Abstract
We previously isolated a pollen factor, ui6.1, which encodes a Cullin1 protein (CUL1) that functions in unilateral interspecific incompatibility (UI) in Solanum. Here we show that CUL1 is also required for pollen function in self-incompatibility (SI). We used RNA interference (RNAi) to reduce CUL1 expression in pollen of Solanum arcanum, a wild SI tomato relative. Hemizygous T0 plants showed little or no transmission of the transfer DNA (T-DNA) through pollen when crossed onto nontransgenic SI plants, indicating that CUL1-deficient pollen are selectively eliminated. When crossed onto a related self-compatible (SC) accession lacking active S-RNase, pollen transmission of the T-DNA followed Mendelian ratios. These results provide further evidence for functional overlap between SI and UI on the pollen side and suggest that CUL1 mutations will reinforce SI-to-SC transitions in natural populations only if preceded by loss of pistil S-RNase expression.
Keywords: Cullin1, unilateral incompatibility, tomato, gametophytic self-incompatibility, Solanum arcanum
SELF-INCOMPATIBILITY (SI) is a widespread genetic mechanism in hermaphroditic plants that allows for the recognition and rejection of closely related pollen to prevent inbreeding. The breakdown of SI to self-compatibility (SC) through mutation occurs frequently (Igic et al. 2008), presumably driven by reproductive assurance under conditions where pollen from compatible mates is limiting. Pollen from SC species or populations is typically rejected on pistils of related SI species or populations, while, in the reciprocal crosses (SC pollinated by SI), no pollen rejection occurs. This pattern of unilateral incompatibility (UI) is known as the “SI × SC rule” (Lewis and Crowe 1958). While the mechanisms underlying SI have been the subject of much investigation, pollen rejection by UI is less well understood.
The cultivated tomato (Solanum lycopersicum) and related wild Solanum species provide a powerful system with which to study these reproductive barriers. They exhibit a wide range of mating systems, including SI-enforced obligate outcrossing, SC with facultative outcrossing, and SC with high levels of inbreeding (Rick 1988). Self-compatible biotypes or accessions of mostly SI species provide a source of natural variation for studying SI-related factors.
Self-incompatibility in Solanum and other Solanaceae is the S-RNase based, gametophytic type, in which S-specificity is determined by S-RNases in the pistil (McClure et al. 1989) and S-locus F-box proteins (SLFs) in pollen (Sijacic et al. 2004). F-box proteins, together with Skp1 and Cullin1 proteins, are components of Skp, Cullin, Fbox type (SCF) ubiquitin E3 ligases that mark proteins for degradation by the 26S proteasome (Zheng et al. 2002; Moon et al. 2004). The ubiquitin–proteasome pathway is thought to regulate pollen-side SI responses in the Solanaceae (Zhang et al. 2009). In the “collaborative non-self-recognition” model (Kubo et al. 2010), the S-locus encodes multiple SLF proteins that together recognize different suites of S-RNases. In compatible pollinations, SLF/S-RNase interactions lead to protection of pollen tubes against cytotoxic S-RNase, while in incompatible pollinations a failure to recognize “self” S-RNase results in pollen-tube inhibition. The absence of any deletions recovered among pollen-part SC mutants in Nicotiana alata is consistent with the presence of an S-RNase inhibitor encoded by the S-locus because pollen lacking SLF expression would be eliminated on pistils expressing S-RNase (Golz et al. 2001). In addition, modifier genes, such as the HT-B and 120-kDa proteins in the pistil, are required for SI function but not for specificity (McClure et al. 1999; Hancock et al. 2005).
The molecular mechanisms of pollen rejection in UI are complex. On the pistil side, S-RNase expression is required for pollen rejection in some crosses, while, in other cases, an S-RNase-independent pollen rejection system is evident (Murfett et al. 1996; Covey et al. 2010). On the pollen side, we showed previously that ui6.1 encodes a Cullin1 protein that is required for pollen to overcome S-RNase-dependent UI (Li and Chetelat 2010). Furthermore, ui6.1 (designated herein CUL1) interacts genetically with another pollen factor, ui1.1, which maps to the S-locus and thus might encode an SLF protein(s) (Li et al. 2010). These results suggested that pollen rejection in UI is controlled by a mechanism biochemically related to SI, either as an independent pathway or as a secondary effect of loss of SI factor(s) in the pollen. We therefore examined whether CUL1 functions directly in SI.
