Abstract
Signal transduction occurs by the reversible assembly of oligomeric protein complexes that include both enzymatic proteins and proteins without known enzymatic activity. These nonenzymatic components can serve as scaffolds or anchors and regulate the efficiency, specificity, and localization of the signaling pathway. Here we report the identification of MORG1 (mitogen-activated protein kinase organizer 1), a member of the WD-40 protein family that was isolated as a binding partner of the extracellular signal-regulated kinase (ERK) pathway scaffold protein MP1. MORG1 specifically associates with several components of the ERK pathway, including MP1, Raf-1, MEK, and ERK, and stabilizes their assembly into an oligomeric complex. MORG1 facilitates ERK activation when cells are stimulated with lysophosphatidic acid, phorbol 12-myristate 13-acetate, or serum, but not in response to epidermal growth factor. Suppression of MORG1 by short interfering RNA leads to a marked reduction in ERK activity when cells are stimulated with serum. We propose that MORG1 is a component of a modular scaffold system that participates in the regulation of agonist-specific ERK signaling.
The mitogen-activated protein kinases (MAPKs) extracellular signal-regulated kinase (ERK)-1 and ERK2 are components of an evolutionarily conserved protein kinase cascade that participates in the regulation of various cellular processes, including gene expression, growth, differentiation, and apoptosis. The ERK cascade displays a three-kinase architecture in which ERKs are phosphorylated and activated by the upstream protein kinases MAPK/ERK kinase (MEK)-1 and MEK2. MEKs in turn are activated by MEKKs (MAPK/ERK kinase kinase) of the Raf family (1–3).
The activation of MAPK cascades is a highly ordered process that involves tight regulation of all components in the pathway. Protein–protein interactions play an important role in many of these regulatory mechanisms and can facilitate appropriate propagation of a signal in time and space (4–6). Because the ERK pathway is ubiquitously activated in response to diverse signals, secondary mechanisms must exist to channel the ERKs in ways that link specific input signals with an appropriate biological response.
A number of so-called scaffolding proteins, both yeast and mammalian, were shown to associate with and enhance functional interaction of the components of MAPK pathways. The prototypical MAPK scaffold protein Ste5 is an essential component of the mating pathway in Saccharomyces cerevisiae. Ste5 tethers protein kinases Ste7 [MAPK kinase (MAPKK)], Ste11 [MAPKK kinase (MAPKKK)], and Fus3 (MAPK) (7, 8), and assembly of this signaling complex increases efficiency of signal flow (9). In addition, Ste5 generates signal specificity by insulating the pheromone response pathway from parallel pathways that use common signaling components (9–11).
Several proteins with adapter/scaffolding function have been identified in mammalian MAPK cascades, although their physiological and molecular functions are incompletely understood. In the ERK and c-Jun N-terminal kinase (JNK) pathways, biochemical and genetic evidence suggests the operation of kinase suppressor of Ras 1 (KSR-1) and JNK-interacting protein 1 (JIP-1), respectively, as scaffold-like proteins. When overexpressed, KSR-1 interacts with the protein kinases Raf-1, MEK, and ERK (12–15). However, at the endogenous level, Raf-1 cannot be detected in complexes containing KSR (16). Depending on expression level, KSR-1 can either inhibit or enhance ERK activity (17–20), and ERK activation is severely compromised in KSR-1-null fibroblasts (16, 21).
JIP-1 interacts with protein kinases of the JNK cascade, including JNK, MKK7, and the MAPKKKs DLK and MLK3. In tissue culture, JIP-1 can enhance JNK activation (22) or, when overexpressed, can act as an inhibitor of JNK signaling (23). Disruption of the Jip1 gene causes severe defects in JNK activation in response to a subset of stress factors in neurons (24), demonstrating that JIP-1 participates in the regulation of specific biological activities of the JNK pathway.
In our laboratory, MEK partner 1 (MP1) was identified as a protein that specifically interacts with MEK1 and ERK1 and facilitates the activation of ERK1 (25). MP1 differs from typical MAPK scaffolds with respect to its small size (only 14.5 kDa) and because it interacts with MEK1 and ERK1, but not with other members of the kinase cascade further upstream (e.g., Raf). We therefore hypothesized that the ability to interact with other components of the ERK cascade might be distributed among several proteins and that this modular assembly of a complete scaffold would allow the generation of combinatorial specificity in the control of signaling (4, 25). Here, we describe the protein MAPK organizer (MORG)-1, which interacts with several components of the ERK cascade and functions as a scaffold protein linking ERK responses to specific agonists.
Materials and Methods
Plasmid Constructs and Two-Hybrid Screen. Full-length MP1 was fused in-frame with the DNA-binding domain of Gal4 to screen a pACT vector human brain library according to standard procedures (Clontech MATCHMAKER GAL4 Two-Hybrid System user manual).
