Abstract
Methylation interference experiments reveal bases involved in three different catalytic functions of the T4-phage derived sunY self-splicing intron. RNA molecules methylated at the N-7 position of the guanine at the cofactor binding site are inactive in cofactor-dependent splicing and 3' splice-site hydrolysis. In contrast, 5' splice-site hydrolysis occurs despite methylation at this position. Specific adenines that have been implicated in docking of the P1 stem to the catalytic core are shown to be important for cofactor-dependent splicing and essential for 5' splice-site hydrolysis. Similarly, methylation of bases in the P9.0 stem, as well as C56 in J5/4, interferes with 3' splice-site hydrolysis and with the splicing reaction. All of the bases identified as important for the overall splicing reaction are also identified as essential for either the 5' or 3' splice-site hydrolysis reactions, and vice versa. It is inferred that the bases implicated in 5' and 3' splice-site hydrolysis are involved in specific interactions of the 5' and 3' splice site, respectively, with the catalytic core.
Full text
PDF
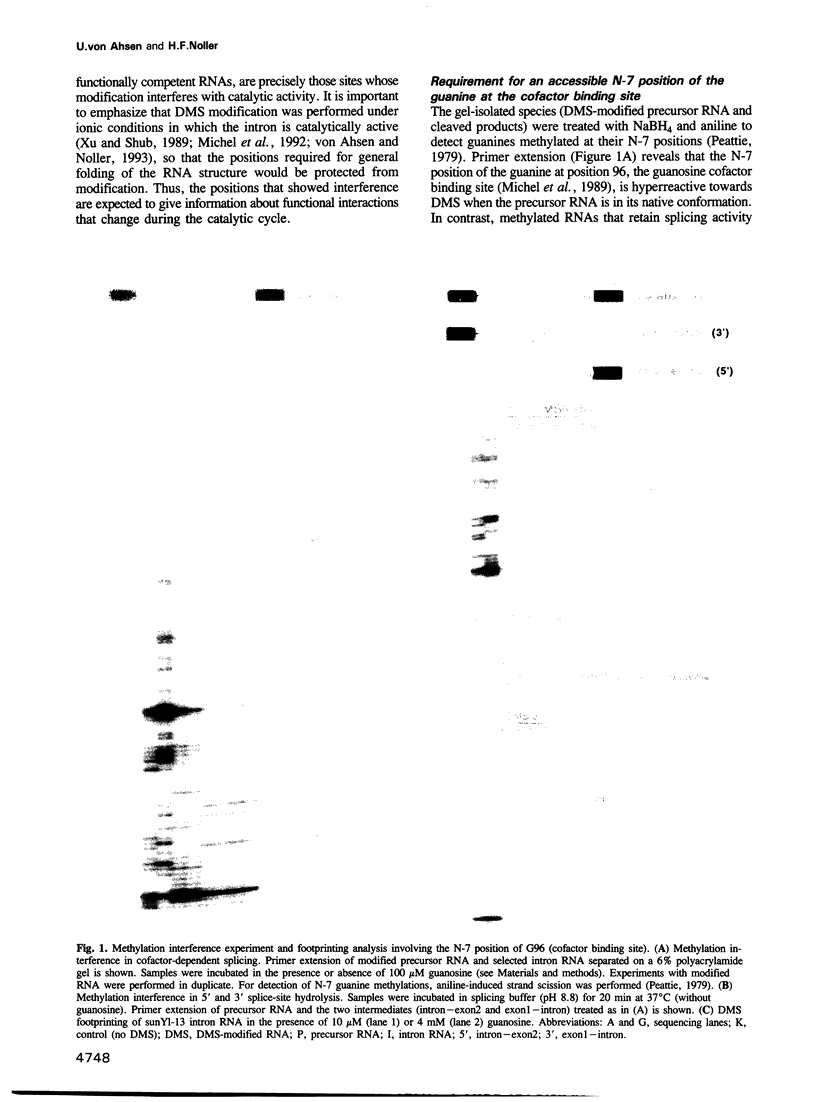






Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Been M. D., Perrotta A. T. Group I intron self-splicing with adenosine: evidence for a single nucleoside-binding site. Science. 1991 Apr 19;252(5004):434–437. doi: 10.1126/science.2017681. [DOI] [PubMed] [Google Scholar]
- Bevilacqua P. C., Kierzek R., Johnson K. A., Turner D. H. Dynamics of ribozyme binding of substrate revealed by fluorescence-detected stopped-flow methods. Science. 1992 Nov 20;258(5086):1355–1358. doi: 10.1126/science.1455230. [DOI] [PubMed] [Google Scholar]
- Burke J. M., Esherick J. S., Burfeind W. R., King J. L. A 3' splice site-binding sequence in the catalytic core of a group I intron. Nature. 1990 Mar 1;344(6261):80–82. doi: 10.1038/344080a0. [DOI] [PubMed] [Google Scholar]
- Caprara M. G., Waring R. B. Important 2'-hydroxyl groups within the core of a group I intron. Biochemistry. 1993 Apr 13;32(14):3604–3610. doi: 10.1021/bi00065a011. [DOI] [PubMed] [Google Scholar]
