Abstract
The rRNAs constitute the catalytic and structural components of the ribosome, the protein synthesis machinery of cells. The level of rRNA synthesis, mediated by Pol I (RNA polymerase I), therefore has a major impact on the life and destiny of a cell. In order to elucidate how cells achieve the stringent control of Pol I transcription, matching the supply of rRNA to demand under different cellular growth conditions, it is essential to understand the components and mechanics of the Pol I transcription machinery. In this review, we discuss: (i) the molecular composition and functions of the Pol I enzyme complex and the two main Pol I transcription factors, SL1 (selectivity factor 1) and UBF (upstream binding factor); (ii) the interplay between these factors during pre-initiation complex formation at the rDNA promoter in mammalian cells; and (iii) the cellular control of the Pol I transcription machinery.
Introduction
The Pol I (RNA polymerase I) transcription machinery comprises three main components dedicated to the transcription of the rRNA genes: the Pol I enzyme, the TBP (TATA-binding protein)–TAF (TBP-associated factor) complex SL1 (selectivity factor 1)/TIF-IB (transcription initiation factor-IB) and the transactivator protein UBF (upstream binding factor) (reviewed in [1–6]). Confined to the nucleolar subcompartment of the nucleus, this machinery directs transcription from the rDNA promoter at some 200–400 rRNA (ribosomal RNA) genes in the nucleolar organizer regions. The Pol I transcription machinery churns out 50% of the nascent RNA in the cell in the form of pre-rRNA (47 S precursor rRNA), which is processed into three mature rRNAs (18 S, 5.8 S and 28 S), and these, together with the 5 S rRNA synthesized by Pol III, comprise the enzymatic and structural scaffold of the ribosome [7]. Transcription of the rRNA genes drives the biogenesis of the several millions of ribosomes needed in an actively growing cell, thereby dictating the overall level of cellular protein synthesis. Thus transcription by Pol I is essential for, and inextricably linked to, cell growth and, hence, to normal cell division (reviewed in [6,8]). Predictably, therefore, Pol I transcription is at the receiving end of many of the signalling pathways implicated in the control of cell growth, nutrient sensing and proliferation [1,6,9–17].
The Pol I enzyme complexes
DNA-dependent RNA polymerases are highly conserved multisubunit enzyme complexes in eukaryotes. The composition of the Pol I enzyme complex has been derived primarily from yeast genetic and biochemical studies [18]. Our understanding of the Pol I enzyme complex in mammalian cells has advanced rapidly in recent years following identification of mammalian subunits both by comparison with yeast Pol I and by MS analysis of purified Pol I complexes. Pol I enzymes from yeast to humans show conservation with Pols II and III and with the α2ββ′ω subunit composition of prokaryotic core RNA polymerase (reviewed in [19]) (Table 1). The largest and second largest subunits of Pol I share substantial similarity with those of Pol II and Pol III and with prokaryotic β′ and β subunits respectively, possessing most of the enzymatic functions (substrate binding, active centre, template movement). The heterodimer AC40–AC19 of Pol I is shared with Pol III, and is homologous with the RPB3–RPB11 heterodimer of Pol II and functionally analogous to the prokaryotic α2 homodimer. The RPB6 subunit of Pol I, shared with Pols II and III, is a structural and functional homologue of the bacterial ω subunit. There are four additional subunits that are required to synthesize RNA from a non-specific DNA template, shared between Pols I, II and III (RPB5, RPB8, RPB10 and RPB12). Subunit A12.2 is unique to Pol I (Table 1). In addition to these 10 subunits, collectively referred to as the ‘core’ complex, Pol I heterodimer A14–A43 shares genetic, biochemical and structural characteristics with heterodimers RPB4–RPB7 and C17–C25 of Pols II and III respectively, and there is sequence similarity between subunits RPB7, A43 and C25 [20–22]. There are two additional subunits, A49 and A34.5, that are specific to Pol I [18,23,24] and hence likely targets of the Pol I cognate transcription factors.
Table 1. Comparison of Pol I subunit composition in yeast and humans.
Most Pol I subunits are essential for viability in Saccharomyces cerevisiae (as determined by null mutant analysis); those subunits not essential are marked viable (v) or conditional mutants (c) (adapted from [18]). ‘Unique or shared’ refers to subunits unique in Pol I, or shared with Pol III, or shared with Pols II and III. Human Pol I subunits are named primarily following yeast nomenclature
| S. cerevisiae Pol I subunits | Unique or shared | Human Pol I subunits | Homologues* | Interactions in mammalian PIC |
|---|---|---|---|---|
| RPA190 | I | hRPA190 (A194) | β′; RPB1 (B220) | |
| RPA135 | I | hRPA127 | β; RPB2 (B140) | |
| RPA49c | I | PAF53 | None | UBF [29] |
| RPA43 | I | hRPA43 | RPB7 | hRRN3 [41] |
| RPA40 (AC40) | I, III | hRPA40 | α; RPB3 (B45) | |
| RPA34.5v | I | CAST (PAF49) | None | UBF and SL1 [25,26]† |
| RPB5 (ABC27) | I, II, III | hRPB5 | – | |
| RPB6 (ABC23) | I, II, III | hRPB6 | ω | |
| RPA19 (AC19) | I, III | hRPA19 | α; RPB11(B12.5) | |
| RPB8 (ABC14.5) | I, II, III | RPB8 | – | |
| RPA14v | I | ? | RPB4 | |
| RPA12c | I | hRPA12.2 | RPB9 | |
| RPB10 (ABC10β) | I, II, III | RPB10 | – | |
| RPB12 (ABC10α) | I, II, III | RPB12 | – |
Structural and/or functional homologues with bacterial RNA polymerase (β′, β, α2, ω) and yeast Pol II (B).
K.I. Panov, T.B. Panova and J.C.B.M. Zomerdijk, unpublished work.