MP1, Raf-1, epitope-tagged MEK, and ERK expression constructs have been described previously (26). The proline-rich sequence-deletion mutant of MEK1 (MEK1Δ270–307) is called MEK1ΔPRS for clarity. For the expression of untagged MORG1 in mammalian cells, cDNA containing the mouse MORG1 coding sequence was subcloned to pcDNA3 vector (Invitrogen). MORG1 was also hemagglutinin (HA)-tagged at the N terminus by PCR and subcloned to pcDNA3. For the generation of HeLa cell lines stably expressing MORG1, the HA-tagged MORG1 cDNA was subcloned into pREP4 (Invitrogen).
Cell Lines and Transfections. NIH 3T3 cells were maintained in DMEM supplemented with 5% FBS and 5% calf serum (all obtained from GIBCO/BRL). CCL39, Cos1, and HeLa cells were maintained in DMEM supplemented with 10% FBS. Transfections were performed by using Lipofectamine (GIBCO/BRL) according to the manufacturer's instructions.
Immunoprecipitations and Immune-Complex Kinase Assays. Cells were lysed 24 h posttransfection in FLAG lysis buffer [50 mM Tris·HCl, 150 mM NaCl, 1% Triton X-100, 10% glycerol, 0.5 mM EDTA, and 0.5 mM EGTA (pH 7.3), supplemented with 50 mM NaF, 5 mM Na4P2O7, 0.2 mM Na3VO4, and protease inhibitors]. Clarified extracts were incubated for 2 h with M2 anti-FLAG affinity resin (Sigma) at 4°C. Immune complexes were washed four times with lysis buffer and resolved by SDS/PAGE. Immunoblotting was carried out as described previously (26).
For immune-complex kinase assays, cells were transfected as described above. Cells were serum-deprived 24 h posttransfection in 0.15% FBS in DMEM for 5 h before stimulation with different agonists. Cells were lysed in FLAG buffer, and expression of either cotransfected Renilla luciferase or β-galactosidase was used to normalize lysates for transfection efficiency before immunoprecipitation. Immune complexes were washed twice in FLAG buffer and twice in 25 mM Hepes/OH (pH 7.5), 10 mM MgCl2, and 1 mM DTT. Kinase assays were performed in the same Hepes buffer supplemented with 0.5 mg/ml myelin basic protein (MBP) and 100 μM [γ-32P]ATP (1,000–3,000 cpm per pmol). After 10 min at 30°C, reactions were terminated with sample buffer, resolved by SDS/PAGE, and transferred to nitrocellulose. MBP bands were excised, and incorporation of labeled phosphate was determined by Cerenkov counting. The amount of ERK precipitated in each experiment was verified by immunoblotting.
Establishment of HeLa Stable Cell Lines. HeLa cells were transfected with either pREP4 or pREP-HA-MORG vector by using Poly-Fect (Qiagen, Valencia, CA). Either 300 or 500 μg/ml hygromycin was used to select HeLa-MORG300 or HeLa-MORG500 cell populations, respectively. Both cell lines have been shown to be uniformly HA epitope-positive by immunofluorescence (data not shown). Western blotting with HA antibody showed significantly higher MORG1 expression levels in the HeLa-MORG500 than in the HeLa-MORG300 cell line. For determination of ERK phosphorylation status, cells were cultured in 10% FBS in the absence of hygromycin for three days. Then, cells were serum-deprived for 5 h and stimulated with agonists for 10 min, and cell lysates were probed as indicated.
Short Interfering RNA (siRNA) Gene-Silencing Assay. A double-stranded siRNA targeting the 21-nt sequence CAAGCTGGACTGCTGCCTGAG of the MORG1 gene and the control siRNA (Non-Specific Control Duplexes-XIII) were obtained from Dharmacon Research (Lafayette, CO). The siRNA was transfected into HeLa cells by using the calcium phosphate protocol, and 24 h posttransfection cell lysates were probed as indicated. Mock control cells received transfection reagent only.
Statistical Methods. Data from four independent experiments were used to analyze agonist specificity of MORG1-mediated enhancement of ERK activity. In each of four experiments, independent triplicate samples were obtained for five different experimental conditions.
Before statistical modeling, use of the Box-Cox method suggested that a logarithmic transformation of ERK activity measurements would scale the data for analysis most appropriately. Two-factor ANOVA was performed by using the transformed ERK activity measurements as the response variable and experiment (four levels) and experimental condition (five levels) as the two factors. Inclusion of experiment as a factor was necessary to adjust for differences in overall ERK activity across the four experiments. To describe the effect of MORG1 on ERK activity, model-based estimates of fold changes and corresponding 95% confidence intervals were calculated for comparisons of interest among the experimental conditions.
