Abstract
Polypyrimidine Tract Binding Protein (PTB) is an intensely studied RNA binding protein involved in several post-transcriptional regulatory events of gene expression. Initially described as a pre-mRNA splicing regulator, PTB is now widely accepted as a multifunctional protein shuttling between nucleus and cytoplasm. Accordingly, PTB can interact with selected RNA targets, structural elements and proteins. There is increasing evidence that PTB and its paralog PTBP2 play a major role as repressors of alternatively spliced exons, whose transcription is tissue-regulated. In addition to alternative splicing, PTB is involved in almost all steps of mRNA metabolism, including polyadenylation, mRNA stability and initiation of protein translation. Furthermore, it is well established that PTB recruitment in internal ribosome entry site (IRES) activates the translation of picornaviral and cellular proteins. Detailed studies of the structural properties of PTB have contributed to our understanding of the mechanism of RNA binding by RNA Recognition Motif (RRM) domains. In the present review, we will describe the structural properties of PTB, its paralogs and co-factors, the role in post-transcriptional regulation and actions in cell differentiation and pathogenesis. Defining the multifunctional roles of PTB will contribute to the understanding of key regulatory events in gene expression.
Keywords: PTB, raver1, alternative splicing, IRES, RRM, RNA binding proteins, ribonucleoproteins, hnRNP, mRNA stability, cell differentiation
1. Introduction
The control of gene expression is a fundamental process required by all living cells. In eukaryotic cells, multiple and coordinated processes contribute to the fine-tuning of the transcription and translation of selected genes in the nucleus and the cytoplasm. The initial steps take place in the nucleus where promoter recognition, mediated by transcription factors, requires chromatin and nucleosome modifications driven by epigenetic events, and allows the regulation of the RNA polymerase transcriptional activity. When pre-mRNAs emerge from the transcription sites, they are associated with trans-acting proteins and RNAs, to form the RNA-protein complexes also referred as ribonucleoprotein (RNP) complexes.
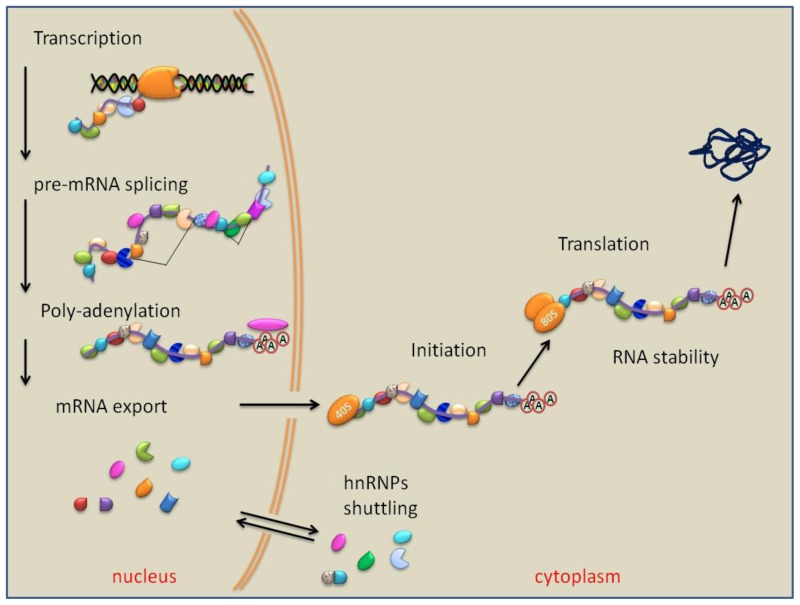
The RNPs assembled on each mRNA diversify their composition by recruiting and reassorting selected components. During mRNA maturation, the RNPs are further recombined and modelled, regulating the subsequent stages of RNA biogenesis, which include the 5′-end capping, splice site recognition, RNA editing, 3′-end cleavage, polyadenylation, nuclear pores export, sub-cellular localization, ending to protein translation and RNA degradation [1–3] (Figure 1). Alternative splicing (AS) of the precursor mRNA (pre-mRNA) occurs in >90% of human genes and represents the principal mechanism responsible for the complexity of genome expression and protein diversity [4–6].
Figure 1.
RNA binding proteins are involved in all steps of RNA biogenesis. Starting from pre-mRNA transcript synthesis, a variety of different RNA binding proteins are associated within the heterogeneous ribonucleoprotein complexes, participating in the processes which result in the formation of mature mRNA, including pre-mRNA splicing and addition of 3′ poly(A) tails in the nucleus and mRNA export through the nuclear pores. In the cytoplasm, they participate in localization, initiation of translation, and mRNA stability.
Two major RNA binding protein families, the heterogenous nuclear ribonucleoproteins (hnRNPs) and the serine/arginine (SR)-rich proteins (SR proteins) contribute to AS events within RNP complexes [7–10]. Both families of RNA binding proteins are associated with nascent transcripts and influence most steps of mRNA biogenesis by synergic or antagonist effects on similar processes [11]. Typical SR proteins bind to enhancers of exon splicing, activating constitutive and alternative splice sites by recruiting spliceosomal components to the pre-mRNA [12]. Most of these splicing factors are ubiquitously expressed, but their relative abundance or activity may vary in the tissues and can be modulated by the induction of specific cellular signaling pathways [13–15]. HnRNPs, instead, influence constitutive and alternative splicing events by preferentially binding to splicing silencers [12]. The hnRNPs form ribonucleoprotein complexes that are distinct from small nuclear ribonucleoprotein particles (snRNPs) of the spliceosomal complexes. The combination of different types of hnRNPs in the ribonucleoprotein complexes bound to the pre-mRNA may, in turn, antagonize or favor splice site selection. Their action supports the structural modeling of pre-mRNA, bringing together regions that are distant hundreds of nucleotides, and assists in the splice site recognition [16].
One of the most investigated exon recognition repressors, belonging to the hnRNPs family, is the polypyrimidine tract-binding protein (PTB) that binds to intron pyrimidine-rich elements and mediates tissue-specific regulation of exon splicing [17,18].
PTB is recruited not only in AS, but in additional steps of RNA processes eliciting various biological effects, including transport, stabilization, and mRNA translation. The cell type and the position of the specific RNA binding sites are determinant factors for the variety of PTB effects on RNA destiny [19]. Starting from its first description in 1992 [20,21], an increasing number of studies have been published, describing structural and functional properties of PTB, which thus became one of the most cited proteins in studies focusing on nuclear localization, alternative splicing, and RNA binding proteins.
This review will focus on the established roles of PTB in splicing regulation and protein translation highlighting the most recent contributions to the understanding of its multi-functionality.
2. PTB Gene Expression
PTB (accredited gene name PTBP1), also known as hnRNP I, is a shuttling protein that moves rapidly between nucleus and cytoplasm, with a predominant nuclear localization [22]. The 57 kDa protein is organized into four RNA recognition motifs (RRM1–4), a bipartite nuclear localization determinant (NLD), which we have demonstrated to be recognized by importin α [23,24], and a nuclear export signal at the N-terminus of the protein [25]. The PTB gene maps on chromosome 19 at position p13.1, and is organized in 15 exons that are subjected to alternative splicing, producing at least four isoforms [26]. PTB1 is the first described isoform and consists of 521 amino acids containing all four RRMs. The alternatively spliced isoforms, PTB3 and PTB4, contain additional 19 or 26 amino acids between the RRM2 and RRM3 domains derived from exon 9 inclusion, which has two alternative 3′ splice sites. The alternative splicing of exons 2–10 produces the fourth isoform, named PTB2, which is translated in a protein lacking the RRM1 and RRM2. PTB is expressed in all human tissues in different isoforms and levels according to the specific cell type [26]. In spite of their high levels of homology, the different PTB proteins show distinct activities in splicing and in IRES-mediated initiation of translation [19].
3. Unique Properties of PTB RNA Recognition Motifs
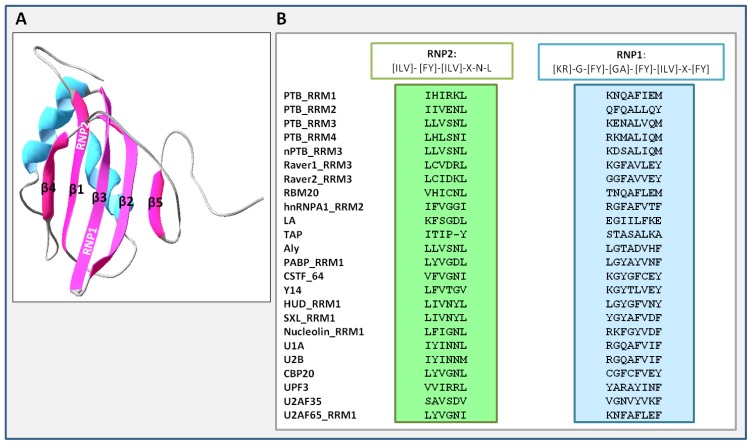
The members of the hnRNP protein family were initially discovered in the late 1980s and were attributed to the same protein family by the presence of the RRM, which is the most conserved domain among the RNA binding domains (RBDs). The RRM is a functional domain broadly distributed in hundreds of proteins encoded by eukaryotic genomes. Compared to other known RBDs, such as the K-homology domain, the RGG (Arg-Gly-Gly) box, the Sm domain, the DEAD/DEAH box, the Pumilio/FBF (PUF domain) and the Piwi/Argonaute/Zwille (PAZ), for review see [27], RRM is structurally quite different. RRMs containing genes represent about 0.5%–1% of the human genes. The RRM is often present in more than one copy and can be associated with additional protein domains, e.g. zinc finger of the CCCH type, the polyadenylate binding protein C-terminal domain (PABP), and the WW domain [28]. The structural and biochemical properties of the RRMs have contributed to the interpretation of RNA recognition capability, but also to the identification of protein-protein interactions between splicing factors [29,30]. Canonical RRMs consist of approximately 90 amino acids, containing two conserved core sequences, named RNP2 and RNP1, directly involved in the RNA interaction (Figure 2A) [28]. In a typical RRM, the peptide folds with a β1α1β2β3α2β4 topology leading to a structural model of four β-sheets packed against two α-helices (Figure 2A). Both RRM1 and RRM4 domains of PTB fold in the canonical βαββαβ RRM topology, whereas RRM2 and RRM3 contain an additional fifth β-strand (Figure 2A) that extends the canonical RNA binding surface [31,32] and contributes to sequence specific recognition.
Figure 2.
PTB RRM3 structure. (A) The PTB RRM3 structure was obtained with Swiss-PdbViewer (http://www.expasy.org/spdbv/); (B) Alignments of ribonucleoprotein (RNP) 2 and RNP1 sequences present in the PTB RRM3 with RRM containing proteins. The alignment was generated by ClustalW and manually optimized. PTB, polypryrimidine tract binding protein; nPTB, neuronal PTB; hnRNPA1, heterogeneous nuclear ribonucleoprotein type A1; LA, La protein; TAP, mRNA export factor Tap; PABP, Poly(A)-binding protein; CSTF64, cleavage stimulation factor 64; Y14, Y14 Magoh complex; HUD, Hu protein; SXL, Sex-Lethal; U1A, U2B, U1 snRNP proteins; CBP20, cap-binding protein 20; UPF3, up-frameshift protein 3; U2AF65, U2 snRNP auxiliary factor; RBM20, RNA binding motif 20.
RRM3 can compete with the pre-mRNA splicing factor U2AF65 (U2 auxiliary factor) at polypyrimidine stretches located at the 3′ splice site of target introns [33]. In addition to the presence of a fifth β-strand in RRM2 and RRM3, all four RRMs present in PTB show divergences from the conserved RRM sequences, represented by the absence of the aromatic side chains typically found in RNPs (Figure 2B).
The peculiarity of the structural features of the RRMs allows PTB to be involved in the splice site choice in alternative splicing events, but also to be recruited in several additional events in post-transcriptional regulation, including RNA stability and translation. The consensus sequences for RNA-PTB binding are represented by sequences containing 15–25 pyrimidines, with a preference for pyrimidine tracts containing UCUU, CUCUCU [34–36]. The structures of PTB complexed with RNA have been intensively investigated by NMR spectroscopy [37]. The solved three dimensional solution structures demonstrate that each RRM of PTB may bind one RNA molecule containing the CUCUCU sequence, and all RRMs bind polypyrimidine tracts by the contribution of their β-sheets. While RRMs 1 and 2 remain independent in solution, RRMs 3 and 4 may interact with each other producing a single globular protein moiety. RRM3 and 4 bind to pyrimidine tracts separated by intervening sequences of 15 nucleotides or longer, inducing RNA looping [37–39].