The ~600 kDa yeast Pol I complex is therefore composed of 14 subunits, and homologues of all of the yeast Pol I subunits are found in Metazoa, with the exception of the Pol I-specific subunit A14, for which a mammalian homologue has yet to be identified. Mammalian homologues of the Pol I-specific subunit A34.5 were discovered only recently: the mouse homologue is PAF49 (polymerase I-associated factor of 49 kDa) [25]. Human Pol I proteomics led us to identify a 72 kDa subunit of Pol I with homology to the yeast Pol I subunit A34.5 (K.I. Panov, T.B. Panova and J.C.B.M. Zomerdijk, unpublished work), previously named ASE-1 [antisense for ERCC-1 (excision repair cross-complementing-1)] [26] or CAST (CD3ε-associated signal transducer) [27] and, curiously, also implicated in T-cell activation (Table 1). The A34.5 and A49 subunits are absent from yeast Pol I isolated under high-salt conditions (referred to as A*), which displays a reduced specific activity in RNA synthesis from calf thymus DNA and a higher sensitivity to α-amanitin, suggesting a role for these subunits in elongation or processivity [28]. PAF49 and PAF53, the mammalian homologues of yeast A34.5 and A49, also dissociate from Pol I under certain conditions, most notably upon serum starvation of cells [25,29].
The structures of yeast Pol I and Pol II complexes have been studied extensively by electron microscopy and crystallography [30–33]. The cryo-electron-microscopic structure of yeast Pol I displays an overall similarity with the crystal structure of yeast Pol II (lacking RPB4 and RPB7), except at the positions of the Pol I-specific subunits A34.5, A49, A43 and A14 [33]. The A34.5 subunit of Pol I is positioned adjacent to the second largest subunit A135, at the entry to the cleft which binds the DNA template [31], where it is proposed to stabilize the interaction of the DNA template with the ‘core’ enzyme. The A49 subunit is located on the head region of the clamp, formed by the largest subunit of Pol I [33,34] and constituting one side of the cleft. A49 could potentially affect the conformation of the clamp [33], moving it inwards to hold downstream DNA more firmly, or interact directly with the DNA, thereby increasing the processivity of the enzyme [30,31]. Another structure specific to Pol I is the stalk, formed by subunits A43 and A14 and postulated to function as an interface for the assembly of initiation factors through its binding to the essential Pol I-associated factor Rrn3p. Interestingly, the stalk protrudes from the Pol I structure at the same site as the C-terminal domain of the largest subunit in Pol II, which has a role in initiation via its interaction with the basal transcription factors [33]. There are different conformational subpopulations of the yeast Pol I enzyme [34], consistent with conformational flexibility in other RNA polymerases, where the clamp maintains an open conformation in the initiation-competent enzyme, being converted into a closed conformation during elongation [19,34].
We have shown previously that the Pol I enzyme in human cells is found in at least two functionally distinct complexes (Pol Iα and Pol Iβ). These complexes are considerably larger than the yeast Pol I complex, at >1 MDa, and contain Pol I-associated factors in addition to the core subunits [35] (K.I. Panov and J.C.B.M. Zomerdijk, unpublished work). Pol Iβ, comprising approx. 10% of the total Pol I extractable from the cell nucleus, is the initiation-competent form of the enzyme, able to direct accurate initiation of transcription from the rDNA promoter. By contrast, Pol Iα, representing the bulk of the Pol I, is unable to direct rDNA promoter-specific transcription, although it does possess Pol activity, catalysing the random synthesis of RNA. Given these properties and its relative abundance compared with Pol Iβ, the Pol Iα complex might represent the elongating Pol I complex, or polymerase with the capacity to be converted readily into initiation-competent Pol Iβ, or spent polymerase complexes following transcription. The Pol Iβ enzyme complex is distinguishable by its associated factors, likely to be involved specifically in initiation. hRRN3 (human RRN3; equivalent to mouse TIF-IA/yeast Rrn3p) [35–39] is one such Pol Iβ-specific factor. RRN3 is tethered to the A43 subunit of Pol I [40,41] and links Pol I to the promoter-bound essential transcription factor SL1. The role of RRN3 in bridging the polymerase and promoter recognition components of the Pol I transcription machinery (SL1 or core factor) is conserved from yeast to humans, and is required for PIC (pre-initiation complex) assembly [35,37,38].
The association of hRRN3/TIF-IA with Pol I, to generate Pol Iβ [35,36], is an important point of control in pre-rRNA synthesis. The hRRN3/TIF-IA–Pol I interaction is impaired and cellular rRNA synthesis is down-regulated in stationary-phase, nutrient-limited or cycloheximide-treated mammalian cells [41,42]. There is evidence that rDNA transcription is regulated by the phosphorylation state of RRN3, which determines the steady-state concentration of the RRN3–Pol I complex Pol Iβ [41]. Perhaps as a result of a reversal of phosphorylation events, Rrn3 is inactivated during the process of rDNA transcription [43]. The phosphorylation of TIF-IA by JNK2 (c-Jun N-terminal kinase 2), which can occur upon cellular stress, leads to inactivation of TIF-IA and down-regulation of rRNA synthesis [44]. RRN3 activity and the RRN3–Pol I interaction are also regulated by the ERK (extracellular-signal-regulated kinase)/MAPK (mitogen-activated protein kinase) and mTOR (mammalian target of rapamycin) signalling pathways, which are involved in growth factor signalling and nutrient sensing. Increased RRN3/TIF-IA activity, rRNA gene transcription and cell proliferation correlate with phosphorylation of RRN3/TIF-IA by ERK and RSK (p90 ribosomal S6 kinase) in growth factor-stimulated mouse NIH 3T3 cells [10]. In separate studies, RRN3 activity has been reported to be down-regulated [12] or unaffected [9] following inactivation of mTOR by rapamycin treatment of NIH 3T3 cells. Perhaps, therefore, the specific physiology of a cell determines the involvement of mTOR in the regulation of RRN3 activity. The mTOR pathway has been implicated in PIC formation, as the down-regulation of RRN3 by rapamycin was correlated with changes in the phosphorylation status of RRN3, an inability of RRN3 to interact with Pol I and SL1, and translocation of TIF-IA/RRN3 to the cytoplasm [12]. Furthermore, inactivation of yeast TOR by rapamycin is accompanied by release of Pol I from the nucleolus and inhibition of rDNA transcription [45], and the TOR signalling pathway has been shown to regulate the Rrn3p–Pol I interaction and the Rrn3-dependent recruitment of Pol I to the promoter [14]. Curiously, in yeast, phosphorylation of Pol I, not Rrn3p, is necessary for the formation of a stable Pol I–Rrn3p complex for efficient transcription initiation in vitro, and a change in the phosphorylation state of Pol I is correlated with the association of Pol I with Rrn3p in vivo [46].