Results
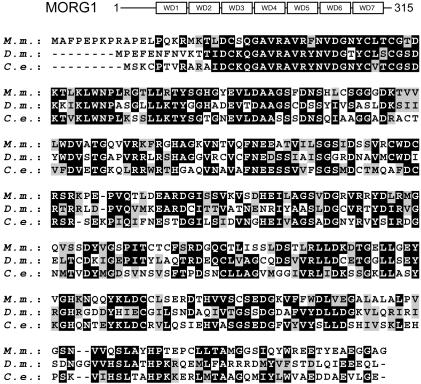
Cloning of MORG1. To identify binding partners of MP1, a yeast two-hybrid screen of a human brain cDNA library was performed with MP1 as bait. The MORG1 protein interacted specifically with MP1. Analysis of the mouse cDNA sequence showed that MORG1 is a protein of 315 aa composed almost entirely of seven WD-40 domains (Fig. 1) with a predicted molecular mass of 34.5 kDa. Mammalian MORG1 shares >50% amino acid sequence identity with uncharacterized proteins of Drosophila melanogaster and Caenorhabditis elegans (Fig. 1). MORG1 is ubiquitously expressed, as judged by multiple-tissue Northern blotting. Abundant 1.3-kbp MORG1 transcripts were detected in heart, brain, liver, kidney, and testis, whereas lower amounts of MORG1 were detected in lung, spleen, and skeletal muscle (data not shown). The expression pattern of MORG1 mRNA mostly overlaps with that of MP1 (25), although in some tissues differences in the relative abundance can be observed.
Fig. 1.
Amino acid sequence, domain structure, and tissue distribution of MORG1. (Upper) Domain structure of full-length MORG1. (Lower) Alignment of the predicted amino acid sequence of mouse MORG1 (M.m.) (GenBank accession no. AY365434), D. melanogaster (D.m.) (GenBank accession no. U41278), and hypothetical protein F33G12.2 C. elegans. (C.e.) (GenBank accession no. AE003650.2). Amino acids in black represent identical residues; those in gray represent conservative changes.
MORG1 Associates with Components of the ERK Cascade. To verify the MORG1–MP1 interaction in mammalian cells, CCL39 fibroblasts were transiently transfected with FLAG-MP1 and HA-MORG1, and FLAG-MP1 was isolated by immunoprecipitation. MP1 effectively coimmunoprecipitated MORG1 (Fig. 2A).
Fig. 2.
MORG1 associates with components of the MAPK/ERK cascade in cells. (A) Coimmunoprecipitation of MORG1 with MP1. CCL39 cells were transfected with either control vector or FLAG-MP1 and HA-MORG1 constructs. MP1 was precipitated and detected with FLAG M2 antibody. Coprecipitated MORG1 was detected with HA antibody. Cell lysates were probed with the HA antibody to verify comparable expression levels of MORG1 protein. (B) Coimmunoprecipitation of MORG1 with MEK1, MEK1ΔPRS, and MEK2. CCL39 cells were transfected with either control vector or various FLAG-MEK constructs and HA-MORG1. MEK precipitation and immunodetection was performed as in A.(C) Coimmunoprecipitation of MORG1 with Raf-1. CCL39 cells were transfected with either control vector or FLAG-Raf-1 and HA-MORG1. Raf-1 precipitation and immunodetection was performed as in A. (D) Coimmunoprecipitation of MORG1 with ERK1 and ERK2. CCL39 cells were transfected with either control vector or FLAG-ERK1 and HA-MORG1 or FLAG-ERK2 and HA-MORG1. Precipitation of ERKs and immunodetection was performed as in A. IP, immunoprecipitation; MEK1/2, MEK1 and MEK2.
MP1 was shown to associate with the protein kinase MEK1 (25), which raises the possibility that MP1 targets MEK1 to MORG1. We therefore examined whether MORG1 is present in MEK1 immune complexes. Analysis of MEK1 immunoprecipitates from cells transfected with HA-MORG1 and FLAG-MEK1 showed that MEK1 coimmunoprecipitated MORG1 (Fig. 2B). However, we found that both MEK1ΔPRS and MEK2, although unable to bind to MP1 (25), coimmunoprecipitated MORG1 to a similar level as MEK1 (Fig. 2B), indicating that MP1-binding is dispensable for the MEK–MORG1 interaction. In addition, MORG1Δ6–7, a deletion mutant that does not interact with MP1, still coprecipitated with MEK1 (data not shown), further supporting the notion that MORG1 associates with MEK independently of MP1.
Extended binding studies with other components of the ERK pathway revealed that MORG1 also coimmunoprecipitated with Raf-1 (Fig. 2C) and B-Raf (data not shown) as well as with both ERK isoforms (Fig. 2D).