Here we show that in Solanum arcanum, an SI wild tomato species, silencing of CUL1 expression in pollen by RNA interference (RNAi) causes pollen rejection in normally compatible sib crosses, whereas the same pollen retain full compatibility on an SC accession expressing an inactive S-RNase. Our results strongly suggest that CUL1 functions to protect pollen from S-RNases in SI as well as in UI and provide further evidence of overlap between intra- and interspecific pollen rejection pathways.
Results
We used a loss-of-function approach to test whether SI and UI are functionally linked on the pollen side. Figure 1A shows the strategy for testing the effects of suppressing ui6.1 CUL1 expression in intraspecific crosses. A similar approach was used to test the function of PhSSK1, another putative pollen SI factor that, like CUL1, is proposed to form part of an SCF complex (Zhao et al. 2010). We used the S. arcanum accessions LA2163 (SI) and LA2157 (SC), which are similar in most respects, apart from mating system (Rick 1986). These two accessions are cross-compatible in either direction (Rick 1986; Kowyama et al. 1994). Importantly, LA2157 expresses a mutant S-RNase that lacks RNase activity (Kowyama et al. 1994; Royo et al. 1994) and confers self-compatibility (Rivers and Bernatzky 1994).
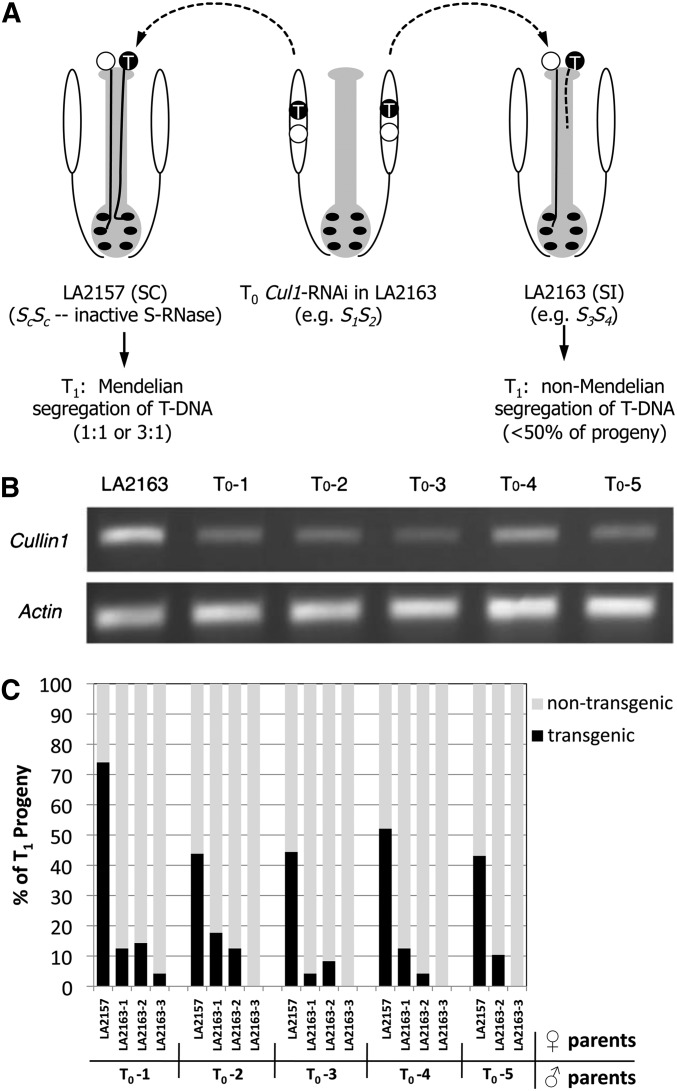
Figure 1.