TBP–TAF complex SL1
RNA polymerase enzymes themselves have no intrinsic ability to recognize and bind specifically to promoter DNA sequences. Therefore PIC formation in transcription calls for the recruitment of the polymerases to the promoter via transcription factors. Sigma factors fulfil this role in bacteria, and basal or general transcription factors co-operate in this function in eukaryotes, where the three classes of highly related enzymes, Pol I, Pol II and Pol III, catalyse the transcription of specific sets of genes. A complex of TBP and TAFs is required for the accurate initiation of transcription by all three eukaryotic RNA polymerases [47], with few exceptions. The complement of TAFs in each complex, although variable in Pol II transcription, is specific to each class of genes. Binding to TBP of TAF subunits specific to one class of polymerases can preclude the binding of TAFs from a different class [48]. The TBP–TAF complexes specific to Pols I, II and III perform distinct roles in mediating a specific interaction between the polymerases and their respective promoters.
In transcription by mammalian Pol II, the TBP–TAF complex TFIID (transcription factor IID) can bind the promoter through interaction of the TBP protein with TATA boxes and of the TAF proteins with other promoter sequence elements. Pol II and other factors are recruited subsequently to form the PIC [49]. In transcription by Pol III, the initial phases converge on the recruitment of TBP–TAF complex TFIIIB and the Pol III enzyme in formation of the PIC [23,50], although the type of Pol III promoter determines whether loading of promoter-type specific TFIIIB at the promoter DNA is facilitated by the specific DNA-binding abilities of TFIIIA and TFIIIC, TFIIIC alone, or the multisubunit complex PBP–PTF–SNAPC.
In transcription by mammalian Pol I (reviewed in [6]), the TBP–TAF complex SL1 (murine TIF-IB) is essential [48,51–53]. This ~300 kDa complex is composed of TBP and at least three TAFs [48,51,52], TAFI48, TAFI63 and TAFI110 (mouse TAFI48, TAFI68 and TAFI95 respectively). Reconstitution of transcription in vitro using recombinant SL1 comprising only these four components is inefficient, suggesting that more components and/or specific modifications might be required to reproduce the full activity of SL1 [54]. We have recent evidence for the existence of another TAFI in SL1 (J.J. Gorski, S. Pathak and J.C.B.M. Zomerdijk, unpublished work). SL1 functions through the essential core element of the rDNA promoter, overlapping the start site of transcription [55]. SL1 alone does not produce a DNase I footprint, but a combination of SL1 and UBF extends the footprint of UBF alone, demonstrating co-operativity between these proteins at the promoter [56,57].
SL1 directs promoter- and species-specific Pol I transcription. The SL1 complexes of humans and mice are not interchangeable between these systems, in contrast with Pol I and the activator UBF. These findings imply a specific interaction between SL1 and its cognate rDNA promoter (reviewed in [58]). Indeed, we have shown recently that human SL1 can bind the human rDNA promoter independently and direct the initiation of transcription specifically from the human rDNA promoter in the absence of UBF [59], consistent with the findings that TIF-IB and rat SL1 can bind and direct transcription from their cognate rDNA promoters independently of UBF [60,61].
In addition to promoter recognition and binding, SL1 has an essential role in the recruitment of Pol I to the start site of transcription [35,59]. SL1 subunits TAFI63 and TAFI110 interact with the Pol I-associated RRN3 [35–37], which in turn is attached to Pol I through subunit A43 in the ‘stalk’ structure of the enzyme complex and, perhaps, PAF67 [41,42].
Another function for SL1 in PIC formation has been reported recently: we found that SL1 can stabilize UBF binding at the rDNA promoter [59]. In the absence of SL1, UBF associates with and dissociates from the rDNA rapidly, whereas the presence of SL1 at the rDNA promoter alters significantly the binding equilibrium of UBF, thereby increasing the residence time of UBF at the promoter. It is possible that the increased lifetime of the rDNA–UBF complex is essential to facilitate the UBF-mediated activation of Pol I transcription.
Collectively, these results support a unified model for SL1 function in the mammalian system, where SL1 drives or nucleates PIC formation specifically at the rDNA promoter, leading ultimately to the productive initiation of transcription by Pol I (Figure 1). SL1 remains stably bound to the rDNA promoter following initiation of transcription and escape by Pol I, and the SL1 complex, together with UBF at the promoter, might serve as a re-initiation scaffold to support multiple rounds of transcription [62].
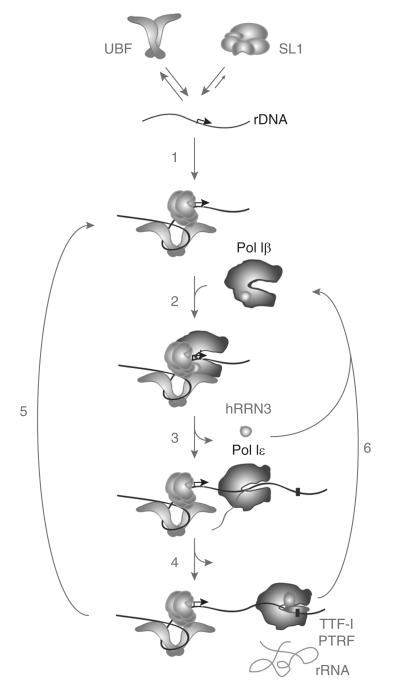
Figure 1. The Pol I transcription cycle.
The TBP–TAF complex SL1 interacts stably with the rDNA promoter, near the start site of transcription (black arrow). Dimers of UBF interact dynamically with rDNA promoter elements (grey arrows), such that UBF–rDNA complexes might be relatively short-lived. UBF and SL1 interact co-operatively at the promoter elements, and SL1 reduces the dissociation of UBF from the rDNA, thereby increasing the residence time of UBF at the promoter (1). rDNA promoter topology is altered dramatically by UBF binding. SL1 in the UBF–SL1–rDNA complex recruits the initiation-competent Pol I–hRRN3 complex Pol Iβ through the interactions of TAFI subunits with hRRN3. This culminates in formation of the complete PIC, poised for initiation of transcription (2). UBF stimulates the promoter escape/clearance of Pol I. In the transition from initiation to elongation, hRRN3 dissociates from Pol I, and the Pol I enzyme is converted into a transcript-elongating enzyme complex, Pol Iε (3). SL1 and UBF remain promoter-bound following promoter clearance by Pol I (4) and, thus, can function as a re-initiation scaffold (5), to allow multiple rounds of transcription from the same rDNA promoter. Pol Iε transcription terminates and rRNA is released at termination sites (black rectangle on rDNA); this process is directed by TTF-I (transcription termination factor-I) and PTRF (polymerase I and transcript release factor). Following termination of transcription, it is possible that the spent Pol I complex is converted back into initiation-competent Pol Iβ and thence recruited to the rDNA promoter by SL1 to initiate a new round of transcription (6).
