Abstract
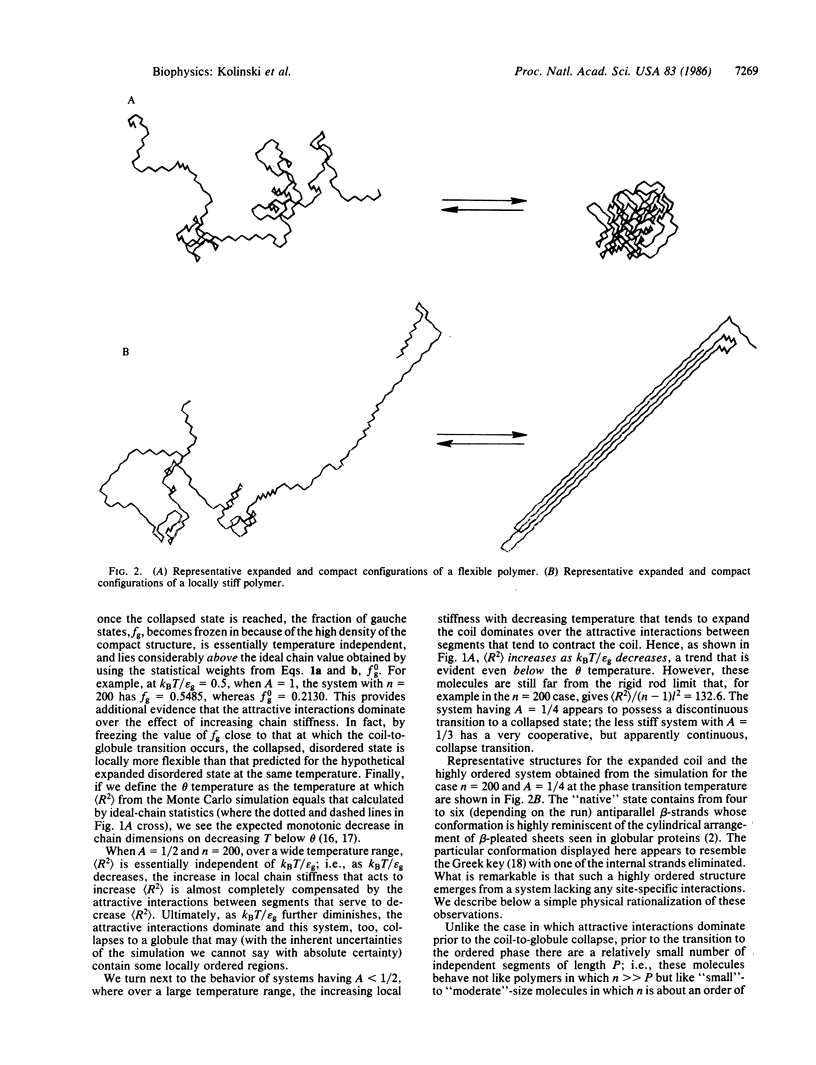
Monte Carlo simulations were performed on a diamond lattice, globular protein model in which the trans conformational state is energetically favored over the gauche states (thereby perhaps favoring a beta-sheet secondary structure) and in which nonspecific nonbonded nearest-neighbor attractive interactions are allowed. If the attractive interactions are sufficiently weak that the molecule possesses a relatively high fraction of trans states in the denatured state, then on collapse, a beta-barrel tertiary structure, highly reminiscent of the "native" structure seen in beta-proteins, spontaneously forms. If, however, the attractive interactions are dominant, a coil-to-random globule collapse transition is observed. The roles of short-, medium-, and long-range interactions and topological constraints in determining the observed tertiary structure are addressed, and the implications and limitations of the simulations for the equilibrium folding process in renal globular proteins are explored.
Full text
PDF



Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Chou P. Y., Fasman G. D. Beta-turns in proteins. J Mol Biol. 1977 Sep 15;115(2):135–175. doi: 10.1016/0022-2836(77)90094-8. [DOI] [PubMed] [Google Scholar]
- Dill K. A. Theory for the folding and stability of globular proteins. Biochemistry. 1985 Mar 12;24(6):1501–1509. doi: 10.1021/bi00327a032. [DOI] [PubMed] [Google Scholar]
- Levitt M., Chothia C. Structural patterns in globular proteins. Nature. 1976 Jun 17;261(5561):552–558. doi: 10.1038/261552a0. [DOI] [PubMed] [Google Scholar]
- Levitt M. Protein conformation, dynamics, and folding by computer simulation. Annu Rev Biophys Bioeng. 1982;11:251–271. doi: 10.1146/annurev.bb.11.060182.001343. [DOI] [PubMed] [Google Scholar]
- Lin S. H., Konishi Y., Denton M. E., Scheraga H. A. Influence of an extrinsic cross-link on the folding pathway of ribonuclease A. Conformational and thermodynamic analysis of cross-linked (lysine7-lysine41)-ribonuclease a. Biochemistry. 1984 Nov 6;23(23):5504–5512. doi: 10.1021/bi00318a019. [DOI] [PubMed] [Google Scholar]
- Miyazawa S., Jernigan R. L. Equilibrium folding and unfolding pathways for a model protein. Biopolymers. 1982 Jul;21(7):1333–1363. doi: 10.1002/bip.360210706. [DOI] [PubMed] [Google Scholar]
- Ptitsyn O. B., Finkelstein A. V. Similarities of protein topologies: evolutionary divergence, functional convergence or principles of folding? Q Rev Biophys. 1980 Aug;13(3):339–386. doi: 10.1017/s0033583500001724. [DOI] [PubMed] [Google Scholar]
- Richardson J. S. beta-Sheet topology and the relatedness of proteins. Nature. 1977 Aug 11;268(5620):495–500. doi: 10.1038/268495a0. [DOI] [PubMed] [Google Scholar]
- Rose G. D., Gierasch L. M., Smith J. A. Turns in peptides and proteins. Adv Protein Chem. 1985;37:1–109. doi: 10.1016/s0065-3233(08)60063-7. [DOI] [PubMed] [Google Scholar]
- Skolnick J. Role of topological constraints in the all-or-none transition of a globular protein model: theory of the helix-coil transition in doubly crosslinked, coiled coils. Biochem Biophys Res Commun. 1985 Jun 28;129(3):848–853. doi: 10.1016/0006-291x(85)91969-2. [DOI] [PubMed] [Google Scholar]
- Tanford C. Protein denaturation. Adv Protein Chem. 1968;23:121–282. doi: 10.1016/s0065-3233(08)60401-5. [DOI] [PubMed] [Google Scholar]
- Venkatachalam C. M. Stereochemical criteria for polypeptides and proteins. V. Conformation of a system of three linked peptide units. Biopolymers. 1968 Oct;6(10):1425–1436. doi: 10.1002/bip.1968.360061006. [DOI] [PubMed] [Google Scholar]


