Abstract
Ewing’s sarcomas are highly aggressive round cell tumors of bone and soft tissues that afflict children and young adults. The majority of these tumors harbor the t(11;22) translocation and express the fusion protein EWS-FLI. Modern molecular profiling experiments indicate that Ewing’s tumors originate from mesenchymal precursors in young individuals. EWS-FLI alters the morphology of mesenchymal cells and prevents lineage specification; however, the molecular mechanisms for differentiation arrest are unclear. We recently showed that EWS-FLI binds Runx2, a master regulator of osteoblast differentiation. In this report, we demonstrate that FLI sequences within EWS-FLI are responsible for interactions with Runx2. EWS-FLI blocks the expression of osteoblastic genes in a multipotent progenitor cell line that requires Runx2 to integrate bone morphogenic protein (Bmp)2 signaling while increasing proliferation and altering cell morphology. These results demonstrate that EWS-FLI blocks the ability of Runx2 to induce osteoblast specification of a mesenchymal progenitor cell. Disrupting interactions between Runx2 and EWS-FLI1 may promote differentiation of the tumor cell.
Keywords: EWSR1, FLI1, Cbfa1, AML-3, PEBP2aA, Ets, Bmp2
Introduction
Ewing’s sarcomas are aggressive, undifferentiated, round cell tumors of bone and soft tissue that cause pain and morbidity because they induce local osteolysis and have a high metastatic rate, which drastically worsens the long-term prognosis. This rare malignancy afflicts approximately three to five per million people under the age of 20. The current long-term survival rate for patients without overt metastases is 50%, but is only 25% for patients presenting with metastatic lesions and just 10% following recurrence [Jedlicka, 2010]. The cellular progenitor of Ewing’s sarcoma has been debated since in 1921 when James Ewing described the malignancy as a diffuse endothelioma [Ewing, 1921]. These tumors have also been historically classified as peripheral primitive neuroectodermal tumors, peripheral neuroepitheliomas, and Askin tumors. All of these entities harbor similar genetic mutations and recent molecular signature analyses suggest that they originate from mesenchymal and/or neural crest progenitors [Hancock and Lessnick, 2008; Kauer et al., 2009; Riggi et al., 2005; Staege et al., 2004; Suva et al., 2009; Tirode et al., 2007; Toomey et al., 2010].
The chromosomal translocation t(11:22)(q24:q212) is the most common genetic mutation detected in Ewing’s sarcomas and occurs in approximately 85% of cases [Turc-Carel et al., 1988]. It fuses EWSR1 (Ewings sarcoma breakpoint region 1) to the Ets transcription factor gene FLI1 (Friend leukemia integration 1) and creates the cancer-specific fusion protein EWS-FLI [Delattre et al., 1992]. Fusions between EWSR1 and other ETS family members, namely ERG, ETV1, ETV4 or FEV, are detected in the majority of the remaining 15% of Ewing’s sarcomas [Law et al., 2006; Tan and Manley, 2009]. The translocations fuse a transcriptional regulatory domain in the N-terminus of EWSR1 to the DNA binding domain of ETS factors. As a consequence, a large portion of FLI1, particularly its DNA binding domain, is now regulated by the EWSR1 promoter and is placed into a new structural context. The EWS-FLI fusion protein directly and indirectly activates and represses the expression of numerous target genes that maintain stem cell phenotypes, promote proliferation, survival and drug resistance, and block differentiation into mesenchymal tissues (reviewed in [Jedlicka, 2010; Toomey et al., 2010]).
Runx2 (Cbfa1, AML3, PEBPa2A) is a transcription factor required for osteoblast development from multipotent mesenchymal and neural crest cells [Komori et al., 1997; Otto et al., 1997]. Runx2 binds DNA via a highly conserved Runt domain and associates with the nuclear matrix [Zeng et al., 1998]. Runx2 is both an activator and repressor of transcription. It associates with numerous transcription factors, including ETS proteins [Fowler et al., 2005; Li et al., 2004; Sato et al., 1998], and transcriptional co-factors (e.g. p300 or Hdacs) to regulate gene expression in a context-dependent fashion [Jensen et al., 2010]. Runx2 is highly expressed in osteosarcomas, which are tumors derived from cells of a mature osteoblast phenotype, and is also easily detected in tumors that metastasize to the skeleton. We recently showed that Runx2 and Runx1 associate with TET family members, EWSR1, EWS-FLI and FUS, but the nature of those interactions remained unresolved [Li et al., 2010].
The recent observations that Ewing’s sarcomas are of mesenchymal and/or neural crest origin and that EWS-FLI associates with Runx2, a protein essential for the differentiation of these progenitor cells into osteoblasts, led to the hypothesis that EWS-FLI prevents osteoblast specification by blocking Runx2 activities. Previous studies demonstrated that EWS-FLI blocks osteocalcin expression in bone marrow stromal cells from Arf-deficient mice [Torchia et al., 2003] and that suppression of the t(11:22) product, EWS-FLI, in an Ewing sarcoma cell line by RNA interference increased the expression of osteoblast differentiation genes (e.g. Runx2 and SPP1/osteopontin) and permitted matrix mineralization [Tirode et al., 2007]. Moreover, transgenic mice expressing EWS-FLI in primitive mesenchymal cells exhibited impaired differentiation of numerous mesenchymal lineages [Lin et al., 2008]. These studies indicate that EWS-FLI promotes the maintenance of an undifferentiated state; however, the effects of EWS-FLI on osteogenic signaling pathways have not been extensively elucidated. In this report, we demonstrate that EWS-FLI associates with Runx2 via amino acids from FLI1 and that sustained EWS-FLI expression in a murine multipotent mesenchymal cell line alters the expression of numerous genes required for osteoblast differentiation.
Materials & Methods
GST Pulldown Assays
The GST-Runx2 proteins were previously described [Jensen et al., 2008; Li et al., 2009]. 35S-labeled Myc-EWSR1, FLAG-FLI, and EWS-FLI proteins were made in vitro from plasmids provided by Ralf Jacknecht [Rossow and Janknecht, 2001], Dennis Watson [Nowling et al., 2008], and Aykut Üren [Erkizan et al., 2009], respectively, using Promega’s TnT system. Pulldown experiments were performed as reported [Jensen et al., 2008; Li et al., 2009].
Immunofluorescence Experiments
U2-OS cells were grown on glass coverslips and transiently transfected with myc-EWSR1 (2–656) or Flag-FLI1 using Lipofectamine (Invitrogen) and analyzed after 48 hours. The cells were fixed in 4% paraformaldehyde for 20 min, permeabilized with 0.3% TritonX-100 in PBS for 5 min, blocked for 30 min in immunofluorescence buffer (3% BSA, 20mM MgCl2, 0.3% Tween 20 in PBS), and incubated with anti-Flag antibody (Sigma, F3165), anti-Myc antibody (Cell Signaling Technology, 2276), and anti-Runx2 in immunofluorescence buffer. Cells were washed three times with 0.1% TritonX-100 in PBS, incubated 30 min with Alexa-conjugated secondary antibodies at 1:800 (Invitrogen), washed three times, and mounted in Permount (Fisher Scientific). Images were obtained using a ZEISS LSM510 confocal microscope.