RRM1 and RRM2 are separated by flexible linkers from the rest of the protein, which allow them to adopt different conformations and to favour contact with specifically structured RNA. A novel interaction with U1 snRNA has been recently attributed to RRM1 and RRM2 and this represents the first demonstration of a direct interaction between PTB and the spliceosome [40]. Detailed analyses described by Clerte and Hall [41] and Kafasla et al. [42], have highlighted a modular function of PTB-RNA binding in which RRM1 and RRM2 preferentially bind to short tracts of U/C located in loop structured RNA sequences, whereas the interacting RRM3 and RRM4 preferentially bind to longer flexible RNA sequences.
In addition to RNA, RRM2 can interact with the amino acid sequence (S/G)(I/L)LGXXP present in the co-repressor proteins Raver1 and Raver2 [43–46], enabling PTB to recruit other regulatory proteins in AS events.
4. PTB Paralogs and Co-Factors
Two PTB paralogs are generally expressed in mammalian tissues: the nPTB/brPTB/PTBP2 which is expressed at high levels in adult brain, muscle and testis [47,48] and ROD1 (PTBP3) which is expressed preferentially in haematopoietic cells [49]. ROD1 has been demonstrated to act prevalently in the nonsense-mediated mRNA decay (NMD) of aberrant spliced isoforms through the interaction with key factors of the RNA surveillance pathway, such as the UPF1 factor, with which it shares the endogenous targets [50]. Rodents, but not humans, express an additional paralog, named smPTB, which is highly expressed in smooth muscle cells [51].
PTB and its paralogs share more than 70% amino acid sequence identity organized in the four RRM-type domains. The most extensively studied is the nPTB, initially described as a neuron specific splicing factor [52]. Further studies have shown that nPTB can be alternatively spliced, leading to different isoforms expressed in a tissue specific manner [48,53]. The expression of PTB and its paralogs is tightly regulated through alternative splicing events. PTB regulates its own expression by repressing the splicing of exon 11, causing NMD of the resultant mRNA [54], but it also promotes the exon 10 exclusion from nPTB transcripts, leading to NMD of nPTB transcripts. Both PTB and nPTB may promote the non-productive splicing of ROD1 [55]. In neuronal cells, nPTB acts by autoregulating its own exon 10 inclusion, leading to an increased expression level of the nPTB protein [56].
In T lymphocytes, both PTB and the alternatively spliced isoform lacking RRM1and RRM2, called PTB-T, participate in humoral and cellular immunity by targeting the CD154 cytokine mRNA. The activity is elicited in the cytoplasm by binding to pyrimidine cis-acting sequences at the 3′ untranslated region (3′UTR) of CD154. In activated T lymphocytes, the alternatively spliced isoforms of PTB expressed at different amounts act as trans-factors competing with each other for binding to the CD154 3′UTR, and modulating CD154 mRNA stability [57]. These data confirm that, although PTB is ubiquitously expressed, its activity is tissue specific and is mediated by the relative amount of alternatively spliced isoforms, paralogs, or competing factors for binding to polypyrimidine tracts.
As a splicing repressor factor, PTB may require the binding to selected factors that act as co-repressors. Raver1 and Raver2 are two well characterized PTB co-repressors [58–60]. Raver1 has been demonstrated to be essential for effective repressor function of PTB in the splicing repression of α-tropomyosin exon 3 [59,61]. A cooperative action in regulating adjacent exons has been confirmed by the structural model of PTB RRM2 bound to the Raver1 peptide [43]. The human Raver1 gene is ubiquitously expressed in tissues at different levels and encodes a 748 amino acid protein containing three RRMs, two nuclear localization signals (NLSs), one nuclear export sequence and a proline-rich region at its C-terminal region [58,62,63]. Similarly to PTB, Raver1 is a shuttle-protein. By analizing the interaction specificity, Raver1 has been demonstrated to interact with the cytoskeletal proteins α-actinin and metavinculin in the cytoplasm [58]. During myotube differentiation, Raver1 migrates to the cytoplasm, co-localizes with the microfilament attachment sites [58,63] and may be involved in the formation of adhesion complexes and intercalated discs in muscle cells [64]. In a raver1 null mutant mouse, the loss of Raver1 is associated with a reduction of synaptic plasticity [65]. A recent report has shown that Raver1 may act also as a co-activator of the RIG-I-like receptor MDA5 (melanoma differentiation-associated gene 5), which is involved in the innate antiviral response [66].
Raver2, the additional PTB co-factor is a 625 aa protein of 72 kDa with a similar overall organization of the functional domains when compared to Raver1. Raver2 expression is restricted to differentiated neurons and glia cells. We have recently contributed to the characterization of Raver1 and Raver2 promoters and to the identification of cis-acting signals [62,67]. In contrast to Raver1, which binds to cytosolic proteins, so far there is no evidence of a Raver2 interaction with cytoplasmic proteins. PTB interacts with Raver2 by recognizing the SLLGEPP peptide motif, which is well conserved in both Raver1 and Raver2 [45].
5. PTB Role in Pre-mRNA Splicing
PTB is implicated in the control of alternative exon selection during mRNA processing of many different transcribed genes, including its own pre-mRNA. The exon typologies include short alternative exons named “cassette exons”, “mutually exclusive exons” or alternative 3′ terminal exons [17,54,61,68] (summarized in Table 1). The polypyrimidine stretches recognized by PTB are frequently clustered in the intron sequences which surround the regulated exon, but they may also be present in the exon sequences. The most investigated mechanism of PTB action is related to its role in repressing the exon inclusion. To summarize the results derived from these studies, four different structural models may represent the dynamic of the different types of PTB splicing repression activity (Figure 3).
Table 1.
Exons regulated by PTB.
| Gene | Regulated Exon | Exon typology | Position of PTB-cis-acting elements | Ref. |
|---|---|---|---|---|
| α-actinin | SM | Mutually exclusive | Upstream | [69,70] |
| α-actinin | NM | Mutually exclusive | Flanking the branch-point | [69,70] |
| α-tropomyosin | 3 | Cassette | Up- and downstream | [71] |
| ATP syntetase γ-subunit | 9 | Cassette | Upstream | [72] |
| β-tropomyosin | 7 | Mutually exclusive | Upstream | [73] |
| c-src tyrosine kinase | N1 | Cassette | Up- and downstream | [74,75] |
| Calcitonin | 4 | Cassette | Downstream | [76] |
| Caspase-2 | 9 | Cassette | Downstream | [77] |
| Cardiac troponin T | 5 | Cassette | Up- and downstream | [78] |
| Clathrin light chain B | EN | Cassette | Upstream | [79] |
| FGF-R1 | α | Cassette | Up- and downstream | [80] |
| FGF-R2 | IIIb | Mutually exclusive | Downstream | [81] |
| GABAAγ2 | 2 | Cassette | Upstream | [79,82] |
| IgM | M1 M2 | Cassette | Exonic | [83] |
| NMDA receptor 1 | 5 | Cassette | Upstream | [79] |
| MEF2 | β | Cassette | Up- and downstream | [84] |
| CaV1.2 calcium channel | 8a-8 | Mutually exclusive | Upstream | [85] |
| PSD-95 | 18 | Cassette | Upstream | [86] |
| PTB | 11 | Cassette | Exonic | [54] |
Figure 3.
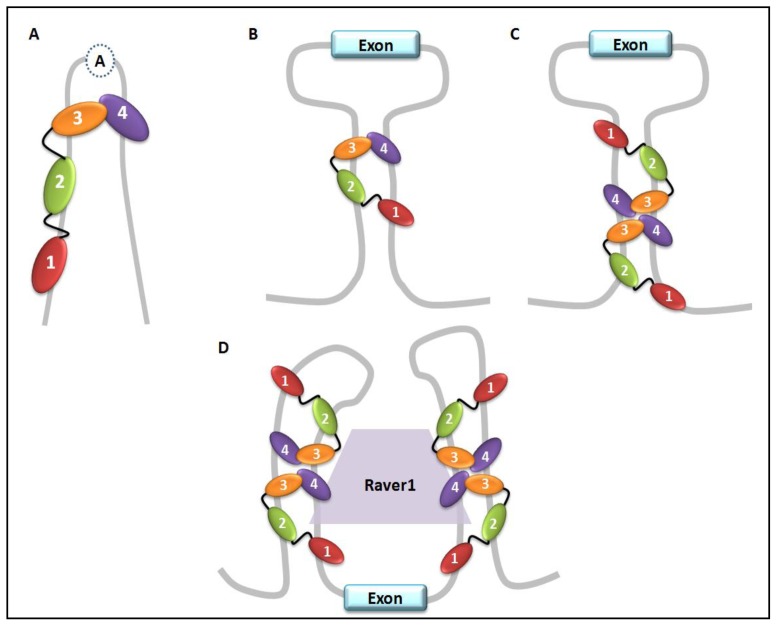
Models of RNA-PTB interaction in exon splicing repression. (A) Repression by PTB looping out a branch-point adenosine; (B) PTB monomer looping-out an exon; (C) Multiple PTB molecules binding cooperatively around an alternative exon; (D) PTB cooperates with the co-factor Raver1 to loop out an alternative exon by binding to distant intronic pyrimidine tracts.
These models describe a looping-out RNA mechanism in which PTB binds distant pyrimidine tracts, resulting in a new position in which the polypyrimidine tracts are close to each other [17,68]. All models require multiple PTB binding sites positioned in the introns. The polypyrimide sequences may be at the branch-point adenosine or at intron sequences, both upstream or downstream to the regulated exon (Figure 3A,B). A slightly different RNA loop model requires the cooperative multimerization of PTB (Figure 3C) that involves RRM3 and RRM4 binding. The repression may occur by preventing the binding of spliceosomal components. These models usually involve the dimerization of PTB and the competition with the splicing factor U2AF65 for binding to polypyrimidine tracts. However, PTB does not always dimerize to repress splicing, as shown for the γ-aminobutyric acid gamma2 (GABAA-γ2) exon 9 regulation, in which a single PTB molecule is required to loop out a branch-point adenosine [87]. In the fourth model, the repression of exon inclusion derives by PTB interaction with sequences upstream and downstream of the regulated exon, building a scaffold structure that may recruit splicing auxiliary elements, such as Raver1 co-factor (Figure 3D).
Additional factors, other than U2AF65, may antagonize the PTB repressive role such as the ETR3-like factor (CELF) proteins CUG-BP, RBM4, TIA-1, Nova, and Fox-1 [68]. Recently, the muscleblind-like (MBNL1) protein that acts as a direct RNA-binding regulator of alternative splicing during development of striate muscle has been demonstrated to directly interact with PTB, promoting the skipping of α-tropomyosin exon 3 in smooth muscle cells. In this model, MBNL1 binding to RNA promotes the interaction with PTB, and consecutively, it induces an RNA conformational change [88].
Microarray-based genome wide approaches and CLIP-seq methods have shown that PTB not only acts as a splicing repressor, but can also enhance the inclusion of exons [89–92]. A global representation of the putative RNA targets containing the cis-acting splicing regulatory elements recognized by PTB has been derived by high-density oligonucleotide splice-sensitive microarray analyses in HeLa cells, where PTB was knocked down [93]. The results showed that the PTB-regulated splicing events were represented by both exon skipping and exon inclusion, and the majority of the events, almost 70%, were PTB-repressed exons’ inclusion. The exon inclusion determined by PTB may be attributable to the position of the PTB-binding motifs, upstream or downstream to the regulated exon [93]. Two different models have been suggested to describe the exon inclusion. One model proposes that PTB inclusion is activated through the binding to polypyrimidine stretches positioned exclusively downstream to the exon to be included [93]. The alternative model suggests that inclusion derives from binding to sequences near the flanking constitutive splice sites of the exon, competing with and antagonizing the action of different splicing repressors [92,94]. The data supporting the different PTB models of action converge on the common interpretation that PTB regulated splicing activities are consequences of a fine-tuning of the dynamic spatial, stoichiometrical PTB binding to RNA polypyrimidine motifs and of protein-protein interactions [95]. Although the mechanism of PTB activity is not completely understood, all models confirm the described motif recognition by PTB and its specific mRNA targets. Most of them have been utilized as validation targets for biophysical models to identify splicing regulatory elements and their interactions [86,96,97].
6. PTB Role in Internal Ribosome Entry Site (IRES)-Mediated Translation Initiation
PTB has the property to shuttle between nucleus and cytoplasm. Whereas in the nucleus PTB acts as an alternative splicing factor, in the cytoplasm it is involved in post-transcriptional regulation processes that require cap-independent translational controls, RNA localization and stability.