Experimental design and sample results testing CUL1 function in self-incompatibility. (A) Diagram of crosses between transgenic CUL1 RNAi lines as pollen donors onto self-compatible (LA2157) or self-incompatible (LA2163) accessions of S. arcanum with inferred S-genotypes of each. (B) RT-PCR analysis of CUL1 expression in pollen of five independent T0 CUL1 RNAi plants (T0-1 to T0-5) and nontransgenic control accession LA2163. (C) The frequency of transgenic and nontransgenic T1 progeny from five independent CUL1 RNAi transformants (T0-1, two T-DNA insertions; T0-2 to T0-5, one insertion) crossed onto SC LA2157 or three independent SI LA2163 plants. The segregation of the T-DNA in the progeny of pollinations onto LA2157 fits Mendelian ratios, whereas progeny from the crosses onto LA2163 show an extreme deficiency of transgenic plants. These data (and full results in Table S2) indicate that CUL1-silenced pollen are rejected on pistils of independent nontransgenic LA2163 plants with different S-genotypes.
We transformed a CUL1 RNAi construct (Supporting Information, File S1, Figure S1, and Table S1) into SI S. arcanum LA2163, reasoning that if CUL1 functions in SI as well as in UI, then suppressing CUL1 expression would disrupt pollen function in “collaborative non-self-recognition,” leading to its rejection by pistils of other LA2163 plants, but not by LA2157 pistils that lack S-RNase. Because plants were grown from a random sample of seed from this obligately outcrossing accession, each independent transgenic plant (designated T0-1, -2, etc.) and each nontransgenic pistil tester plant (LA2163-1, -2, -3) was expected to carry different S-genotypes. This was confirmed by controlled crosses between different LA2163 plants (transgenic or nontransgenic), which were all compatible (Figure S3), as expected. Thus, for CUL1 RNAi plants with a single T-DNA insertion, the specific expectation is a 1:1 transgene segregation in crosses onto LA2157 pistils and little or no transgene transmission in crosses onto LA2163 (Figure 1A). Plants with two unlinked insertions should show a 3:1 ratio in crosses to LA2157 and little or no transmission in crosses to LA2163.
To avoid cross-silencing of other Cullin1 gene family members, the RNAi construct was built from the combined 5′ + 3′ untranslated regions (UTRs) of CUL1 from LA2163 (Figure S1). The UTR sequences showed no homology to other Cullin1 genes in the tomato genome. Pollen-specific expression was driven by the LAT52 promoter (Twell et al. 1990). Stably transformed plants (T0) of LA2163 were obtained by Agrobacterium-mediated transformation. Figure 1B shows semiquantitative RT-PCR results for five T0 plants with reduced CUL1 messenger RNA (mRNA) levels (T0-1, with two insertions; T0-2 to T0-5 with single insertions). Since the T0 plants are hemizygous, the RNAi construct is present in 50% (single insertion) or 75% (two insertions) of pollen; thus, overall CUL1 mRNA levels in pollen from T0 plants was expected to be at least 25–50% of normal.
Observations of pollen tube growth in styles (Figure 2; full results in Figure S2 and Figure S3) showed that T0 plants behaved like nontransgenic LA2163 SI plants: self-pollinations are incompatible, and sib pollinations onto other LA2163 plants with different S-genotypes (transgenic or nontransgenic) are compatible, although some pollen tubes are arrested (Figure 2). These results were expected because hemizygous T0 plants produce both transgenic and nontransgenic pollen, which should manifest as at least 25–50% compatible pollen in crosses onto other LA2163 plants and 100% incompatible pollen in self-pollinations. The observation of some incompatible pollen tubes in the crosses onto LA2163 (transgenic or nontransgenic) is consistent with this prediction.