PIC assembly could perhaps be regulated through SL1, influencing the level of transcription by Pol I. Indeed, an increase in SL1 occupancy of the rDNA promoter correlates with increased Pol I transcription in IGF-1 (insulin-like growth factor-1)-stimulated HEK293 cells [13]. Although phosphorylation of SL1 in response to growth stimulatory factors has not yet been reported, some of the SL1 subunits are known to be phosphoproteins. SL1 activity can be controlled by acetylation, as acetylation of the mouse SL1 (TIF-IB) subunit TAFI68 by P/CAF [p300/CBP (CREB-binding protein)-associated factor] stimulates binding of SL1 to the rDNA promoter and Pol I-mediated transcription [63], and it is possible that this acetylation is a regulated, rather than constitutive, event in cells. Furthermore, SL1 could be a limiting component of rRNA gene expression in cells, and therefore an increase in its abundance would stimulate Pol I transcription. Although tissue-specific or differential expression of the TAFs of SL1 has not been reported as yet, regulated expression of these Pol I-specific TAFI subunits could impact on transcription by Pol I. Up-regulation of TBP expression, following activation of the MAPK signalling pathway, can lead to stimulation of transcription from Pol I and Pol III promoters in certain cell types, implying that overexpression of this single subunit can drive the biogenesis of the separate TBP–TAF complexes required in the co-ordinated transcription of rRNA and tRNA genes [64,65].
Transcription activator UBF
Reconstituted transcription assays have revealed the components minimally required for transcription initiation. ‘Basal’ levels of transcription from the rDNA promoter can be supported by SL1 and initiation-competent Pol Iβ alone [60,61]. UBF efficiently activates rDNA transcription above this basal level. Stimulated transcription by UBF in the reconstituted system constitutes ‘true’ or ‘net’ activation, as distinct from stimulation of transcription through the potential anti-repression function of UBF in the context of chromatin transcription in vivo [66–68].
UBF is proposed to function as an architectural protein at the rDNA promoter [69,70], where it binds as a dimer to the UCE (upstream control element), which has a modulatory role in Pol I transcription, and to the essential core element. UBF (97 kDa) is composed of several distinct domains [69,70]. The N-terminal dimerization domain of UBF is essential for its transactivation function [69,71]. The large central portion contains six HMG (high-mobility group) boxes [72,73] which exhibit a relaxed sequence specificity of DNA binding and display the ability to bend DNA [69,74–79]. In UBF, these HMG boxes do not contribute equally to DNA binding; rather, it is thought that HMG boxes 1–3 interact with the DNA minor groove in the interaction of UBF with the rDNA promoter [74,79]. Binding of a UBF dimer induces bending of the DNA, to create a loop of some 140 bp of DNA in a single 360° turn, generating a structure termed the enhancesome [78]. It has been proposed that tandem enhancesomes at the rDNA promoter would juxtapose the UCE and core promoter elements [78,79]. There is co-operativity of binding of UBF and SL1 to the UCE and the core elements of the rDNA promoter [56,57], although the precise arrangement and stoichiometry of UBF and SL1 at the rDNA promoter remain unknown. Interaction between UBF and SL1 [69,80,81] requires the highly acidic C-terminal domain of UBF [69,82,83] and may involve subunits TAFI48 and TBP of SL1 [84,85]. UBF also interacts with Pol I [29,86] through Pol I subunits ASE1/CAST and PAF53 [29,87] (K.I. Panov, T.B. Panova and J.C.B.M. Zomerdijk, unpublished work) (Table 1).
These interactions and the co-operativity demonstrated between UBF and SL1 at the rDNA promoter led to the proposal that UBF recruits SL1 and Pol I to the rDNA promoter, activating transcription by facilitating PIC assembly. This is the principal mechanism used by activators of Pol II transcription [88]. However, in contrast with the sequence-specific Pol II transcriptional activators, binding of UBF occurs rather indiscriminately [75,76] and extensively throughout the rDNA repeats in vivo [89,90], inconsistent with a role for UBF in nucleating PIC assembly specifically at the rDNA promoter. Moreover, as discussed, SL1 alone can nucleate PIC formation [59–61], stabilizing UBF binding at the rDNA promoter [59] and recruiting initiation-competent Pol I to the core promoter [35].
Recently, using a highly purified and reconstituted transcription system, combined with an immobilized ribosomal promoter DNA template, we have defined a specific and novel function for UBF in activating the rate of RNA synthesis at promoter escape or clearance by Pol I (K.I. Panov, J.K. Friedrich, J. Russell and J.C.B.M. Zomerdijk, unpublished work), a step we had identified previously as the rate-limiting step for Pol I transcription in vitro [62]. This mechanism enables UBF to activate transcription both from previously inactive promoters following PIC assembly and from SL1-engaged promoters at each successive round of transcription following re-initiation (Figure 1). Stimulation of promoter escape/clearance by activators might also make an important contribution to efficient transcription by Pols II and III.
Given its role in activating transcription, any variation in UBF expression levels or activity would be anticipated to have a dramatic effect on the level of transcription by Pol I. Indeed, UBF, limiting in some cells [91], is expressed at relatively high levels in cardiac hypertrophy and liver cancer cells, which support high levels of rRNA synthesis [92,93]. In mitogen-stimulated lymphocytes, UBF overexpression is apparent prior to increased rRNA synthesis, and a decrease in UBF expression occurs upon induced differentiation [94]. UBF expression can be up-regulated by serum induction [9], operating through the mTOR pathway, and also by the myc proto-oncogene [95].