PTB participation in internal ribosome entry site (IRES)-mediated translation initiation of both cellular and viral mRNAs is extensively documented. IRESs require the formation of specific RNA structural conformations at the 5′UTR of the mRNA that help the recruitment of the translational machinery. Additional cellular factors, known as initiation of translation accessory factors (ITAFs), may bind the IRESs and lead to a more efficient translation initiation [98]. PTB has been frequently described as ITAF of IRES-mediated translation in human viruses including hepatitis viruses, several picornaviruses (alphaviruses and cardioviruses), flaviviruses such as hepatitis C (HCV), and noroviruses [99–101]. PTB and its paralog nPTB are key factors to allow the functional conformation of viral IRESs sequences [102,103], acting prevalently as mRNA chaperones [104]. In poliovirus infected cells, PTB is extensively re-localized to the cytoplasm, as a consequence of the proteolysis of the nuclear pore complex caused by the poliovirus.
PTB is a determinant of tissue and host tropism for rhinovirus and poliovirus [105–107]. PTB not only is involved in viral RNA translation but also in viral replication, as demonstrated for HCV [108] and dengue viruses (DENV) [109,110]. Interestingly, PTB may be cleaved during viral infection as a consequence of the shut off mechanism of host cell protein synthesis [105,111]. PTB plays the role of ITAF not exclusively in viral infected cells, but also in the translation of several host cellular factors, as listed in Table 2.
Table 2.
IRESs interacting with PTB.
| mRNA | PTB effect in facilitating translational initiation | Ref. |
|---|---|---|
| HAV | + | [112] |
| EMCV | + | [42] |
| Poliovirus | + | [113] |
| HVC | + | [108] |
| Norovirus | + | [114] |
| DENV | + | [115] |
| FMDV | + | [116] |
| CVB3 (coxsackievirus B3) | + | [100] |
| CDK11 (p58) | + | [117] |
| EGR2 | + | [118] |
| INSULIN | + | [103] |
| p53 | + | [119] |
| Cat-1 | + | [120] |
| APAF-1 | + | [102] |
| HIF-1alpha | + | [121] |
| p27Kip1 | + | [122] |
| IRF2 | + | [123] |
| Rev-erb α | + | [124] |
| c-myc | + | [125] |
| VEGF | + | [126,127] |
| IGFR1 | n.d. | [128] |
| IR (insulin receptor) | + | [103] |
| UNR | − | [129] |
The mouse homologue of PTB is involved in the regulation of early mouse development through IRES dependent translation of CDK11 (p58) protein kinase and modulates the cell cycle in mouse embryonic stem cells [117]. In the context of inflammatory responsive gene expression regulation, PTB together with additional ITAFs targets the early growth response 2 (egr2) 5′UTR, enhancing IRES-dependent translation under proinflammatory conditions [118]. It has been demonstrated in vitro that PTB binds specifically to the insulin mRNA 5′UTR, when cap-independent insulin biosynthesis is produced in response to nitrosative stress [130]. PTB positively regulates the IRES activities of p53 isoforms, relocating them from the nucleus to the cytoplasm during stress conditions [119] and regulates the IRES activity of Cat-1 (cationic amino acid transporter) during nutritional stress [120].
As a chaperone, PTB influences apoptotic peptidase activating factor 1 (Apaf1) translation by recognizing the Apaf1-IRES structure [102] and positively regulates the translation of hypoxia-inducible factor 1 alpha (HIF-1 alpha) and cyclin-dependent kinase inhibitor 1B (p27Kip1) [121,122]. For most of the mRNAs, PTB positively regulates the IRES-mediated translation [131], but not always, as in the case of PTB effect on UNR (upstream N-ras) translation regulatory factor [129]. Cytoplasmic levels of PTB are increased by apoptosis or in response to toxic drugs, leading to modifications in the activity of selected IRESs containing transcripts [132,133].
7. PTB Role in RNA Polyadenylation, Transport and Stability
PTB has been found to be associated with pyrimidine-tracts present at 5′ and/or 3′UTRs of mRNAs. By binding to UTR sequences, PTB may be involved in the 3′-end processing, localization, and stability of several mRNAs [132,134–138]. In the neurite growth process, the export of PTB from the nucleus to neurite growth terminals has been associated with the localization of α-actin mRNA bound to PTB at the neurite terminals [136]. A similar role in facilitating mRNA export has been proposed for PTB as a co-factor in nuclear export of hepatitis B virus RNA by recognition of a post-transcriptional regulatory element [139]. Additional studies have highlighted the cytoplasmic role of PTB in the regulation of mRNA stability. PTB binding to pyrimidine-rich sequences downstream of the termination codon and upstream to the polyadenylation signal in preproinsulin mRNA determines increased preproinsulin mRNA stability and levels [140]. PTB promotes the stabilization of additional insulin granule components, such as islet cell autoantigen (ICA)512, prohormone convertases 1/3 (PC1/3), and 2 (PC2) mRNAs [141]. A similar effect of PTB on mRNA stability has been also demonstrated for vascular endothelial growth factor, HIF-1α and inducible NO synthase transcripts [126,135,142–144]. Stability of mRNA in activated T and B cells is regulated by a pathway that also involves PTB, as demonstrated for CD40 ligand and Rab8A [145–148]. PTB has also been found to be associated with the 3′UTR of the ATP synthase β-subunit, where it enhances translation in a cap-independent manner [149]. 5′-end processing efficiency is enhanced in the pre-mRNA of complement 2 (C2), protrombin (F2) and cyclooxygenase-2 (COX-2) [150–152], whereas PTB binding to the GU-rich sequences downstream to the polyadenylation signal, decreased the efficiency of 3′-end cleavage of the human α-globin and β-globin pre-mRNAs [134,137]. An interesting PTB role derives from the mouse model of circadian mRNA oscillation, in which PTB mediates the mRNA decay of expressed mper2, a circadian core clock gene, by binding to the 3′UTR CU-rich sequences [153].
An additional PTB functional role that deserves to be highlighted derives from the studies on Drosophila and Xenopus development. These studies demonstrate that PTB plays a role in the remodeling of RNP, promoting RNA localization and translation regulation of critical determinants of the embryo development. In the cytoplasm, Drosophila PTB, known as the product of the hephaestus locus, binds to multiple sites in the oskar 3′UTR leading to mRNA oligomerization that causes oskar translation repression. The model proposed in this study is that PTB may promote the formation of densely packed RNP particles with oskar RNAs, thereby preventing access for the translation machinery [154]. A similar role in RNP remodeling has been attributed to the Xenopus PTB ortholog. In Xenopus oocytes, PTB is required for transport and localization of the Vg1 RNA, a critical factor for the correct patterning of the embryo. Lewis et al. [155] have demonstrated that PTB is associated with RNP complexes including Vg1 RNA in the nucleus and in the cytoplasm. During cytoplasmic translocation PTB promotes the remodeling of the Vg1 RNP by modulating protein-protein interaction within the Vg1 RNP.
8. PTB Role in Cell Type Differentiation
Based on the extensive literature supporting the relevance of PTB in multiple biological processes, including development, as mentioned in section seven for germ cell differentiation in Drosophila and in development of Xenopus [154,155], it is not surprising to find evidence for an essential role of PTB also in mammalian development. In fact, in ptb knockout mice the absence of PTB results in embryonic lethality and severe defects in proliferation and differentiation of embryonic stem cells [156]. As well, a key role of PTB in cell differentiation is revealed by recent studies on muscle and neuronal cell differentiation [86,157]. Wide gene expression analyses demonstrate that PTB and nPTB expression is associated with repression of various neuronal and striate muscle specific exons in genes encoding cytoskeletal components and proteins involved in cell signalling [55,92,93]. Both PTB and nPTB protein levels decrease during myogenesis, affecting exon splicing of several transcripts regulated during myoblast differentiation [89]. Splicing-sensitive microarray analyses have demonstrated a combinatorial control of the myogenic splicing program by PTB and additional proteins that may compete for RNA binding. This is the case for the Quaking (QK) protein whose expression is raised during differentiation of myoblasts to myotubes, whereas PTB levels are reduced [158]. An estimation of the exons regulated by both QK and PTB, indicates that nearly 39% of the QK splicing network is also controlled by PTB [156]. An interesting model of PTB action in competition with an additional splicing regulatory factor during muscle cell differentiation, has been proposed by Lin and Tarn [159,160]. The authors demonstrate that the RNA binding motif 4 protein (RBM4) competes for common targets with PTB and nPTB, antagonizing their effects in muscle cell-specific alternative splicing. RBM4 contributes, at least in part, to the reduction of PTB and nPTB levels during the C2C12 cells differentiation, involving an AS-NMD pathway by activating exon skipping of PTB/nPTB trancripts. PTB contributes to post-transcriptional regulation of gene expression also during cardiomyocyte differentiation. As well as in myoblast differentiation, PTB expression is reduced in heart development. In differentiating cardiomyocytes, PTB regulates the expression of apoptotic genes [161]. Ye et al. [84] have demonstrated the direct involvement of PTB in the AS regulation of transcription factor myocyte enhancer-factor 2 (MEF2) and the cardiac structural proteins tropomyosin 1 (TPM1) and tropomyosin 2 (TPM2) during cardiomyocyte differentiation. In vivo and in vitro results demonstrate that the reduced expression of PTB derives from a caspase-dependent cleavage of the protein during the myocardium development. The process is regulated by histone deacetylases and the caspase inhibitor FLICE-like protein (cFLIP), which induce the PTB degradation during cardiac muscle differentiation.
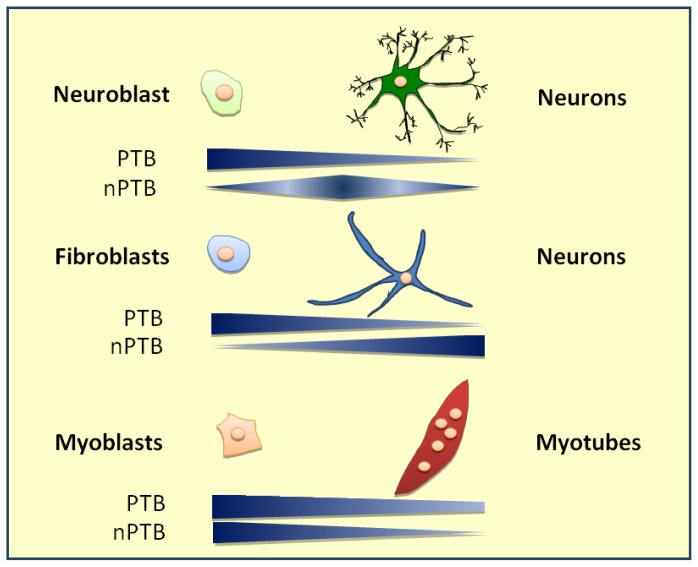
A relevant role played by PTB in splicing regulation during cell differentiation is highlighted by studies on neuronal differentiation, reviewed in [18]. PTB and its homolog nPTB are subject to a programmed switch during neuronal development [89,162]. In post-mitotic neurons a reduced expression of PTB coincides with an increased expression of nPTB, which modifies the neuronal alternative splicing program [47,52,90,163]. A similar switch in the expression of PTB versus nPTB seems to occur in post-mitotic lens fibre cells [164]. A recent study described by Zheng et al. [86] in a mouse model reveals new clues for the understanding of the PTB role during neuronal development. The authors describe three defined phases during neuronal differentiation in which different levels of PTB and nPTB expression mediate splicing regulation. These three phases are characterized by the expression of PTB regulated alternative splice isoforms of the postsynaptic density protein 95 (PSD-95). PSD-95 is an essential factor required for synaptic maturation and plasticity. Both PTB and nPTB repress PSD-95 exon 18 inclusion, leading to premature translation termination and subsequent NMD of the spliced isoform. The reduced expression of PSD-95 protein prevents a precocious stabilization of early generated synapses. The results show that during the initial phase occurring in neuronal stem cells, PTB expression is associated with low levels of PSD-95. The transition to the second phase, represented by the start of neuronal differentiation, comes along with the sequential shift to an increased expression of nPTB and downregulation of PTB, but maintaining a sufficient amount of the two proteins to repress the PSD-95 protein expression. Finally, in the third phase, after mouse birth in cells undergoing synaptic maturation, both PTB and nPTB expression is reduced, causing the stability of the PSD-95 splicing isoforms and consequently an increase of the protein levels and the start of synaptic maturation. Further proof of the functional role of the switch in PTB/nPTB expression in controlling alternative splicing programs required during neuronal development is given by recent results showing the conversion of PTB-depleted fibroblast into neuronal-like cells [165]. A comprehensive view of the regulated expression of PTB and nPTB during cell differentiation is presented in Figure 4.
