Abstract
Recent developments in multidimensional heteronuclear NMR spectroscopy and large-scale synthesis of uniformly 13C- and 15N-labeled oligonucleotides have greatly improved the prospects for determination of the solution structure of RNA. However, there are circumstances in which it may be advantageous to label only a segment of the entire RNA chain. For example, in a larger RNA molecule the structural question of interest may reside in a localized domain. Labeling only the corresponding nucleotides simplifies the spectrum and resonance assignments because one can filter proton spectra for coupling to 13C and 15N. Another example is in resolving alternative secondary structure models that are indistinguishable in imino proton connectivities. Here we report a general method for enzymatic synthesis of quantities of segmentally labeled RNA molecules required for NMR spectroscopy. We use the method to distinguish definitively two competing secondary structure models for the 5' half of Caenorhabditis elegans spliced leader RNA by comparison of the two-dimensional [15N] 1H heteronuclear multiple quantum correlation spectrum of the uniformly labeled sample with that of a segmentally labeled sample. The method requires relatively small samples; solutions in the 200-300 microM concentration range, with a total of 30 nmol or approximately 40 micrograms of RNA in approximately 150 microliters, give strong NMR signals in a short accumulation time. The method can be adapted to label an internal segment of a larger RNA chain for study of localized structural problems. This definitive approach provides an alternative to the more common enzymatic and chemical footprinting methods for determination of RNA secondary structure.
Full text
PDF


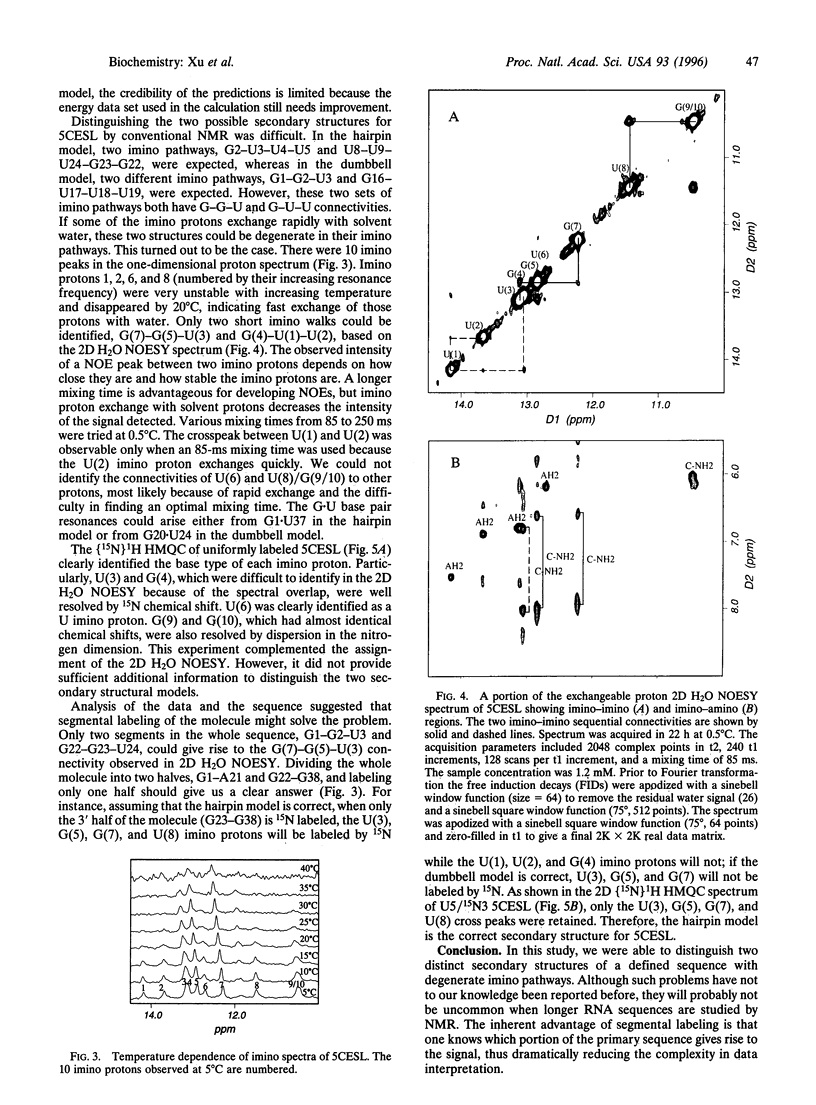

Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Agabian N. Trans splicing of nuclear pre-mRNAs. Cell. 1990 Jun 29;61(7):1157–1160. doi: 10.1016/0092-8674(90)90674-4. [DOI] [PubMed] [Google Scholar]
- Batey R. T., Inada M., Kujawinski E., Puglisi J. D., Williamson J. R. Preparation of isotopically labeled ribonucleotides for multidimensional NMR spectroscopy of RNA. Nucleic Acids Res. 1992 Sep 11;20(17):4515–4523. doi: 10.1093/nar/20.17.4515. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bhattacharyya A., Blackburn E. H. Architecture of telomerase RNA. EMBO J. 1994 Dec 1;13(23):5721–5731. doi: 10.1002/j.1460-2075.1994.tb06910.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bruzik J. P., Van Doren K., Hirsh D., Steitz J. A. Trans splicing involves a novel form of small nuclear ribonucleoprotein particles. Nature. 1988 Oct 6;335(6190):559–562. doi: 10.1038/335559a0. [DOI] [PubMed] [Google Scholar]
- Grodberg J., Dunn J. J. ompT encodes the Escherichia coli outer membrane protease that cleaves T7 RNA polymerase during purification. J Bacteriol. 1988 Mar;170(3):1245–1253. doi: 10.1128/jb.170.3.1245-1253.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hayase Y., Inoue H., Ohtsuka E. Secondary structure in formylmethionine tRNA influences the site-directed cleavage of ribonuclease H using chimeric 2'-O-methyl oligodeoxyribonucleotides. Biochemistry. 1990 Sep 18;29(37):8793–8797. doi: 10.1021/bi00489a041. [DOI] [PubMed] [Google Scholar]