MORG1 associates less stably with MEK than it does with Raf-1 or MP1 as estimated from the amount of HA-MORG1 in MP1, Raf-1, and MEK precipitates (data not shown). Because simultaneous association of several proteins that interact weakly with each other can result in more stable oligomeric complexes, we wanted to analyze MEK–MORG1 interactions in the presence of Raf-1, ERK, and MP1, respectively. Cells were transfected with FLAG-MEK1 and HA-MORG1 together with control vector, Raf-1, HA-ERK1, or MP1, and MEK1 was immunoprecipitated. Comparison of the level of HA-MORG1 in MEK1 precipitates showed that the amount of HA-MORG1 was increased significantly when Raf-1, ERK1, or MP1 was coexpressed (Fig. 3A). However, when the Raf-1–MORG1 interaction was examined in analogous experiments, little or no effect was seen when MEK1, ERK1, and MP1 were coexpressed (Fig. 3B). These results indicate that the relatively weak MORG1–MEK1 interaction can be stabilized in ternary complexes when MP1, Raf-1, or ERK is present. Consequently, one would predict that MEK proteins that are unable to interact with MP1 (such as MEK1ΔPRS or MEK2) should no longer be stabilized in complexes with MORG1 when MP1 is overexpressed. Indeed, we find that both MEK1ΔPRS and MEK2 precipitated comparable amounts of MORG1 in the presence or absence of MP1 (Fig. 3C and data not shown). Taken together, these data are consistent with the idea that distinct protein–protein interactions facilitate the recruitment of MEK1 to a multimeric protein complex, consisting of at least Raf, ERK, MORG1, and MP1.
Fig. 3.
Enhanced association of MORG1 with MEK1 mediated by components of the MAPK pathway. (A) Raf-1, ERK1, and MP1 enhance MORG1-MEK1 association. CCL39 cells were transfected with FLAG-MEK1and HA-MORG1 together with Raf-1, HA-ERK1, or MP1 as indicated. Precipitation of MEK and immunodetection were performed as in Fig. 2 A. Cell lysates were probed with antibodies as indicated to verify expression of cotransfected proteins. (B) MEK1, ERK1, and MP1 do not enhance MORG1-Raf-1 association. CCL39 cells were transfected with FLAG-Raf-1 and HA-MORG1 together with HA-MEK1, HA-ERK1, or MP1. Precipitation of Raf-1 and immunodetection was performed as in A. The additional band detected below HA-MORG1 represents the degradation product of HA-ERK1, which is occasionally seen. (C) Prolinerich sequence of MEK1 is necessary for MP1-mediated enhancement of MEK1–MORG1 interactions. CCL39 cells were transfected with various FLAG-MEK constructs and HA-MORG1 either with or without MP1. Precipitation of MEKs and immunodetection were performed as in A.
MORG1 Facilitates ERK Activation. Because MORG1 participates in the assembly of ERK cascade signaling complexes, we investigated whether MORG1 was able to modulate this signaling system. Cells were transfected with FLAG-ERK1 and increasing amounts of MORG1. After serum starvation, cells were stimulated with FBS, and ERK1 kinase activity was assessed in immune-complex kinase assays. MORG1 enhanced ERK1 activity in a dose-dependent manner at low to medium concentrations (Fig. 4A), but this effect was reversed at higher levels of MORG1 expression. This observation led us to speculate that, as with MP1 (25), a functional and productive signaling unit can only be assembled when the relative stoichiometry of the respective components is optimal. Consequently, very high concentrations of MORG1 should lead to the formation of nonproductive complexes that interfere with signal transduction. Consistent with this idea, transfection of very high concentrations of MORG1 cDNA resulted in significant inhibition of FBS-mediated ERK activation, as assessed by ERK immunecomplex kinase assays (Fig. 4B).
Fig. 4.