Figure 4.
Relative ratio of PTB and nPTB expression during cell differentiation.
9. PTB Regulation Mediated by miRNA
The levels of PTB and nPTB expression are subject to microRNA control. Given the estabilished role of microRNAs (miRNAs) in post-transcriptional gene expression regulation, their involvement in PTB and nPTB expression during cell differentiation was expected to be demonstrated. A number of pieces of evidence are now available supporting the function of tissue specific microRNAs in the regulation of PTB expression. The reduced level of PTB and the almost complete absence of nPTB during myoblast differentiation, have been demonstrated to be associated with the induction of the muscle-specific microRNA miR-133 expression [89]. During neurogenesis PTB is downregulated by the activity of the neuron specific microRNA miR-124 [162]. MicroRNAs roles as regulators of PTB levels in specific tissue contexts have also been supported by studies in human pancreatic islets responding to the presence of high levels of glucose. MiR-133a is induced by high amounts of glucose leading to lower PTB levels and lower insulin biosynthesis rates by the effect of mRNA destabilization [166]. Xue et al. [165] demonstrate the power of PTB in reprogramming the differentiation of fibroblast cells types into neuronal-like cells, involving the activity of PTB regulated microRNA circuits. The stable knockdown of PTB obtained by small hairpin RNAs in fibroblast cells leads to the switch to neuronal-like cells. Thus, the absence of PTB leads to the reprogramming of non-neuronal cells into neuronal types by removing a PTB mediated block of microRNAs action. The authors showed that miR-124 represses neuronal genes in non-neuronal cells by an autoregulatory loop mechanism. PTB may positively or negatively regulate the microRNA targeting by two alternative actions. One activity is obtained by competing with microRNA targeting the same transcripts, the other one by promoting microRNA targeting additional transcripts and altering the local RNA structures. Engels et al. [167] have shown that PTB interacts with miRNAs and human Argonaute 2 (Ago2) in the RNA induced silencing complex. Their results showed that miRNAs may be co-regulated by PTB and Ago2 in human cells. We expect that in future studies analysing the global expression in PTB downregulated-reprogrammed cells, new interactions between PTB, additional RBPs and microRNA programs will be revealed.
10. PTB and Cancer
PTB expression may affect cancer initiation and progression [168,169]. Perinucleolar compartments (PNCs) that get formed in malignant cells during transformation, are enriched with RNA polymerase III RNAs and RNA binding proteins, including PTB [170]. PTB differentially affects cancer cell malignancy, depending on the cell type [169]. Increased levels of PTB expression have been associated with growth and maintenance of transformed properties of ovarian tumor cells [168,171]. Suppression of PTB expression by siRNA in ovarian cancer cells leads to inhibition of cell proliferation, anchorage-dependent growth, and cell invasion [168]. PTB may affect malignant cell growth by inducing aberrant splicing events in transcripts involved in tumor progression. PTB in fact induces exon skipping in genes that are involved in proliferation (FGFR1, FGFR2), apoptosis (FAS, CASP2), invasion (CSRC), motility (ACTN, FBN), and multi-drug resistance (ABCC1). In the cytoplasm, PTB increases IRES-mediated translation or RNA stability of transcribed genes involved in proliferation (IGFR1), angiogenesis (VEGF), and apoptosis (APAF1) [102,126–128]. The relevance of the PTB involvement in post-transcriptional regulation of key factors in tumor progression has been recently supported by the demonstration of the effect of a single nucleotide polymorphism (SNP) in the 5′UTR of the p53 tumor suppressor gene expressed in human melanoma tumors. In the cytoplasm PTB positively regulates the translation of p53 mRNA and its N-terminal truncated isoform by binding to IRESs cis-acting elements [119]. The presence of the SNP leads to a weaker binding to PTB and thereafter to a reduced cap-independent translation of p53 [172].
PTB forms complexes with transcripts encoding focal adhesion scaffolding proteins which may affect cell spreading [173,174]. Knockdown of PTB in glioma cell lines modifies the alternative splicing of the transmembrane RNT4 (reticulon 4) factor and decreases cell proliferation and migration, whereas cell adhesion is increased [175]. Recently, it has been reported that PTB promotes the switch of expression of two forms of pyruvate kinase, PKM1 and PKM2, resulting in an increase of the aerobic glycolysis and consequently in a rapid proliferation of cancer cells [176–178]. PTB, together with hnRNPA1 and hnRNPA2, acts in regulating the inclusion of exon 10 and the repression of exon 9 of the pyruvate kinase, resulting in the PKM2 isoform. In human gliomas PTB is overexpressed and the highest level of the protein correlates with the level of PKM2 expression [178]. An opposite effect of PTB overexpression has been demonstrated in the growth of non-small-cell lung cancer cell types. PTB inhibits the proliferation of H1299 and A549 cells and induces the expression of cyclin-dependent kinase inhibitors in H1299 cells [179]. In cells derived from patients with multiple myeloma, c-myc overexpression correlates with an increased interaction of PTB with factor YB-1, as well as with an increased expression of both factors, supporting the role of PTB in stimulating IRES-mediated translation [180]. Taken together, mis-regulation of PTB could cause multiple epigenetic changes in gene expression promoting tumorigenesis, which might be tissue-dependent.
11. Conclusions
The contributions derived from the intense studies on RNA binding proteins confirm that the unique properties of PTB RRM domains are the key elements enabling its multiple cellular functions. The relevant role played by PTB in most post-transcriptional processes is indeed based on its ability to recognize specific RNA sequences, modifying the local RNA structural arrangements. Competition in RNA binding between PTB and other regulators of the RNA metabolism appears critical to its diverse functions. As a consequence, the levels of PTB expression as well as the rate of nuclear import and export may become censorious to the cell fate. Thus, while the identification of new PTB targets is rapidly progressing by means of genome-wide investigations [92], an effort is now being made to identify chemical PTB modulators [181]. It has been demonstrated that tannic acid induces the expression of PTB by activating its promoter region. In congenital myasthenic syndrome, the aberrant inclusion of muscle nicotinic acetylcholine receptor α subunit (CHRNA1) exon P3A is caused by an intronic mutation. PTB overexpression by tannic acid has been demonstrated to foster CHRNA1exon P3A skipping, opening new possibilities of therapeutic approaches based on PTB expression regulation [182]. We expect that on-going and future studies on PTB activities will reveal additional aspects of the complex molecular machinery underlying tissue differentiation events, with a particular focus on the concurrent roles played by miRNAs.
Acknowledgments
This work was supported by the AIRC 2008-Cariverona Regional grant and by the University of Verona Joint Project research funds to MGR. ED was supported in part by AIRC-Cariverona Regional fellowship.
Because of the considerable number of articles published on PTB we have to make a selection. We apologize to those colleagues whose work has not been cited directly.
Conflicts of Interest
The authors declare no conflict of interest.
References
- 1.Moore M.J. From birth to death: The complex lives of eukaryotic mRNAs. Science. 2005;309:1514–1518. doi: 10.1126/science.1111443. [DOI] [PubMed] [Google Scholar]
- 2.Glisovic T., Bachorik J.L., Yong J., Dreyfuss G. RNA-binding proteins and post-transcriptional gene regulation. FEBS Lett. 2008;582:1977–1986. doi: 10.1016/j.febslet.2008.03.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Halbeisen R.E., Galgano A., Scherrer T., Gerber A.P. Post-transcriptional gene regulation: From genome-wide studies to principles. Cell. Mol. Life Sci. 2008;65:798–813. doi: 10.1007/s00018-007-7447-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Pan Q., Shai O., Lee L.J., Frey B.J., Blencowe B.J. Deep surveying of alternative splicing complexity in the human transcriptome by high-throughput sequencing. Nat. Genet. 2008;40:1413–1415. doi: 10.1038/ng.259. [DOI] [PubMed] [Google Scholar]
- 5.Wang E.T., Sandberg R., Luo S., Khrebtukova I., Zhang L., Mayr C., Kingsmore S.F., Schroth G.P., Burge C.B. Alternative isoform regulation in human tissue transcriptomes. Nature. 2008;456:470–476. doi: 10.1038/nature07509. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Nilsen T.W., Graveley B.R. Expansion of the eukaryotic proteome by alternative splicing. Nature. 2010;463:457–463. doi: 10.1038/nature08909. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Wang Z., Burge C.B. Splicing regulation: From a parts list of regulatory elements to an integrated splicing code. RNA. 2008;14:802–813. doi: 10.1261/rna.876308. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Chen M., Manley J.L. Mechanisms of alternative splicing regulation: Insights from molecular and genomics approaches. Nat. Rev. Mol. Cell Biol. 2009;10:741–754. doi: 10.1038/nrm2777. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Wahl M.C., Will C.L., Lührmann R. The spliceosome: Design principles of a dynamic RNP machine. Cell. 2009;136:701–718. doi: 10.1016/j.cell.2009.02.009. [DOI] [PubMed] [Google Scholar]
- 10.Tejedor J.R., Valcárcel J. Gene regulation: Breaking the second genetic code. Nature. 2010;465:45–46. doi: 10.1038/465045a. [DOI] [PubMed] [Google Scholar]
- 11.Singh R., Valcárcel J. Building specificity with nonspecific RNA-binding proteins. Nat. Struct. Mol. Biol. 2005;12:645–653. doi: 10.1038/nsmb961. [DOI] [PubMed] [Google Scholar]
- 12.Busch A., Hertel K.J. Evolution of SR protein and hnRNP splicing regulatory factors. Wiley Interdiscip. Rev. RNA. 2012;3:1–12. doi: 10.1002/wrna.100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Lynch K.W. Regulation of alternative splicing by signal transduction pathways. Adv. Exp. Med. Biol. 2007;623:161–174. doi: 10.1007/978-0-387-77374-2_10. [DOI] [PubMed] [Google Scholar]
- 14.Long J.C., Caceres J.F. The SR protein family of splicing factors: Master regulators of gene expression. Biochem. J. 2009;417:15–27. doi: 10.1042/BJ20081501. [DOI] [PubMed] [Google Scholar]
- 15.Licatalosi D.D., Darnell R.B. RNA processing and its regulation: Global insights into biological networks. Nat. Rev. Genet. 2010;11:75–87. doi: 10.1038/nrg2673. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Smith C.W., Valcárcel J. Alternative pre-mRNA splicing: The logic of combinatorial control. Trends Biochem. Sci. 2000;25:381–388. doi: 10.1016/s0968-0004(00)01604-2. [DOI] [PubMed] [Google Scholar]
- 17.Auweter S.D., Allain F.H.-T. Structure-function relationships of the polypyrimidine tract binding protein. Cell. Mol. Life Sci. 2008;65:516–527. doi: 10.1007/s00018-007-7378-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Keppetipola N., Sharma S., Li Q., Black D.L. Neuronal regulation of pre-mRNA splicing by polypyrimidine tract binding proteins, PTBP1 and PTBP2. Crit. Rev. Biochem. Mol. Biol. 2012;47:360–378. doi: 10.3109/10409238.2012.691456. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Sawicka K., Bushell M., Spriggs K.A., Willis A.E. Polypyrimidine-tract-binding protein: A multifunctional RNA-binding protein. Biochem. Soc. Trans. 2008;36:641–647. doi: 10.1042/BST0360641. [DOI] [PubMed] [Google Scholar]
