Abstract
Tethering is a screening technique for discovering small-molecule fragments that bind to pre-determined sites via formation of a disulphide bond. Tethering screens traditionally rely upon mass spectrometry to detect disulphide bind formation, which requires a time-consuming liquid chromatography step. Here we show that Tethering can be performed rapidly and inexpensively using a homogenous fluorescence polarization (FP) assay that detects displacement of a peptide ligand from the protein target as an indirect readout of disulphide formation. We apply this method, termed FP Tethering, to identify fragments that disrupt the protein-protein interaction between the KIX domain of the transcriptional coactivator CBP and the transcriptional activator peptide pKID.
Introduction
Protein-protein interactions (PPIs) underpin all cellular processes, and dysregulation of PPI networks is strongly correlated with human disease.1 For this reason, synthetic molecules that modulate PPIs are highly sought tools. Despite their importance, small molecule PPIs modulators are difficult to obtain through either screening or design.2 This is largely due to the intrinsic features of PPIs, most of which are comprised of significantly larger surface area (average ~1949 ± 760 Å2) than typical protein-ligand interfaces and are often flatter with few interaction features.3 Indeed, PPIs have often been described as ‘undruggable’ due to the challenges associated with identifying small molecules that can effectively engage these binding interfaces.
Tethering is a screening strategy that circumvents many of the difficulties associated with PPIs and has been successfully used to discover a range of small molecule modulators.4, 5 It is a fragment discovery method in which a library of disulphide-containing fragments (MW <300 Da) are screened under reversible conditions against a protein target bearing a native or engineered cysteine adjacent to the binding site of interest (Figure 1a).6, 7 Fragments that interact favourably with the protein target bias the equilibrium towards the mixed disulphide, which is subsequently detected by liquid chromatography-mass spectrometry (LC-MS) (Figure 1b). The resulting fragment molecules can be converted to non-covalent inhibitors by growing the fragment or used as covalent binders for functional and biophysical studies.8
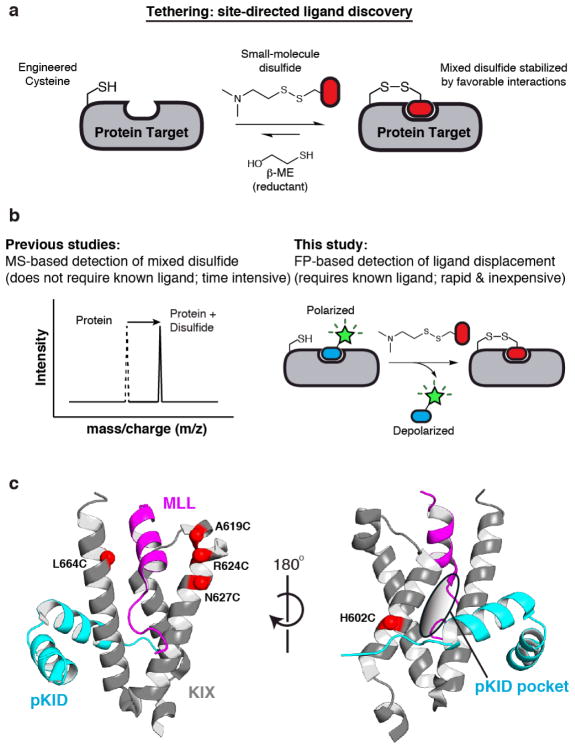
Figure 1.
Tethering combined with a ligand displacement assay is used to identify chemical probes that disrupt the interaction between KIX and its binding partners. A) Schematic of the Tethering methodology for site-directed ligand discovery. A reversible disulphide-trapping reaction is set up such that the mixed disulphide only persists when the ligand makes favourable interactions with the protein. Increasing the β-ME concentration results in a more stringent assay. B) Comparison between traditional liquid chromatography-mass spectrometry (LC-MS)-based Tethering with the newly devised fluorescence polarization (FP)-based Tethering. C) Cartoon representation of the KIX domain (grey) bound to two coactivator peptides MLL (magenta) and pKID (cyan). (PDB IDs: 2AGH, 1SB0) Previously, four cysteines lining the MLL-binding pocket (left) were screened using the LC-MS-based Tethering. Here, we use FP-based Tethering to target the pKID-binding pocket by introducing a cysteine at position H602 (right).
