Abstract
We have cloned and sequenced a large portion of the hunchback (hb) locus from Drosophila virilis. Comparison with the Drosophila melanogaster hb sequence shows multiple strong homologies in the upstream and downstream regions of the gene, including most of the known functional parts. The coding sequence is highly conserved within the presumptive DNA-binding finger regions, but more diverged outside of them. The regions of high divergence are correlated with regions which are rich in short direct repeats (regions of high 'cryptic simplicity'), suggesting a significant influence of slippage-like mechanisms in the evolutionary divergence of the two genes. Staining of early D.virilis embryos with an hb antibody reveals conserved and divergent features of the spatial expression pattern at blastoderm stage. It appears that the basic expression pattern, which serves as the gap gene function of hb, is conserved, while certain secondary expression patterns, which have separate functions for the segmentation process, are partly diverged. Thus, both slippage driven mutations in the coding region, which are likely to occur at higher rates than point mutations and the evolutionary divergence of secondary expression patterns may contribute to the evolution of regulatory genes.
Full text
PDF



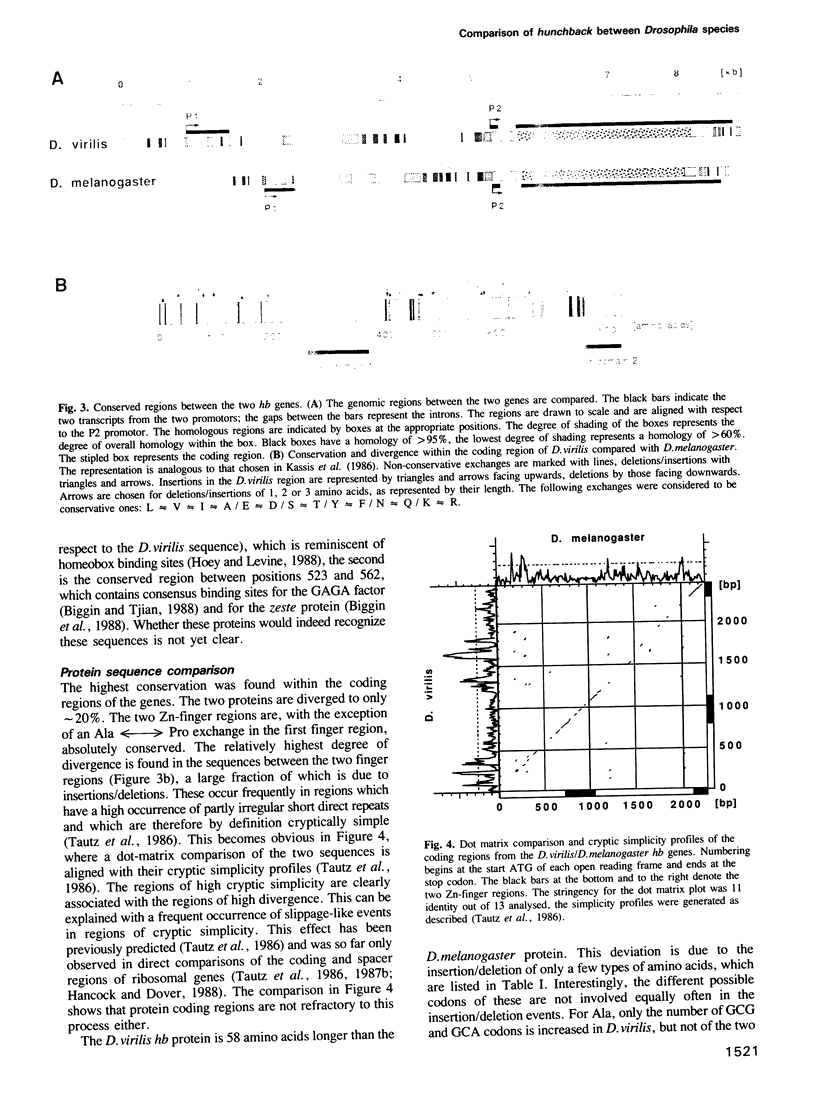




Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Akam M. The molecular basis for metameric pattern in the Drosophila embryo. Development. 1987 Sep;101(1):1–22. [PubMed] [Google Scholar]
- Bender M., Horikami S., Cribbs D., Kaufman T. C. Identification and expression of the gap segmentation gene hunchback in Drosophila melanogaster. Dev Genet. 1988;9(6):715–732. doi: 10.1002/dvg.1020090604. [DOI] [PubMed] [Google Scholar]
- Bender M., Turner F. R., Kaufman T. C. A development genetic analysis of the gene regulator of postbithorax in Drosophila melanogaster. Dev Biol. 1987 Feb;119(2):418–432. doi: 10.1016/0012-1606(87)90046-7. [DOI] [PubMed] [Google Scholar]
- Bienz M., Tremml G. Domain of Ultrabithorax expression in Drosophila visceral mesoderm from autoregulation and exclusion. Nature. 1988 Jun 9;333(6173):576–578. doi: 10.1038/333576a0. [DOI] [PubMed] [Google Scholar]
- Biggin M. D., Bickel S., Benson M., Pirrotta V., Tjian R. Zeste encodes a sequence-specific transcription factor that activates the Ultrabithorax promoter in vitro. Cell. 1988 Jun 3;53(5):713–722. doi: 10.1016/0092-8674(88)90089-x. [DOI] [PubMed] [Google Scholar]
- Biggin M. D., Tjian R. Transcription factors that activate the Ultrabithorax promoter in developmentally staged extracts. Cell. 1988 Jun 3;53(5):699–711. doi: 10.1016/0092-8674(88)90088-8. [DOI] [PubMed] [Google Scholar]
