Abstract
To determine whether a mammalian cytochrome c could efficiently replace iso-1-cytochrome c, which is encoded by the yeast CYC1 gene, the coding sequence of RC9 (a nondefective processed gene from rat) was cloned in both single- and multiple-copy expression vectors under the direction of the yeast alcohol dehydrogenase 1 (ADC1) promoter. Upon transformation of a CYC1 deletion strain, the multiple-copy construct restored a wild-type growth rate on lactate medium; such growth normally requires derepressed amounts of iso-1-cytochrome c. These transformants expressed a level of hybrid ADC1/RC9 mRNA approximately 5- to 10-fold greater than the amount of message from the endogenous ADC1 gene and produced a steady-state level of rat cytochrome c equivalent to that of the wild-type yeast protein. A requirement for the vector was evidenced by its absence in all transformants that lost the lactate growth phenotype after propagation in nonselective medium. In contrast, the level of vector-specific message in single copy was equivalent to that of the endogenous ADC1 mRNA, but transformants exhibited no significant growth on lactate. Constructions having a small deletion or a mammalian intron within the rat cytochrome c coding region failed to support lactate-dependent growth, indicating that complementation depends upon proper translation of the correct rat coding sequence. Therefore, the rat polypeptide, when expressed at normal physiological levels, is recognized by the yeast machinery involved in the multiple steps required for the processing and transport of an active cytochrome c as well as its functional interaction with the respiratory apparatus.
Full text
PDF

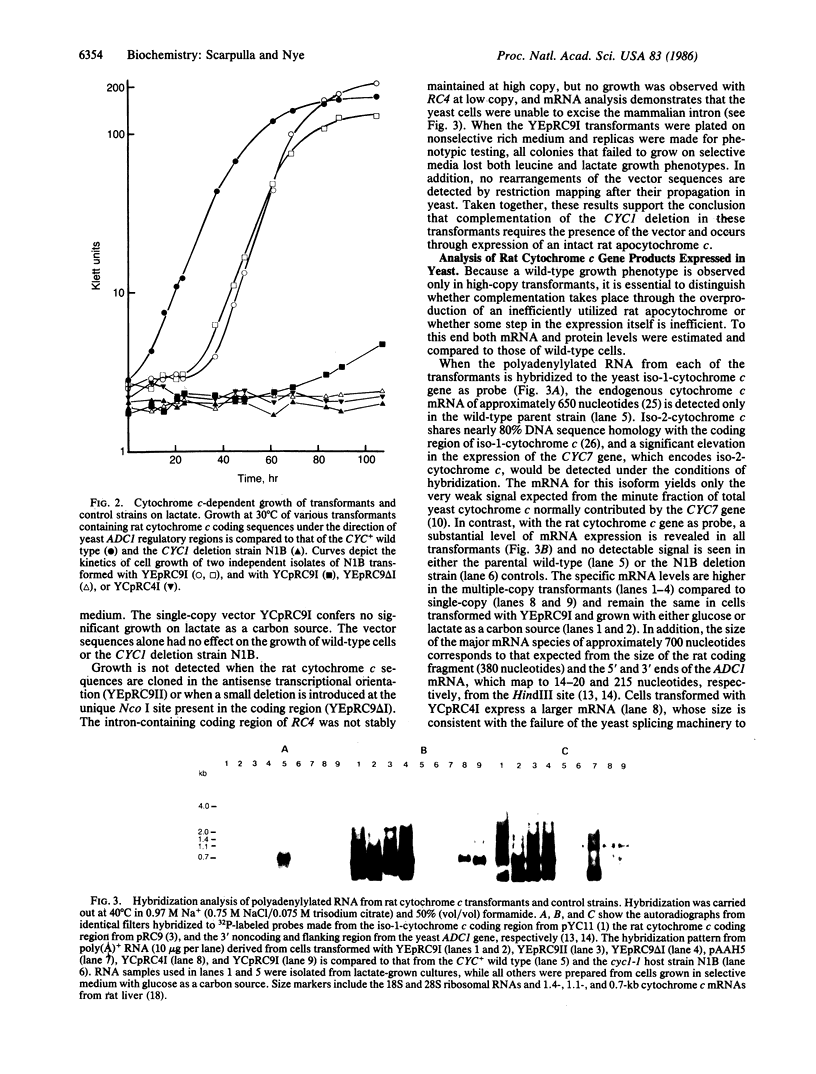


Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Ammerer G. Expression of genes in yeast using the ADCI promoter. Methods Enzymol. 1983;101:192–201. doi: 10.1016/0076-6879(83)01014-9. [DOI] [PubMed] [Google Scholar]
- Aviv H., Leder P. Purification of biologically active globin messenger RNA by chromatography on oligothymidylic acid-cellulose. Proc Natl Acad Sci U S A. 1972 Jun;69(6):1408–1412. doi: 10.1073/pnas.69.6.1408. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Baum S. G., Horwitz M. S., Maizel J. V., Jr Studies of the mechanism of enhancement of human adenovirus infection in monkey cells by simian virus 40. J Virol. 1972 Aug;10(2):211–219. doi: 10.1128/jvi.10.2.211-219.1972. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bennetzen J. L., Hall B. D. The primary structure of the Saccharomyces cerevisiae gene for alcohol dehydrogenase. J Biol Chem. 1982 Mar 25;257(6):3018–3025. [PubMed] [Google Scholar]