We recently reported the application of Tethering to discover small molecule ligands for the KIX domain of the master coactivator CBP/p300.9 Several fragments identified from this screen proved to be excellent inhibitors of the PPI formed between KIX and the transcriptional activator MLL as well as enhancers of the complex that KIX forms with the transcriptional activation domain of CREB (pKID). Nonetheless, we noted that some of the fragments that bound to KIX efficiently did not affect KIX PPIs. In retrospect this is not surprising since the readout of the screen is binding, not inhibition. We hypothesized that combining Tethering with a binding readout would provide a more direct route to PPI modulators (Figure 1b). Here we show that fluorescence polarization (FP) Tethering is a rapid method for the direct discovery of PPI modulators and use this method for the identification of effective inhibitors of the complex formed between KIX and pKID.
Results and Discussion
Fluorescence Polarisation (FP) is a homogenous, mix-and-read method used to directly measure the fraction of a small fluorescently-labelled peptide tracer that is bound to a larger protein.10 Binding of the tracer to the protein results in high fluorescence polarization, but the tracer’s fluorescence is rapidly depolarized following displacement by an unlabelled ligand (Figure 1b).11 This type of ligand displacement assay provides a way to measure the affinity between the protein target and an unlabelled ligand. However, FP ligand displacement assays are less sensitive than x-ray crystallography or NMR for detecting the low affinity interactions of fragments, which typically occur in the hundreds of micromolar to millimolar range.12, 13 Since Tethering of fragments via a disulphide bond enhances their affinity 10- to 100-fold, we reasoned that these binding events should be detectable by FP.
To test this approach we first used the results from our earlier Tethering screen against cysteine mutants of the KIX domain. The KIX domain is a small (90 residue) three-helix bundle containing two binding sites that can be occupied simultaneously by different transcriptional activators (Figure 1c).14–18 These two binding sites are in allosteric communication, with the binding of MLL in the smaller and deeper of the two sites leading to up to 2-fold enhancement of binding with activators such as pKID (the transcriptional activation domain of CREB) at the second, more shallow binding site.16, 19 Two disulphide fragments isolated from the original Tethering screen bind well to a N627C mutant of KIX, fragments 1A10 and 1C11, and also were effective inhibitors of MLL binding when covalently associated with KIX N627C.9 Rather than measuring binding inhibition using a fully labeled protein, we reasoned a competitive inhibition readout could report on fragment-labelling of the KIX N627C mutant during the Tethering reaction. Indeed, in the presence of 5 mM BME, 82% of the KIX N627C mutant was labelled in the presence of 250 μM of 1A10, and inhibition of a fluorescently-labeled MLL peptide was observed in a dose-dependent fashion as a functional consequence of protein labeling (IC50 68 μM, see SI Figure S1). The ability to increase or decrease percent inhibition levels by changing the BME concentration affords a control for ruling out fluorescent impurities or other false positives.
Next, 80 disulphide-containing fragments were screened at two different β-ME concentrations against the KIX N627C mutant (Figure 2b). The FP Tethering screen identified the same set of ligands that were previously identified in the MS-based screen (Figure 2c). Notably the hits that resulted in the largest inhibition of the KIX domain and MLL interaction were also the most potent hits from our MS-based screen (data not shown). These results suggested that the FP Tethering assay was suitable for high-throughput identification of fragments that disrupt the interaction between KIX and its coactivators.