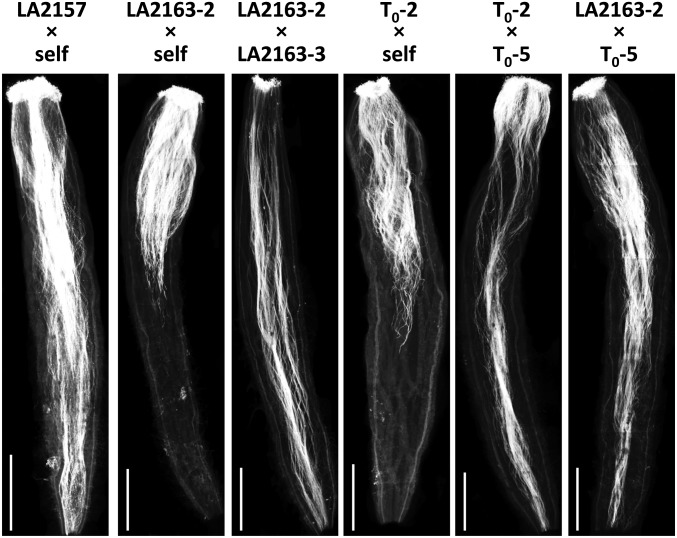
Figure 2.
Representative images showing pollen tube growth in pistils of (left to right) LA2157 self-pollinated; nontransgenic LA2163 plants selfed or crossed; T0 CUL1 RNAi plants selfed or crossed; and nontransgenic LA2163 pollinated by a T0 plant. Styles were fixed 24 hr after pollination, and pollen tubes were stained with aniline blue and visualized under UV light. Bar, 1 mm.
Transgene segregation ratios show that CUL1 is required for SI. When the four single-insertion hemizygous transformants (T0-2 to T0-5) are crossed onto SC LA2157, segregation of the transgene in the T1 progeny is consistent with the predicted 1:1 ratio (Figure 1C; full results in Table S2). These results also establish that pollen containing the CUL1 RNAi construct are viable. In contrast, the CUL1 RNAi transgene transmits poorly or not at all in crosses onto four different LA2163 tester plants (Figure 1C and Table S2), and the segregation ratios deviate significantly from 1:1 in every case (P < 0.0001). A double-insertion transformant (T0-1) also shows the predicted (3:1) segregation of the CUL1 RNAi transgene when crossed onto LA2157 but little transmission of the T-DNA in crosses onto other LA2163 plants (Figure 1C and Table S2). Since all five independent T0 plants, including one with two insertion loci, exhibited highly significant elimination of transgenic pollen on LA2163 pistils, we can rule out insertion-site effects or linkage to other genes under selection.
The best explanation for the transmission ratio differences is that CUL1 is required for pollen-side SI function. If CUL1 expression is necessary for pollen resistance to pistil S-RNases, then pollen that harbor the RNAi transgene should be rejected on pistils of any LA2163 plant, regardless of its S-genotype, leading to a deficiency of transgene transmission to the T1 progeny, as was observed. On the other hand, progeny from the crosses onto LA2157, which lacks active S-RNase, should segregate for the transgene in predicted Mendelian ratios, which again is consistent with our results.
Discussion
We previously reported that most accessions of the red-fruited tomato species display CUL1 mutations and proposed that CUL1 forms part of a pollen-resistance mechanism for S-RNase-based UI between red-fruited SC species and green-fruited SI species (Li and Chetelat 2010). Here we demonstrate that CUL1-deficient pollen are selectively eliminated on pistils following intraspecific crosses in SI S. arcanum. In contrast, crosses onto an S-RNase-deficient SC accession do not exhibit pollen elimination. These observations provide direct evidence that CUL1 function is limited to S-RNase-based pollen rejection mechanisms.
Our results also provide an explanation for earlier observations of segregation distortion near ui6.1 in certain interspecific mapping populations. Specifically, F2 progeny from hybrids between SC and SI species show preferential transmission of ui6.1-linked markers from the SI parent (Graham 2005; Trujillo-Moya et al. 2011). In contrast, F2 progeny from interspecific SC × SC crosses (i.e., lacking functional S-RNase) show normal Mendelian segregation ratios around ui6.1 (van Heusden et al. 1999; Li et al. 2010). The simplest interpretation of these results is that CUL1 activity is necessary for protecting growing pollen tubes against pistil S-RNases in compatible crosses. These findings establish that CUL1 functions in self- as well as interspecific incompatibility and provide further evidence of overlap, on the pollen side, between these forms of incompatibility.
