Abstract
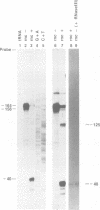
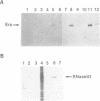
RNase III has been implicated in the control of gene expression by the processing of mRNA. We have found that the rnc operon is autoregulated; rnc- mutant strains oversynthesize the operon's mRNA and protein products. A site in the 5'-noncoding region of the operon's message is cleaved by RNase III. This site-specific cleavage appears to be the initial step in the functional inactivation of the message, since the half-life of the cut message is dramatically shorter than that of the uncut message.
Full text
PDF





Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Ahnn J., March P. E., Takiff H. E., Inouye M. A GTP-binding protein of Escherichia coli has homology to yeast RAS proteins. Proc Natl Acad Sci U S A. 1986 Dec;83(23):8849–8853. doi: 10.1073/pnas.83.23.8849. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Aiba H., Adhya S., de Crombrugghe B. Evidence for two functional gal promoters in intact Escherichia coli cells. J Biol Chem. 1981 Nov 25;256(22):11905–11910. [PubMed] [Google Scholar]
- Altuvia S., Locker-Giladi H., Koby S., Ben-Nun O., Oppenheim A. B. RNase III stimulates the translation of the cIII gene of bacteriophage lambda. Proc Natl Acad Sci U S A. 1987 Sep;84(18):6511–6515. doi: 10.1073/pnas.84.18.6511. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Apirion D., Gitelman D. R. Decay of RNA in RNA processing mutants of Escherichia coli. Mol Gen Genet. 1980 Jan;177(2):339–343. doi: 10.1007/BF00267448. [DOI] [PubMed] [Google Scholar]
- Barry G., Squires C., Squires C. L. Attenuation and processing of RNA from the rplJL--rpoBC transcription unit of Escherichia coli. Proc Natl Acad Sci U S A. 1980 Jun;77(6):3331–3335. doi: 10.1073/pnas.77.6.3331. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bechhofer D. H., Dubnau D. Induced mRNA stability in Bacillus subtilis. Proc Natl Acad Sci U S A. 1987 Jan;84(2):498–502. doi: 10.1073/pnas.84.2.498. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Belasco J. G., Beatty J. T., Adams C. W., von Gabain A., Cohen S. N. Differential expression of photosynthesis genes in R. capsulata results from segmental differences in stability within the polycistronic rxcA transcript. Cell. 1985 Jan;40(1):171–181. doi: 10.1016/0092-8674(85)90320-4. [DOI] [PubMed] [Google Scholar]
- Blundell M., Kennell D. Evidence for endonucleolytic attack in decay of lac messenger RNA in Escherichia coli. J Mol Biol. 1974 Feb 25;83(2):143–161. doi: 10.1016/0022-2836(74)90385-4. [DOI] [PubMed] [Google Scholar]
- Bram R. J., Young R. A., Steitz J. A. The ribonuclease III site flanking 23S sequences in the 30S ribosomal precursor RNA of E. coli. Cell. 1980 Feb;19(2):393–401. doi: 10.1016/0092-8674(80)90513-9. [DOI] [PubMed] [Google Scholar]
- Brock M. L., Shapiro D. J. Estrogen stabilizes vitellogenin mRNA against cytoplasmic degradation. Cell. 1983 Aug;34(1):207–214. doi: 10.1016/0092-8674(83)90151-4. [DOI] [PubMed] [Google Scholar]
- Chen C. Y., Beatty J. T., Cohen S. N., Belasco J. G. An intercistronic stem-loop structure functions as an mRNA decay terminator necessary but insufficient for puf mRNA stability. Cell. 1988 Feb 26;52(4):609–619. doi: 10.1016/0092-8674(88)90473-4. [DOI] [PubMed] [Google Scholar]
- Cowing D. W., Bardwell J. C., Craig E. A., Woolford C., Hendrix R. W., Gross C. A. Consensus sequence for Escherichia coli heat shock gene promoters. Proc Natl Acad Sci U S A. 1985 May;82(9):2679–2683. doi: 10.1073/pnas.82.9.2679. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Daniels D. L., Subbarao M. N., Blattner F. R., Lozeron H. A. Q-mediated late gene transcription of bacteriophage lambda: RNA start point and RNase III processing sites in vivo. Virology. 1988 Dec;167(2):568–577. [PubMed] [Google Scholar]
- Deutscher M. P. E. coli RNases: making sense of alphabet soup. Cell. 1985 Apr;40(4):731–732. doi: 10.1016/0092-8674(85)90330-7. [DOI] [PubMed] [Google Scholar]
- Downing W. L., Dennis P. P. Transcription products from the rplKAJL-rpoBC gene cluster. J Mol Biol. 1987 Apr 20;194(4):609–620. doi: 10.1016/0022-2836(87)90238-5. [DOI] [PubMed] [Google Scholar]