- Inoue H., Hayase Y., Iwai S., Ohtsuka E. Sequence-dependent hydrolysis of RNA using modified oligonucleotide splints and RNase H. FEBS Lett. 1987 May 11;215(2):327–330. doi: 10.1016/0014-5793(87)80171-0. [DOI] [PubMed] [Google Scholar]
- Jaeger J. A., SantaLucia J., Jr, Tinoco I., Jr Determination of RNA structure and thermodynamics. Annu Rev Biochem. 1993;62:255–287. doi: 10.1146/annurev.bi.62.070193.001351. [DOI] [PubMed] [Google Scholar]
- Jaeger J. A., Turner D. H., Zuker M. Improved predictions of secondary structures for RNA. Proc Natl Acad Sci U S A. 1989 Oct;86(20):7706–7710. doi: 10.1073/pnas.86.20.7706. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kumar A., Ernst R. R., Wüthrich K. A two-dimensional nuclear Overhauser enhancement (2D NOE) experiment for the elucidation of complete proton-proton cross-relaxation networks in biological macromolecules. Biochem Biophys Res Commun. 1980 Jul 16;95(1):1–6. doi: 10.1016/0006-291x(80)90695-6. [DOI] [PubMed] [Google Scholar]
- LeCuyer K. A., Crothers D. M. The Leptomonas collosoma spliced leader RNA can switch between two alternate structural forms. Biochemistry. 1993 May 25;32(20):5301–5311. doi: 10.1021/bi00071a004. [DOI] [PubMed] [Google Scholar]
- Milligan J. F., Groebe D. R., Witherell G. W., Uhlenbeck O. C. Oligoribonucleotide synthesis using T7 RNA polymerase and synthetic DNA templates. Nucleic Acids Res. 1987 Nov 11;15(21):8783–8798. doi: 10.1093/nar/15.21.8783. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Milligan J. F., Uhlenbeck O. C. Synthesis of small RNAs using T7 RNA polymerase. Methods Enzymol. 1989;180:51–62. doi: 10.1016/0076-6879(89)80091-6. [DOI] [PubMed] [Google Scholar]
- Moore M. J., Sharp P. A. Site-specific modification of pre-mRNA: the 2'-hydroxyl groups at the splice sites. Science. 1992 May 15;256(5059):992–997. doi: 10.1126/science.1589782. [DOI] [PubMed] [Google Scholar]
- Nikonowicz E. P., Pardi A. Three-dimensional heteronuclear NMR studies of RNA. Nature. 1992 Jan 9;355(6356):184–186. doi: 10.1038/355184a0. [DOI] [PubMed] [Google Scholar]
- Nikonowicz E. P., Sirr A., Legault P., Jucker F. M., Baer L. M., Pardi A. Preparation of 13C and 15N labelled RNAs for heteronuclear multi-dimensional NMR studies. Nucleic Acids Res. 1992 Sep 11;20(17):4507–4513. doi: 10.1093/nar/20.17.4507. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nilsen T. W. Trans-splicing of nematode premessenger RNA. Annu Rev Microbiol. 1993;47:413–440. doi: 10.1146/annurev.mi.47.100193.002213. [DOI] [PubMed] [Google Scholar]
- Romaniuk P. J., Uhlenbeck O. C. Joining of RNA molecules with RNA ligase. Methods Enzymol. 1983;100:52–59. doi: 10.1016/0076-6879(83)00045-2. [DOI] [PubMed] [Google Scholar]
- Szewczak A. A., Kellogg G. W., Moore P. B. Assignment of NH resonances in nucleic acids using natural abundance 15N-1H correlation spectroscopy with spin-echo and gradient pulses. FEBS Lett. 1993 Aug 2;327(3):261–264. doi: 10.1016/0014-5793(93)81000-p. [DOI] [PubMed] [Google Scholar]
- Zuker M. On finding all suboptimal foldings of an RNA molecule. Science. 1989 Apr 7;244(4900):48–52. doi: 10.1126/science.2468181. [DOI] [PubMed] [Google Scholar]