Luciferase Assays
C2C12 cells were transiently transfected with p6OSE2-luciferase and indicated expression plasmids using Lipofectamine as previously described [Jensen et al., 2008; Li et al., 2009].
Retroviral Transduction
HEK 293T cells were transfected with MSCV-Hygro or MSCV-EWS-FLI [Kinsey et al., 2009] and resulting retrovirus-containing supernatants were incubated with C2C12 cells using previously reported protocols [Hoeppner et al., 2009]. C2C12 cells were incubated with 800 μg/ml hygromycin for seven days to select for stably transduced cells. C2C12 cells were propagated and maintained in DMEM containing 10% FBS.
Immunoblotting
C2C12 cells were rinsed with cold PBS and then incubated in cold modified RIPA buffer (50 mM Tris-HCl, pH 7.4, 150 mM NaCl, 1% NP-40, 0.25% sodium deoxycholate, 1 mM EDTA, complete protease inhibitors (Roche)) for 5 minutes on ice. Crude lysates were sonicated and cleared by centrifugation at 12,000 rpm at 4°. Total protein was quantified using the detergent-compatible protein assay (BioRad). Total protein (100 μg) was resolved by SDS-PAGE and transferred to Immobilon-P membranes (Millipore). Membranes were blotted with antibodies against Runx2, Flag (Sigma-Aldrich, clone M2), or actin (Santa Cruz, clone I-19). After extensive washing, membranes were incubated with HRP-conjugated secondary antibodies. Proteins were visualized using Supersignal Chemiluminescent Substrate (Fisher Scientific).
Proliferation Assays
C2C12 cells were placed in wells of a 12 well plate at 5×104 cells/well. Cultures were incubated in DMEM containing 10% FBS for the indicated number of days, with fresh medium added every 3 days. Cell number was measured by hemocytometer enumeration. Reported values represent the mean of four samples from each culture. The assay was repeated on four separate occasions. For MTT assays, 2×104 cells were plated per well of a 96 well plate. Reported data are the mean of four wells and are representative of three independent experiments.
Cell Shape Measurements
Confluent cells were imaged at 200X magnification, and cell morphology was determined with an image analysis program (Bioquant OSTEO, BIOQUANT Image Analysis Corporation, Nashville, TN). Cell perimeter (Ce.Pm, μm), cell area (Ce.Ar, μm^2), and cell shape factor (Ce.SF) were measured from 50 cells within each culture. Cell shape factor was automatically calculated as the ratio of each cell’s area to the area of a perfect circle with an identical perimeter, yielding a measurement of roundness on a scale from 0 (straight line) to 1 (perfect circle).
Osteoblast Differentiation Assays
Confluent cultures of stably transduced C2C12 cells were incubated in DMEM supplemented with 10% fetal bovine serum, 300 ng/mL BMP2, 50 μg/ml ascorbic acid, and 4 mM beta-glycerolphosphate. This differentiation medium was replaced every two to three days.
Q-PCR
RNA was extracted and purified from C2C12 cells with TRIzol (Invitrogen) according to the manufacturer’s protocol. Messenger RNA was reverse transcribed into cDNA using the SuperScript III First-Strand Synthesis System (Invitrogen). Expression levels of mRNAs associated with osteoblast proliferation and differentiation (osteopontin, type I collagen, alkaline phosphatase, parathyroid hormone related protein (PTHrP), p21), as well as several EWS-FLI regulated genes (cysteine-rich protein 61 (Cyr61), dickkoff 1 (Dkk1), insulin growth factor binding proteins (Igfbp5 and Igfbp7) [Tirode et al., 2007]) were quantified using real-time PCR. Primer sequences are reported in Table 1. Real time reactions were performed using 25 ng of cDNA per 15 μl with Bio-Rad iQ SYBR Green Supermix and the Bio-Rad MyiQ Single Color Real-Time PCR Detection System. Transcript levels were normalized to the reference gene YWHAZ. Gene expression levels were quantified using the 2−ΔΔCt method [Pfaffl, 2001].
Table 1.
Primer sequences used in Q-PCRs.
| Gene name | ID | Forward primer sequence | Reverse primer sequence |
|---|---|---|---|
| Col1a1 | NM_007742 | GCTTCACCTACAGCACCCTTGT | TGACTGTCTTGCCCCAAGTTC |
| Alkaline phosphatase | NM_007431 | CACAGATTCCCAAAGCACCT | GGGATGGAGGAGAGAAGGTC |
| Osteopontin/Spp1 | NM_009263 | CCCGGTGAAAGTGACTGATTCT | GATCTGGGTGCAGGCTGTAAA |
| PTHrP | NM_008970 | AAGGGCAAGTCCATCCAAGAC | GCGATCAGATGGTGGAGGAA |
| Dkk1 | NM_010051 | TGCCTCCGATCATCAGACTGT | CTTGGACCAGAAGTGTCTTGCA |
| Cyr61 | NM_010516 | CAGCTCACTGAAGAGGCTTCCT | GCGTGCAGAGGGTTGAAAAG |
| Igfbp5 | NM_010518 | CAGAAAGAAGCTGACCCAGTC | TTCGGATTCCTGTCTCATCTC |
| Igfpb7 | NM_008048 | GAGGACGCTGGAGAGTATGAG | GAGGGCATCAACCACTGTAAT |
| p21/Cdkn1a | NM_007669 | GAACATCTCAGGGCCGAAAA | TGCGCTTGGAGTGATAGAAATC |
Results
FLI sequences are necessary for EWS-FLI interactions with Runx2
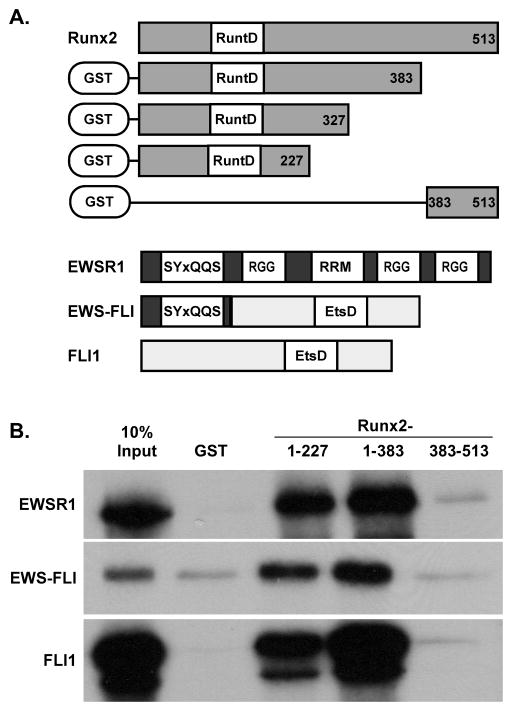
We recently reported that EWSR1 and EWS-FLI interact with the Runt domains of Runx2 and Runx1 [Li et al., 2010]. To determine the region of EWS-FLI that binds Runx2, GST-pulldown assays were performed. GST-Runx2 fusion proteins (Fig 1A) were incubated with in vitro transcribed and translated EWSR1, EWS-FLI or FLI. Similar to our previous observations [Li et al., 2010], EWSR1 and EWS-FLI interacted with the N-terminal sequences of Runx2 (amino acids 1–227 and 1–383) but not to the last 130 residues (383–513) of Runx2 (Fig 1B). Interestingly, FLI1 bound to the same Runx2 sequences. These results suggested that interactions between the Ewing’s sarcoma fusion protein, EWS-FLI, and Runx2 were mediated by amino acids derived from either EWSR1 or FLI1, or both proteins.