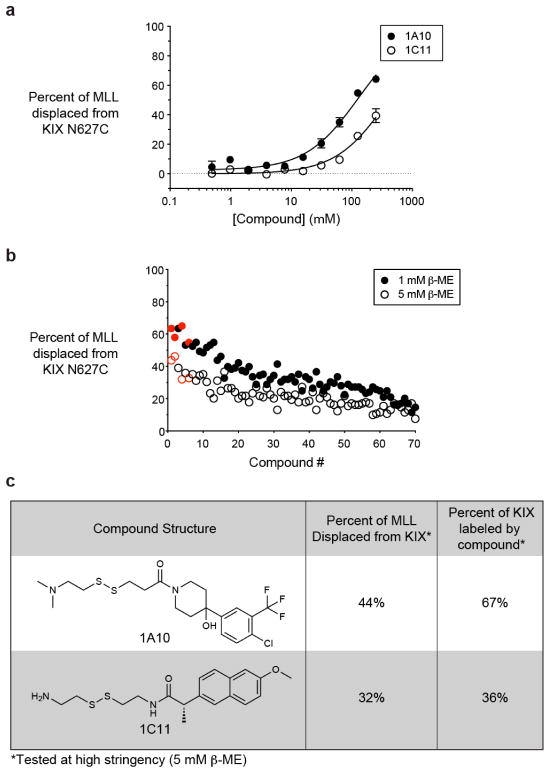
Figure 2.
FP- and MS-based Tethering identify overlapping ligand sets for the MLL-binding site. A) Dose response of previously identified small-molecule disulphides in a FP displacement assay with KIX N627C and fluorophore-tagged MLL. B) The percent of MLL displaced from KIX N627C is shown for 70 small-molecule disulphides. The screen was performed at medium and high stringency (1 mM and 5 mM β-ME, respectively). The top 4 fragments that were previously identified by MS-based Tethering are coloured in red. C) The structures and properties of the two fragments that were rediscovered by FP-based Tethering.
To test the FP Tethering approach in a pure discovery mode, we targeted the second binding site within the KIX domain, that used by pKID in addition to the activator c-Myb. This is the considerably more challenging of the two sites, with a larger surface area (1480 Å2) and a less defined surface.20 In preparation for the screen, we mutated several residues lining the binding site to cysteine and determined the affinity of the mutant KIX domains for both pKID and c-Myb tracers (see SI Table S1). The KIX H602C mutant retained its affinity for both tracers and it was thus chosen for the FP Tethering screen. The competition assay was developed with 25 nM FITC-labelled pKID and 4 μM of KIX H602C. These conditions were sufficient to detect displacement of FITC-labelled pKID tracer from the KIX domain by the unlabelled pKID peptide.
With these assay conditions in hand, we screened 960 disulphide-containing fragments at three different β-ME concentrations against a fluorescent pKID tracer in complex with KIX H602C (Figure 3a). Fragments that yielded total fluorescence intensities (TFI) greater than 150% of the DMSO control wells were flagged as fluorescent artifacts and excluded from further analysis. Next the remaining fragments were ranked according to percent inhibition of pKID binding across the three β-ME concentrations (Figure 3b). As expected, the highest stringency of 5 mM β-ME resulted in decreased inhibition relative to the lower β-ME concentrations. The two lower β-ME concentrations, 0.2 mM and 1 mM, deviated from this trend in which the lowest percent inhibition fluctuated between 1 mM β-ME and 0.2 mM β-ME for the different fragments.
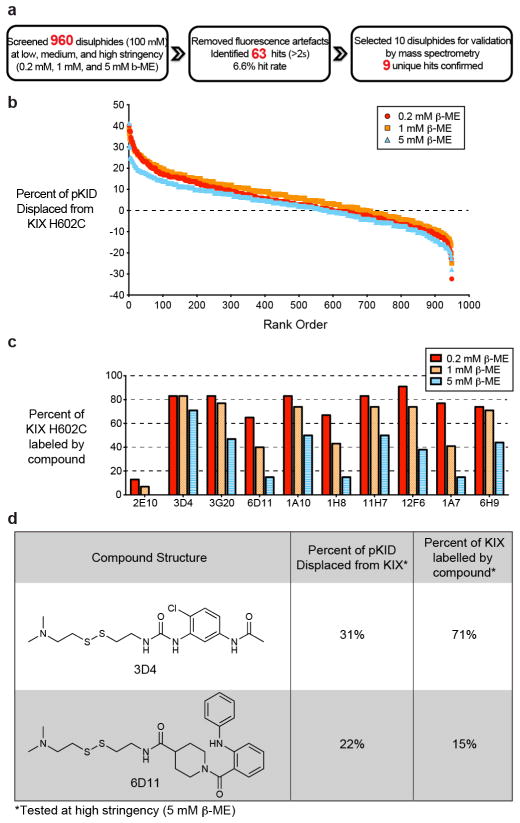
Figure 3.