- 20.Patton J.G., Mayer S.A., Tempst P., Nadal-Ginard B. Characterization and molecular cloning of polypyrimidine tract-binding protein: A component of a complex necessary for pre-mRNA splicing. Genes Dev. 1991;5:1237–1251. doi: 10.1101/gad.5.7.1237. [DOI] [PubMed] [Google Scholar]
- 21.Ghetti A., Piñol-Roma S., Michael W.M., Morandi C., Dreyfuss G. hnRNP I, the polypyrimidine tract-binding protein: Distinct nuclear localization and association with hnRNAs. Nucleic Acids Res. 1992;20:3671–3678. doi: 10.1093/nar/20.14.3671. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Kamath R.V., Leary D.J., Huang S. Nucleocytoplasmic shuttling of polypyrimidine tract-binding protein is uncoupled from RNA export. Mol. Biol. Cell. 2001;12:3808–3820. doi: 10.1091/mbc.12.12.3808. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Romanelli M.G., Weighardt F., Biamonti G., Riva S., Morandi C. Sequence determinants for hnRNP I protein nuclear localization. Exp. Cell Res. 1997;235:300–304. doi: 10.1006/excr.1997.3677. [DOI] [PubMed] [Google Scholar]
- 24.Romanelli M.G., Morandi C. Importin alpha binds to an unusual bipartite nuclear localization signal in the heterogeneous ribonucleoprotein type I. Eur. J. Biochem. 2002;269:2727–2734. doi: 10.1046/j.1432-1033.2002.02942.x. [DOI] [PubMed] [Google Scholar]
- 25.Li B., Yen T.S.B. Characterization of the nuclear export signal of polypyrimidine tract-binding protein. J. Biol. Chem. 2002;277:10306–10314. doi: 10.1074/jbc.M109686200. [DOI] [PubMed] [Google Scholar]
- 26.Romanelli M.G., Lorenzi P., Morandi C. Organization of the human gene encoding heterogeneous nuclear ribonucleoprotein type I (hnRNP I) and characterization of hnRNP I related pseudogene. Gene. 2000;255:267–272. doi: 10.1016/s0378-1119(00)00331-0. [DOI] [PubMed] [Google Scholar]
- 27.Lunde B.M., Moore C., Varani G. RNA-binding proteins: Modular design for efficient function. Nat. Rev. Mol. Cell Biol. 2007;8:479–490. doi: 10.1038/nrm2178. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Maris C., Dominguez C., Allain F.H.-T. The RNA recognition motif, a plastic RNA-binding platform to regulate post-transcriptional gene expression. FEBS J. 2005;272:2118–2131. doi: 10.1111/j.1742-4658.2005.04653.x. [DOI] [PubMed] [Google Scholar]
- 29.Dreyfuss G., Kim V.N., Kataoka N. Messenger-RNA-binding proteins and the messages they carry. Nat. Rev. Mol. Cell Biol. 2002;3:195–205. doi: 10.1038/nrm760. [DOI] [PubMed] [Google Scholar]
- 30.Cléry A., Blatter M., Allain F.H.-T. RNA recognition motifs: Boring? Not quite. Curr. Opin. Struct. Biol. 2008;18:290–298. doi: 10.1016/j.sbi.2008.04.002. [DOI] [PubMed] [Google Scholar]
- 31.Conte M.R., Grüne T., Ghuman J., Kelly G., Ladas A., Matthews S., Curry S. Structure of tandem RNA recognition motifs from polypyrimidine tract binding protein reveals novel features of the RRM fold. EMBO J. 2000;19:3132–3141. doi: 10.1093/emboj/19.12.3132. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Vitali F., Henning A., Oberstrass F.C., Hargous Y., Auweter S.D., Erat M., Allain F.H.-T. Structure of the two most C-terminal RNA recognition motifs of PTB using segmental isotope labeling. EMBO J. 2006;25:150–162. doi: 10.1038/sj.emboj.7600911. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Izquierdo J.M., Majós N., Bonnal S., Martínez C., Castelo R., Guigó R., Bilbao D., Valcárcel J. Regulation of Fas alternative splicing by antagonistic effects of TIA-1 and PTB on exon definition. Mol. Cell. 2005;19:475–484. doi: 10.1016/j.molcel.2005.06.015. [DOI] [PubMed] [Google Scholar]
- 34.Pérez I., McAfee J.G., Patton J.G. Multiple RRMs contribute to RNA binding specificity and affinity for polypyrimidine tract binding protein. Biochemistry. 1997;36:11881–11890. doi: 10.1021/bi9711745. [DOI] [PubMed] [Google Scholar]
- 35.Ray D., Kazan H., Chan E.T., Peña Castillo L., Chaudhry S., Talukder S., Blencowe B.J., Morris Q., Hughes T.R. Rapid and systematic analysis of the RNA recognition specificities of RNA-binding proteins. Nat. Biotechnol. 2009;27:667–670. doi: 10.1038/nbt.1550. [DOI] [PubMed] [Google Scholar]
- 36.Reid D.C., Chang B.L., Gunderson S.I., Alpert L., Thompson W.A., Fairbrother W.G. Next-generation SELEX identifies sequence and structural determinants of splicing factor binding in human pre-mRNA sequence. RNA. 2009;15:2385–2397. doi: 10.1261/rna.1821809. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Oberstrass F.C., Auweter S.D., Erat M., Hargous Y., Henning A., Wenter P., Reymond L., Amir-Ahmady B., Pitsch S., Black D.L., et al. Structure of PTB bound to RNA: Specific binding and implications for splicing regulation. Science. 2005;309:2054–2057. doi: 10.1126/science.1114066. [DOI] [PubMed] [Google Scholar]
- 38.Lamichhane R., Daubner G.M., Thomas-Crusells J., Auweter S.D., Manatschal C., Austin K.S., Valniuk O., Allain F.H.-T., Rueda D. RNA looping by PTB: Evidence using FRET and NMR spectroscopy for a role in splicing repression. Proc. Natl. Acad. Sci. USA. 2010;107:4105–4110. doi: 10.1073/pnas.0907072107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Maynard C.M., Hall K.B. Interactions between PTB RRMs induce slow motions and increase RNA binding affinity. J. Mol. Biol. 2010;397:260–277. doi: 10.1016/j.jmb.2009.12.051. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Sharma S., Maris C., Allain F.H.-T., Black D.L. U1 snRNA directly interacts with polypyrimidine tract-binding protein during splicing repression. Mol. Cell. 2011;41:579–588. doi: 10.1016/j.molcel.2011.02.012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Clerte C., Hall K.B. The domains of polypyrimidine tract binding protein have distinct RNA structural preferences. Biochemistry. 2009;48:2063–2074. doi: 10.1021/bi8016872. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Kafasla P., Morgner N., Pöyry T.A.A., Curry S., Robinson C.V., Jackson R.J. Polypyrimidine tract binding protein stabilizes the encephalomyocarditis virus IRES structure via binding multiple sites in a unique orientation. Mol. Cell. 2009;34:556–568. doi: 10.1016/j.molcel.2009.04.015. [DOI] [PubMed] [Google Scholar]
- 43.Rideau A.P., Gooding C., Simpson P.J., Monie T.P., Lorenz M., Hüttelmaier S., Singer R.H., Matthews S., Curry S., Smith C.W.J. A peptide motif in Raver1 mediates splicing repression by interaction with the PTB RRM2 domain. Nat. Struct. Mol. Biol. 2006;13:839–848. doi: 10.1038/nsmb1137. [DOI] [PubMed] [Google Scholar]
- 44.Lorenz M. Visualizing protein-RNA interactions inside cells by fluorescence resonance energy transfer. RNA. 2009;15:97–103. doi: 10.1261/rna.1307809. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Henneberg B., Swiniarski S., Becke S., Illenberger S. A conserved peptide motif in Raver2 mediates its interaction with the polypyrimidine tract-binding protein. Exp. Cell Res. 2010;316:966–979. doi: 10.1016/j.yexcr.2009.11.023. [DOI] [PubMed] [Google Scholar]
- 46.Joshi A., Coelho M.B., Kotik-Kogan O., Simpson P.J., Matthews S.J., Smith C.W.J., Curry S. Crystallographic analysis of polypyrimidine tract-binding protein-Raver1 interactions involved in regulation of alternative splicing. Structure. 2011;19:1816–1825. doi: 10.1016/j.str.2011.09.020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Polydorides A.D., Okano H.J., Yang Y.Y., Stefani G., Darnell R.B. A brain-enriched polypyrimidine tract-binding protein antagonizes the ability of Nova to regulate neuron-specific alternative splicing. Proc. Natl. Acad. Sci. USA. 2000;97:6350–6355. doi: 10.1073/pnas.110128397. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Romanelli M.G., Lorenzi P., Morandi C. Identification and analysis of the human neural polypyrimidine tract binding protein (nPTB) gene promoter region. Gene. 2005;356:11–18. doi: 10.1016/j.gene.2005.04.031. [DOI] [PubMed] [Google Scholar]
- 49.Coutinho-Mansfield G.C., Xue Y., Zhang Y., Fu X.-D. PTB/nPTB switch: A post-transcriptional mechanism for programming neuronal differentiation. Genes Dev. 2007;21:1573–1577. doi: 10.1101/gad.1575607. [DOI] [PubMed] [Google Scholar]
- 50.Brazão T.F., Demmers J., van IJcken W., Strouboulis J., Fornerod M., Romão L., Grosveld F.G. A new function of ROD1 in nonsense-mediated mRNA decay. FEBS Lett. 2012;586:1101–1110. doi: 10.1016/j.febslet.2012.03.015. [DOI] [PubMed] [Google Scholar]
- 51.Gooding C., Roberts G.C., Moreau G., Nadal-Ginard B., Smith C.W. Smooth muscle-specific switching of alpha-tropomyosin mutually exclusive exon selection by specific inhibition of the strong default exon. EMBO J. 1994;13:3861–3872. doi: 10.1002/j.1460-2075.1994.tb06697.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Markovtsov V., Nikolic J.M., Goldman J.A., Turck C.W., Chou M.Y., Black D.L. Cooperative assembly of an hnRNP complex induced by a tissue-specific homolog of polypyrimidine tract binding protein. Mol. Cell. Biol. 2000;20:7463–7479. doi: 10.1128/mcb.20.20.7463-7479.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Rahman L., Bliskovski V., Reinhold W., Zajac-Kaye M. Alternative splicing of brain-specific PTB defines a tissue-specific isoform pattern that predicts distinct functional roles. Genomics. 2002;80:245–249. doi: 10.1006/geno.2002.6826. [DOI] [PubMed] [Google Scholar]
- 54.Wollerton M.C., Gooding C., Wagner E.J., Garcia-Blanco M.A., Smith C.W.J. Autoregulation of polypyrimidine tract binding protein by alternative splicing leading to nonsense-mediated decay. Mol. Cell. 2004;13:91–100. doi: 10.1016/s1097-2765(03)00502-1. [DOI] [PubMed] [Google Scholar]
- 55.Spellman R., Llorian M., Smith C.W.J. Crossregulation and functional redundancy between the splicing regulator PTB and its paralogs nPTB and ROD1. Mol. Cell. 2007;27:420–434. doi: 10.1016/j.molcel.2007.06.016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Ni J.Z., Grate L., Donohue J.P., Preston C., Nobida N., O’Brien G., Shiue L., Clark T.A., Blume J.E., Ares M. Ultraconserved elements are associated with homeostatic control of splicing regulators by alternative splicing and nonsense-mediated decay. Genes Dev. 2007;21:708–718. doi: 10.1101/gad.1525507. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Hamilton B.J., Genin A., Cron R.Q., Rigby W.F. Delineation of a novel pathway that regulates CD154 (CD40 ligand) expression. Mol. Cell. Biol. 2003;23:510–525. doi: 10.1128/MCB.23.2.510-525.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Hüttelmaier S., Illenberger S., Grosheva I., Rüdiger M., Singer R.H., Jockusch B.M. Raver1, a dual compartment protein, is a ligand for PTB/hnRNPI and microfilament attachment protein. J. Cell Biol. 2001;155:775–786. doi: 10.1083/jcb.200105044. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Gromak N., Rideau A., Southby J., Scadden A.D.J., Gooding C., Hüttelmaier S., Singer R.H., Smith C.W.J. The PTB interacting protein raver1 regulates alpha-tropomyosin alternative splicing. EMBO J. 2003;22:6356–6364. doi: 10.1093/emboj/cdg609. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Kleinhenz B., Fabienke M., Swiniarski S., Wittenmayer N., Kirsch J., Jockusch B.M., Arnold H.H., Illenberger S. Raver2, a new member of the hnRNP family. FEBS Lett. 2005;579:4254–4258. doi: 10.1016/j.febslet.2005.07.001. [DOI] [PubMed] [Google Scholar]
- 61.Spellman R., Smith C.W.J. Novel modes of splicing repression by PTB. Trends Biochem. Sci. 2006;31:73–76. doi: 10.1016/j.tibs.2005.12.003. [DOI] [PubMed] [Google Scholar]