Figure 1. EWSR1, EWS-FLI and FLI1 Bind to the N-terminus of Runx2.
A. Schematic of proteins used in this study. GST: glutathione S transferase tag; RuntD: Runt domain; RRM: RNA recognition motif; EtsD: Ets DNA binding domain. B. GST pulldown assays with GST-Runx2 fusion proteins and radiolabeled EWSR1, EWS-FLI and FLI1.
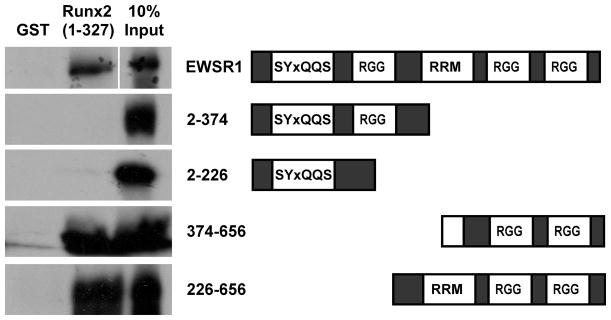
To identify the region of EWSR1 that binds Runx2, we tested several EWSR1 deletion constructs [Rossow and Janknecht, 2001] for their ability to bind the N-terminal half of Runx2 in the GST pulldown assays. EWSR1 is a 96 kD protein that contains 31 copies of the repeat sequence SYxQQS in its N-terminus, three RGG-rich regions, and an RNA recognition motif (RRM). Myc-tagged EWSR1 proteins containing the C-terminus of EWSR1 (amino acids 2–656, 226–656 and 374–656) bound to Runx2 (residues 1–327) (Fig 2). In contrast, the first 226 and 374 amino acids of EWSR1 did not bind to Runx2. These data demonstrate that the C-terminal portion of EWSR1 that contains two RGG-rich motifs is required for interactions with Runx2. They also show that the regions of EWSR1 that are present in the EWS-FLI fusion protein do not interact with Runx2. Therefore, amino acids contributed to the fusion protein by FLI1 mediate the interactions between EWS-FLI and Runx2.
Figure 2. The C-terminus of EWSR1 Binds Runx2.
GST pulldown assays with GST or the GST-Runx2 (1–327) fusion proteins and radiolabeled EWSR1 proteins.
EWSR1, FLI1 and EWS-FLI Affect Runx2-Dependent Transcription
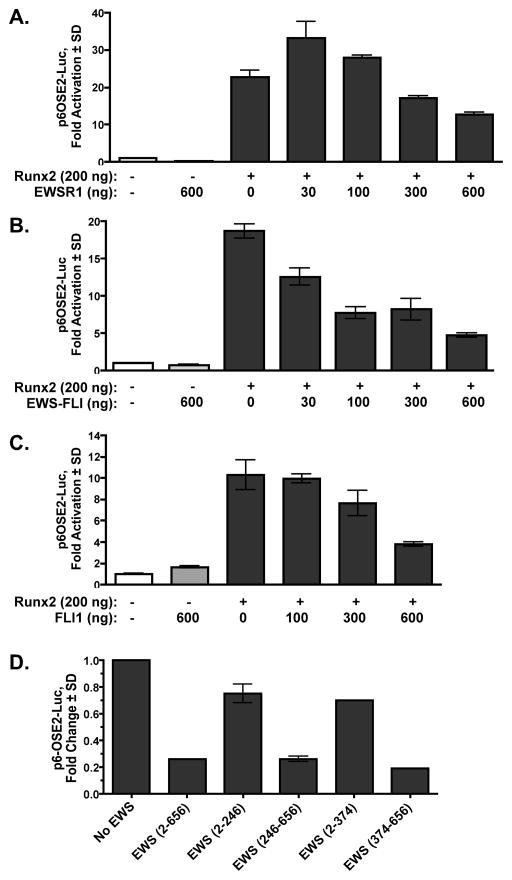
Runx2 is a DNA binding factor that regulates gene expression. Our GST-pulldown assays (Fig 1 and [Li et al., 2010]) indicated that EWSR1, FLI and EWS-FLI interact with the conserved Runt domain of Runx factors. To verify these interactions in a functional assay, C2C12 cells were transfected with the artificial Runx reporter, p6OSE-luciferase, which contains six Runx binding sites, and expression plasmids for Runx2, EWSR1, EWS-FLI or FLI1. As expected, Runx2 activated the reporter 10 to 20-fold in these assays (Fig 3). At low levels, EWSR1 modestly augmented Runx2 activation, but at higher levels it repressed reporter activity in a concentration-dependent manner. FLI and the EWS-FLI fusion protein each blocked Runx2 dependent luciferase expression in a concentration dependent manner (Fig 3A–C). All EWSR1 proteins ((2–656), (226–656) and (374–656)) that associated with Runx2 in GST pulldown assays also inhibited endogenous Runx2 activity, but the EWSR1 N-terminal proteins ((2–226) and (2–394)) that did not bind Runx2, did not repress p6OSE-luciferase expression (Fig 3D). These data demonstrate that EWS-FLI and its parent proteins bind Runx2 and inhibit its ability to regulate gene expression.
Figure 3. EWSR1, EWS-FLI and FLI1 Inhibit Runx2-Transactivation.
C2C12 cells were transiently transfected with p6OSE-Luc, pRL-Luc, pCMV-Runx2 (MASNSL isoform) and the indicated amounts of CMV-Myc-EWSR1 (A), -Flag-EWS-FLI (B), or –Flag-FLI (C). In (D) EWSR1 and Runx2 expression plasmids (300 ng each) were added as indicated. An empty CMV expression vector was added as necessary to equalize plasmid levels during the transfection. Firefly luciferase expression was normalized to Renilla luciferase levels. Values represent the mean of triplicate samples ± SD. Data are representative of at least three independent experiments.