- Boss J. M., Gillam S., Zitomer R. S., Smith M. Sequence of the yeast iso-1-cytochrome c mRNA. J Biol Chem. 1981 Dec 25;256(24):12958–12961. [PubMed] [Google Scholar]
- Carlson M., Botstein D. Two differentially regulated mRNAs with different 5' ends encode secreted with intracellular forms of yeast invertase. Cell. 1982 Jan;28(1):145–154. doi: 10.1016/0092-8674(82)90384-1. [DOI] [PubMed] [Google Scholar]
- Carlson S. S., Mross G. A., Wilson A. C., Mead R. T., Wolin L. D., Bowers S. F., Foley N. T., Muijsers A. O., Margoliash E. Primary structure of mouse, rat, and guinea pig cytochrome c. Biochemistry. 1977 Apr 5;16(7):1437–1442. doi: 10.1021/bi00626a031. [DOI] [PubMed] [Google Scholar]
- Denis C. L., Ferguson J., Young E. T. mRNA levels for the fermentative alcohol dehydrogenase of Saccharomyces cerevisiae decrease upon growth on a nonfermentable carbon source. J Biol Chem. 1983 Jan 25;258(2):1165–1171. [PubMed] [Google Scholar]
- Hawkes R., Niday E., Gordon J. A dot-immunobinding assay for monoclonal and other antibodies. Anal Biochem. 1982 Jan 1;119(1):142–147. doi: 10.1016/0003-2697(82)90677-7. [DOI] [PubMed] [Google Scholar]
- Hennig B., Koehler H., Neupert W. Receptor sites involved in posttranslational transport of apocytochrome c into mitochondria: specificity, affinity, and number of sites. Proc Natl Acad Sci U S A. 1983 Aug;80(16):4963–4967. doi: 10.1073/pnas.80.16.4963. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hinnen A., Hicks J. B., Fink G. R. Transformation of yeast. Proc Natl Acad Sci U S A. 1978 Apr;75(4):1929–1933. doi: 10.1073/pnas.75.4.1929. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Johnston M., Davis R. W. Sequences that regulate the divergent GAL1-GAL10 promoter in Saccharomyces cerevisiae. Mol Cell Biol. 1984 Aug;4(8):1440–1448. doi: 10.1128/mcb.4.8.1440. [DOI] [PMC free article] [PubMed] [Google Scholar]
- LOWRY O. H., ROSEBROUGH N. J., FARR A. L., RANDALL R. J. Protein measurement with the Folin phenol reagent. J Biol Chem. 1951 Nov;193(1):265–275. [PubMed] [Google Scholar]
- Laemmli U. K. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature. 1970 Aug 15;227(5259):680–685. doi: 10.1038/227680a0. [DOI] [PubMed] [Google Scholar]
- Lutstorf U., Megnet R. Multiple forms of alcohol dehydrogenase in Saccharomyces cerevisiae. I. Physiological control of ADH-2 and properties of ADH-2 and ADH-4. Arch Biochem Biophys. 1968 Sep 10;126(3):933–944. doi: 10.1016/0003-9861(68)90487-6. [DOI] [PubMed] [Google Scholar]