Our results also suggest that mutations that suppress pollen SI function by blocking CUL1 expression will not be transmitted to the next generation unless preceded by loss of S-RNase expression in the pistil. In our study, CUL1-deficient pollen were selectively eliminated on pistils of other LA2163 plants and were transmitted at a low rate, or not at all, to the next generation. In natural populations, mutations that block CUL1 expression or activity in pollen are unlikely to become fixed unless expression of S-RNase in the pistil has already been lost. Thus CUL1 mutations are likely to be secondary mutational events that reinforce a prior loss of pistil-side SI. This prediction is consistent with the presence of a CUL1 loss-of-function mutant in cultivated tomato and other red- or orange-fruited species (Li and Chetelat 2010), a clade that is entirely SC and lacks both S-RNase and HT expression in the pistil (Kondo et al. 2002; Covey et al. 2010). Furthermore, to our knowledge, SC tomato species or populations lacking pollen SI function while retaining pistil function have not been reported. In contrast, SC mutations affecting only the pistil-side are known; for example, both Solanum pennellii LA0716 and S. arcanum LA2157 lack functional S-RNase in the pistil yet produce pollen that functions on pistils of conspecific SI accessions (Hardon 1967; Rick 1986; Royo et al. 1994; Covey et al. 2010).
Supplementary Material
Acknowledgments
We thank the C. M. Rick Tomato Genetics Resource Center staff for supplying seed stocks, Marcus Tamura for composing style images, Kim Carney and David Tricoli at the Parsons Plant Transformation Facility for producing transgenic plants, and Bruce McClure and Pat Bedinger for comments on the manuscript. The project was supported by National Science Foundation grant MCB 1127059.
Footnotes
Communicating editor: D. Charlesworth
Literature Cited
- Covey P., Kondo K., Welch L., Frank E., Sianta S., et al. , 2010. Multiple features that distinguish unilateral incongruity and self-incompatibility in the tomato clade. Plant J. 64: 367–378. [DOI] [PubMed] [Google Scholar]
- Graham, E. B., 2005 Genetic diversity and crossing relationships of Lycopersicon chilense. Ph.D. Thesis, University of California, Davis: 1–157. [Google Scholar]
- Hancock C. N., Kent L., McClure B. A., 2005. The stylar 120kDa glycoprotein is required for S-specific pollen rejection in Nicotiana. Plant J. 43: 716–723. [DOI] [PubMed] [Google Scholar]
- Hardon J. J., 1967. Unilateral incompatibility between Solanum pennellii and Lycopersicon esculentum. Genetics 57: 795–808. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Golz J. F., Oh H., Su V., Kusaba M., Newbigin E., 2001. Genetic analysis of Nictotiana pollen-part mutants is consistent with the presence of an S-ribonuclease inhibitor at the S-locus. Genetics 98: 15372–15376. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Igic B., Lande R., Kohn J. R., 2008. Loss of self-incompatibility and its evolutionary consequences. Int. J. Plant Sci. 169: 93–104. [Google Scholar]
- Kondo K., Yamamoto M., Matton D. P., Sato T., Hirai M., et al. , 2002. Cultivated tomato has defects in both S-RNase and HT genes required for stylar function of self-incompatibility. Plant J. 29: 627–636. [DOI] [PubMed] [Google Scholar]
- Kowyama Y., Kunz C., Lewis I., Newbigin E., Clarke A. E., et al. , 1994. Self-incompatibility in a Lycopersicon peruvianum variant (LA2157) is associated with a lack of style S-RNase activity. Theor. Appl. Genet. 88: 859–864. [DOI] [PubMed] [Google Scholar]
- Kubo K., Entani T., Takara A., Wang N., Fields A. M., et al. , 2010. Collaborative non-self recognition system in S-RNAse-based self-incompatibility. Science 330: 796–799. [DOI] [PubMed] [Google Scholar]
- Lewis D., Crowe L. K., 1958. Unilateral interspecific incompatibility in flowering plants. Heredity 12: 233–256. [Google Scholar]
