis DR, Grusak MA. Identification and characterization of several new members of the ZIP family of metal ion transporters in Medicago truncatula. Plant Mol Biol. 2004;54(4):583–596. doi: 10.1023/B:PLAN.0000038271.96019.aa. [DOI] [PubMed] [Google Scholar]
- Stephens BW, Cook DR, Grusak MA. Characterization of zinc transport by divalent metal transporters of the ZIP family from the model legume Medicago truncatula. Biometals. 2011;24(1):51–58. doi: 10.1007/s10534-010-9373-6. [DOI] [PubMed] [Google Scholar]
- Ramesh SA, Choimes S, Schachtman DP. Over-expression of an Arabidopsis zinc transporter in hordeum vulgare increases short-term zinc uptake after zinc deprivation and seed zinc content. Plant Mol Biol. 2004;54(3):373–385. doi: 10.1023/B:PLAN.0000036370.70912.34. [DOI] [PubMed] [Google Scholar]
- Vert G, Barberon M, Zelazny E, Seguela M, Briat JF, Curie C. Arabidopsis IRT2 cooperates with the high-affinity iron uptake system to maintain iron homeostasis in root epidermal cells. Planta. 2009;229(6):1171–1179. doi: 10.1007/s00425-009-0904-8. [DOI] [PubMed] [Google Scholar]
- Yang X, Huang J, Jiang Y, Zhang HS. Cloning and functional identification of two members of the ZIP (Zrt, Irt-like protein) gene family in rice (Oryza sativa L.) Mol Biol Rep. 2009;36(2):281–287. doi: 10.1007/s11033-007-9177-0. [DOI] [PubMed] [Google Scholar]
- Gainza-Cortes F, Perez-Diaz R, Perez-Castro R, Tapia J, Casaretto JA, Gonzalez S, Pena-Cortes H, Ruiz-Lara S, Gonzalez E. Characterization of a putative grapevine Zn transporter, VvZIP3, suggests its involvement in early reproductive development in Vitis vinifera L. BMC Plant Biol. 2012;12:111. doi: 10.1186/1471-2229-12-111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pedas P, Schjoerring JK, Husted S. Identification and characterization of zinc-starvation-induced ZIP transporters from barley roots. Plant Physiol Biochem. 2009;47(5):377–383. doi: 10.1016/j.plaphy.2009.01.006. [DOI] [PubMed] [Google Scholar]
- Narayanan NN, Vasconcelos MW, Grusak MA. Expression profiling of Oryza sativa metal homeostasis genes in different rice cultivars using a cDNA macroarray. Plant Physiol Biochem. 2007;45(5):277–286. doi: 10.1016/j.plaphy.2007.03.021. [DOI] [PubMed] [Google Scholar]
- Sperotto RA, Boff T, Duarte GL, Santos LS, Grusak MA, Fett JP. Identification of putative target genes to manipulate Fe and Zn concentrations in rice grains. J Plant Physiol. 2010;167(17):1500–1506. doi: 10.1016/j.jplph.2010.05.003. [DOI] [PubMed] [Google Scholar]
- Lin YX, Jiang HY, Chu ZX, Tang XL, Zhu SW, Cheng BJ. Genome-wide identification, classification and analysis of heat shock transcription factor family in maize. BMC Genomics. 2011;12:76. doi: 10.1186/1471-2164-12-76. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xing H, Pudake RN, Guo G, Xing G, Hu Z, Zhang Y, Sun Q, Ni Z. Genome-wide identification and expression profiling of auxin response factor (ARF) gene family in maize. BMC Genomics. 2011;12:178. doi: 10.1186/1471-2164-12-178. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhou X, Li S, Zhao Q, Liu X, Zhang S, Sun C, Fan Y, Zhang C, Chen R. Genome-wide identification, classification and expression profiling of nicotianamine synthase (NAS) gene family in maize. BMC Genomics. 2013;14(1):238. doi: 10.1186/1471-2164-14-238. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Burleigh SH, Kristensen BK, Bechmann IE. A plasma membrane zinc transporter from Medicago truncatula is up-regulated in roots by Zn fertilization, yet down-regulated by arbuscular mycorrhizal colonization. Plant Mol Biol. 2003;52(5):1077–1088. doi: 10.1023/A:1025479701246. [DOI] [PubMed] [Google Scholar]
- Assuncao AG, Herrero E, Lin YF, Huettel B, Talukdar S, Smaczniak C, Immink RG, van Eldik M, Fiers M, Schat H. et al. Arabidopsis thaliana transcription factors bZIP19 and bZIP23 regulate the adaptation to zinc deficiency. Proc Natl Acad Sci U S A. 2010;107(22):10296–10301. doi: 10.1073/pnas.1004788107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG. The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res. 1997;25(24):4876–4882. doi: 10.1093/nar/25.24.4876. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Krogh A, Larsson B, von Heijne G, Sonnhammer EL. Predicting transmembrane protein topology with a hidden Markov model: application to complete genomes. J Mol Biol. 2001;305(3):567–580. doi: 10.1006/jmbi.2000.4315. [DOI] [PubMed] [Google Scholar]