How is the activity of the UBF protein regulated? Phosphorylation of UBF can affect its interaction with other PIC components or DNA. C-terminal phosphorylation of UBF increases its ability to interact with SL1 [69,82,83]. The protein kinase CK2 phosphorylates UBF at serine residues within the C-terminal acidic domain, and this contributes to transcriptional activation [96]. Phosphoinositide 3-kinase phosphorylates UBF predominantly in the C-terminus upon IGF-1 (insulin-like growth factor 1) treatment of serum-starved mouse cells, where IRS-1 (insulin receptor substrate-1) targets phosphoinositide 3-kinase to UBF following translocation of IRS-1 to the nucleolus [15], and this correlates with increased rRNA synthesis. C-terminal phosphorylation of UBF can also be mediated by the mTOR signalling pathway and requires S6 kinase 1. This occurs upon serum induction of rDNA transcription in NIH 3T3 cells and promotes the interaction between UBF and SL1 [9]. The ERK/MAPK pathway targets UBF upon stimulation of Pol I transcription by epidermal growth factor in human neuroepithelioma cells. ERK1/2-mediated phosphorylation in HMG boxes 1 and 2 alters the interaction of UBF with DNA [11]. UBF activity is also regulated by competitive acetylation/deacetylation. In this case, the ability of UBF to bind DNA is not affected [68]. Rather, binding of CBP acetyltransferase to UBF might up-regulate UBF by preventing the binding of the tumour suppressor Rb (retinoblastoma) (and associated histone deacetylases) to UBF, precluding the negative effect of Rb on the binding of UBF to SL1 [97].
Concluding remarks
From the evidence obtained so far, we conclude that the Pol I TBP–TAF complex SL1 performs three important functions that, together, drive PIC formation at the rDNA promoter. SL1 recognizes and binds the core promoter elements through its TAFI proteins. The SL1 complex, positioned accurately at the rDNA promoter, recruits the initiation-competent form of Pol I, the Pol I–RRN3 complex Pol Iβ, to the start site of transcription, through the interactions of TAFI110 and TAFI63 with RRN3. SL1 also stabilizes UBF at the rDNA promoter, and the consequential increase in residence time of UBF at the promoter is predicted to enhance the efficacy of UBF in its role as an activator of Pol I transcription. The dramatic alterations in DNA architecture created by UBF at the rDNA promoter are likely to be important for its activation function, although the significance of these has yet to be realized. We consider that the interaction of UBF with subunits of Pol I is crucial for the ability of UBF to stimulate the escape of Pol I from the promoter in the transition state between initiation and elongation of transcription.
Pol I transcription can be regulated at the levels of PIC formation and UBF activity. These are influenced by the downstream effectors of growth factor-responsive signalling pathways (including the MAPK, mTOR, and ERK pathways), the availability of nutrients (mTOR) and/or the cellular stress response pathways (JNK2). The level of rRNA transcription activity has a direct effect on the overall level of cellular protein synthesis, thereby influencing the growth of the cell and cell division (reviewed in [6]). There is a correlation between the level of cellular rRNA transcription and the growth status of cells, in that slow-growing or differentiated cells display low levels of rRNA transcription, whereas rapidly growing and cancerous cells maintain high levels of rRNA synthesis. Down-regulation of Pol I transcription can lead to the activation of the p53 cellular response pathway, and therefore directly influence the cell’s decision to arrest or exit from the cell cycle in response to DNA damage, stress or malfunction [98]. Up-regulation of rRNA transcription, with the growth and proliferative advantage it bestows on cells, could represent an early event in oncogenesis; significantly, the oncoprotein Myc has been shown recently to control cellular rRNA synthesis [99–101].
Acknowledgments
We thank present and past members of our laboratory for their research and interest. We apologize to our colleagues whose work could not be cited due to space limitations. The Wellcome Trust funds research in the laboratory of J.C.B.M.Z., and J.C.B.M.Z. is a Wellcome Trust Senior Research Fellow in the Basic Biomedical Sciences.
References
- 1.Grummt I. Genes Dev. 2003;17:1691–1702. doi: 10.1101/gad.1098503R. [DOI] [PubMed] [Google Scholar]
- 2.Moss T, Stefanovsky VY. Cell. 2002;109:545–548. doi: 10.1016/s0092-8674(02)00761-4. [DOI] [PubMed] [Google Scholar]
- 3.Reeder RH. Prog. Nucleic Acid Res. Mol. Biol. 1999;62:293–327. doi: 10.1016/s0079-6603(08)60511-5. [DOI] [PubMed] [Google Scholar]
- 4.Hannan KM, Hannan RD, Rothblum LI. Front. Biosci. 1998;3:d376–d398. doi: 10.2741/a282. [DOI] [PubMed] [Google Scholar]
- 5.Comai L. Adv. Protein Chem. 2004;67:123–155. doi: 10.1016/S0065-3233(04)67005-7. [DOI] [PubMed] [Google Scholar]