- Dunn J. J., Studier F. W. Complete nucleotide sequence of bacteriophage T7 DNA and the locations of T7 genetic elements. J Mol Biol. 1983 Jun 5;166(4):477–535. doi: 10.1016/s0022-2836(83)80282-4. [DOI] [PubMed] [Google Scholar]
- GROS F., HIATT H., GILBERT W., KURLAND C. G., RISEBROUGH R. W., WATSON J. D. Unstable ribonucleic acid revealed by pulse labelling of Escherichia coli. Nature. 1961 May 13;190:581–585. doi: 10.1038/190581a0. [DOI] [PubMed] [Google Scholar]
- Gitelman D. R., Apirion D. The synthesis of some proteins is affected in RNA processing mutants of Escherichia coli. Biochem Biophys Res Commun. 1980 Oct 16;96(3):1063–1070. doi: 10.1016/0006-291x(80)90060-1. [DOI] [PubMed] [Google Scholar]
- Gorski K., Roch J. M., Prentki P., Krisch H. M. The stability of bacteriophage T4 gene 32 mRNA: a 5' leader sequence that can stabilize mRNA transcripts. Cell. 1985 Dec;43(2 Pt 1):461–469. doi: 10.1016/0092-8674(85)90176-x. [DOI] [PubMed] [Google Scholar]
- Greenberg M. E., Ziff E. B. Stimulation of 3T3 cells induces transcription of the c-fos proto-oncogene. Nature. 1984 Oct 4;311(5985):433–438. doi: 10.1038/311433a0. [DOI] [PubMed] [Google Scholar]
- Harland R., Misher L. Stability of RNA in developing Xenopus embryos and identification of a destabilizing sequence in TFIIIA messenger RNA. Development. 1988 Apr;102(4):837–852. doi: 10.1242/dev.102.4.837. [DOI] [PubMed] [Google Scholar]
- Hayashi M. N., Hayashi M. Cloned DNA sequences that determine mRNA stability of bacteriophage phi X174 in vivo are functional. Nucleic Acids Res. 1985 Aug 26;13(16):5937–5948. doi: 10.1093/nar/13.16.5937. [DOI] [PMC free article] [PubMed] [Google Scholar]
- King T. C., Sirdeskmukh R., Schlessinger D. Nucleolytic processing of ribonucleic acid transcripts in procaryotes. Microbiol Rev. 1986 Dec;50(4):428–451. doi: 10.1128/mr.50.4.428-451.1986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Krinke L., Wulff D. L. OOP RNA, produced from multicopy plasmids, inhibits lambda cII gene expression through an RNase III-dependent mechanism. Genes Dev. 1987 Nov;1(9):1005–1013. doi: 10.1101/gad.1.9.1005. [DOI] [PubMed] [Google Scholar]
- Mackie G. A. Posttranscriptional regulation of ribosomal protein S20 and stability of the S20 mRNA species. J Bacteriol. 1987 Jun;169(6):2697–2701. doi: 10.1128/jb.169.6.2697-2701.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- March P. E., Ahnn J., Inouye M. The DNA sequence of the gene (rnc) encoding ribonuclease III of Escherichia coli. Nucleic Acids Res. 1985 Jul 11;13(13):4677–4685. doi: 10.1093/nar/13.13.4677. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Melefors O., von Gabain A. Site-specific endonucleolytic cleavages and the regulation of stability of E. coli ompA mRNA. Cell. 1988 Mar 25;52(6):893–901. doi: 10.1016/0092-8674(88)90431-x. [DOI] [PubMed] [Google Scholar]
- Morrison P. T., Lovett S. T., Gilson L. E., Kolodner R. Molecular analysis of the Escherichia coli recO gene. J Bacteriol. 1989 Jul;171(7):3641–3649. doi: 10.1128/jb.171.7.3641-3649.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mott J. E., Galloway J. L., Platt T. Maturation of Escherichia coli tryptophan operon mRNA: evidence for 3' exonucleolytic processing after rho-dependent termination. EMBO J. 1985 Jul;4(7):1887–1891. doi: 10.1002/j.1460-2075.1985.tb03865.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nashimoto H., Uchida H. DNA sequencing of the Escherichia coli ribonuclease III gene and its mutations. Mol Gen Genet. 1985;201(1):25–29. doi: 10.1007/BF00397981. [DOI] [PubMed] [Google Scholar]
- Newbury S. F., Smith N. H., Higgins C. F. Differential mRNA stability controls relative gene expression within a polycistronic operon. Cell. 1987 Dec 24;51(6):1131–1143. doi: 10.1016/0092-8674(87)90599-x. [DOI] [PubMed] [Google Scholar]