MORG1 enhances ERK1 activation in a concentration-dependent manner. (A) MORG1 effect on ERK activation is concentration-dependent. Cos1 cells were transfected with FLAG-ERK1 and with either empty vector or 1× (0.03 μg), 1.3× (0.04 μg), 1.6× (0.05 μg), 2× (0.06 μg), or 3.3× (0.1 μg) the amount of HA-MORG1. After serum starvation for 5 h, cells were either left untreated or stimulated with 10% FBS for 10 min, and ERK1 kinase activity was assayed in ERK1 immune complexes. Immunoprecipitates were probed with ERK1 antibody to verify comparable levels of ERK protein precipitated (results from duplicates are shown). Equal amounts of cell lysates from duplicate experiments were pooled, and MORG1 expression levels were verified with HA antibody. Comparable results have been obtained when ERK was stimulated with Raf-1, PMA, or LPA. (B) High level of MORG1 expression inhibits ERK activation. Cos1 cells were transfected with FLAG-ERK1 and either empty vector or 1× (0.04 μg), 8× (0.3 μg), or 64× (2.4 μg) the amount of HA-MORG1, and ERK kinase activity was determined as described in A. (C) Ectopically expressed MORG1 modulates activity of endogenous ERK. Either control cells (HeLa-pREP) or HeLa cells stably expressing low (HeLa-MORG300) and high (HeLa-MORG500) levels of HA-MORG1 were serum-starved for 5 h and stimulated with 10% FBS for 10 min. Cell lysates were probed with antibody recognizing the doubly phosphorylated form of ERK (pp-ERK) and with antibody directed against tubulin to confirm equal loading of proteins. HA-MORG1 expression was determined by probing lysates with HA antibody.
To determine whether ectopically expressed MORG1 can modulate activation of endogenous ERK, we established HeLa-MORG300 and HeLa-MORG500, cell lines stably expressing low and high levels of HA-MORG1, respectively (data not shown). After cell stimulation with FBS, ERK activation was assessed by probing the cell lysates with antibody that specifically recognizes doubly phosphorylated, activated ERK. Analogous to results described above, low levels of MORG1 expression (HeLa-MORG300) enhanced endogenous ERK phosphorylation whereas high MORG1 expression levels (HeLa-MORG500) markedly reduced ERK phosphorylation (Fig. 4C).
Because MORG1 associates with all of the kinases of the Raf/MEK/ERK cascade, it is conceivable that MORG1 facilitates the efficiency of signal propagation from Raf-1 to ERK. We therefore tested whether MORG1 was able to enhance Raf-1-mediated activation of ERK. Cells were transfected with FLAG-ERK1, with or without Raf-1 and increasing amounts of MORG1 as indicated (Fig. 5A). After serum deprivation, ERK1 was immunoprecipitated, and its kinase activity was assessed. The basal activity of ectopically expressed Raf-1 was sufficient to stimulate ERK1 kinase activity, and, when MORG1 was coexpressed, a moderate, dose-dependent enhancement of Raf-1-induced ERK1 activation was observed (Fig. 5A). Similarly, B-Raf was also found to stimulate ERK1 activation, and cotransfection of MORG1 resulted in a further increase in ERK1 activity (data not shown).
Fig. 5.
MORG1 enhances ERK1 enzymatic activation in response to specific stimuli. (A) MORG1 enhances Raf-1-mediated activation of ERK1. NIH 3T3 cells were transfected with either control vector and FLAG-ERK1 or Raf-1 and FLAG-ERK1 with 1× (0.3 μg), 3× (0.9 μg), or 9× (2.7 μg) the amount of untagged MORG1. After serum starvation for 5 h, ERK1 kinase activity was determined as described in Fig. 4A. (B) MORG1 enhances ERK1 activation by LPA. NIH 3T3 cells were transfected with FLAG-ERK1 and with either empty vector or with 1× (0.3 μg), 3× (0.9 μg), or 9× (2.7 μg) the amount of untagged MORG1. After serum starvation for 5 h, cells were either left untreated or stimulated with 3 μM LPA for 10 min. ERK1 activity and expression levels were determined as described in Fig. 4A. (C) MORG1 enhances ERK1 activation by PMA. Cos1 cells were transfected, and ERK activity was determined as described in B with the exception that cells were stimulated with 5 nM PMA. (D) MORG1-mediated enhancement of ERK1 activity is agonist-specific. (Only one of four experiments is shown.) Cos1 cells were transfected, and ERK activity was determined as described in B with the exception that, in parallel, transfected dishes were stimulated with either LPA (1 μM) or EGF (1 ng/ml) for 10 min. (E) Ectopically expressed HA-MORG1 modulates activity of endogenous ERK in an agonist-specific manner. Either control cells (HeLa-pREP) or HeLa cells stably expressing low (HeLa-MORG300) and high (HeLa-MORG500) levels of HA-MORG1 were treated as described in Fig. 4C with the exception that cells were stimulated in parallel with either 10% FBS or 0.5 ng/ml EGF. Cell lysates were probed as described in Fig. 4C. pp-ERK, doubly phosphorylated ERK.