- Blackman R. K., Meselson M. Interspecific nucleotide sequence comparisons used to identify regulatory and structural features of the Drosophila hsp82 gene. J Mol Biol. 1986 Apr 20;188(4):499–515. doi: 10.1016/s0022-2836(86)80001-8. [DOI] [PubMed] [Google Scholar]
- Carroll S. B., Scott M. P. Localization of the fushi tarazu protein during Drosophila embryogenesis. Cell. 1985 Nov;43(1):47–57. doi: 10.1016/0092-8674(85)90011-x. [DOI] [PubMed] [Google Scholar]
- Dover G. A., Tautz D. Conservation and divergence in multigene families: alternatives to selection and drift. Philos Trans R Soc Lond B Biol Sci. 1986 Jan 29;312(1154):275–289. doi: 10.1098/rstb.1986.0007. [DOI] [PubMed] [Google Scholar]
- Dover G. Molecular drive: a cohesive mode of species evolution. Nature. 1982 Sep 9;299(5879):111–117. doi: 10.1038/299111a0. [DOI] [PubMed] [Google Scholar]
- Driever W., Nüsslein-Volhard C. The bicoid protein is a positive regulator of hunchback transcription in the early Drosophila embryo. Nature. 1989 Jan 12;337(6203):138–143. doi: 10.1038/337138a0. [DOI] [PubMed] [Google Scholar]
- Hancock J. M., Dover G. A. Molecular coevolution among cryptically simple expansion segments of eukaryotic 26S/28S rRNAs. Mol Biol Evol. 1988 Jul;5(4):377–391. doi: 10.1093/oxfordjournals.molbev.a040505. [DOI] [PubMed] [Google Scholar]
- Hoey T., Levine M. Divergent homeo box proteins recognize similar DNA sequences in Drosophila. Nature. 1988 Apr 28;332(6167):858–861. doi: 10.1038/332858a0. [DOI] [PubMed] [Google Scholar]
- Ingham P. W. The molecular genetics of embryonic pattern formation in Drosophila. Nature. 1988 Sep 1;335(6185):25–34. doi: 10.1038/335025a0. [DOI] [PubMed] [Google Scholar]
- Kassis J. A., Poole S. J., Wright D. K., O'Farrell P. H. Sequence conservation in the protein coding and intron regions of the engrailed transcription unit. EMBO J. 1986 Dec 20;5(13):3583–3589. doi: 10.1002/j.1460-2075.1986.tb04686.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Krause H. M., Klemenz R., Gehring W. J. Expression, modification, and localization of the fushi tarazu protein in Drosophila embryos. Genes Dev. 1988 Aug;2(8):1021–1036. doi: 10.1101/gad.2.8.1021. [DOI] [PubMed] [Google Scholar]
- Lehmann R., Nüsslein-Volhard C. hunchback, a gene required for segmentation of an anterior and posterior region of the Drosophila embryo. Dev Biol. 1987 Feb;119(2):402–417. doi: 10.1016/0012-1606(87)90045-5. [DOI] [PubMed] [Google Scholar]
- Levinson G., Gutman G. A. Slipped-strand mispairing: a major mechanism for DNA sequence evolution. Mol Biol Evol. 1987 May;4(3):203–221. doi: 10.1093/oxfordjournals.molbev.a040442. [DOI] [PubMed] [Google Scholar]
- Macdonald P. M., Struhl G. A molecular gradient in early Drosophila embryos and its role in specifying the body pattern. Nature. 1986 Dec 11;324(6097):537–545. doi: 10.1038/324537a0. [DOI] [PubMed] [Google Scholar]