EWS-FLI Co-localizes with Runx2 in Nuclei
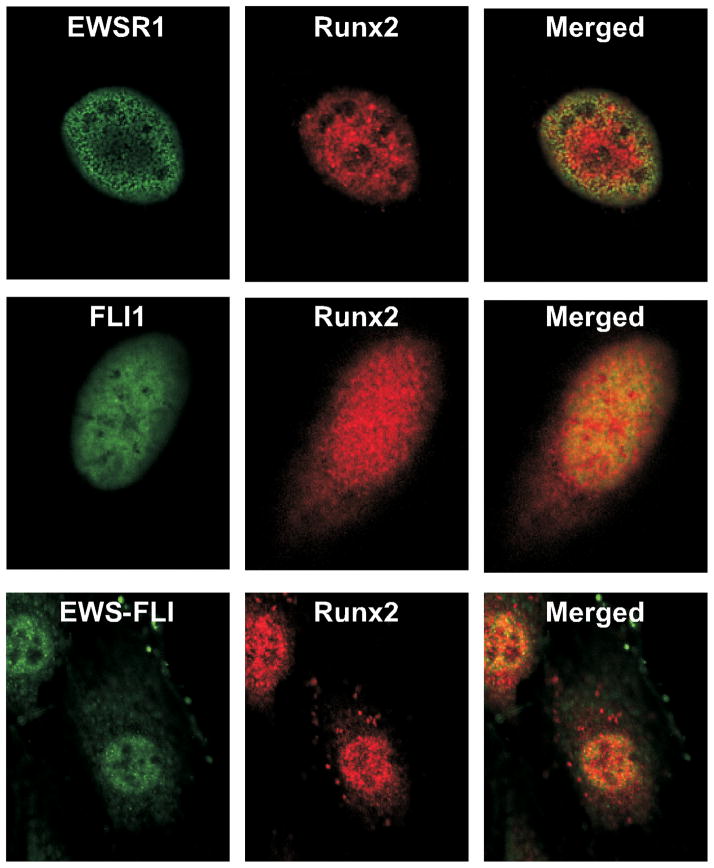
To verify that the interactions observed in vitro also occurred in situ, the co-localization of endogenous Runx2 with Myc-EWSR1, Flag-FLI1 and Flag-EWS-FLI was examined by confocal microscopy and immunofluorescence. All proteins were predominantly expressed in punctate foci within nuclei (Fig 4). Interestingly, EWSR1 was present on the nuclear periphery and formed a shell around Runx2 nuclear domains in merged images (Fig 4A). Careful examination of the boundary between the Runx2 and EWSR1 stained regions revealed a few foci of co-localization. By comparison, FLI1 showed more central nuclear localization and co-localized with Runx2 to a great extent than EWSR1 (Fig 4B). EWS-FLI had a nuclear distribution pattern similar to FLI1 and localized with Runx2 to a similar degree as FLI1 (Fig 4C). These data support the conclusion that FLI1 contributes Runx2 binding sequences to the EWS-FLI fusion protein. They also show that interactions between Runx2 and EWSR1 are possible but rare in cell nuclei because the proteins are spatially separated.
Figure 4. EWS-FLI and FLI1, but not EWSR1, co-localize with Runx2 in Cell Nuclei.
A and B) U2-OS cells were transiently transfected with pCMV-Myc-EWSR1 (A) or –FLAG-FLI1. The exogenous EWSR1 or FLI1 protein and endogenous Runx2 were detected by confocal microscopy. C. C2C12 cells stably expressing FLAG-EWS-FLI and endogenous Runx2 were examined by confocal microscopy.
EWS-FLI Alters Cell Shape, Increases Proliferation, and Suppresses Osteoblast Differentiation of Mesenchymal Progenitor Cells
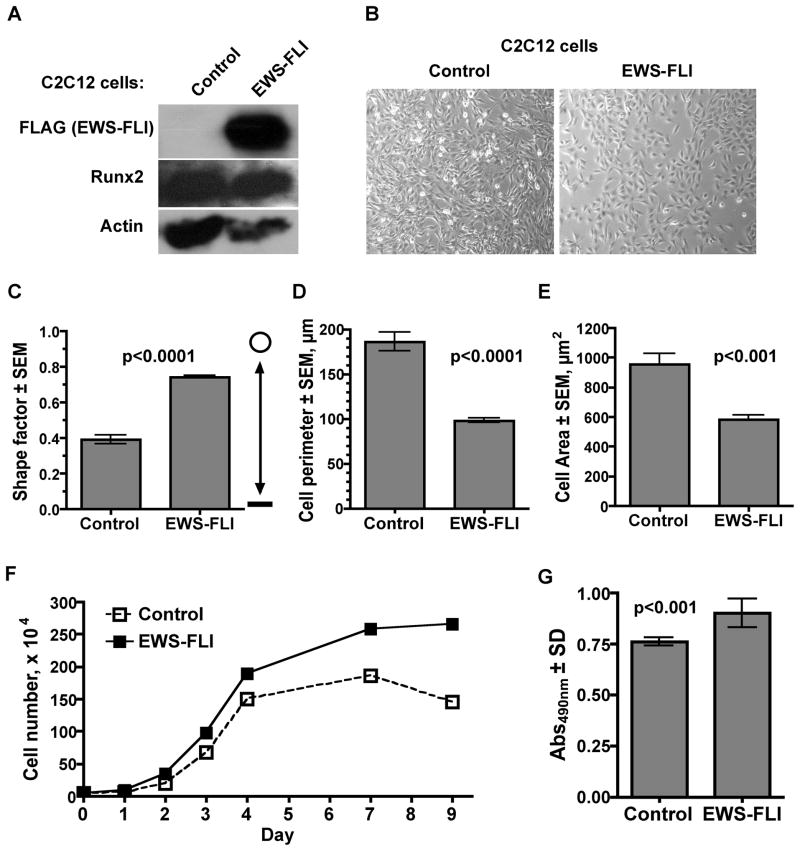
Previous studies demonstrated that EWS-FLI blocks the differentiation of tumor cells, cell lines and primary bone marrow cells towards the osteoblastic lineage [Eliazer et al., 2003; Kovar, 2005; Tirode et al., 2007; Torchia et al., 2003]. In light of our observations that EWS-FLI interacts with and represses Runx2 and with knowledge of the fact that Runx2 integrates numerous signaling pathways, particularly BMP2 signaling, to control osteoblast differentiation [Afzal et al., 2005; Javed et al., 2009; Javed et al., 2008], we asked whether the t(11;22) product could block BMP2-induced osteoblastic maturation of multipotent C2C12 cells. EWS-FLI was expressed at high levels in the stably transduced C2C12 cells and did not appear to drastically affect Runx2 levels (Fig 5A). As was observed in other laboratories [Eliazer et al., 2003; Torchia et al., 2003; Riggi et al., 2005], EWS-FLI expression altered the morphology of transfected cells (Fig 5B). Thus, EWS-FLI cells appeared rounder than C2C12 cells transduced with a control retrovirus. Image analysis confirmed that EWS-FLI cells were indeed rounder (Fig 5C), had a smaller cell perimeter (Fig 5D) and reduced cellular area (Fig 5E) than control cells. The EWS-FLI expressing cells proliferated at a faster rate than control cells in normal growth medium (Fig 5F). MTT assays demonstrated that metabolic activity was also elevated in EWS-FLI transduced C2C12 cells (Fig 5G). Together these data demonstrate that EWS-FLI alters the morphology and behavior of C2C12 cells to mimic the properties of Ewing’s sarcoma cells.