MORG1 Facilitates Agonist-Specific ERK Activation. To gain insight into the biological functions of MORG1, we examined its utility in response to different agonists. Ectopic expression of MORG1 resulted in a dose-dependent enhancement of ERK1 activity when cells were treated with lysophosphatidic acid (LPA; Fig. 5B) or phorbol 12-myristate 13-acetate (PMA; Fig. 5C). In contrast, little or no MORG1-mediated enhancement of ERK activity was seen in initial experiments, when cells were stimulated with epidermal growth factor (EGF) or platelet-derived growth factor (data not shown). This finding raised the possibility that MORG1 functions in an agonist-specific manner. To further explore this concept, we performed side-by-side comparisons of MORG1 function in cells stimulated with either LPA or EGF (four independent experiments, each in triplicate). MORG1 enhanced ERK activity in cells stimulated with LPA with an estimated fold change of 1.74 (P = 0.018) but had no statistically significant effect on ERK activation (P = 0.60) in cells stimulated with EGF (Fig. 5D; for detailed statistical analysis see Table 1, which is published as supporting information on the PNAS web site). Consistent with this finding, we observed that in HeLa cells endogenous ERK phosphorylation was only marginally affected by HA-MORG1 expression when cells were stimulated with EGF. However, when identical experiments were performed after FBS stimulation, low expression levels of MORG (HeLa-MORG300) noticeably enhanced ERK phosphorylation (Fig. 5E).
Knockdown of MORG1 Interferes with ERK Activation in an Agonist-Specific Manner. To assess whether loss of MORG1 affects ERK activation, MORG1 mRNA was knocked down with siRNA oligonucleotides. We targeted H A-MORG1 in HeLa-MORG300 cells, engineered to express HA-tagged MORG1, and determined with anti-HA antibody that siRNA knocked down MORG1 protein to the limits of detection (Fig. 6A). We also targeted unengineered parental HeLa cells and determined that siRNA oligonucleotides also down-regulated endogenous MORG1 mRNA by ≈70%, as determined by quantitative realtime PCR (data not shown), confirming the ability of the siRNA duplex to knock down both exogenous and endogenous MORG1. When parental HeLa cells were treated with MORG1 siRNA, ERK activation was noticeably reduced in response to FBS (Fig. 6B). To test whether siRNA-mediated suppression of MORG1 also affects ERK activation in response to EGF, parental HeLa cells were stimulated with different EGF concentrations or with 10% FBS in the presence or absence of siRNA. Knockdown of endogenous MORG1 had little or no effect on ERK activation after EGF stimulation but caused a significant reduction of ERK activity when cells were stimulated with FBS (Fig. 6C).
Fig. 6.
siRNA-mediated suppression of MORG1 affects ERK activation. (A) MORG1 siRNA efficiently down-regulates MORG1. HeLa-MORG300 cells were transfected with siRNA duplexes directed against MORG1, and HA-MORG1 expression levels were determined with HA antibody. ERK2 expression levels served as a loading control. (B) Down-regulation of endogenous MORG1 protein inhibits ERK phosphorylation in response to FBS. Parental HeLa cells were transfected with siRNA for 24 h, serum-starved for 5 h, and stimulated with 10% FBS for 10 min. Endogenous MORG1 mRNA was knocked down by 70%, as determined by quantitative real-time PCR (data not shown). Cell lysates were probed with antibody recognizing doubly phosphorylated ERK (pp-ERK) and with antibody directed against ERK2 and tubulin to confirm equal loading of proteins. (C) Down-regulation of endogenous MORG1 protein inhibits ERK phosphorylation in an agonist-specific manner. Parental HeLa cells were treated as described in B with the exception that cells were stimulated in parallel with either 10% FBS or an increasing concentration of EGF (0.1, 0.3, and 1.0 ng/ml EGF). Cell lysates were probed as described in B.
Discussion
Taken together, the data presented here show that MORG1 acts as a module in the assembly of a multicomponent scaffold for the ERK pathway. This WD-40 protein was isolated as a binding partner of MP1, which also displays scaffold properties but is likely to be too small (14.5 kDa) to function independently in that capacity. MORG1 associates with multiple protein kinase components of the MAPK pathway, ERK1, ERK2, MEK1, MEK2, Raf-1, and B-Raf, and selectively facilitates the activation of ERK1 and ERK2 in response to a subset of agonists. Importantly, siRNA knockdown studies provide convincing evidence that MORG1 is a significant component in the activation of ERKs in response to FBS but is dispensable for EGF-mediated ERK activation.
In agreement with the theoretical model of MAPK scaffolds (27), MORG1 biphasically modulates the activation of the ERK cascade: at low concentrations MORG1 enhances ERK activation, whereas high MORG1 concentrations lead to the inhibition of ERK activation. Likewise, overexpression of both MP1 and KSR-1 can result in different responses, depending on the relative stoichiometry of the individual components (17–20, 25, 28). Thus, the stoichiometry of components participating in a given scaffolding complex is critical for optimal signal processing. Low scaffold expression levels favor formation of productive signaling complexes, whereas overexpression of scaffolds would result in forming incomplete, nonfunctional entities (27, 29).