FP-based Tethering screen to identify new ligands that disrupt the interaction between KIX and pKID. A) Overview of the workflow for screening, hit selection, and validation. B) The percent of pKID displaced from KIX H602C is shown for 960 small-molecule disulphides at low, medium, and high stringency (0.2 mM, 1 mM, and 5 mM β-ME, respectively). C) The percent of KIX H602C that is labelled by the top 10 small-molecule disulphides was measured by mass spectrometry (MS). The reaction was performed with increasing stringency to permit assessment of relative affinities. D) The structures and properties of ligands that displace pKID from KIX H602C.
From the top hits, 10 disulphides were cherry-picked for validation by LC-MS analysis (Figure 3c). All of these fragments covalently labelled the KIX domain. However 2E10 was likely a false positive given its high percent inhibition by FP yet a low degree of labelling by MS. Comparison of labelling across the three β-ME concentrations reveals the relative potencies of the fragments. From the ten hits, two fragments, 3D4 and 6D11 were chosen for further characterisation. The fragment 3D4 labelled KIX H602C more effectively than any other fragments and maintained a high percent inhibition of pKID even at 5 mM β-ME (31%) (Figure 3d). The fragment 6D11 had the highest average percent inhibition of pKID for the three β-ME concentration. Notably, percent labelling did not always correlate with inhibition of pKID binding. While fragment 3D4 labelled KIX ~70% at 5 mM β-ME, this only resulted in 24% inhibition of pKID binding. The larger fragment 6D11 labelled KIX only ~15% at 5 mM β-ME, but this translated to nearly equivalent inhibition (22%) of pKID binding. This finding suggests that although 6D11 may exhibit weaker overall potency, it is the more efficient inhibitor. Other top hits replaced the diphenylamine moiety in 6D11 for a diphenylmethane or 2-chloro-2-N-phenylaniline. These changes did not improve their ability to displace the pKID tracer, but their ability to bind to the KIX H602C mutant improved slightly. The disulphide fragment library contained several variants of the core scaffolds of 3D4 and 6D11 and a comparison revealed that both structures are sensitive to substituent placement and identity; this is particularly true for 6D11 where modifications of the distal aromatic ring leads to dramatic changes in inhibition (SI Figure S2).
For comparison to previous LC-MS Tethering screens, dose-response curves were determined for 3D4 and 6D11 using LC-MS to determine the percent of KIX H602C mutant bound to the fragment. (Figure 4a). The concentration at which 50% of the KIX H602C mutant is bound by the fragment (DR50) was lower with the fragment 3D4 than 6D11. This finding was consistent with single point LC-MS measurements in which the fragment 3D4 was more potent than the fragment 6D11.
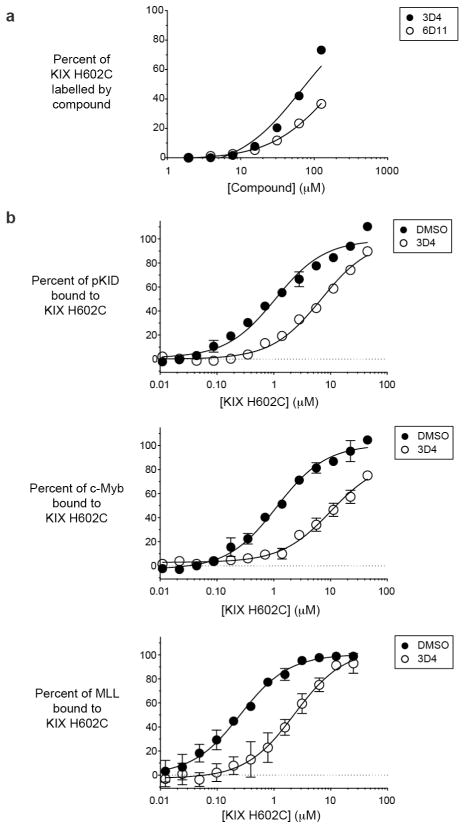
Figure 4.