- Montgomery D. L., Leung D. W., Smith M., Shalit P., Faye G., Hall B. D. Isolation and sequence of the gene for iso-2-cytochrome c in Saccharomyces cerevisiae. Proc Natl Acad Sci U S A. 1980 Jan;77(1):541–545. doi: 10.1073/pnas.77.1.541. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nolan C., Margoliash E. Comparative aspects of primary structures of proteins. Annu Rev Biochem. 1968;37:727–790. doi: 10.1146/annurev.bi.37.070168.003455. [DOI] [PubMed] [Google Scholar]
- Ohashi A., Gibson J., Gregor I., Schatz G. Import of proteins into mitochondria. The precursor of cytochrome c1 is processed in two steps, one of them heme-dependent. J Biol Chem. 1982 Nov 10;257(21):13042–13047. [PubMed] [Google Scholar]
- Scarpulla R. C., Agne K. M., Wu R. Cytochrome c gene-related sequences in mammalian genomes. Proc Natl Acad Sci U S A. 1982 Feb;79(3):739–743. doi: 10.1073/pnas.79.3.739. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Scarpulla R. C., Agne K. M., Wu R. Isolation and structure of a rat cytochrome c gene. J Biol Chem. 1981 Jun 25;256(12):6480–6486. [PubMed] [Google Scholar]
- Scarpulla R. C. Processed pseudogenes for rat cytochrome c are preferentially derived from one of three alternate mRNAs. Mol Cell Biol. 1984 Nov;4(11):2279–2288. doi: 10.1128/mcb.4.11.2279. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Scarpulla R. C., Wu R. Nonallelic members of the cytochrome c multigene family of the rat may arise through different messenger RNAs. Cell. 1983 Feb;32(2):473–482. doi: 10.1016/0092-8674(83)90467-1. [DOI] [PubMed] [Google Scholar]
- Sherman F., Stewart J. W., Parker J. H., Inhaber E., Shipman N. A., Putterman G. J., Gardisky R. L., Margoliash E. The mutational alteration of the primary structure of yeast iso-1-cytochrome c. J Biol Chem. 1968 Oct 25;243(20):5446–5456. [PubMed] [Google Scholar]
- Sherman F., Taber H., Campbell W. Genetic determination of iso-cytochromes c in yeast. J Mol Biol. 1965 Aug;13(1):21–39. doi: 10.1016/s0022-2836(65)80077-8. [DOI] [PubMed] [Google Scholar]
- Struhl K. The new yeast genetics. 1983 Sep 29-Oct 5Nature. 305(5933):391–397. doi: 10.1038/305391a0. [DOI] [PubMed] [Google Scholar]
- Thomas P. S. Hybridization of denatured RNA and small DNA fragments transferred to nitrocellulose. Proc Natl Acad Sci U S A. 1980 Sep;77(9):5201–5205. doi: 10.1073/pnas.77.9.5201. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Towbin H., Staehelin T., Gordon J. Electrophoretic transfer of proteins from polyacrylamide gels to nitrocellulose sheets: procedure and some applications. Proc Natl Acad Sci U S A. 1979 Sep;76(9):4350–4354. doi: 10.1073/pnas.76.9.4350. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zuker M., Stiegler P. Optimal computer folding of large RNA sequences using thermodynamics and auxiliary information. Nucleic Acids Res. 1981 Jan 10;9(1):133–148. doi: 10.1093/nar/9.1.133. [DOI] [PMC free article] [PubMed] [Google Scholar]