- Mueller P. P., Hinnebusch A. G. Multiple upstream AUG codons mediate translational control of GCN4. Cell. 1986 Apr 25;45(2):201–207. doi: 10.1016/0092-8674(86)90384-3. [DOI] [PubMed] [Google Scholar]
- Nüsslein-Volhard C., Frohnhöfer H. G., Lehmann R. Determination of anteroposterior polarity in Drosophila. Science. 1987 Dec 18;238(4834):1675–1681. doi: 10.1126/science.3686007. [DOI] [PubMed] [Google Scholar]
- Nüsslein-Volhard C., Wieschaus E. Mutations affecting segment number and polarity in Drosophila. Nature. 1980 Oct 30;287(5785):795–801. doi: 10.1038/287795a0. [DOI] [PubMed] [Google Scholar]
- Pustell J., Kafatos F. C. A high speed, high capacity homology matrix: zooming through SV40 and polyoma. Nucleic Acids Res. 1982 Aug 11;10(15):4765–4782. doi: 10.1093/nar/10.15.4765. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sanger F., Nicklen S., Coulson A. R. DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci U S A. 1977 Dec;74(12):5463–5467. doi: 10.1073/pnas.74.12.5463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schröder C., Tautz D., Seifert E., Jäckle H. Differential regulation of the two transcripts from the Drosophila gap segmentation gene hunchback. EMBO J. 1988 Sep;7(9):2881–2887. doi: 10.1002/j.1460-2075.1988.tb03145.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tautz D. Regulation of the Drosophila segmentation gene hunchback by two maternal morphogenetic centres. Nature. 1988 Mar 17;332(6161):281–284. doi: 10.1038/332281a0. [DOI] [PubMed] [Google Scholar]
- Tautz D., Renz M. Simple sequences are ubiquitous repetitive components of eukaryotic genomes. Nucleic Acids Res. 1984 May 25;12(10):4127–4138. doi: 10.1093/nar/12.10.4127. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tautz D., Tautz C., Webb D., Dover G. A. Evolutionary divergence of promoters and spacers in the rDNA family of four Drosophila species. Implications for molecular coevolution in multigene families. J Mol Biol. 1987 Jun 5;195(3):525–542. doi: 10.1016/0022-2836(87)90181-1. [DOI] [PubMed] [Google Scholar]
- Tautz D., Trick M., Dover G. A. Cryptic simplicity in DNA is a major source of genetic variation. Nature. 1986 Aug 14;322(6080):652–656. doi: 10.1038/322652a0. [DOI] [PubMed] [Google Scholar]
- Wharton K. A., Yedvobnick B., Finnerty V. G., Artavanis-Tsakonas S. opa: a novel family of transcribed repeats shared by the Notch locus and other developmentally regulated loci in D. melanogaster. Cell. 1985 Jan;40(1):55–62. doi: 10.1016/0092-8674(85)90308-3. [DOI] [PubMed] [Google Scholar]
- Wilson A. C., Maxson L. R., Sarich V. M. Two types of molecular evolution. Evidence from studies of interspecific hybridization. Proc Natl Acad Sci U S A. 1974 Jul;71(7):2843–2847. doi: 10.1073/pnas.71.7.2843. [DOI] [PMC free article] [PubMed] [Google Scholar]