Labelling of KIX H602C with 3D4 decreases affinity for both pKID and MLL. A) The dose-response binding experiment was performed with KIX H602C and fragments 3D4 and 6D11 using LC-MS analysis. B) Dose response of KIX H602C (closed circles) or KIX H602C labelled with 3D4 (open circles) against fixed concentration of fluorophore-tagged pKID, c-Myb, or MLL.
Finally, we labelled greater than 90% of KIX H602C with the 3D4 fragment and used this protein-ligand conjugate for direct binding experiments with the MLL, pKID, and c-Myb tracers. The equilibrium dissociation constant (Kd) of each tracer for the KIX-3D4 conjugate was assessed by a binding isotherm that accounts for ligand depletion.10, 21 Surprisingly, 3D4 not only increases the Kd for binding to pKID (6.5-fold) and c-Myb (9-fold) but also MLL (9-fold), which binds on the opposite face of the KIX domain. This finding demonstrates a negative allosteric coupling between the pKID- and MLL-binding sites, the first observation of this phenomenon. Given that 3D4 disrupts the binding of all three coactivator peptides, we are pursuing irreversible covalent analogues as chemical probes for studying CBP/p300 biology.
In conclusion, FP Tethering is an efficient and accessible screening format for identifying fragments that disrupt a PPI. This method was successfully benchmarked against a previous MS-based Tethering screen on the MLL-binding site of KIX. Application of this method in discovery mode against a more challenging surface within KIX, the broad interface of the KIX:pKID complex, was also a success. FP Tethering lead to 9 fragments confirmed to displace pKID from the KIX domain and by covalently labelling the protein. These molecules represent promising leads for follow-up studies. More broadly, these results suggest that Tethering could be combined with any binding assay that is compatible with disulphide-containing molecules and β-ME,22, 23 increasingly the accessibility of this important technology.
Supplementary Material
Acknowledgments
T.J.R was supported by a predoctoral fellowship from the NIH (F31 CA180378-01). W.C.P. is thankful to the NIH for a postdoctoral fellowship (F32 GM090550). Aspects of this work were supported by NIH 2R01 GM65330 (to A.K.M.) and NIH R01AI070292 (J.W.) We thank Dr. Mark Burlingame for maintaining the Tethering library, Dr. Chris Wilson for assisting with automation equipment, and Drs. Olivier Julien and Zachary Hill for helpful discussions. We also thank Laura C. Cesa for the providing several KIX cysteine mutant plasmids and Drs. Ningkun Wang and Chinmay Y. Majmudar for helpful discussions.