- 62.Romanelli M.G., Lorenzi P., Avesani F., Morandi C. Functional characterization of the ribonucleoprotein, PTB-binding 1/Raver1 promoter region. Gene. 2007;405:79–87. doi: 10.1016/j.gene.2007.09.004. [DOI] [PubMed] [Google Scholar]
- 63.Zieseniss A., Schroeder U., Buchmeier S., Schoenenberger C.-A., van den Heuvel J., Jockusch B.M., Illenberger S. Raver1 is an integral component of muscle contractile elements. Cell Tissue Res. 2007;327:583–594. doi: 10.1007/s00441-006-0322-1. [DOI] [PubMed] [Google Scholar]
- 64.Lee J.H., Rangarajan E.S., Yogesha S.D., Izard T. Raver1 interactions with vinculin and RNA suggest a feed-forward pathway in directing mRNA to focal adhesions. Structure. 2009;17:833–842. doi: 10.1016/j.str.2009.04.010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Lahmann I., Fabienke M., Henneberg B., Pabst O., Vauti F., Minge D., Illenberger S., Jockusch B.M., Korte M., Arnold H.-H. The hnRNP and cytoskeletal protein raver1 contributes to synaptic plasticity. Exp. Cell Res. 2008;314:1048–1060. doi: 10.1016/j.yexcr.2007.10.022. [DOI] [PubMed] [Google Scholar]
- 66.Chen H., Li Y., Zhang J., Ran Y., Wei J., Yang Y., Shu H.-B. RAVER1 is a coactivator of MDA5-mediated cellular antiviral response. J. Mol. Cell Biol. 2013;5:111–119. doi: 10.1093/jmcb/mjt006. [DOI] [PubMed] [Google Scholar]
- 67.Romanelli M.G., Lorenzi P., Diani E., Filippello A., Avesani F., Morandi C. Transcriptional regulation of the human Raver2 ribonucleoprotein gene. Gene. 2012;493:243–252. doi: 10.1016/j.gene.2011.11.036. [DOI] [PubMed] [Google Scholar]
- 68.Spellman R., Rideau A., Matlin A., Gooding C., Robinson F., McGlincy N., Grellscheid S.N., Southby J., Wollerton M., Smith C.W.J. Regulation of alternative splicing by PTB and associated factors. Biochem. Soc. Trans. 2005;33:457–460. doi: 10.1042/BST0330457. [DOI] [PubMed] [Google Scholar]
- 69.Southby J., Gooding C., Smith C.W.J. Polypyrimidine tract binding protein functions as a repressor to regulate alternative splicing of alpha -actinin mutally exclusive exons. Mol. Cell. Biol. 1999;19:2699–2711. doi: 10.1128/mcb.19.4.2699. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Matlin A.J., Southby J., Gooding C., Smith C.W.J. Repression of α-actinin SM exon splicing by assisted binding of PTB to the polypyrimidine tract. RNA. 2007;13:1214–1223. doi: 10.1261/rna.219607. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Gooding C., Roberts G.C., Smith C.W.J. Role of an inhibitory pyrimidine element and polypyrimidine tract binding protein in repression of a regulated alpha-tropomyosin exon. RNA. 1998;4:85–100. [PMC free article] [PubMed] [Google Scholar]
- 72.Hayakawa M., Sakashita E., Ueno E., Tominaga S., Hamamoto T., Kagawa Y., Endo H. Muscle-specific exonic splicing silencer for exon exclusion in human ATP synthase gamma-subunit pre-mRNA. J. Biol. Chem. 2002;277:6974–6984. doi: 10.1074/jbc.M110138200. [DOI] [PubMed] [Google Scholar]
- 73.Saulière J., Sureau A., Expert-Bezançon A., Marie J. The polypyrimidine tract binding protein (PTB) represses splicing of exon 6B from the beta-tropomyosin pre-mRNA by directly interfering with the binding of the U2AF65 subunit. Mol. Cell. Biol. 2006;26:8755–8769. doi: 10.1128/MCB.00893-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Chan R.C., Black D.L. The polypyrimidine tract binding protein binds upstream of neural cell-specific c-src exon N1 to repress the splicing of the intron downstream. Mol. Cell. Biol. 1997;17:4667–4676. doi: 10.1128/mcb.17.8.4667. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Chou M.Y., Underwood J.G., Nikolic J., Luu M.H., Black D.L. Multisite RNA binding and release of polypyrimidine tract binding protein during the regulation of c-src neural-specific splicing. Mol. Cell. 2000;5:949–957. doi: 10.1016/s1097-2765(00)80260-9. [DOI] [PubMed] [Google Scholar]
- 76.Lou H., Helfman D.M., Gagel R.F., Berget S.M. Polypyrimidine tract-binding protein positively regulates inclusion of an alternative 3′-terminal exon. Mol. Cell. Biol. 1999;19:78–85. doi: 10.1128/mcb.19.1.78. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Côté J., Dupuis S., Wu J.Y. Polypyrimidine track-binding protein binding downstream of caspase-2 alternative exon 9 represses its inclusion. J. Biol. Chem. 2001;276:8535–8543. doi: 10.1074/jbc.M008924200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Charlet-B N., Logan P., Singh G., Cooper T.A. Dynamic antagonism between ETR-3 and PTB regulates cell type-specific alternative splicing. Mol. Cell. 2002;9:649–658. doi: 10.1016/s1097-2765(02)00479-3. [DOI] [PubMed] [Google Scholar]
- 79.Zhang L., Liu W., Grabowski P.J. Coordinate repression of a trio of neuron-specific splicing events by the splicing regulator PTB. RNA. 1999;5:117–130. doi: 10.1017/s1355838299981530. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Jin W., Bruno I.G., Xie T.-X., Sanger L.J., Cote G.J. Polypyrimidine tract-binding protein down-regulates fibroblast growth factor receptor 1 alpha-exon inclusion. Cancer Res. 2003;63:6154–6157. [PubMed] [Google Scholar]
- 81.Carstens R.P., Wagner E.J., Garcia-Blanco M.A. An intronic splicing silencer causes skipping of the IIIb exon of fibroblast growth factor receptor 2 through involvement of polypyrimidine tract binding protein. Mol. Cell. Biol. 2000;20:7388–7400. doi: 10.1128/mcb.20.19.7388-7400.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Ashiya M., Grabowski P.J. A neuron-specific splicing switch mediated by an array of pre-mRNA repressor sites: Evidence of a regulatory role for the polypyrimidine tract binding protein and a brain-specific PTB counterpart. RNA. 1997;3:996–1015. [PMC free article] [PubMed] [Google Scholar]
- 83.Shen H., Kan J.L.C., Ghigna C., Biamonti G., Green M.R. A single polypyrimidine tract binding protein (PTB) binding site mediates splicing inhibition at mouse IgM exons M1 and M2. RNA. 2004;10:787–794. doi: 10.1261/rna.5229704. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Ye J., Llorian M., Cardona M., Rongvaux A., Moubarak R.S., Comella J.X., Bassel-Duby R., Flavell R.A., Olson E.N., Smith C.W.J., et al. A pathway involving HDAC5, cFLIP and caspases regulates expression of the splicing regulator polypyrimidine tract binding protein in the heart. J. Cell Sci. 2013;126:1682–1691. doi: 10.1242/jcs.121384. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 85.Tang Z.Z., Sharma S., Zheng S., Chawla G., Nikolic J., Black D.L. Regulation of the mutually exclusive exons 8a and 8 in the CaV1.2 calcium channel transcript by polypyrimidine tract-binding protein. J. Biol. Chem. 2011;286:10007–10016. doi: 10.1074/jbc.M110.208116. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86.Zheng S., Gray E.E., Chawla G., Porse B.T., O’Dell T.J., Black D.L. PSD-95 is post-transcriptionally repressed during early neural development by PTBP1 and PTBP2. Nat. Neurosci. 2012;15:381–388. doi: 10.1038/nn.3026. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87.Liu H., Zhang W., Reed R.B., Liu W., Grabowski P.J. Mutations in RRM4 uncouple the splicing repression and RNA-binding activities of polypyrimidine tract binding protein. RNA. 2002;8:137–149. doi: 10.1017/s1355838202015029. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 88.Gooding C., Edge C., Lorenz M., Coelho M.B., Winters M., Kaminski C.F., Cherny D., Eperon I.C., Smith C.W.J. MBNL1 and PTB cooperate to repress splicing of Tpm1 exon 3. Nucleic Acids Res. 2013;41:4765–4782. doi: 10.1093/nar/gkt168. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 89.Boutz P.L., Chawla G., Stoilov P., Black D.L. MicroRNAs regulate the expression of the alternative splicing factor nPTB during muscle development. Genes Dev. 2007;21:71–84. doi: 10.1101/gad.1500707. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 90.Boutz P.L., Stoilov P., Li Q., Lin C.-H., Chawla G., Ostrow K., Shiue L., Ares M., Black D.L. A post-transcriptional regulatory switch in polypyrimidine tract-binding proteins reprograms alternative splicing in developing neurons. Genes Dev. 2007;21:1636–1652. doi: 10.1101/gad.1558107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91.Corrionero A., Valcárcel J. RNA processing: Redrawing the map of charted territory. Mol. Cell. 2009;36:918–919. doi: 10.1016/j.molcel.2009.12.004. [DOI] [PubMed] [Google Scholar]
- 92.Xue Y., Zhou Y., Wu T., Zhu T., Ji X., Kwon Y.-S., Zhang C., Yeo G., Black D.L., Sun H., Fu X.-D., Zhang Y. Genome-wide analysis of PTB-RNA interactions reveals a strategy used by the general splicing repressor to modulate exon inclusion or skipping. Mol. Cell. 2009;36:996–1006. doi: 10.1016/j.molcel.2009.12.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 93.Llorian M., Schwartz S., Clark T.A., Hollander D., Tan L.-Y., Spellman R., Gordon A., Schweitzer A.C., de la Grange P., Ast G., et al. Position-dependent alternative splicing activity revealed by global profiling of alternative splicing events regulated by PTB. Nat. Struct. Mol. Biol. 2010;17:1114–1123. doi: 10.1038/nsmb.1881. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 94.Paradis C., Cloutier P., Shkreta L., Toutant J., Klarskov K., Chabot B. hnRNP I/PTB can antagonize the splicing repressor activity of SRp30c. RNA. 2007;13:1287–1300. doi: 10.1261/rna.403607. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 95.Kafasla P., Mickleburgh I., Llorian M., Coelho M., Gooding C., Cherny D., Joshi A., Kotik-Kogan O., Curry S., Eperon I.C., et al. Defining the roles and interactions of PTB. Biochem. Soc. Trans. 2012;40:815–820. doi: 10.1042/BST20120044. [DOI] [PubMed] [Google Scholar]
- 96.Xing Y., Stoilov P., Kapur K., Han A., Jiang H., Shen S., Black D.L., Wong W.H. MADS: A new and improved method for analysis of differential alternative splicing by exon-tiling microarrays. RNA. 2008;14:1470–1479. doi: 10.1261/rna.1070208. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 97.Wen J., Chen Z., Cai X. A biophysical model for identifying splicing regulatory elements and their interactions. PLoS One. 2013;8:e54885. doi: 10.1371/journal.pone.0054885. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 98.Stoneley M., Willis A.E. Cellular internal ribosome entry segments: Structures, trans-acting factors and regulation of gene expression. Oncogene. 2004;23:3200–3207. doi: 10.1038/sj.onc.1207551. [DOI] [PubMed] [Google Scholar]
- 99.Martínez-Salas E., Pacheco A., Serrano P., Fernandez N. New insights into internal ribosome entry site elements relevant for viral gene expression. J. Gen. Virol. 2008;89:611–626. doi: 10.1099/vir.0.83426-0. [DOI] [PubMed] [Google Scholar]
- 100.Verma B., Bhattacharyya S., Das S. Polypyrimidine tract-binding protein interacts with coxsackievirus B3 RNA and influences its translation. J. Gen. Virol. 2010;91:1245–1255. doi: 10.1099/vir.0.018507-0. [DOI] [PubMed] [Google Scholar]
- 101.Vashist S., Urena L., Chaudhry Y., Goodfellow I. Identification of RNA-protein interaction networks involved in the norovirus life cycle. J. Virol. 2012;86:11977–11990. doi: 10.1128/JVI.00432-12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 102.Mitchell S.A., Spriggs K.A., Coldwell M.J., Jackson R.J., Willis A.E. The Apaf-1 internal ribosome entry segment attains the correct structural conformation for function via interactions with PTB and unr. Mol. Cell. 2003;11:757–771. doi: 10.1016/s1097-2765(03)00093-5. [DOI] [PubMed] [Google Scholar]