- Nilsson G., Belasco J. G., Cohen S. N., von Gabain A. Effect of premature termination of translation on mRNA stability depends on the site of ribosome release. Proc Natl Acad Sci U S A. 1987 Jul;84(14):4890–4894. doi: 10.1073/pnas.84.14.4890. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nilsson G., Belasco J. G., Cohen S. N., von Gabain A. Growth-rate dependent regulation of mRNA stability in Escherichia coli. Nature. 1984 Nov 1;312(5989):75–77. doi: 10.1038/312075a0. [DOI] [PubMed] [Google Scholar]
- Plunkett G., 3rd, Echols H. Retroregulation of the bacteriophage lambda int gene: limited secondary degradation of the RNase III-processed transcript. J Bacteriol. 1989 Jan;171(1):588–592. doi: 10.1128/jb.171.1.588-592.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Portier C., Dondon L., Grunberg-Manago M., Régnier P. The first step in the functional inactivation of the Escherichia coli polynucleotide phosphorylase messenger is a ribonuclease III processing at the 5' end. EMBO J. 1987 Jul;6(7):2165–2170. doi: 10.1002/j.1460-2075.1987.tb02484.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rosenberg A. H., Lade B. N., Chui D. S., Lin S. W., Dunn J. J., Studier F. W. Vectors for selective expression of cloned DNAs by T7 RNA polymerase. Gene. 1987;56(1):125–135. doi: 10.1016/0378-1119(87)90165-x. [DOI] [PubMed] [Google Scholar]
- Régnier P., Portier C. Initiation, attenuation and RNase III processing of transcripts from the Escherichia coli operon encoding ribosomal protein S15 and polynucleotide phosphorylase. J Mol Biol. 1986 Jan 5;187(1):23–32. doi: 10.1016/0022-2836(86)90403-1. [DOI] [PubMed] [Google Scholar]
- Saito H., Richardson C. C. Processing of mRNA by ribonuclease III regulates expression of gene 1.2 of bacteriophage T7. Cell. 1981 Dec;27(3 Pt 2):533–542. doi: 10.1016/0092-8674(81)90395-0. [DOI] [PubMed] [Google Scholar]
- Sarmientos P., Sylvester J. E., Contente S., Cashel M. Differential stringent control of the tandem E. coli ribosomal RNA promoters from the rrnA operon expressed in vivo in multicopy plasmids. Cell. 1983 Apr;32(4):1337–1346. doi: 10.1016/0092-8674(83)90314-8. [DOI] [PubMed] [Google Scholar]
- Schmeissner U., McKenney K., Rosenberg M., Court D. Removal of a terminator structure by RNA processing regulates int gene expression. J Mol Biol. 1984 Jun 15;176(1):39–53. doi: 10.1016/0022-2836(84)90381-4. [DOI] [PubMed] [Google Scholar]
- Schneider E., Blundell M., Kennell D. Translation and mRNA decay. Mol Gen Genet. 1978 Apr 6;160(2):121–129. doi: 10.1007/BF00267473. [DOI] [PubMed] [Google Scholar]
- Stanssens P., Remaut E., Fiers W. Inefficient translation initiation causes premature transcription termination in the lacZ gene. Cell. 1986 Mar 14;44(5):711–718. doi: 10.1016/0092-8674(86)90837-8. [DOI] [PubMed] [Google Scholar]
- Steege D. A., Cone K. C., Queen C., Rosenberg M. Bacteriophage lambda N gene leader RNA. RNA processing and translational initiation signals. J Biol Chem. 1987 Dec 25;262(36):17651–17658. [PubMed] [Google Scholar]
- Studier F. W. Genetic mapping of a mutation that causes ribonucleases III deficiency in Escherichia coli. J Bacteriol. 1975 Oct;124(1):307–316. doi: 10.1128/jb.124.1.307-316.1975. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Takata R., Mukai T., Hori K. RNA processing by RNase III is involved in the synthesis of Escherichia coli polynucleotide phosphorylase. Mol Gen Genet. 1987 Aug;209(1):28–32. doi: 10.1007/BF00329832. [DOI] [PubMed] [Google Scholar]
- Takiff H. E., Chen S. M., Court D. L. Genetic analysis of the rnc operon of Escherichia coli. J Bacteriol. 1989 May;171(5):2581–2590. doi: 10.1128/jb.171.5.2581-2590.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Young R. A., Steitz J. A. Complementary sequences 1700 nucleotides apart form a ribonuclease III cleavage site in Escherichia coli ribosomal precursor RNA. Proc Natl Acad Sci U S A. 1978 Aug;75(8):3593–3597. doi: 10.1073/pnas.75.8.3593. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zuker M., Stiegler P. Optimal computer folding of large RNA sequences using thermodynamics and auxiliary information. Nucleic Acids Res. 1981 Jan 10;9(1):133–148. doi: 10.1093/nar/9.1.133. [DOI] [PMC free article] [PubMed] [Google Scholar]