- Li W., Chetelat R. T., 2010. A pollen factor linking inter- and intraspecific pollen rejection. Science 330: 1827–1830. [DOI] [PubMed] [Google Scholar]
- Li W., Royer S., Chetelat R. T., 2010. Fine mapping of ui6.1, a gametophytic factor controlling pollen-side unilateral incompatibility in interspecific Solanum hybrids. Genetics 185: 1069–1080. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McClure B. A., Haring V., Ebert P. R., Anderson M. A., Simpson R. J., et al. , 1989. Style self-incompatibility gene products of Nicotiana alata are ribonucleases. Nature 342: 955–957. [DOI] [PubMed] [Google Scholar]
- McClure B. A., Mou B., Canevascini S., Bernatzky R., 1999. A small asparagine-rich protein required for S-allele-specific pollen rejection in Nicotiana. Proc. Natl. Acad. Sci. USA 96: 13548–13553. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Moon J., Parry G., Estelle M., 2004. The ubiquitin-proteasome pathway and plant development. Plant Cell 16: 3181–3195. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Murfett J., Strabala T. J., Zurek D. M., Mou B., Beecher B., et al. , 1996. S RNase and interspecific pollen rejection in the genus Nicotiana: multiple pollen-rejection pathways contribute to unilateral incompatibility between self-incompatible and self-compatible species. Plant Cell 8: 943–958. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rick C. M., 1986. Reproductive isolation in the Lycopersicon peruvianum complex, pp. 477–496 in Solanaceae Biology and Systematics, edited by D’Arcy W. G. Columbia University Press, New York. [Google Scholar]
- Rick C. M., 1988. Evolution of mating systems in cultivated plants, pp. 133–147 in Plant Evolutionary Biology, edited by Gottlieb L. D., Jain S. K. Chapman and Hall, London. [Google Scholar]
- Rivers B. A., Bernatzky R., 1994. Protein expression of a self-compatible allele from Lycopersicon peruvianum: introgression and behavior in a self-incompatible background. Sex. Plant Reprod. 7: 357–362. [Google Scholar]
- Royo J., Kunz C., Kowyama Y., Anderson M., Clarke A. E., et al. , 1994. Loss of a histidine residue at the active site of S-locus ribonuclease is associated with self-compatibility in Lycopersicon peruvianum. Proc. Natl. Acad. Sci. USA 91: 6511–6514. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sijacic P., Wang W., Skirpan A. L., Wang Y., Dowd P. E., et al. , 2004. Identification of the pollen determinant of S-RNase-mediated self-incompatibility. Nature 429: 302–305. [DOI] [PubMed] [Google Scholar]
- Trujillo-Moya C., Gisbert C., Vilanova S., Nuez F., 2011. Localization of QTLs for in vitro plant regeneration in tomato. BMC Plant Biol. 11: 140. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Twell D., Yamaguchi J., McCormick S., 1990. Pollen-specific gene expression in transgenic plants: coordinate regulation of two different tomato gene promoters during microsporogenesis. Development 109: 705–713. [DOI] [PubMed] [Google Scholar]
- van Heusden A. W., Koorneef M., Voorrips R. E., Brüggemann W., Pet G., et al. , 1999. Three QTLs from Lycopersicon peruvianum confer a high level of resistance to Clavibacter michiganensis ssp. michiganensis. Theor. Appl. Genet. 99: 1068–1074. [Google Scholar]
- Zhang Y., Zhao Z., Xue Y., 2009. Roles of proteolysis in plant self-incompatibility. Annu. Rev. Plant Biol. 60: 21–42. [DOI] [PubMed] [Google Scholar]
- Zhao L., Huang J., Zhao Z., Li Q., Sims T. L., et al. , 2010. The Skp1-like protein SSK1 is required for cross-pollen compatibility in S-RNase-based self-incompatibility. Plant J. 62: 52–63. [DOI] [PubMed] [Google Scholar]
- Zheng N., Schulman B. A., Song L., Miller J. J., Jeffrey P. D., et al. , 2002. Structure of the Cul1-Rbx1-Skp1-F boxSkp2 SCF ubiquitin ligase complex. Nature 416: 703–709. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.