Figure 5. EWS-FLI Alters C2C12 Cell Morphology and Increases Proliferation.
A. Western blot analysis of C2C12 cells transduced with MSCV-Hygro (Control) or MSCV-FLAG-EWS-FLI. Blots were incubated with antibodies recognizing FLAG, Runx2 or Actin. B. Light microscopy images of cells transduced with the Control MSCV virus or EWS-FLI after hygromycin selection. C. The Shape Factor of 50 cells in populations of Control and EWS-FLI C2C12 cells was determined by image analysis. A value of 1 represents a perfect circle and 0 equals a straight line. The perimeters (D) and areas (E) of 50 cells were also measured with image analysis software. F. EWS-FLI increase cells proliferation. C2C12 cells were plated at a starting density of 5×104 cells/well and counted at the indicated times. Values represent the mean of four wells and are representative of three experiments. Standard error bars are too small to be seen. G. MTT assays demonstrate that metabolic activity is increased in EWS-FLI expressing cells. Values represent the mean of four samples and are representative of three experiments. P values were determined with unpaired student’s t tests.
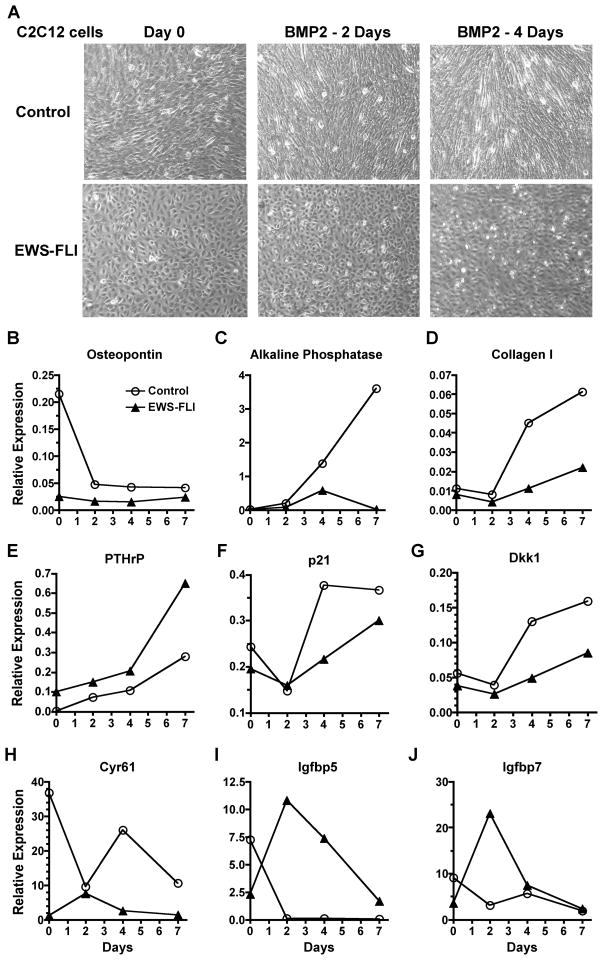
To determine the effects of EWS-FLI on the ability of C2C12 cells to differentiate towards the osteoblast lineage, we treated the control and EWS-FLI transduced cells with BMP2, ascorbic acid and beta-glycerolphosphate for one week and assessed the effects of EWS-FLI on the temporal expression of several genes linked to osteoblast maturation. The cells retained a rounder morphology throughout differentiation (Fig 6A). Osteopontin, a marker of osteoblast progenitors, was expressed at high levels in undifferentiated control C2C12 cells and then decreased during differentiation (Fig 6B). EWS-FLI reduced osteopontin levels in undifferentiated and differentiated cells. Alkaline phosphatase and type I collagen levels rose steadily in control cells beginning after two days of exposure to BMP2 but EWS-FLI suppressed the expression of these early markers of osteoblast differentiation (Fig 6C and 6D). Runx2 mRNA levels were not significantly altered by EWS-FLI (data not shown). Rankl and Opg levels were very low in all cells (data not shown); however, PTHrP levels were elevated in undifferentiated EWS-FLI expressing cells and remained high throughout BMP2-induced differentiation (Fig 6E). Consistent with the increased proliferation rate of the EWS-FLI expressing cells, p21 levels were reduced in undifferentiated and differentiated cells (Fig 6F). Dkk1, an inhibitor of Lrp5/6 and Wnt signaling and a known EWS-FLI regulated gene [Miyagawa et al., 2009], was also suppressed in differentiating EWS-FLI expressing C2C12 cells (Fig 6G). Finally, the expression of three genes that were highly differentially regulated by EWS-FLI in a microarray profiling experiment were assessed [Tirode et al., 2007]. Consistent with those reported results, Cyr61 (Fig 6H), Igfbp5 (Fig 6I) and Igfbp7 (Fig 6J) were repressed by EWS-FLI in undifferentiated C2C12 cells. Cyr61 remained lower in EWS-FLI cells as compared to control cells throughout BMP2-induced differentiation; however, Igfbp5 and Igfbp7 rose dramatically in BMP2-stimulated EWS-FLI cells while decreasing in control cells during differentiation. Together these data demonstrate that EWS-FLI alters the responses of C2C12 multipotent progenitor cells to BMP2.
Figure 6. EWS-FLI Delays BMP2-Induced C2C12 Osteoblast Differentiation.
A. Light microscopy images of confluent C2C12 Control and EWS-FLI expressing cells that were treated with osteogenic medium and BMP2 for 0, 2 or 4 days. The EWS-FLI cells retain the rounder shape throughout differentiation. B-J. The effects of EWS-FLI on the gene expression during the course of BMP-2 induced osteogenic differentiation were measured by RT-PCR. Transcript levels for the specified genes were normalized to the reference gene YWHAZ with the 2−ΔΔCT method.
Discussion
Ewing’s sarcomas are rare but aggressive tumors that afflict children and young adults. A significant advance in our molecular understanding of the pathogenesis of Ewing’s sarcoma tumors occurred in the late 1980’s and early 1990’s when the t(11;22) translocation was described and shown to encode an abnormal fusion protein, EWS-FLI [Delattre et al., 1992; Turc-Carel et al., 1988]. The approximately 15% of Ewing sarcomas that do not contain EWS-FLI harbor fusions between other TET and ETS family members [Law et al., 2006; Tan and Manley, 2009]. A more recent advancement is that Ewing’s sarcomas are derived from multipotent progenitor cells of mesenchymal and/or neural crest origin [Hancock and Lessnick, 2008; Kauer et al., 2009; Riggi et al., 2005; Staege et al., 2004; Suva et al., 2009; Tirode et al., 2007; Toomey et al., 2010]. Tumorigenesis occurs when crucial pathways controlling the proliferation, differentiation, and/or survival of cells are disrupted. In the case of Ewing’s sarcomas, the plasticity of the progenitor cell is also likely to be suppressed. In this report we show that EWS-FLI can interact with Runx2, a crucial regulator of osteoblast lineage specification from mesenchymal and neural crest progenitor cells. Furthermore, we show that EWS-FLI alters the morphology and augments the proliferation of a multipotent cell line, while suppressing osteoblast differentiation. These data provide new insights into the mechanisms whereby EWS-FLI maintains the tumor cell in an undifferentiated state.