- 6.Russell J, Zomerdijk JC. Trends Biochem. Sci. 2005;30:87–96. doi: 10.1016/j.tibs.2004.12.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Moore PB, Steitz TA. Nature (London) 2002;418:229–235. doi: 10.1038/418229a. [DOI] [PubMed] [Google Scholar]
- 8.Warner JR. Trends Biochem. Sci. 1999;24:437–440. doi: 10.1016/s0968-0004(99)01460-7. [DOI] [PubMed] [Google Scholar]
- 9.Hannan KM, Brandenburger Y, Jenkins A, Sharkey K, Cavanaugh A, Rothblum L, Moss T, Poortinga G, McArthur GA, Pearson RB, et al. Mol. Cell. Biol. 2003;23:8862–8877. doi: 10.1128/MCB.23.23.8862-8877.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Zhao J, Yuan X, Frodin M, Grummt I. Mol. Cell. 2003;11:405–413. doi: 10.1016/s1097-2765(03)00036-4. [DOI] [PubMed] [Google Scholar]
- 11.Stefanovsky VY, Pelletier G, Hannan R, Gagnon-Kugler T, Rothblum LI, Moss T. Mol. Cell. 2001;8:1063–1073. doi: 10.1016/s1097-2765(01)00384-7. [DOI] [PubMed] [Google Scholar]
- 12.Mayer C, Zhao J, Yuan X, Grummt I. Genes Dev. 2004;18:423–434. doi: 10.1101/gad.285504. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.James MJ, Zomerdijk JC. J. Biol. Chem. 2004;279:8911–8918. doi: 10.1074/jbc.M307735200. [DOI] [PubMed] [Google Scholar]
- 14.Claypool JA, French SL, Johzuka K, Eliason K, Vu L, Dodd JA, Beyer AL, Nomura M. Mol. Biol. Cell. 2004;15:946–956. doi: 10.1091/mbc.E03-08-0594. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Drakas R, Tu X, Baserga R. Proc. Natl. Acad. Sci. U.S.A. 2004;101:9272–9276. doi: 10.1073/pnas.0403328101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Sun H, Tu X, Prisco M, Wu A, Casiburi I, Baserga R. Mol. Endocrinol. 2003;17:472–486. doi: 10.1210/me.2002-0276. [DOI] [PubMed] [Google Scholar]
- 17.Moss T. Curr. Opin. Genet. Dev. 2004;14:210–217. doi: 10.1016/j.gde.2004.02.005. [DOI] [PubMed] [Google Scholar]
- 18.Carles C, Riva M. In: Transcription of Ribosomal RNA Genes by Eukaryotic RNA Polymerase I. Paule MR, editor. Springer-Verlag; Berlin, Heidelberg, New York: 1998. pp. 9–38. [Google Scholar]
- 19.Cramer P. Curr. Opin. Struct. Biol. 2002;12:89–97. doi: 10.1016/s0959-440x(02)00294-4. [DOI] [PubMed] [Google Scholar]
- 20.Peyroche G, Levillain E, Siaut M, Callebaut I, Schultz P, Sentenac A, Riva M, Carles C. Proc. Natl. Acad. Sci. U.S.A. 2002;99:14670–14675. doi: 10.1073/pnas.232580799. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Meka H, Daoust G, Arnvig KB, Werner F, Brick P, Onesti S. Nucleic Acids Res. 2003;31:4391–4400. doi: 10.1093/nar/gkg652. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Siaut M, Zaros C, Levivier E, Ferri ML, Court M, Werner M, Callebaut I, Thuriaux P, Sentenac A, Conesa C. Mol. Cell. Biol. 2003;23:195–205. doi: 10.1128/MCB.23.1.195-205.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Geiduschek EP, Kassavetis GA. J. Mol. Biol. 2001;310:1–26. doi: 10.1006/jmbi.2001.4732. [DOI] [PubMed] [Google Scholar]
- 24.Chedin S, Ferri ML, Peyroche G, Andrau JC, Jourdain S, Lefebvre O, Werner M, Carles C, Sentenac A. Cold Spring Harbor Symp. Quant. Biol. 1998;63:381–389. doi: 10.1101/sqb.1998.63.381. [DOI] [PubMed] [Google Scholar]
- 25.Yamamoto K, Yamamoto M, Hanada K, Nogi Y, Matsuyama T, Muramatsu M. Mol. Cell. Biol. 2004;24:6338–6349. doi: 10.1128/MCB.24.14.6338-6349.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Whitehead CM, Winkfein RJ, Fritzler MJ, Rattner JB. Chromosoma. 1997;106:493–502. doi: 10.1007/s004120050271. [DOI] [PubMed] [Google Scholar]
- 27.Yamazaki T, Hamano Y, Tashiro H, Itoh K, Nakano H, Miyatake S, Saito T. J. Biol. Chem. 1999;274:18173–18180. doi: 10.1074/jbc.274.26.18173. [DOI] [PubMed] [Google Scholar]
- 28.Huet J, Buhler JM, Sentenac A, Fromageot P. Proc. Natl. Acad. Sci. U.S.A. 1975;72:3034–3038. doi: 10.1073/pnas.72.8.3034. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Hanada K, Song CZ, Yamamoto K, Yano K, Maeda Y, Yamaguchi K, Muramatsu M. EMBO J. 1996;15:2217–2226. [PMC free article] [PubMed] [Google Scholar]
- 30.Cramer P, Bushnell DA, Kornberg RD. Science. 2001;292:1863–1876. doi: 10.1126/science.1059493. [DOI] [PubMed] [Google Scholar]
- 31.Gnatt AL, Cramer P, Fu J, Bushnell DA, Kornberg RD. Science. 2001;292:1876–1882. doi: 10.1126/science.1059495. [DOI] [PubMed] [Google Scholar]
- 32.Kettenberger H, Armache KJ, Cramer P. Mol. Cell. 2004;16:955–965. doi: 10.1016/j.molcel.2004.11.040. [DOI] [PubMed] [Google Scholar]
- 33.Bischler N, Brino L, Carles C, Riva M, Tschochner H, Mallouh V, Schultz P. EMBO J. 2002;21:4136–4144. doi: 10.1093/emboj/cdf392. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.De Carlo S, Carles C, Riva M, Schultz P. J. Mol. Biol. 2003;329:891–902. doi: 10.1016/s0022-2836(03)00510-2. [DOI] [PubMed] [Google Scholar]