It has been suggested that assembly of stable signaling complexes is a consequence of the association of several proteins with multiple relatively weak contacts (4). Therefore, low-affinity binding of a single protein could be shifted to a higher binding affinity by assembling multidomain interactions among multiple proteins, thus excluding irrelevant proteins and obtaining a high degree of specificity in complex assembly. The stabilization of the relatively weak association of MEK with MORG1 through Raf-1, ERK, and MP1 and the observation that MP1 is not a “bridge” for MEK binding to MORG1 provide the potential of MORG1 signaling complexes to achieve, through combinatorial effects, a high degree of stability, signaling specificity, and regulatory flexibility.
The MAPK cascade is activated in response to a broad spectrum of extracellular signals. However, it is unclear how the signal from the ubiquitously activated MAPK cascade is transmitted to create a unique biological response. Scaffold proteins can represent important regulatory components that selectively assemble signaling modules and link them to specific stimuli. Indeed, in neurons isolated from Jip1-/- mice, JIP-1 is necessary for JNK activation when cells are exposed to anoxia or excitotoxic stress, but JIP-1 is dispensable for JNK activation when cells are treated with anisomycin and UV light (24). The fact that MORG1 is used in an agonist-specific manner to facilitate ERK activation raises the possibility that, analogous to JIP-1, MORG1 selectively controls a subset of ERK-dependent biological responses.
The molecular mechanism of MORG1 specificity remains unknown; however, as with Ste5 (10), it is reasonable to predict that MORG1 recruits signaling complexes to a specific upstream activator. In this context it is interesting to note that the MORG1 partner MP1 localizes to late endosomal/lysozomal compartments through interaction with protein p14 (30, 31). The precise intracellular localization of MORG1 is unknown; however, MP1 and MORG1 fused to fluorescent proteins colocalized in vesicle-like structures (unpublished data), supporting the idea that the MORG1-MP1 scaffold participates in ERK signaling at specific intracellular locations.
In conclusion, we propose that MORG1 is a nonenzymatic scaffold-like protein in the MAPK/ERK pathway linking the cascade to specific agonists. The interaction with MP1 suggests that scaffolds could be formed from standardized, modular units, which would allow a high degree of stability, signaling specificity, and regulatory flexibility.
Supplementary Material
Acknowledgments
We thank Kim Wong for expert technical assistance, Mike Harding for help with quantitative real-time PCR, and all of the members of the Parsons–Weber–Parsons laboratories for helpful discussions. This work was supported by National Institutes of Health Grants CA 39076, CA 40042, and GM 47332.
This paper was submitted directly (Track II) to the PNAS office.
Abbreviations: MAPK, mitogen-activated protein kinase; ERK, extracellular signal-regulated kinase; MEK, MAPK/ERK kinase; MP1, MEK partner 1; MORG, MAPK organizer; KSR, kinase suppressor of Ras; JNK, c-Jun N-terminal kinase; JIP, JNK-interacting protein; HA, hemagglutinin; MBP, myelin basic protein; siRNA, short-interfering RNA; LPA, lysophosphatidic acid; PMA, phorbol 12-myristate 13-acetate; EGF, epidermal growth factor.
Data deposition: The sequence reported in this paper has been deposited in the GenBank database (accession no. AY365434).
References
- 1.Lewis, T. S., Shapiro, P. S. & Ahn, N. G. (1998) Adv. Cancer Res. 74, 49-139. [DOI] [PubMed] [Google Scholar]
- 2.Chang, L. & Karin, M. (2001) Nature 410, 37-40. [DOI] [PubMed] [Google Scholar]
- 3.Pearson, G., Robinson, F., Beers, G. T., Xu, B. E., Karandikar, M., Berman, K. & Cobb, M. H. (2001) Endocr. Rev. 22, 153-183. [DOI] [PubMed] [Google Scholar]
- 4.Schaeffer, H. J. & Weber, M. J. (1999) Mol. Cell. Biol. 19, 2435-2444. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Smith, F. D. & Scott, J. D. (2002) Curr. Biol. 12, R32-R40. [DOI] [PubMed] [Google Scholar]