- 103.Spriggs K.A., Cobbold L.C., Ridley S.H., Coldwell M., Bottley A., Bushell M., Willis A.E., Siddle K. The human insulin receptor mRNA contains a functional internal ribosome entry segment. Nucleic Acids Res. 2009;37:5881–5893. doi: 10.1093/nar/gkp623. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 104.Song Y., Tzima E., Ochs K., Bassili G., Trusheim H., Linder M., Preissner K.T., Niepmann M. Evidence for an RNA chaperone function of polypyrimidine tract-binding protein in picornavirus translation. RNA. 2005;11:1809–1824. doi: 10.1261/rna.7430405. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 105.Back S.H., Kim Y.K., Kim W.J., Cho S., Oh H.R., Kim J.-E., Jang S.K. Translation of polioviral mRNA is inhibited by cleavage of polypyrimidine tract-binding proteins executed by polioviral 3C(pro) J. Virol. 2002;76:2529–2542. doi: 10.1128/jvi.76.5.2529-2542.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 106.Florez P.M., Sessions O.M., Wagner E.J., Gromeier M., Garcia-Blanco M.A. The polypyrimidine tract binding protein is required for efficient picornavirus gene expression and propagation. J. Virol. 2005;79:6172–6179. doi: 10.1128/JVI.79.10.6172-6179.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 107.Jahan N., Wimmer E., Mueller S. Polypyrimidine tract binding protein-1 (PTB1) is a determinant of the tissue and host tropism of a human rhinovirus/poliovirus chimera PV1(RIPO) PLoS One. 2013;8:e60791. doi: 10.1371/journal.pone.0060791. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 108.Chang K.-S., Luo G. The polypyrimidine tract-binding protein (PTB) is required for efficient replication of hepatitis C virus (HCV) RNA. Virus Res. 2006;115:1–8. doi: 10.1016/j.virusres.2005.06.012. [DOI] [PubMed] [Google Scholar]
- 109.Agis-Juárez R.A., Galván I., Medina F., Daikoku T., Padmanabhan R., Ludert J.E., del Angel R.M. Polypyrimidine tract-binding protein is relocated to the cytoplasm and is required during dengue virus infection in Vero cells. J. Gen. Virol. 2009;90:2893–2901. doi: 10.1099/vir.0.013433-0. [DOI] [PubMed] [Google Scholar]
- 110.Jiang L., Yao H., Duan X., Lu X., Liu Y. Polypyrimidine tract-binding protein influences negative strand RNA synthesis of dengue virus. Biochem. Biophys. Res. Commun. 2009;385:187–192. doi: 10.1016/j.bbrc.2009.05.036. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 111.Kanda T., Gauss-Müller V., Cordes S., Tamura R., Okitsu K., Shuang W., Nakamoto S., Fujiwara K., Imazeki F., Yokosuka O. Hepatitis A virus (HAV) proteinase 3C inhibits HAV IRES-dependent translation and cleaves the polypyrimidine tract-binding protein. J. Viral Hepat. 2010;17:618–623. doi: 10.1111/j.1365-2893.2009.01221.x. [DOI] [PubMed] [Google Scholar]
- 112.Venkatramana M., Ray P.S., Chadda A., Das S. A 25 kDa cleavage product of polypyrimidine tract binding protein (PTB) present in mouse tissues prevents PTB binding to the 5′ untranslated region and inhibits translation of hepatitis A virus RNA. Virus Res. 2003;98:141–149. doi: 10.1016/j.virusres.2003.09.004. [DOI] [PubMed] [Google Scholar]
- 113.Kafasla P., Morgner N., Robinson C.V., Jackson R.J. Polypyrimidine tract-binding protein stimulates the poliovirus IRES by modulating eIF4G binding. EMBO J. 2010;29:3710–3722. doi: 10.1038/emboj.2010.231. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 114.Bailey D., Karakasiliotis I., Vashist S., Chung L.M.W., Rees J., Reese J., McFadden N., Benson A., Yarovinsky F., Simmonds P., et al. Functional analysis of RNA structures present at the 3′ extremity of the murine norovirus genome: The variable polypyrimidine tract plays a role in viral virulence. J. Virol. 2010;84:2859–2870. doi: 10.1128/JVI.02053-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 115.Anwar A., Leong K.M., Ng M.L., Chu J.J.H., Garcia-Blanco M.A. The polypyrimidine tract-binding protein is required for efficient dengue virus propagation and associates with the viral replication machinery. J. Biol. Chem. 2009;284:17021–17029. doi: 10.1074/jbc.M109.006239. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 116.Niepmann M., Petersen A., Meyer K., Beck E. Functional involvement of polypyrimidine tract-binding protein in translation initiation complexes with the internal ribosome entry site of foot-and-mouth disease virus. J. Virol. 1997;71:8330–8339. doi: 10.1128/jvi.71.11.8330-8339.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 117.Ohno S., Shibayama M., Sato M., Tokunaga A., Yoshida N. Polypyrimidine tract-binding protein regulates the cell cycle through IRES-dependent translation of CDK11(p58) in mouse embryonic stem cells. Cell Cycle. 2011;10:3706–3713. doi: 10.4161/cc.10.21.17903. [DOI] [PubMed] [Google Scholar]
- 118.Rübsamen D., Blees J.S., Schulz K., Döring C., Hansmann M.-L., Heide H., Weigert A., Schmid T., Brüne B. IRES-dependent translation of egr2 is induced under inflammatory conditions. RNA. 2012;18:1910–1920. doi: 10.1261/rna.033019.112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 119.Grover R., Ray P.S., Das S. Polypyrimidine tract binding protein regulates IRES-mediated translation of p53 isoforms. Cell Cycle. 2008;7:2189–2198. doi: 10.4161/cc.7.14.6271. [DOI] [PubMed] [Google Scholar]
- 120.Majumder M., Yaman I., Gaccioli F., Zeenko V.V., Wang C., Caprara M.G., Venema R.C., Komar A.A., Snider M.D., Hatzoglou M. The hnRNA-binding proteins hnRNP L and PTB are required for efficient translation of the Cat-1 arginine/lysine transporter mRNA during amino acid starvation. Mol. Cell. Biol. 2009;29:2899–2912. doi: 10.1128/MCB.01774-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 121.Schepens B., Tinton S.A., Bruynooghe Y., Beyaert R., Cornelis S. The polypyrimidine tract-binding protein stimulates HIF-1alpha IRES-mediated translation during hypoxia. Nucleic Acids Res. 2005;33:6884–6894. doi: 10.1093/nar/gki1000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 122.Cho S., Kim J.H., Back S.H., Jang S.K. Polypyrimidine tract-binding protein enhances the internal ribosomal entry site-dependent translation of p27Kip1 mRNA and modulates transition from G1 to S phase. Mol. Cell. Biol. 2005;25:1283–1297. doi: 10.1128/MCB.25.4.1283-1297.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 123.Dhar D., Venkataramana M., Ponnuswamy A., Das S. Role of polypyrimidine tract binding protein in mediating internal initiation of translation of interferon regulatory factor 2 RNA. PLoS One. 2009;4:e7049. doi: 10.1371/journal.pone.0007049. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 124.Kim D.-Y., Woo K.-C., Lee K.-H., Kim T.-D., Kim K.-T. hnRNP Q and PTB modulate the circadian oscillation of mouse Rev-erb alpha via IRES-mediated translation. Nucleic Acids Res. 2010;38:7068–7078. doi: 10.1093/nar/gkq569. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 125.Cobbold L.C., Spriggs K.A., Haines S.J., Dobbyn H.C., Hayes C., de Moor C.H., Lilley K.S., Bushell M., Willis A.E. Identification of internal ribosome entry segment (IRES)-trans-acting factors for the Myc family of IRESs. Mol. Cell. Biol. 2008;28:40–49. doi: 10.1128/MCB.01298-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 126.Coles L.S., Bartley M.A., Bert A., Hunter J., Polyak S., Diamond P., Vadas M.A., Goodall G.J. A multi-protein complex containing cold shock domain (Y-box) and polypyrimidine tract binding proteins forms on the vascular endothelial growth factor mRNA. Potential role in mRNA stabilization. Eur. J. Biochem. 2004;271:648–660. doi: 10.1111/j.1432-1033.2003.03968.x. [DOI] [PubMed] [Google Scholar]
- 127.Huez I., Créancier L., Audigier S., Gensac M.C., Prats A.C., Prats H. Two independent internal ribosome entry sites are involved in translation initiation of vascular endothelial growth factor mRNA. Mol. Cell. Biol. 1998;18:6178–6190. doi: 10.1128/mcb.18.11.6178. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 128.Giraud S., Greco A., Brink M., Diaz J.J., Delafontaine P. Translation initiation of the insulin-like growth factor I receptor mRNA is mediated by an internal ribosome entry site. J. Biol. Chem. 2001;276:5668–5675. doi: 10.1074/jbc.M005928200. [DOI] [PubMed] [Google Scholar]
- 129.Cornelis S., Tinton S.A., Schepens B., Bruynooghe Y., Beyaert R. UNR translation can be driven by an IRES element that is negatively regulated by polypyrimidine tract binding protein. Nucleic Acids Res. 2005;33:3095–3108. doi: 10.1093/nar/gki611. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 130.Fred R.G., Bang-Berthelsen C.H., Mandrup-Poulsen T., Grunnet L.G., Welsh N. High glucose suppresses human islet insulin biosynthesis by inducing miR-133a leading to decreased polypyrimidine tract binding protein-expression. PLoS One. 2010;5:e10843. doi: 10.1371/journal.pone.0010843. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 131.Mitchell S.A., Spriggs K.A., Bushell M., Evans J.R., Stoneley M., Le Quesne J.P.C., Spriggs R.V., Willis A.E. Identification of a motif that mediates polypyrimidine tract-binding protein-dependent internal ribosome entry. Genes Dev. 2005;19:1556–1571. doi: 10.1101/gad.339105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 132.Bushell M., Stoneley M., Kong Y.W., Hamilton T.L., Spriggs K.A., Dobbyn H.C., Qin X., Sarnow P., Willis A.E. Polypyrimidine tract binding protein regulates IRES-mediated gene expression during apoptosis. Mol. Cell. 2006;23:401–412. doi: 10.1016/j.molcel.2006.06.012. [DOI] [PubMed] [Google Scholar]
- 133.Dobbyn H.C., Hill K., Hamilton T.L., Spriggs K.A., Pickering B.M., Coldwell M.J., de Moor C.H., Bushell M., Willis A.E. Regulation of BAG-1 IRES-mediated translation following chemotoxic stress. Oncogene. 2008;27:1167–1174. doi: 10.1038/sj.onc.1210723. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 134.Castelo-Branco P., Furger A., Wollerton M., Smith C., Moreira A., Proudfoot N. Polypyrimidine tract binding protein modulates efficiency of polyadenylation. Mol. Cell. Biol. 2004;24:4174–4183. doi: 10.1128/MCB.24.10.4174-4183.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 135.Pautz A., Linker K., Hubrich T., Korhonen R., Altenhöfer S., Kleinert H. The polypyrimidine tract-binding protein (PTB) is involved in the post-transcriptional regulation of human inducible nitric oxide synthase expression. J. Biol. Chem. 2006;281:32294–32302. doi: 10.1074/jbc.M603915200. [DOI] [PubMed] [Google Scholar]
- 136.Ma S., Liu G., Sun Y., Xie J. Relocalization of the polypyrimidine tract-binding protein during PKA-induced neurite growth. Biochim. Biophys. Acta. 2007;1773:912–923. doi: 10.1016/j.bbamcr.2007.02.006. [DOI] [PubMed] [Google Scholar]
- 137.Millevoi S., Decorsière A., Loulergue C., Iacovoni J., Bernat S., Antoniou M., Vagner S. A physical and functional link between splicing factors promotes pre-mRNA 3′ end processing. Nucleic Acids Res. 2009;37:4672–4683. doi: 10.1093/nar/gkp470. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 138.Gama-Carvalho M., Barbosa-Morais N.L., Brodsky A.S., Silver P.A., Carmo-Fonseca M. Genome-wide identification of functionally distinct subsets of cellular mRNAs associated with two nucleocytoplasmic-shuttling mammalian splicing factors. Genome Biol. 2006;7:R113. doi: 10.1186/gb-2006-7-11-r113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 139.Zang W.Q., Li B., Huang P.Y., Lai M.M., Yen T.S. Role of polypyrimidine tract binding protein in the function of the hepatitis B virus posttranscriptional regulatory element. J. Virol. 2001;75:10779–10786. doi: 10.1128/JVI.75.22.10779-10786.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 140.Fred R.G., Welsh N. The importance of RNA binding proteins in preproinsulin mRNA stability. Mol. Cell. Endocrinol. 2009;297:28–33. doi: 10.1016/j.mce.2008.06.007. [DOI] [PubMed] [Google Scholar]
- 141.Knoch K.-P., Bergert H., Borgonovo B., Saeger H.-D., Altkrüger A., Verkade P., Solimena M. Polypyrimidine tract-binding protein promotes insulin secretory granule biogenesis. Nat. Cell Biol. 2004;6:207–214. doi: 10.1038/ncb1099. [DOI] [PubMed] [Google Scholar]