- 35.Miller G, Panov KI, Friedrich JK, Trinkle-Mulcahy L, Lamond AI, Zomerdijk JC. EMBO J. 2001;20:1373–1382. doi: 10.1093/emboj/20.6.1373. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Bodem J, Dobreva G, Hoffmann-Rohrer U, Iben S, Zentgraf H, Delius H, Vingron M, Grummt I. EMBO Rep. 2000;1:171–175. doi: 10.1093/embo-reports/kvd032. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Moorefield B, Greene EA, Reeder RH. Proc. Natl. Acad. Sci. U.S.A. 2000;97:4724–4729. doi: 10.1073/pnas.080063997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Peyroche G, Milkereit P, Bischler N, Tschochner H, Schultz P, Sentenac A, Carles C, Riva M. EMBO J. 2000;19:5473–5482. doi: 10.1093/emboj/19.20.5473. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Yamamoto RT, Nogi Y, Dodd JA, Nomura M. EMBO J. 1996;15:3964–3973. [PMC free article] [PubMed] [Google Scholar]
- 40.Thuriaux P, Mariotte S, Buhler JM, Sentenac A, Vu L, Lee BS, Nomura M. J. Biol. Chem. 1995;270:24252–24257. doi: 10.1074/jbc.270.41.24252. [DOI] [PubMed] [Google Scholar]
- 41.Cavanaugh AH, Hirschler-Laszkiewicz I, Hu Q, Dundr M, Smink T, Misteli T, Rothblum LI. J. Biol. Chem. 2002;277:27423–27432. doi: 10.1074/jbc.M201232200. [DOI] [PubMed] [Google Scholar]
- 42.Yuan X, Zhao J, Zentgraf H, Hoffmann-Rohrer U, Grummt I. EMBO Rep. 2002;3:1082–1087. doi: 10.1093/embo-reports/kvf212. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Hirschler-Laszkiewicz I, Cavanaugh AH, Mirza A, Lun M, Hu Q, Smink T, Rothblum LI. J. Biol. Chem. 2003;278:18953–18959. doi: 10.1074/jbc.M301093200. [DOI] [PubMed] [Google Scholar]
- 44.Mayer C, Bierhoff H, Grummt I. Genes Dev. 2005;19:933–941. doi: 10.1101/gad.333205. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Tsang CK, Bertram PG, Ai W, Drenan R, Zheng XF. EMBO J. 2003;22:6045–6056. doi: 10.1093/emboj/cdg578. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Fath S, Milkereit P, Peyroche G, Riva M, Carles C, Tschochner H. Proc. Natl. Acad. Sci. U.S.A. 2001;98:14334–14339. doi: 10.1073/pnas.231181398. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Hernandez N. Genes Dev. 1993;7:1291–1308. doi: 10.1101/gad.7.7b.1291. [DOI] [PubMed] [Google Scholar]
- 48.Comai L, Zomerdijk JC, Beckmann H, Zhou S, Admon A, Tjian R. Science. 1994;266:1966–1972. doi: 10.1126/science.7801123. [DOI] [PubMed] [Google Scholar]
- 49.Verrijzer CP, Tjian R. Trends Biochem. Sci. 1996;21:338–342. [PubMed] [Google Scholar]
- 50.Schramm L, Hernandez N. Genes Dev. 2002;16:2593–2620. doi: 10.1101/gad.1018902. [DOI] [PubMed] [Google Scholar]
- 51.Comai L, Tanese N, Tjian R. Cell. 1992;68:965–976. doi: 10.1016/0092-8674(92)90039-f. [DOI] [PubMed] [Google Scholar]
- 52.Zomerdijk JCBM, Beckmann H, Comai L, Tjian R. Science. 1994;266:2015–2018. doi: 10.1126/science.7801130. [DOI] [PubMed] [Google Scholar]
- 53.Zomerdijk JCBM, Tjian R. In: Transcription of Eukaryotic Ribosomal RNA Genes by RNA Polymerase I. Paule MR, editor. Springer-Verlag; Berlin, Heidelberg, New York: 1998. pp. 67–73. [Google Scholar]
- 54.Heix J, Zomerdijk J, Ravanpay A, Tjian R, Grummt I. Proc. Natl. Acad. Sci. U.S.A. 1997;94:1733–1738. doi: 10.1073/pnas.94.5.1733. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Zomerdijk JCBM, Tjian R. In: Transcription of Ribosomal RNA Genes by Eukaryotic RNA Polymerase I. Paule MR, editor. Springer-Verlag; Berlin, Heidelberg, New York: 1998. pp. 121–134. [Google Scholar]
- 56.Learned RM, Learned TK, Haltiner MM, Tjian RT. Cell. 1986;45:847–857. doi: 10.1016/0092-8674(86)90559-3. [DOI] [PubMed] [Google Scholar]
- 57.Bell SP, Learned RM, Jantzen HM, Tjian R. Science. 1988;241:1192–1197. doi: 10.1126/science.3413483. [DOI] [PubMed] [Google Scholar]
- 58.Heix J, Grummt I. Curr. Opin. Genet. Dev. 1995;5:652–656. doi: 10.1016/0959-437x(95)80035-2. [DOI] [PubMed] [Google Scholar]
- 59.Friedrich JK, Panov KI, Cabart P, Russell J, Zomerdijk JC. J. Biol. Chem. 2005;280:29551–29558. doi: 10.1074/jbc.M501595200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Schnapp A, Grummt I. J. Biol. Chem. 1991;266:24588–24595. [PubMed] [Google Scholar]
- 61.Smith SD, O’Mahony DJ, Kinsella BT, Rothblum LI. Gene Expression. 1993;3:229–236. [PMC free article] [PubMed] [Google Scholar]
- 62.Panov KI, Friedrich JK, Zomerdijk JC. Mol. Cell. Biol. 2001;21:2641–2649. doi: 10.1128/MCB.21.8.2641-2649.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Muth V, Nadaud S, Grummt I, Voit R. EMBO J. 2001;20:1353–1362. doi: 10.1093/emboj/20.6.1353. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Zhong S, Zhang C, Johnson DL. Mol. Cell. Biol. 2004;24:5119–5129. doi: 10.1128/MCB.24.12.5119-5129.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.White RJ. Oncogene. 2004;23:3208–3216. doi: 10.1038/sj.onc.1207547. [DOI] [PubMed] [Google Scholar]
- 66.Kuhn A, Grummt I. Proc. Natl. Acad. Sci. U.S.A. 1992;89:7340–7344. doi: 10.1073/pnas.89.16.7340. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Brou C, Kuhn A, Staub A, Chaudhary S, Grummt I, Davidson I, Tora L. Nucleic Acids Res. 1993;21:4011–4018. doi: 10.1093/nar/21.17.4011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Pelletier G, Stefanovsky VY, Faubladier M, Hirschler-Laszkiewicz I, Savard J, Rothblum LI, Cote J, Moss T. Mol. Cell. 2000;6:1059–1066. doi: 10.1016/s1097-2765(00)00104-0. [DOI] [PubMed] [Google Scholar]
- 69.Jantzen HM, Chow AM, King DS, Tjian R. Genes Dev. 1992;6:1950–1963. doi: 10.1101/gad.6.10.1950. [DOI] [PubMed] [Google Scholar]