We previously showed that EWS, EWSR1 and another TET family member, FUS, bind Runx1 and Runx2 [Li et al., 2010]. The Runx DNA binding domain, commonly referred to as the Runt domain, was necessary and sufficient for interactions with EWS, FUS and EWSR1 in vitro. In this project, we sought to further characterize these interactions. Using a set of EWSR1 deletions constructs, we found that the C-terminal region of EWSR1 was sufficient to bind Runx2, but the N-terminus of EWSR1 did not interact with Runx2. These results negated our hypothesis that the repeated SYxQQS motif in the N-terminus TET proteins was responsible for the interactions with Runx2. This hypothesis was formulated from data showing that CoAA, which contains similar repeat sequences (YxxQ), binds the Runt domain of Runx proteins [Li et al., 2009]. These results also suggested that EWS-FLI associated with Runx2 via different mechanisms than EWSR1. Indeed, FLI1 bound the N-terminus of Runx2. FLI1 also physically associates with RUNX1 to regulate gene expression in certain contexts [Huang et al., 2009]. Numerous interactions between Runx2 and Ets transcription factors have been described [Fowler et al., 2005; Li et al., 2004; Sato et al., 1998]. These associations synergistically augment expression of the target gene of interest because the spatial arrangement of Runx and Ets sites allow for cooperative recruitment of transcription co-activators and RNA polymerase II. In our studies, we used a promoter containing multiple Runx binding elements to verify that FLI1, EWSR1 and EWS-FLI affect Runx activity. This artificial context does not necessarily predict that FLI1, EWSR1, and EWS-FLI will always repress Runx factors. The transcriptional consequences of such interactions will be contextual and dependent on promoter/enhancer architecture as well as the cellular environment.
Chromosomal translocations have multiple consequences. They can create fusion proteins, like EWS-FLI, which fuses an activation domain of EWSR1 to the DNA binding domain of FLI1. These fusion proteins may have altered activity and/or localization. Another, not necessarily mutually exclusive, consequence is that a gene is aberrantly expressed, sometimes at very high levels, in a cell because its regulation is under the control of another gene’s promoter or enhancer. The normative levels of EWSR1 and FLI1 in mesenchymal progenitor cells and osteoblast lineage cells have not yet been rigorously examined; however, EWSR1 and FLI1 are both present in mesenchymal cells during development [Kovar, 2005; Lin et al., 2008]. The EWS-FLI fusion protein has a subnuclear localization pattern more similar to FLI1, than to EWSR1. EWS-FLI and FLI1 both co-localize with Runx2 within subnuclear punctate foci to a greater extent than EWSR1 and Runx2 do. In contrast, the EWSR1 domain incases Runx2 regions within nuclei, with interaction taking place only at the interface of these nuclear compartments. These results demonstrate that EWSR1 may affect Runx2 activity, and vice-versa, but these effects are tightly regulated and would only occur when the nuclear compartments are disrupted. Our data also suggest that EWS-FLI is capable of inhibiting Runx2 and suppressing osteoblast differentiation because it assumes properties of FLI1.
The physical association of EWS-FLI with Runx2 provides a molecular explanation for how Ewing’s sarcomas maintain an undifferentiated phenotype. Suppressing EWS-FLI expression allowed tumor cells to differentiate into osteoblast, neural, adipocyte and chondrocyte lineages [Tirode et al., 2007]. On the other hand, expressing EWS-FLI in Arf1-deficient murine bone marrow stromal cells prevented this multilineage differentiation [Torchia et al., 2003]. There are inherent and empirical limitations to all cell models in which these and other experiments were done [Kovar, 2005] and finding the perfect in vitro model in which to study any tumor is perhaps impossible. We chose the C2C12 cell line because it is multipotent and begins expressing osteoblast genes when exposed to BMP2. Runx2 interacts with SMADs to integrate the BMP2 signaling and is required for osteoblast differentiation of these cells [Afzal et al., 2005; Javed et al., 2009; Javed et al., 2008]. EWS-FLI suppressed the expression of many early osteoblast differentiation genes, including osteopontin, alkaline phosphatase and type I collagen. Runx2 mRNA and protein levels were not significantly changed. It is possible that EWS-FLI blocks other proteins required for osteoblast differentiation, such as SMADs and osterix, but by altering the activity of the master regulator of osteoblast differentiation, Runx2, EWS-FLI controls an essential factor to lineage specification. A previous study showed that overexpressing EWS-FLI in C2C12 cells arrested myogenic differentiation by blocking lineage-defining transcription factors MyoD and myogenin, but the effects of EWS-FLI on BMP2 induced osteoblast differentiation in this model were not tested [Eliazer et al., 2003]. Together these data demonstrate EWS-FLI blocks lineage-determining transcription factors to control progenitor cell fate.
Our data provide insights into therapeutic strategies for Ewing’s sarcomas. First, relieving the interactions between EWS-FLI and Runx2 may promote the differentiation of tumor cells. Although disrupting protein-protein interactions in the nucleus is difficult, new technologies may make this possible and there is precedence for success in Ewing’s sarcomas where a peptide that blocked EWS-FLI interactions with RNA helicase A proved efficacious [Erkizan et al., 2009; Erkizan et al., 2010]. We also showed that EWS-FLI induces the expression of PTHrP, which indirectly induces osteolysis and may be a major mediator for bone destruction and metastasis, as it is in other tumors [Liao and McCauley, 2006]. The IGF pathway was identified as a potential therapeutic target in Ewing’s sarcomas more than two decades ago and blocking agents (e.g. small molecules and antibodies) are now being intensely pursued in clinical trials [Olmos et al., 2010; Yee et al., 1990]. EWS-FLI suppresses the expression of IGF binding proteins (Igfbp), which are natural inhibitors of IGF [Prieur et al., 2004; Tirode et al., 2007]. Our results confirm that EWS-FLI suppresses Igfbp5 and Igfbp7 in undifferentiated C2C12 cells, but interestingly BMP2 increased their expression. Thus, lineage-inducing agents may augment the potency of IGF blocking therapeutics by inducing natural antagonists.
Acknowledgments
Funding sources: R01 AR48147, T32 AR056950
Contract grant sponsor: NIH; Contract grant numbers: AR048147 and AR056950
We thank Drs Stephen Lessnick, Ralf Jacknecht, Aykut Üren, Dennis Watson for generously sharing their EWS-FLI, EWSR1, and FLI1 expression plasmids. The NIH partially supported this work through grants R01 AR048147 and T32 AR056950.
Contributor Information
Xiaodong Li, Email: li.xiaodong@mayo.edu.