- 142.Galbán S., Kuwano Y., Pullmann R., Martindale J.L., Kim H.H., LaI A., Abdelmohsen K., Yang X., Dang Y., Liu J.O., Lewis S.M., et al. RNA-binding proteins HuR and PTB promote the translation of hypoxia-inducible factor 1alpha. Mol. Cell. Biol. 2008;28:93–107. doi: 10.1128/MCB.00973-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 143.Wang M.J., Lin S. A region within the 5′-untranslated region of hypoxia-inducible factor-1alpha mRNA mediates its turnover in lung adenocarcinoma cells. J. Biol. Chem. 2009;284:36500–36510. doi: 10.1074/jbc.M109.008904. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 144.Gorospe M., Tominaga K., Wu X., Fähling M., Ivan M. Post-transcriptional control of the hypoxic response by RNA-binding proteins and microRNAs. Front. Mol. Neurosci. 2011;4:7. doi: 10.3389/fnmol.2011.00007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 145.Kosinski P.A., Laughlin J., Singh K., Covey L.R. A complex containing polypyrimidine tract-binding protein is involved in regulating the stability of CD40 ligand (CD154) mRNA. J. Immunol. 2003;170:979–988. doi: 10.4049/jimmunol.170.2.979. [DOI] [PubMed] [Google Scholar]
- 146.Porter J.F., Vavassori S., Covey L.R. A polypyrimidine tract-binding protein-dependent pathway of mRNA stability initiates with CpG activation of primary B cells. J. Immunol. 2008;181:3336–3345. doi: 10.4049/jimmunol.181.5.3336. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 147.Vavassori S., Covey L.R. Post-transcriptional regulation in lymphocytes: The case of CD154. RNA Biol. 2009;6:259–265. doi: 10.4161/rna.6.3.8581. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 148.Matus-Nicodemos R., Vavassori S., Castro-Faix M., Valentin-Acevedo A., Singh K., Marcelli V., Covey L.R. Polypyrimidine tract-binding protein is critical for the turnover and subcellular distribution of CD40 ligand mRNA in CD4+ T cells. J. Immunol. 2011;186:2164–2171. doi: 10.4049/jimmunol.1003236. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 149.Reyes R., Izquierdo J.M. The RNA-binding protein PTB exerts translational control on 3′-untranslated region of the mRNA for the ATP synthase beta-subunit. Biochem. Biophys. Res. Commun. 2007;357:1107–1112. doi: 10.1016/j.bbrc.2007.04.088. [DOI] [PubMed] [Google Scholar]
- 150.Moreira A., Takagaki Y., Brackenridge S., Wollerton M., Manley J.L., Proudfoot N.J. The upstream sequence element of the C2 complement poly(A) signal activates mRNA 3′ end formation by two distinct mechanisms. Genes Dev. 1998;12:2522–2534. doi: 10.1101/gad.12.16.2522. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 151.Danckwardt S., Kaufmann I., Gentzel M., Foerstner K.U., Gantzert A.-S., Gehring N.H., Neu-Yilik G., Bork P., Keller W., Wilm M., et al. Splicing factors stimulate polyadenylation via USEs at non-canonical 3′ end formation signals. EMBO J. 2007;26:2658–2669. doi: 10.1038/sj.emboj.7601699. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 152.Hall-Pogar T., Liang S., Hague L.K., Lutz C.S. Specific trans-acting proteins interact with auxiliary RNA polyadenylation elements in the COX-2 3′-UTR. RNA. 2007;13:1103–1115. doi: 10.1261/rna.577707. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 153.Woo K.-C., Kim T.-D., Lee K.-H., Kim D.-Y., Kim W., Lee K.-Y., Kim K.-T. Mouse period 2 mRNA circadian oscillation is modulated by PTB-mediated rhythmic mRNA degradation. Nucleic Acids Res. 2009;37:26–37. doi: 10.1093/nar/gkn893. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 154.Besse F., López de Quinto S., Marchand V., Trucco A., Ephrussi A. Drosophila PTB promotes formation of high-order RNP particles and represses oskar translation. Genes Dev. 2009;23:195–207. doi: 10.1101/gad.505709. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 155.Lewis R.A., Gagnon J.A., Mowry K.L. PTB/hnRNP I is required for RNP remodeling during RNA localization in Xenopus oocytes. Mol. Cell. Biol. 2008;28:678–686. doi: 10.1128/MCB.00999-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 156.Shibayama M., Ohno S., Osaka T., Sakamoto R., Tokunaga A., Nakatake Y., Sato M., Yoshida N. Polypyrimidine tract-binding protein is essential for early mouse development and embryonic stem cell proliferation. FEBS J. 2009;276:6658–6668. doi: 10.1111/j.1742-4658.2009.07380.x. [DOI] [PubMed] [Google Scholar]
- 157.Llorian M., Smith C.W.J. Decoding muscle alternative splicing. Curr. Opin. Genet. Dev. 2011;21:380–387. doi: 10.1016/j.gde.2011.03.006. [DOI] [PubMed] [Google Scholar]
- 158.Hall M.P., Nagel R.J., Fagg W.S., Shiue L., Cline M.S., Perriman R.J., Donohue J.P., Ares M. Quaking and PTB control overlapping splicing regulatory networks during muscle cell differentiation. RNA. 2013;19:627–638. doi: 10.1261/rna.038422.113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 159.Lin J.-C., Tarn W.-Y. Exon selection in alpha-tropomyosin mRNA is regulated by the antagonistic action of RBM4 and PTB. Mol. Cell. Biol. 2005;25:10111–10121. doi: 10.1128/MCB.25.22.10111-10121.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 160.Lin J.-C., Tarn W.-Y. RBM4 down-regulates PTB and antagonizes its activity in muscle cell-specific alternative splicing. J. Cell Biol. 2011;193:509–520. doi: 10.1083/jcb.201007131. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 161.Zhang J., Bahi N., Llovera M., Comella J.X., Sanchis D. Polypyrimidine tract binding proteins (PTB) regulate the expression of apoptotic genes and susceptibility to caspase-dependent apoptosis in differentiating cardiomyocytes. Cell Death Differ. 2009;16:1460–1468. doi: 10.1038/cdd.2009.87. [DOI] [PubMed] [Google Scholar]
- 162.Makeyev E.V., Zhang J., Carrasco M.A., Maniatis T. The MicroRNA miR-124 promotes neuronal differentiation by triggering brain-specific alternative pre-mRNA splicing. Mol. Cell. 2007;27:435–448. doi: 10.1016/j.molcel.2007.07.015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 163.Lilleväli K., Kulla A., Ord T. Comparative expression analysis of the genes encoding polypyrimidine tract binding protein (PTB) and its neural homologue (brPTB) in prenatal and postnatal mouse brain. Mech. Dev. 2001;101:217–220. doi: 10.1016/s0925-4773(00)00566-9. [DOI] [PubMed] [Google Scholar]
- 164.Bitel C.L., Perrone-Bizzozero N.I., Frederikse P.H. HuB/C/D, nPTB, REST4, and miR-124 regulators of neuronal cell identity are also utilized in the lens. Mol. Vis. 2010;16:2301–2316. [PMC free article] [PubMed] [Google Scholar]
- 165.Xue Y., Ouyang K., Huang J., Zhou Y., Ouyang H., Li H., Wang G., Wu Q., Wei C., Bi Y., et al. Direct conversion of fibroblasts to neurons by reprogramming PTB-regulated microRNA circuits. Cell. 2013;152:82–96. doi: 10.1016/j.cell.2012.11.045. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 166.Fred R.G., Sandberg M., Pelletier J., Welsh N. The human insulin mRNA is partly translated via a cap- and eIF4A-independent mechanism. Biochem. Biophys. Res. Commun. 2011;412:693–698. doi: 10.1016/j.bbrc.2011.08.030. [DOI] [PubMed] [Google Scholar]
- 167.Engels B., Jannot G., Remenyi J., Simard M.J., Hutvagner G. Polypyrimidine tract binding protein (hnRNP I) is possibly a conserved modulator of miRNA-mediated gene regulation. PLoS One. 2012;7:e33144. doi: 10.1371/journal.pone.0033144. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 168.He X., Pool M., Darcy K.M., Lim S.B., Auersperg N., Coon J.S., Beck W.T. Knockdown of polypyrimidine tract-binding protein suppresses ovarian tumor cell growth and invasiveness in vitro. Oncogene. 2007;26:4961–4968. doi: 10.1038/sj.onc.1210307. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 169.Wang C., Norton J.T., Ghosh S., Kim J., Fushimi K., Wu J.Y., Stack M.S., Huang S. Polypyrimidine tract-binding protein (PTB) differentially affects malignancy in a cell line-dependent manner. J. Biol. Chem. 2008;283:20277–20287. doi: 10.1074/jbc.M803682200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 170.Pollock C., Daily K., Nguyen V.T., Wang C., Lewandowska M.A., Bensaude O., Huang S. Characterization of MRP RNA-protein interactions within the perinucleolar compartment. Mol. Biol. Cell. 2011;22:858–867. doi: 10.1091/mbc.E10-09-0768. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 171.He X., Arslan A.D., Pool M.D., Ho T.-T., Darcy K.M., Coon J.S., Beck W.T. Knockdown of splicing factor SRp20 causes apoptosis in ovarian cancer cells and its expression is associated with malignancy of epithelial ovarian cancer. Oncogene. 2011;30:356–365. doi: 10.1038/onc.2010.426. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 172.Khan D., Sharathchandra A., Ponnuswamy A., Grover R., Das S. Effect of a natural mutation in the 5′ untranslated region on the translational control of p53 mRNA. Oncogene. 2013;32:4148–4159. doi: 10.1038/onc.2012.422. [DOI] [PubMed] [Google Scholar]
- 173.De Hoog C.L., Foster L.J., Mann M. RNA and RNA binding proteins participate in early stages of cell spreading through spreading initiation centers. Cell. 2004;117:649–662. doi: 10.1016/s0092-8674(04)00456-8. [DOI] [PubMed] [Google Scholar]
- 174.Babic I., Sharma S., Black D.L. A role for polypyrimidine tract binding protein in the establishment of focal adhesions. Mol. Cell. Biol. 2009;29:5564–5577. doi: 10.1128/MCB.00590-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 175.Cheung H.C., Hai T., Zhu W., Baggerly K.A., Tsavachidis S., Krahe R., Cote G.J. Splicing factors PTBP1 and PTBP2 promote proliferation and migration of glioma cell lines. Brain. 2009;132:2277–2288. doi: 10.1093/brain/awp153. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 176.Chen M., Zhang J., Manley J.L. Turning on a fuel switch of cancer: hnRNP proteins regulate alternative splicing of pyruvate kinase mRNA. Cancer Res. 2010;70:8977–8980. doi: 10.1158/0008-5472.CAN-10-2513. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 177.Clower C.V., Chatterjee D., Wang Z., Cantley L.C., vander Heiden M.G., Krainer A.R. The alternative splicing repressors hnRNP A1/A2 and PTB influence pyruvate kinase isoform expression and cell metabolism. Proc. Natl. Acad. Sci. USA. 2010;107:1894–1899. doi: 10.1073/pnas.0914845107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 178.David C.J., Chen M., Assanah M., Canoll P., Manley J.L. HnRNP proteins controlled by c-Myc deregulate pyruvate kinase mRNA splicing in cancer. Nature. 2010;463:364–368. doi: 10.1038/nature08697. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 179.Lin S., Wang M.J., Tseng K.-Y. Polypyrimidine tract-binding protein induces p19(Ink4d) expression and inhibits the proliferation of H1299 cells. PLoS One. 2013;8:e58227. doi: 10.1371/journal.pone.0058227. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 180.Cobbold L.C., Wilson L.A., Sawicka K., King H.A., Kondrashov A.V., Spriggs K.A., Bushell M., Willis A.E. Upregulated c-myc expression in multiple myeloma by internal ribosome entry results from increased interactions with and expression of PTB-1 and YB-1. Oncogene. 2010;29:2884–2891. doi: 10.1038/onc.2010.31. [DOI] [PubMed] [Google Scholar]
- 181.Arslan A.D., He X., Wang M., Rumschlag-Booms E., Rong L., Beck W.T. A high-throughput assay to identify small-molecule modulators of alternative pre-mRNA splicing. J. Biomol. Screen. 2013;18:180–190. doi: 10.1177/1087057112459901. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 182.Bian Y., Masuda A., Matsuura T., Ito M., Okushin K., Engel A.G., Ohno K. Tannic acid facilitates expression of the polypyrimidine tract binding protein and alleviates deleterious inclusion of CHRNA1 exon P3A due to an hnRNP H-disrupting mutation in congenital myasthenic syndrome. Hum. Mol. Genet. 2009;18:1229–1237. doi: 10.1093/hmg/ddp023. [DOI] [PMC free article] [PubMed] [Google Scholar]