- 70.Reeder RH, Pikaard CS, McStay B. In: Nucleic Acids and Molecular Biology. Eckstein F, Lilley DMJ, editors. vol. 9. Springer-Verlag; Berlin, Heidelberg, New York: 1995. pp. 251–263. [Google Scholar]
- 71.McStay B, Frazier MW, Reeder RH. Genes Dev. 1991;5:1957–1968. doi: 10.1101/gad.5.11.1957. [DOI] [PubMed] [Google Scholar]
- 72.Bachvarov D, Moss T. Nucleic Acids Res. 1991;19:2331–2335. doi: 10.1093/nar/19.9.2331. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Jantzen HM, Admon A, Bell SP, Tjian R. Nature (London) 1990;344:830–836. doi: 10.1038/344830a0. [DOI] [PubMed] [Google Scholar]
- 74.Leblanc B, Read C, Moss T. EMBO J. 1993;12:513–525. doi: 10.1002/j.1460-2075.1993.tb05683.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Copenhaver GP, Putnam CD, Denton ML, Pikaard CS. Nucleic Acids Res. 1994;22:2651–2657. doi: 10.1093/nar/22.13.2651. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Hu CH, McStay B, Jeong SW, Reeder RH. Mol. Cell. Biol. 1994;14:2871–2882. doi: 10.1128/mcb.14.5.2871. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Putnam CD, Copenhaver GP, Denton ML, Pikaard CS. Mol. Cell. Biol. 1994;14:6476–6488. doi: 10.1128/mcb.14.10.6476. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Bazett Jones DP, Leblanc B, Herfort M, Moss T. Science. 1994;264:1134–1137. doi: 10.1126/science.8178172. [DOI] [PubMed] [Google Scholar]
- 79.Stefanovsky VY, Pelletier G, Bazett-Jones DP, Crane-Robinson C, Moss T. Nucleic Acids Res. 2001;29:3241–3247. doi: 10.1093/nar/29.15.3241. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Hempel WM, Cavanaugh AH, Hannan RD, Taylor L, Rothblum LI. Mol. Cell. Biol. 1996;16:557–563. doi: 10.1128/mcb.16.2.557. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Bodeker M, Cairns C, McStay B. Mol. Cell. Biol. 1996;16:5572–5578. doi: 10.1128/mcb.16.10.5572. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Kihm AJ, Hershey JC, Haystead TA, Madsen CS, Owens GK. Proc. Natl. Acad. Sci. U.S.A. 1998;95:14816–14820. doi: 10.1073/pnas.95.25.14816. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.Tuan JC, Zhai W, Comai L. Mol. Cell. Biol. 1999;19:2872–2879. doi: 10.1128/mcb.19.4.2872. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Beckmann H, Chen JL, O’Brien T, Tjian R. Science. 1995;270:1506–1509. doi: 10.1126/science.270.5241.1506. [DOI] [PubMed] [Google Scholar]
- 85.Kwon H, Green MR. J. Biol. Chem. 1994;269:30140–30146. [PubMed] [Google Scholar]
- 86.Schnapp G, Santori F, Carles C, Riva M, Grummt I. EMBO J. 1994;13:190–199. doi: 10.1002/j.1460-2075.1994.tb06248.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87.Seither P, Zatsepina O, Hoffmann M, Grummt I. Chromosoma. 1997;106:216–225. doi: 10.1007/s004120050242. [DOI] [PubMed] [Google Scholar]
- 88.Ptashne M, Gann A. Nature (London) 1997;386:569–577. doi: 10.1038/386569a0. [DOI] [PubMed] [Google Scholar]
- 89.O’Sullivan AC, Sullivan GJ, McStay B. Mol. Cell. Biol. 2002;22:657–668. doi: 10.1128/MCB.22.2.657-668.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 90.Mais C, Wright JE, Prieto JL, Raggett SL, McStay B. Genes Dev. 2005;19:50–64. doi: 10.1101/gad.310705. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91.Hannan R, Stefanovsky V, Arino T, Rothblum L, Moss T. Nucleic Acids Res. 1999;27:1205–1213. doi: 10.1093/nar/27.4.1205. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92.Brandenburger Y, Jenkins A, Autelitano DJ, Hannan RD. FASEB J. 2001;15:2051–2053. doi: 10.1096/fj.01-0853fje. [DOI] [PubMed] [Google Scholar]
- 93.Huang R, Wu T, Xu L, Liu A, Ji Y, Hu G. FASEB J. 2002;16:293–301. doi: 10.1096/fj.01-0687com. [DOI] [PubMed] [Google Scholar]
- 94.Cabart P, Kalousek I. Cell Mol. Biol. (Noisy-Le-Grand) 1998;44:343–350. [PubMed] [Google Scholar]
- 95.Poortinga G, Hannan KM, Snelling H, Walkley CR, Jenkins A, Sharkey K, Wall M, Brandenburger Y, Palatsides M, Pearson RB, et al. EMBO J. 2004;23:3325–3335. doi: 10.1038/sj.emboj.7600335. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 96.Voit R, Kuhn A, Sander EE, Grummt I. Nucleic Acids Res. 1995;23:2593–2599. doi: 10.1093/nar/23.14.2593. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 97.Hannan KM, Kennedy BK, Cavanaugh AH, Hannan RD, Hirschler-Laszkiewicz I, Jefferson LS, Rothblum LI. Oncogene. 2000;19:3487–3497. doi: 10.1038/sj.onc.1203690. [DOI] [PubMed] [Google Scholar]
- 98.Yuan X, Zhou Y, Casanova E, Chai M, Kiss E, Grone H-J, Schutz G, Grummt I. Mol. Cell. 2005;19:77–87. doi: 10.1016/j.molcel.2005.05.023. [DOI] [PubMed] [Google Scholar]
- 99.Grandori C, Gomez-Roman N, Felton-Edkins ZA, Ngouenet C, Galloway DA, Eisenman RN, White RJ. Nat. Cell Biol. 2005;7:311–318. doi: 10.1038/ncb1224. [DOI] [PubMed] [Google Scholar]
- 100.Arabi A, Wu S, Ridderstrale K, Bierhoff H, Shiue C, Fatyol K, Fahlen S, Hydbring P, Soderberg O, Grummt I, et al. Nat. Cell Biol. 2005;7:303–310. doi: 10.1038/ncb1225. [DOI] [PubMed] [Google Scholar]
- 101.Grewal SS, Li L, Orian A, Eisenman RN, Edgar BA. Nat. Cell Biol. 2005;7:295–302. doi: 10.1038/ncb1223. [DOI] [PubMed] [Google Scholar]