Meghan E. McGee-Lawrence, Email: mcgeelawrence.meghan@mayo.edu.
Matthew Decker, Email: mndecker@stanford.edu.
Jennifer J. Westendorf, Email: westendorf.jennifer@mayo.edu.
References
- Afzal F, Pratap J, Ito K, Ito Y, Stein JL, van Wijnen AJ, Stein GS, Lian JB, Javed A. Smad function and intranuclear targeting share a Runx2 motif required for osteogenic lineage induction and BMP2 responsive transcription. J Cell Physiol. 2005;204:63–72. doi: 10.1002/jcp.20258. [DOI] [PubMed] [Google Scholar]
- Delattre O, Zucman J, Plougastel B, Desmaze C, Melot T, Peter M, Kovar H, Joubert I, de Jong P, Rouleau G, et al. Gene fusion with an ETS DNA-binding domain caused by chromosome translocation in human tumours. Nature. 1992;359:162–5. doi: 10.1038/359162a0. [DOI] [PubMed] [Google Scholar]
- Eliazer S, Spencer J, Ye D, Olson E, Ilaria RL., Jr Alteration of mesodermal cell differentiation by EWS/FLI-1, the oncogene implicated in Ewing’s sarcoma. Mol Cell Biol. 2003;23:482–92. doi: 10.1128/MCB.23.2.482-492.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Erkizan HV, Kong Y, Merchant M, Schlottmann S, Barber-Rotenberg JS, Yuan L, Abaan OD, Chou TH, Dakshanamurthy S, Brown ML, Uren A, Toretsky JA. A small molecule blocking oncogenic protein EWS-FLI1 interaction with RNA helicase A inhibits growth of Ewing’s sarcoma. Nat Med. 2009;15:750–6. doi: 10.1038/nm.1983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Erkizan HV, Uversky VN, Toretsky JA. Oncogenic Partnerships: EWS-FLI1 Protein Interactions Initiate Key Pathways of Ewing’s Sarcoma. Clin Cancer Res. 2010 doi: 10.1158/1078-0432.CCR-09-2261. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ewing J. Diffuse Endothelioma of Bone. Proc New York Pathol Soc. 1921;21:17–24. [Google Scholar]
- Fowler M, Borazanci E, McGhee L, Pylant SW, Williams BJ, Glass J, Davis JN, Meyers S. RUNX1 (AML-1) and RUNX2 (AML-3) cooperate with prostate-derived Ets factor to activate transcription from the PSA upstream regulatory region. J Cell Biochem. 2005 doi: 10.1002/jcb.20664. [DOI] [PubMed] [Google Scholar]
- Hancock JD, Lessnick SL. A transcriptional profiling meta-analysis reveals a core EWS-FLI gene expression signature. Cell Cycle. 2008;7:250–6. doi: 10.4161/cc.7.2.5229. [DOI] [PubMed] [Google Scholar]
- Hoeppner LH, Secreto F, Jensen ED, Li X, Kahler RA, Westendorf JJ. Runx2 and bone morphogenic protein 2 regulate the expression of an alternative Lef1 transcript during osteoblast maturation. J Cell Physiol. 2009;221:480–9. doi: 10.1002/jcp.21879. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huang H, Yu M, Akie TE, Moran TB, Woo AJ, Tu N, Waldon Z, Lin YY, Steen H, Cantor AB. Differentiation-dependent interactions between RUNX-1 and FLI-1 during megakaryocyte development. Mol Cell Biol. 2009;29:4103–15. doi: 10.1128/MCB.00090-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Javed A, Afzal F, Bae JS, Gutierrez S, Zaidi K, Pratap J, van Wijnen AJ, Stein JL, Stein GS, Lian JB. Specific residues of RUNX2 are obligatory for formation of BMP2-induced RUNX2-SMAD complex to promote osteoblast differentiation. Cells Tissues Organs. 2009;189:133–7. doi: 10.1159/000151719. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Javed A, Bae JS, Afzal F, Gutierrez S, Pratap J, Zaidi SK, Lou Y, van Wijnen AJ, Stein JL, Stein GS, Lian JB. Structural coupling of Smad and Runx2 for execution of the BMP2 osteogenic signal. J Biol Chem. 2008;283:8412–22. doi: 10.1074/jbc.M705578200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jedlicka P. Ewing Sarcoma, an enigmatic malignancy of likely progenitor cell origin, driven by transcription factor oncogenic fusions. Int J Clin Exp Pathol. 2010;3:338–47. [PMC free article] [PubMed] [Google Scholar]
- Jensen ED, Gopalakrishnan R, Westendorf JJ. Regulation of gene expression in osteoblasts. Biofactors. 2010;36:25–32. doi: 10.1002/biof.72. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jensen ED, Niu L, Caretti G, Nicol SM, Teplyuk N, Stein GS, Sartorelli V, van Wijnen AJ, Fuller-Pace FV, Westendorf JJ. p68 (Ddx5) interacts with Runx2 and regulates osteoblast differentiation. J Cell Biochem. 2008;103:1438–51. doi: 10.1002/jcb.21526. [DOI] [PubMed] [Google Scholar]
- Kauer M, Ban J, Kofler R, Walker B, Davis S, Meltzer P, Kovar H. A molecular function map of Ewing’s sarcoma. PLoS One. 2009;4:e5415. doi: 10.1371/journal.pone.0005415. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kinsey M, Smith R, Iyer AK, McCabe ER, Lessnick SL. EWS/FLI and Its Downstream Target NR0B1 Interact Directly to Modulate Transcription and Oncogenesis in Ewing’s Sarcoma. Cancer Res. 2009 doi: 10.1158/0008-5472.CAN-09-1540. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Komori T, Yagi H, Nomura S, Yamaguchi A, Sasaki K, Deguchi K, Shimizu Y, Bronson RT, Gao YH, Inada M, Sato M, Okamoto R, Kitamura Y, Yoshiki S, Kishimoto T. Targeted disruption of Cbfa1 results in a complete lack of bone formation owing to maturational arrest of osteoblasts. Cell. 1997;89:755–64. doi: 10.1016/s0092-8674(00)80258-5. [DOI] [PubMed] [Google Scholar]
- Kovar H. Context matters: the hen or egg problem in Ewing’s sarcoma. Semin Cancer Biol. 2005;15:189–96. doi: 10.1016/j.semcancer.2005.01.004. [DOI] [PubMed] [Google Scholar]
- Law WJ, Cann KL, Hicks GG. TLS, EWS and TAF15: a model for transcriptional integration of gene expression. Brief Funct Genomic Proteomic. 2006;5:8–14. doi: 10.1093/bfgp/ell015. [DOI] [PubMed] [Google Scholar]
- Li V, Raouf A, Kitching R, Seth A. Ets2 transcription factor inhibits mineralization and affects target gene expression during osteoblast maturation. In Vivo. 2004;18:517–24. [PubMed] [Google Scholar]
