Abstract
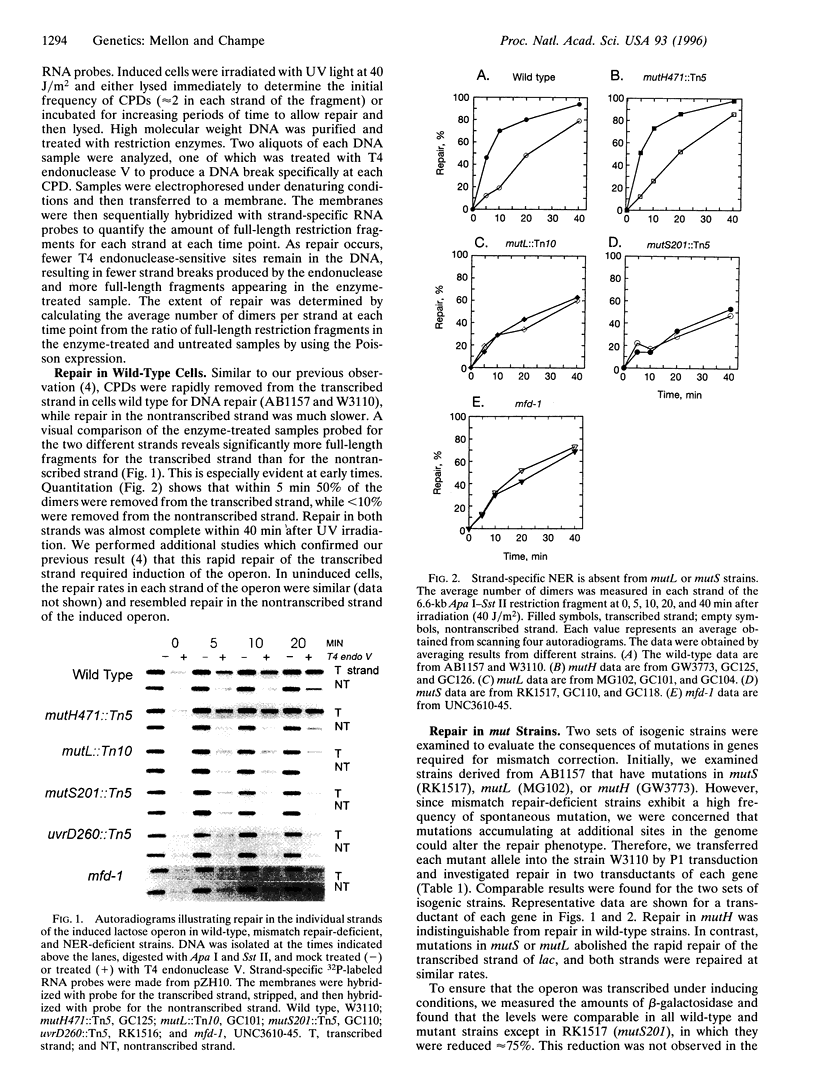
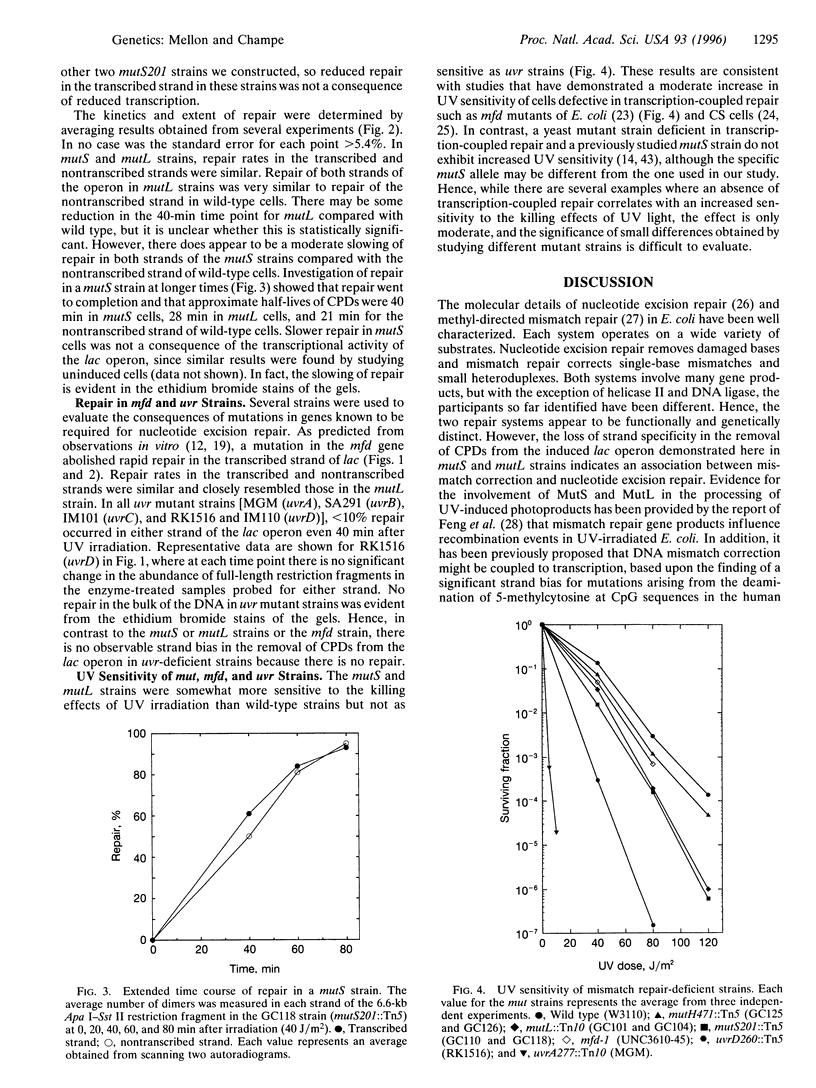
To improve our understanding of the mechanism that couples nucleotide-excision repair to transcription in expressed genes, we have examined the effects of mutations in several different DNA repair genes on the removal of cyclobutane pyrimidine dimers from the individual strands of the induced lactose operon in UV-irradiated Escherichia coli. As expected, we found little repair in either strand of the lactose operon in strains with mutations in established nucleotide excision-repair genes (uvrA, uvrB, uvrC, or uvrD). In contrast, we found that mutations in either of two genes required for DNA-mismatch correction (mutS and mutL) selectively abolish rapid repair in the transcribed strand and render the cells moderately sensitive to UV irradiation. Similar results were found in a strain with a mutation in the mfd gene, the product of which has been previously shown to be required for transcription-coupled repair in vitro. Our results demonstrate an association between mismatch-correction and nucleotide-excision repair and implicate components of DNA-mismatch repair in transcription-coupled repair. In addition, they may have important consequences for human disease and may enhance our understanding of the etiology of certain cancers which have been associated with defects in mismatch correction.