Footnotes
Electronic Supplementary Information (ESI) available: this includes experimental details of protein expression and screening. See DOI: 10.1039/b000000x/
References
- 1.Berg T. Angew Chem Int Ed Engl. 2003;42:2462–2481. doi: 10.1002/anie.200200558. [DOI] [PubMed] [Google Scholar]
- 2.Wells JA, McClendon CL. Nat Rev Drug Discov. 2007;240:1001–1009. [Google Scholar]
- 3.Thompson AD, Dugan A, Gestwicki JE, Mapp AK. ACS Chem Biol. 2012;7:1311–1320. doi: 10.1021/cb300255p. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Braisted AC, Oslob JD, Delano WL, Hyde RS, Waal N, Yu C, Arkin MR, Raimundo BC. J Am Chem Soc. 2003;125:3714–3715. doi: 10.1021/ja034247i. [DOI] [PubMed] [Google Scholar]
- 5.Sadowsky JD, Burlingame MA, Wolan DW, McClendon CL, Jacobson MP, Wells JA. Proc Natl Acad Sci USA. 2011;108:6056–6061. doi: 10.1073/pnas.1102376108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Erlanson DA, Braisted AC, Raphael DR, Randal M, Stroud RM, Gordon EM, Wells JA. Proc Natl Acad Sci USA. 2000;19:9367–9372. doi: 10.1073/pnas.97.17.9367. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Erlanson DA, Wells JA, Braisted AC. Annu Rev Biophys and Biomol Struct. 2004;33:199–223. doi: 10.1146/annurev.biophys.33.110502.140409. [DOI] [PubMed] [Google Scholar]
- 8.Nonoo RH, Armstrong A, Mann DJ. ChemMedChem. 2012;7:2082–2086. doi: 10.1002/cmdc.201200404. [DOI] [PubMed] [Google Scholar]
- 9.Wang N, Majmudar CY, Pomerantz WC, Gagnon JK, Sadowsky JD, Meagher JL, Johnson TK, Stuckey JA, Brooks CL, III, Wells JA, Mapp AK. J Am Chem Soc. 2013;125:3363–3366. doi: 10.1021/ja3122334. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Lakowicz JR. Principles of Fluorescence Spectroscopy. Springer; 2007. [Google Scholar]
- 11.Owicki JC. J Biomol Screen. 2000;5:297–306. doi: 10.1177/108705710000500501. [DOI] [PubMed] [Google Scholar]
- 12.Huang X. J Biomol Screen. 2003;8:34–38. doi: 10.1177/1087057102239666. [DOI] [PubMed] [Google Scholar]
- 13.Kobayashi M, Retra K, Figaroa F, Hollander JG, Ab E, Heetebrij RJ, Irth H, Siegal G. J Biomol Screen. 2010;15:978–989. doi: 10.1177/1087057110375614. [DOI] [PubMed] [Google Scholar]
- 14.Radhakrishnan I, Perez-Alvarado GC, Parker D, Dyson HJ, Montminy MR, Wright PE. Cell. 1997;91:741–752. doi: 10.1016/s0092-8674(00)80463-8. [DOI] [PubMed] [Google Scholar]
- 15.Radhakrishnan I, Perez-Alvarado GC, Parker D, Dyson HJ, Montminy MR, Wright PE. J Mol Biol. 1999;287:859–865. doi: 10.1006/jmbi.1999.2658. [DOI] [PubMed] [Google Scholar]
- 16.Goto NK, Zor T, Martinez-Yamout M, Dyson HJ, Wright PE. J Biol Chem. 2002;277:43168–43174. doi: 10.1074/jbc.M207660200. [DOI] [PubMed] [Google Scholar]
- 17.Campbell KM, Lumb KJ. Biochemistry. 2002;41:13956–13964. doi: 10.1021/bi026222m. [DOI] [PubMed] [Google Scholar]
- 18.Vendel AC, McBryant SJ, Lumb KJ. Biochemistry. 2003;42:12481–12487. doi: 10.1021/bi0353023. [DOI] [PubMed] [Google Scholar]
- 19.De Guzman RN, Goto NK, Dyson HJ, Wright PE. J Mol Biol. 2006;355:1005–1013. doi: 10.1016/j.jmb.2005.09.059. [DOI] [PubMed] [Google Scholar]
- 20.Zor T, De Guzman RN, Dyson HJ, Wright PE. J Mol Biol. 2004;337:521–534. doi: 10.1016/j.jmb.2004.01.038. [DOI] [PubMed] [Google Scholar]
- 21.Hulme EC, Trevethick MA. Br J Pharmacol. 2010;161:1219–1237. doi: 10.1111/j.1476-5381.2009.00604.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Buck E, Bourne H, Wells JA. J Biol Chem. 2005;280:4009–4012. doi: 10.1074/jbc.C400500200. [DOI] [PubMed] [Google Scholar]
- 23.Buck E, Wells JA. Proc Natl Acad Sci USA. 2005;102:2719–2724. doi: 10.1073/pnas.0500016102. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.