- Li X, Decker M, Westendorf JJ. TEThered to Runx: novel binding partners for runx factors. Blood Cells Mol Dis. 2010;45:82–5. doi: 10.1016/j.bcmd.2010.03.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li X, Hoeppner LH, Jensen ED, Gopalakrishnan R, Westendorf JJ. Co-activator activator (CoAA) prevents the transcriptional activity of Runt domain transcription factors. J Cell Biochem. 2009;108:378–87. doi: 10.1002/jcb.22263. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liao J, McCauley LK. Skeletal metastasis: Established and emerging roles of parathyroid hormone related protein (PTHrP) Cancer Metastasis Rev. 2006;25:559–71. doi: 10.1007/s10555-006-9033-z. [DOI] [PubMed] [Google Scholar]
- Lin PP, Pandey MK, Jin F, Xiong S, Deavers M, Parant JM, Lozano G. EWS-FLI1 induces developmental abnormalities and accelerates sarcoma formation in a transgenic mouse model. Cancer Res. 2008;68:8968–75. doi: 10.1158/0008-5472.CAN-08-0573. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Miyagawa Y, Okita H, Itagaki M, Toyoda M, Katagiri YU, Fujimoto J, Hata J, Umezawa A, Kiyokawa N. EWS/ETS regulates the expression of the Dickkopf family in Ewing family tumor cells. PLoS One. 2009;4:e4634. doi: 10.1371/journal.pone.0004634. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nowling TK, Fulton JD, Chike-Harris K, Gilkeson GS. Ets factors and a newly identified polymorphism regulate Fli1 promoter activity in lymphocytes. Mol Immunol. 2008;45:1–12. doi: 10.1016/j.molimm.2007.05.018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Olmos D, Tan DS, Jones RL, Judson IR. Biological rationale and current clinical experience with anti-insulin-like growth factor 1 receptor monoclonal antibodies in treating sarcoma: twenty years from the bench to the bedside. Cancer J. 2010;16:183–94. doi: 10.1097/PPO.0b013e3181dbebf9. [DOI] [PubMed] [Google Scholar]
- Otto F, Thornell AP, Crompton T, Denzel A, Gilmour KC, Rosewell IR, Stamp GW, Beddington RS, Mundlos S, Olsen BR, Selby PB, Owen MJ. Cbfa1, a candidate gene for cleidocranial dysplasia syndrome, is essential for osteoblast differentiation and bone development. Cell. 1997;89:765–71. doi: 10.1016/s0092-8674(00)80259-7. [DOI] [PubMed] [Google Scholar]
- Pfaffl MW. A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Res. 2001;29:e45. doi: 10.1093/nar/29.9.e45. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Prieur A, Tirode F, Cohen P, Delattre O. EWS/FLI-1 silencing and gene profiling of Ewing cells reveal downstream oncogenic pathways and a crucial role for repression of insulin-like growth factor binding protein 3. Mol Cell Biol. 2004;24:7275–83. doi: 10.1128/MCB.24.16.7275-7283.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Riggi N, Cironi L, Provero P, Suva ML, Kaloulis K, Garcia-Echeverria C, Hoffmann F, Trumpp A, Stamenkovic I. Development of Ewing’s sarcoma from primary bone marrow-derived mesenchymal progenitor cells. Cancer Res. 2005;65:11459–68. doi: 10.1158/0008-5472.CAN-05-1696. [DOI] [PubMed] [Google Scholar]
- Rossow KL, Janknecht R. The Ewing’s sarcoma gene product functions as a transcriptional activator. Cancer Res. 2001;61:2690–5. [PubMed] [Google Scholar]
- Sato M, Morii E, Komori T, Kawahata H, Sugimoto M, Terai K, Shimizu H, Yasui T, Ogihara H, Yasui N, Ochi T, Kitamura Y, Ito Y, Nomura S. Transcriptional regulation of osteopontin gene in vivo by PEBP2alphaA/CBFA1 and ETS1 in the skeletal tissues. Oncogene. 1998;17:1517–25. doi: 10.1038/sj.onc.1202064. [DOI] [PubMed] [Google Scholar]
- Staege MS, Hutter C, Neumann I, Foja S, Hattenhorst UE, Hansen G, Afar D, Burdach SE. DNA microarrays reveal relationship of Ewing family tumors to both endothelial and fetal neural crest-derived cells and define novel targets. Cancer Res. 2004;64:8213–21. doi: 10.1158/0008-5472.CAN-03-4059. [DOI] [PubMed] [Google Scholar]
- Suva ML, Riggi N, Stehle JC, Baumer K, Tercier S, Joseph JM, Suva D, Clement V, Provero P, Cironi L, Osterheld MC, Guillou L, Stamenkovic I. Identification of cancer stem cells in Ewing’s sarcoma. Cancer Res. 2009;69:1776–81. doi: 10.1158/0008-5472.CAN-08-2242. [DOI] [PubMed] [Google Scholar]
- Tan AY, Manley JL. The TET Family of Proteins: Functions and Roles in Disease. J Mol Cell Biol. 2009 doi: 10.1093/jmcb/mjp025. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tirode F, Laud-Duval K, Prieur A, Delorme B, Charbord P, Delattre O. Mesenchymal stem cell features of Ewing tumors. Cancer Cell. 2007;11:421–9. doi: 10.1016/j.ccr.2007.02.027. [DOI] [PubMed] [Google Scholar]
- Toomey EC, Schiffman JD, Lessnick SL. Recent advances in the molecular pathogenesis of Ewing’s sarcoma. Oncogene. 2010 doi: 10.1038/onc.2010.205. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Torchia EC, Jaishankar S, Baker SJ. Ewing tumor fusion proteins block the differentiation of pluripotent marrow stromal cells. Cancer Res. 2003;63:3464–8. [PubMed] [Google Scholar]
- Turc-Carel C, Aurias A, Mugneret F, Lizard S, Sidaner I, Volk C, Thiery JP, Olschwang S, Philip I, Berger MP, et al. Chromosomes in Ewing’s sarcoma. I. An evaluation of 85 cases of remarkable consistency of t(11;22)(q24;q12) Cancer Genet Cytogenet. 1988;32:229–38. doi: 10.1016/0165-4608(88)90285-3. [DOI] [PubMed] [Google Scholar]
- Yee D, Favoni RE, Lebovic GS, Lombana F, Powell DR, Reynolds CP, Rosen N. Insulin-like growth factor I expression by tumors of neuroectodermal origin with the t(11;22) chromosomal translocation. A potential autocrine growth factor. J Clin Invest. 1990;86:1806–14. doi: 10.1172/JCI114910. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zeng C, McNeil S, Pockwinse S, Nickerson J, Shopland L, Lawrence JB, Penman S, Hiebert S, Lian JB, van Wijnen AJ, Stein JL, Stein GS. Intranuclear targeting of AML/CBFalpha regulatory factors to nuclear matrix-associated transcriptional domains. Proc Natl Acad Sci U S A. 1998;95:1585–9. doi: 10.1073/pnas.95.4.1585. [DOI] [PMC free article] [PubMed] [Google Scholar]