Full text
PDF



Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Bhattacharyya N. P., Skandalis A., Ganesh A., Groden J., Meuth M. Mutator phenotypes in human colorectal carcinoma cell lines. Proc Natl Acad Sci U S A. 1994 Jul 5;91(14):6319–6323. doi: 10.1073/pnas.91.14.6319. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bohr V. A., Smith C. A., Okumoto D. S., Hanawalt P. C. DNA repair in an active gene: removal of pyrimidine dimers from the DHFR gene of CHO cells is much more efficient than in the genome overall. Cell. 1985 Feb;40(2):359–369. doi: 10.1016/0092-8674(85)90150-3. [DOI] [PubMed] [Google Scholar]
- Bronner C. E., Baker S. M., Morrison P. T., Warren G., Smith L. G., Lescoe M. K., Kane M., Earabino C., Lipford J., Lindblom A. Mutation in the DNA mismatch repair gene homologue hMLH1 is associated with hereditary non-polyposis colon cancer. Nature. 1994 Mar 17;368(6468):258–261. doi: 10.1038/368258a0. [DOI] [PubMed] [Google Scholar]
- Caillet-Fauquet P., Maenhaut-Michel G., Radman M. SOS mutator effect in E. coli mutants deficient in mismatch correction. EMBO J. 1984 Apr;3(4):707–712. doi: 10.1002/j.1460-2075.1984.tb01873.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Christians F. C., Hanawalt P. C. Inhibition of transcription and strand-specific DNA repair by alpha-amanitin in Chinese hamster ovary cells. Mutat Res. 1992 Aug;274(2):93–101. doi: 10.1016/0921-8777(92)90056-9. [DOI] [PubMed] [Google Scholar]
- Donahue B. A., Yin S., Taylor J. S., Reines D., Hanawalt P. C. Transcript cleavage by RNA polymerase II arrested by a cyclobutane pyrimidine dimer in the DNA template. Proc Natl Acad Sci U S A. 1994 Aug 30;91(18):8502–8506. doi: 10.1073/pnas.91.18.8502. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Feng W. Y., Lee E. H., Hays J. B. Recombinagenic processing of UV-light photoproducts in nonreplicating phage DNA by the Escherichia coli methyl-directed mismatch repair system. Genetics. 1991 Dec;129(4):1007–1020. doi: 10.1093/genetics/129.4.1007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fishel R., Lescoe M. K., Rao M. R., Copeland N. G., Jenkins N. A., Garber J., Kane M., Kolodner R. The human mutator gene homolog MSH2 and its association with hereditary nonpolyposis colon cancer. Cell. 1993 Dec 3;75(5):1027–1038. doi: 10.1016/0092-8674(93)90546-3. [DOI] [PubMed] [Google Scholar]
- Hanawalt P. C., Donahue B. A., Sweder K. S. Repair and transcription. Collision or collusion? Curr Biol. 1994 Jun 1;4(6):518–521. doi: 10.1016/s0960-9822(00)00112-3. [DOI] [PubMed] [Google Scholar]
- Kantor G. J., Barsalou L. S., Hanawalt P. C. Selective repair of specific chromatin domains in UV-irradiated cells from xeroderma pigmentosum complementation group C. Mutat Res. 1990 May;235(3):171–180. doi: 10.1016/0921-8777(90)90071-c. [DOI] [PubMed] [Google Scholar]
- Leach F. S., Nicolaides N. C., Papadopoulos N., Liu B., Jen J., Parsons R., Peltomäki P., Sistonen P., Aaltonen L. A., Nyström-Lahti M. Mutations of a mutS homolog in hereditary nonpolyposis colorectal cancer. Cell. 1993 Dec 17;75(6):1215–1225. doi: 10.1016/0092-8674(93)90330-s. [DOI] [PubMed] [Google Scholar]
- Leadon S. A., Cooper P. K. Preferential repair of ionizing radiation-induced damage in the transcribed strand of an active human gene is defective in Cockayne syndrome. Proc Natl Acad Sci U S A. 1993 Nov 15;90(22):10499–10503. doi: 10.1073/pnas.90.22.10499. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Leadon S. A., Lawrence D. A. Preferential repair of DNA damage on the transcribed strand of the human metallothionein genes requires RNA polymerase II. Mutat Res. 1991 Jul;255(1):67–78. doi: 10.1016/0921-8777(91)90019-l. [DOI] [PubMed] [Google Scholar]