- Tamura K, Dudley J, Nei M, Kumar S. MEGA4: Molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol Biol Evol. 2007;24(8):1596–1599. doi: 10.1093/molbev/msm092. [DOI] [PubMed] [Google Scholar]
- Han MJ, Jung KH, Yi G, Lee DY, An G. Rice Immature Pollen 1 (RIP1) is a regulator of late pollen development. Plant Cell Physiol. 2006;47(11):1457–1472. doi: 10.1093/pcp/pcl013. [DOI] [PubMed] [Google Scholar]
- Yoo S-D, Cho Y-H, Sheen J. Arabidopsis mesophyll protoplasts: a versatile cell system for transient gene expression analysis. Nat Protocols. 2007;2(7):1565–1572. doi: 10.1038/nprot.2007.199. [DOI] [PubMed] [Google Scholar]
- Nelson BK, Cai X, Nebenfuhr A. A multicolored set of in vivo organelle markers for co-localization studies in Arabidopsis and other plants. Plant J. 2007;51(6):1126–1136. doi: 10.1111/j.1365-313X.2007.03212.x. [DOI] [PubMed] [Google Scholar]
- Minet M, Dufour ME, Lacroute F. Complementation of Saccharomyces cerevisiae auxotrophic mutants by Arabidopsis thaliana cDNAs. Plant J. 1992;2(3):417–422. doi: 10.1111/j.1365-313x.1992.00417.x. [DOI] [PubMed] [Google Scholar]
- Zhao H, Eide D. The yeast ZRT1 gene encodes the zinc transporter protein of a high-affinity uptake system induced by zinc limitation. Proc Natl Acad Sci U S A. 1996;93(6):2454–2458. doi: 10.1073/pnas.93.6.2454. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhao H, Eide D. The ZRT2 gene encodes the low affinity zinc transporter in Saccharomyces cerevisiae. J Biol Chem. 1996;271(38):23203–23210. doi: 10.1074/jbc.271.38.23203. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Amino acid alignment of ZmIRT1 with other IRT proteins. Sequences were aligned using Clustal X Version 2.0 and the identical or similar amino acids are shaded by BOXSHADE (http://www.ch.embnet.org/software/BOX_form.html). The residue D100 in ZmIRT1 is marked with an asterisk.
Expression patterns of the nine ZmZIP genes in maize seedlings. The shoots (S) and roots (R) of two weeks old hydroponically cultured maize seedlings in Hoagland nutrient solution were harvested, respectively. Relative mRNA abundance of each gene was normalized with ZmActin1 gene. Data from real-time RT-PCR experiments were analyzed according to the 2-∆∆Ct method. The error bars indicate standard deviations.
Expression patterns of the nine ZmZIP genes in maize under Cu-, Mn- deficiency. Two-week old shoots (S) and roots (R) of maize seedlings, under Cu-, Mn-deficiency treated (Cu-), (Mn-) were harvested respectively at 0 h, 6 h, 12 h, 24 h, 48 h and 96 h. Relative mRNA abundance of each gene was normalized with ZmActin1 gene. Data from real-time RT-PCR experiments were analyzed according to the 2-∆∆Ct method. The error bars indicate standard deviations.
Subcellular localization of ZmZIPs in onion epidermal cells. Full-length coding regions without stop codon of the ZmZIP genes were cloned into the pRTL2GFP vector and the resulting plasmid was transiently transformed into onion epidermal cells by bombardment. The fluorescence was observed using a confocal laser scanning microscopy. GFP was imaged using 488 nm excitation and a 500-530 nm bandpass emission filter. The scale bar represents 100 μm.
Subcellular localization of ZmZIP-GFP fusion proteins in plasmolyzed onion epidermal cells. The ZmZIP-GFP fusion proteins were transiently expressed in onion epidermal cells by bombardment, and a set of representative images are shown. The plasmolysis was performed for 15 min in 30% sucrose. The Z-stack of optical sections and single optical slice of GFP fluorescence are shown. The cytoplasm localization of GFP is used as a control. GFP was imaged using 488 nm excitation and a 500-530 nm bandpass emission filter. The scale bar represents 100 μm.
Functional complementation of the Fe transportation yeast mutant by ZmZIPs under different pH values. The Fe transportation mutant fet3fet4 were transformed with the expression vector pFL61 carrying ZmZIP1-ZmZIP8, ZmIRT1 and the functional characterized ZIP genes, OsZIP5, OsZIP8, and OsIRT1. The transformed yeast cells were grown under different pH conditions (A) pH 5.5 and (B) pH 5.8. Cell concentration was adjusted to OD600 = 1 and serial dilutions (1.0, 0.1, 0.01 and 0.001) were made. For assay, 5-μL of each dilution was spotted on plates and grown for 6 days at 30°C.
Putative cis-elements in the 2-Kb upstream promoter region of translation start site in ZmZIP genes.
Primers used in gene cloning, vector construction and real-time RT-PCR analysis.