Abstract
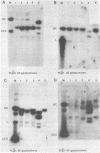
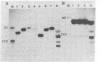
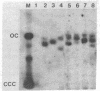
Dicentric plasmids containing either two copies of centromere 4 or one copy of centromere 4 and one copy of centromere 3 in the yeast plasmid vector YRp17 were constructed in vitro and introduced into yeast cells by DNA transformation. The resulting colonies were heterogeneous for a mixed population of rearranged plasmids. The rearrangements always involved deletion of one or both centromere sequences originally present on the plasmid. Heterogeneity was due to the continued production of deleted plasmids from a pool of unrearranged dicentric plasmids maintained within some of the yeast cells in the colony. The RAD52 gene product is known to be required for the repair of DNA double-strand breaks in yeast. Transformation of rad52 mutant yeast cells with dicentric plasmids gave rearranged plasmids similar to those observed with RAD+ yeast cells, but the transformation frequency was only 5-10% compared to transformation with monocentric plasmids. Also, the ratio of unrearranged dicentric plasmid to deleted plasmids was greatly reduced in the rad52-transformed cells. These observations are consistent with a model in which centromeric DNA sequences can interact independently with the yeast cell spindle apparatus. Occasional movement of centromeres to opposite poles may result in mechanical breakage of plasmid sequences. Plasmids deleted for one or both centromere sequences can be obtained from these broken molecules and are resistant to further rearrangement.
Full text
PDF
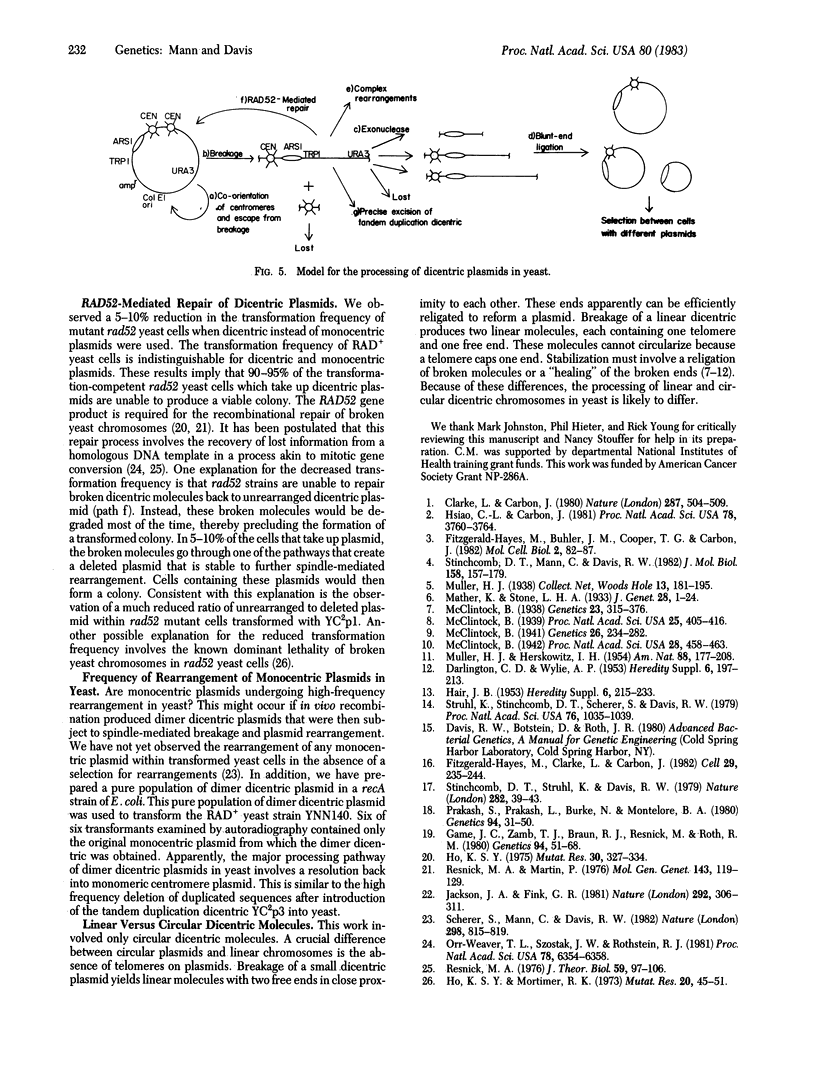



Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Clarke L., Carbon J. Isolation of a yeast centromere and construction of functional small circular chromosomes. Nature. 1980 Oct 9;287(5782):504–509. doi: 10.1038/287504a0. [DOI] [PubMed] [Google Scholar]
- Fitzgerald-Hayes M., Buhler J. M., Cooper T. G., Carbon J. Isolation and subcloning analysis of functional centromere DNA (CEN11) from Saccharomyces cerevisiae chromosome XI. Mol Cell Biol. 1982 Jan;2(1):82–87. doi: 10.1128/mcb.2.1.82. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fitzgerald-Hayes M., Clarke L., Carbon J. Nucleotide sequence comparisons and functional analysis of yeast centromere DNAs. Cell. 1982 May;29(1):235–244. doi: 10.1016/0092-8674(82)90108-8. [DOI] [PubMed] [Google Scholar]
- Game J. C., Zamb T. J., Braun R. J., Resnick M., Roth R. M. The Role of Radiation (rad) Genes in Meiotic Recombination in Yeast. Genetics. 1980 Jan;94(1):51–68. doi: 10.1093/genetics/94.1.51. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ho K. S. Induction of DNA double-strand breaks by X-rays in a radiosensitive strain of the yeast Saccharomyces cerevisiae. Mutat Res. 1975 Dec;30(3):327–334. [PubMed] [Google Scholar]
- Ho K. S., Mortimer R. K. Induction of dominant lethality by x-rays in radiosensitive strain of yeast. Mutat Res. 1973 Oct;20(1):45–51. doi: 10.1016/0027-5107(73)90096-1. [DOI] [PubMed] [Google Scholar]