- Leadon S. A., Lawrence D. A. Strand-selective repair of DNA damage in the yeast GAL7 gene requires RNA polymerase II. J Biol Chem. 1992 Nov 15;267(32):23175–23182. [PubMed] [Google Scholar]
- Loeb L. A. Microsatellite instability: marker of a mutator phenotype in cancer. Cancer Res. 1994 Oct 1;54(19):5059–5063. [PubMed] [Google Scholar]
- Mellon I., Hanawalt P. C. Induction of the Escherichia coli lactose operon selectively increases repair of its transcribed DNA strand. Nature. 1989 Nov 2;342(6245):95–98. doi: 10.1038/342095a0. [DOI] [PubMed] [Google Scholar]
- Mellon I., Spivak G., Hanawalt P. C. Selective removal of transcription-blocking DNA damage from the transcribed strand of the mammalian DHFR gene. Cell. 1987 Oct 23;51(2):241–249. doi: 10.1016/0092-8674(87)90151-6. [DOI] [PubMed] [Google Scholar]
- Modrich P. Mechanisms and biological effects of mismatch repair. Annu Rev Genet. 1991;25:229–253. doi: 10.1146/annurev.ge.25.120191.001305. [DOI] [PubMed] [Google Scholar]
- Oller A. R., Fijalkowska I. J., Dunn R. L., Schaaper R. M. Transcription-repair coupling determines the strandedness of ultraviolet mutagenesis in Escherichia coli. Proc Natl Acad Sci U S A. 1992 Nov 15;89(22):11036–11040. doi: 10.1073/pnas.89.22.11036. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Papadopoulos N., Nicolaides N. C., Wei Y. F., Ruben S. M., Carter K. C., Rosen C. A., Haseltine W. A., Fleischmann R. D., Fraser C. M., Adams M. D. Mutation of a mutL homolog in hereditary colon cancer. Science. 1994 Mar 18;263(5153):1625–1629. doi: 10.1126/science.8128251. [DOI] [PubMed] [Google Scholar]
- Parker B. O., Marinus M. G. Repair of DNA heteroduplexes containing small heterologous sequences in Escherichia coli. Proc Natl Acad Sci U S A. 1992 Mar 1;89(5):1730–1734. doi: 10.1073/pnas.89.5.1730. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Parsons R., Li G. M., Longley M. J., Fang W. H., Papadopoulos N., Jen J., de la Chapelle A., Kinzler K. W., Vogelstein B., Modrich P. Hypermutability and mismatch repair deficiency in RER+ tumor cells. Cell. 1993 Dec 17;75(6):1227–1236. doi: 10.1016/0092-8674(93)90331-j. [DOI] [PubMed] [Google Scholar]
- Pu W. T., Kahn R., Munn M. M., Rupp W. D. UvrABC incision of N-methylmitomycin A-DNA monoadducts and cross-links. J Biol Chem. 1989 Dec 5;264(34):20697–20704. [PubMed] [Google Scholar]
- Selby C. P., Sancar A. Mechanisms of transcription-repair coupling and mutation frequency decline. Microbiol Rev. 1994 Sep;58(3):317–329. doi: 10.1128/mr.58.3.317-329.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Selby C. P., Sancar A. Molecular mechanism of transcription-repair coupling. Science. 1993 Apr 2;260(5104):53–58. doi: 10.1126/science.8465200. [DOI] [PubMed] [Google Scholar]
- Selby C. P., Sancar A. Transcription preferentially inhibits nucleotide excision repair of the template DNA strand in vitro. J Biol Chem. 1990 Dec 5;265(34):21330–21336. [PubMed] [Google Scholar]
- Selby C. P., Witkin E. M., Sancar A. Escherichia coli mfd mutant deficient in "mutation frequency decline" lacks strand-specific repair: in vitro complementation with purified coupling factor. Proc Natl Acad Sci U S A. 1991 Dec 15;88(24):11574–11578. doi: 10.1073/pnas.88.24.11574. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shibata D., Peinado M. A., Ionov Y., Malkhosyan S., Perucho M. Genomic instability in repeated sequences is an early somatic event in colorectal tumorigenesis that persists after transformation. Nat Genet. 1994 Mar;6(3):273–281. doi: 10.1038/ng0394-273. [DOI] [PubMed] [Google Scholar]