- 6.Weston, C. R., Lambright, D. G. & Davis, R. J. (2002) Science 296, 2345-2347. [DOI] [PubMed] [Google Scholar]
- 7.Choi, K. Y., Satterberg, B., Lyons, D. M. & Elion, E. A. (1994) Cell 78, 499-512. [DOI] [PubMed] [Google Scholar]
- 8.Marcus, S., Polverino, A., Barr, M. & Wigler, M. (1994) Proc. Natl. Acad. Sci. USA 91, 7762-7766. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Park, S. H., Zarrinpar, A. & Lim, W. A. (2003) Science 299, 1061-1064. [DOI] [PubMed] [Google Scholar]
- 10.Elion, E. A. (2000) Curr. Opin. Microbiol. 3, 573-581. [DOI] [PubMed] [Google Scholar]
- 11.Harris, K., Lamson, R. E., Nelson, B., Hughes, T. R., Marton, M. J., Roberts, C. J., Boone, C. & Pryciak, P. M. (2001) Curr. Biol. 11, 1815-1824. [PubMed] [Google Scholar]
- 12.Michaud, N. R., Therrien, M., Cacace, A., Edsall, L. C., Spiegel, S., Rubin, G. M. & Morrison, D. K. (1997) Proc. Natl. Acad. Sci. USA 94, 12792-12796. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Xing, H., Kornfeld, K. & Muslin, A. J. (1997) Curr. Biol. 7, 294-300. [DOI] [PubMed] [Google Scholar]
- 14.Denouel-Galy, A., Douville, E. M., Warne, P. H., Papin, C., Laugier, D., Calothy, G., Downward, J. & Eychene, A. (1998) Curr. Biol. 8, 46-55. [DOI] [PubMed] [Google Scholar]
- 15.Yu, W., Fantl, W. J., Harrowe, G. & Williams, L. T. (1998) Curr. Biol. 8, 56-64. [DOI] [PubMed] [Google Scholar]
- 16.Nguyen, A., Burack, W. R., Stock, J. L., Kortum, R., Chaika, O. V., Afkarian, M., Muller, W. J., Murphy, K. M., Morrison, D. K., Lewis, R. E., et al. (2002) Mol. Cell. Biol. 22, 3035-3045. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Joneson, T., Fulton, J. A., Volle, D. J., Chaika, O. V., Bar-Sagi, D. & Lewis, R. E. (1998) J. Biol. Chem. 273, 7743-7748. [DOI] [PubMed] [Google Scholar]
- 18.Cacace, A. M., Michaud, N. R., Therrien, M., Mathes, K., Copeland, T., Rubin, G. M. & Morrison, D. K. (1999) Mol. Cell. Biol. 19, 229-240. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Stewart, S., Sundaram, M., Zhang, Y., Lee, J., Han, M. & Guan, K. L. (1999) Mol. Cell. Biol. 19, 5523-5534. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Roy, F., Laberge, G., Douziech, M., Ferland-McCollough, D. & Therrien, M. (2002) Genes Dev. 16, 427-438. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Lozano, J., Xing, R., Cai, Z., Jensen, H. L., Trempus, C., Mark, W., Cannon, R. & Kolesnick, R. (2003) Cancer Res. 63, 4232-4238. [PubMed] [Google Scholar]
- 22.Whitmarsh, A. J., Cavanagh, J., Tournier, C., Yasuda, J. & Davis, R. J. (1998) Science 281, 1671-1674. [DOI] [PubMed] [Google Scholar]
- 23.Dickens, M., Rogers, J. S., Cavanagh, J., Raitano, A., Xia, Z., Halpern, J. R., Greenberg, M. E., Sawyers, C. L. & Davis, R. J. (1997) Science 277, 693-696. [DOI] [PubMed] [Google Scholar]
- 24.Whitmarsh, A. J., Kuan, C. Y., Kennedy, N. J., Kelkar, N., Haydar, T. F., Mordes, J. P., Appel, M., Rossini, A. A., Jones, S. N., Flavell, R. A., et al. (2001) Genes Dev. 15, 2421-2432. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Schaeffer, H. J., Catling, A. D., Eblen, S. T., Collier, L. S., Krauss, A. & Weber, M. J. (1998) Science 281, 1668-1671. [DOI] [PubMed] [Google Scholar]
- 26.Catling, A. D., Schaeffer, H. J., Reuter, C. W., Reddy, G. R. & Weber, M. J. (1995) Mol. Cell. Biol. 15, 5214-5225. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Levchenko, A., Bruck, J. & Sternberg, P. W. (2000) Proc. Natl. Acad. Sci. USA 97, 5818-5823. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Catling, A. D., Eblen, S. T., Schaeffer, H. J. & Weber, M. J. (2001) Methods Enzymol. 332, 368-387. [DOI] [PubMed] [Google Scholar]
- 29.Burack, W. R. & Shaw, A. S. (2000) Curr. Opin. Cell Biol. 12, 211-216. [DOI] [PubMed] [Google Scholar]
- 30.Wunderlich, W., Fialka, I., Teis, D., Alpi, A., Pfeifer, A., Parton, R. G., Lottspeich, F. & Huber, L. A. (2001) J. Cell Biol. 152, 765-776. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Teis, D., Wunderlich, W. & Huber, L. A. (2002) Dev. Cell 3, 803-814. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.