- Hsiao C. L., Carbon J. Direct selection procedure for the isolation of functional centromeric DNA. Proc Natl Acad Sci U S A. 1981 Jun;78(6):3760–3764. doi: 10.1073/pnas.78.6.3760. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jackson J. A., Fink G. R. Gene conversion between duplicated genetic elements in yeast. Nature. 1981 Jul 23;292(5821):306–311. doi: 10.1038/292306a0. [DOI] [PubMed] [Google Scholar]
- McClintock B. The Behavior in Successive Nuclear Divisions of a Chromosome Broken at Meiosis. Proc Natl Acad Sci U S A. 1939 Aug;25(8):405–416. doi: 10.1073/pnas.25.8.405. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McClintock B. The Production of Homozygous Deficient Tissues with Mutant Characteristics by Means of the Aberrant Mitotic Behavior of Ring-Shaped Chromosomes. Genetics. 1938 Jul;23(4):315–376. doi: 10.1093/genetics/23.4.315. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McClintock B. The Stability of Broken Ends of Chromosomes in Zea Mays. Genetics. 1941 Mar;26(2):234–282. doi: 10.1093/genetics/26.2.234. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Orr-Weaver T. L., Szostak J. W., Rothstein R. J. Yeast transformation: a model system for the study of recombination. Proc Natl Acad Sci U S A. 1981 Oct;78(10):6354–6358. doi: 10.1073/pnas.78.10.6354. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Prakash S., Prakash L., Burke W., Montelone B. A. Effects of the RAD52 Gene on Recombination in SACCHAROMYCES CEREVISIAE. Genetics. 1980 Jan;94(1):31–50. doi: 10.1093/genetics/94.1.31. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Resnick M. A., Martin P. The repair of double-strand breaks in the nuclear DNA of Saccharomyces cerevisiae and its genetic control. Mol Gen Genet. 1976 Jan 16;143(2):119–129. doi: 10.1007/BF00266917. [DOI] [PubMed] [Google Scholar]
- Resnick M. A. The repair of double-strand breaks in DNA; a model involving recombination. J Theor Biol. 1976 Jun;59(1):97–106. doi: 10.1016/s0022-5193(76)80025-2. [DOI] [PubMed] [Google Scholar]
- Scherer S., Mann C., Davis R. W. Reversion of a promoter deletion in yeast. Nature. 1982 Aug 26;298(5877):815–819. doi: 10.1038/298815a0. [DOI] [PubMed] [Google Scholar]
- Stinchcomb D. T., Mann C., Davis R. W. Centromeric DNA from Saccharomyces cerevisiae. J Mol Biol. 1982 Jun 25;158(2):157–190. doi: 10.1016/0022-2836(82)90427-2. [DOI] [PubMed] [Google Scholar]
- Stinchcomb D. T., Struhl K., Davis R. W. Isolation and characterisation of a yeast chromosomal replicator. Nature. 1979 Nov 1;282(5734):39–43. doi: 10.1038/282039a0. [DOI] [PubMed] [Google Scholar]
- Struhl K., Stinchcomb D. T., Scherer S., Davis R. W. High-frequency transformation of yeast: autonomous replication of hybrid DNA molecules. Proc Natl Acad Sci U S A. 1979 Mar;76(3):1035–1039. doi: 10.1073/pnas.76.3.1035. [DOI] [PMC free article] [PubMed] [Google Scholar]