- Skandalis A., Ford B. N., Glickman B. W. Strand bias in mutation involving 5-methylcytosine deamination in the human hprt gene. Mutat Res. 1994 Jan;314(1):21–26. doi: 10.1016/0921-8777(94)90057-4. [DOI] [PubMed] [Google Scholar]
- Smerdon M. J., Thoma F. Site-specific DNA repair at the nucleosome level in a yeast minichromosome. Cell. 1990 May 18;61(4):675–684. doi: 10.1016/0092-8674(90)90479-x. [DOI] [PubMed] [Google Scholar]
- Su S. S., Lahue R. S., Au K. G., Modrich P. Mispair specificity of methyl-directed DNA mismatch correction in vitro. J Biol Chem. 1988 May 15;263(14):6829–6835. [PubMed] [Google Scholar]
- Sweder K. S., Hanawalt P. C. Preferential repair of cyclobutane pyrimidine dimers in the transcribed strand of a gene in yeast chromosomes and plasmids is dependent on transcription. Proc Natl Acad Sci U S A. 1992 Nov 15;89(22):10696–10700. doi: 10.1073/pnas.89.22.10696. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Terleth C., van Sluis C. A., van de Putte P. Differential repair of UV damage in Saccharomyces cerevisiae. Nucleic Acids Res. 1989 Jun 26;17(12):4433–4439. doi: 10.1093/nar/17.12.4433. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Troelstra C., van Gool A., de Wit J., Vermeulen W., Bootsma D., Hoeijmakers J. H. ERCC6, a member of a subfamily of putative helicases, is involved in Cockayne's syndrome and preferential repair of active genes. Cell. 1992 Dec 11;71(6):939–953. doi: 10.1016/0092-8674(92)90390-x. [DOI] [PubMed] [Google Scholar]
- Umar A., Boyer J. C., Thomas D. C., Nguyen D. C., Risinger J. I., Boyd J., Ionov Y., Perucho M., Kunkel T. A. Defective mismatch repair in extracts of colorectal and endometrial cancer cell lines exhibiting microsatellite instability. J Biol Chem. 1994 May 20;269(20):14367–14370. [PubMed] [Google Scholar]
- Venema J., Mullenders L. H., Natarajan A. T., van Zeeland A. A., Mayne L. V. The genetic defect in Cockayne syndrome is associated with a defect in repair of UV-induced DNA damage in transcriptionally active DNA. Proc Natl Acad Sci U S A. 1990 Jun;87(12):4707–4711. doi: 10.1073/pnas.87.12.4707. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Venema J., van Hoffen A., Karcagi V., Natarajan A. T., van Zeeland A. A., Mullenders L. H. Xeroderma pigmentosum complementation group C cells remove pyrimidine dimers selectively from the transcribed strand of active genes. Mol Cell Biol. 1991 Aug;11(8):4128–4134. doi: 10.1128/mcb.11.8.4128. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Verhage R., Zeeman A. M., de Groot N., Gleig F., Bang D. D., van de Putte P., Brouwer J. The RAD7 and RAD16 genes, which are essential for pyrimidine dimer removal from the silent mating type loci, are also required for repair of the nontranscribed strand of an active gene in Saccharomyces cerevisiae. Mol Cell Biol. 1994 Sep;14(9):6135–6142. doi: 10.1128/mcb.14.9.6135. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wade M. H., Chu E. H. Effects of DNA damaging agents on cultured fibroblasts derived from patients with Cockayne syndrome. Mutat Res. 1979 Jan;59(1):49–60. doi: 10.1016/0027-5107(79)90194-5. [DOI] [PubMed] [Google Scholar]
- van Gool A. J., Verhage R., Swagemakers S. M., van de Putte P., Brouwer J., Troelstra C., Bootsma D., Hoeijmakers J. H. RAD26, the functional S. cerevisiae homolog of the Cockayne syndrome B gene ERCC6. EMBO J. 1994 Nov 15;13(22):5361–5369. doi: 10.1002/j.1460-2075.1994.tb06871.x. [DOI] [PMC free article] [PubMed] [Google Scholar]