- Cech T. R. Self-splicing of group I introns. Annu Rev Biochem. 1990;59:543–568. doi: 10.1146/annurev.bi.59.070190.002551. [DOI] [PubMed] [Google Scholar]
- Cech T. R. The chemistry of self-splicing RNA and RNA enzymes. Science. 1987 Jun 19;236(4808):1532–1539. doi: 10.1126/science.2438771. [DOI] [PubMed] [Google Scholar]
- Herr W., Chapman N. M., Noller H. F. Mechanism of ribosomal subunit association: discrimination of specific sites in 16 S RNA essential for association activity. J Mol Biol. 1979 Jun 5;130(4):433–449. doi: 10.1016/0022-2836(79)90433-9. [DOI] [PubMed] [Google Scholar]
- Herschlag D. Evidence for processivity and two-step binding of the RNA substrate from studies of J1/2 mutants of the Tetrahymena ribozyme. Biochemistry. 1992 Feb 11;31(5):1386–1399. doi: 10.1021/bi00120a015. [DOI] [PubMed] [Google Scholar]
- Heuer T. S., Chandry P. S., Belfort M., Celander D. W., Cech T. R. Folding of group I introns from bacteriophage T4 involves internalization of the catalytic core. Proc Natl Acad Sci U S A. 1991 Dec 15;88(24):11105–11109. doi: 10.1073/pnas.88.24.11105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Inoue T., Sullivan F. X., Cech T. R. Intermolecular exon ligation of the rRNA precursor of Tetrahymena: oligonucleotides can function as 5' exons. Cell. 1985 Dec;43(2 Pt 1):431–437. doi: 10.1016/0092-8674(85)90173-4. [DOI] [PubMed] [Google Scholar]
- Inoue T., Sullivan F. X., Cech T. R. New reactions of the ribosomal RNA precursor of Tetrahymena and the mechanism of self-splicing. J Mol Biol. 1986 May 5;189(1):143–165. doi: 10.1016/0022-2836(86)90387-6. [DOI] [PubMed] [Google Scholar]
- Kay P. S., Menzel P., Inoue T. Two guanosine binding sites exist in group I self-splicing IVS RNAs. EMBO J. 1988 Nov;7(11):3531–3537. doi: 10.1002/j.1460-2075.1988.tb03229.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Legault P., Herschlag D., Celander D. W., Cech T. R. Mutations at the guanosine-binding site of the Tetrahymena ribozyme also affect site-specific hydrolysis. Nucleic Acids Res. 1992 Dec 25;20(24):6613–6619. doi: 10.1093/nar/20.24.6613. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Michel F., Ellington A. D., Couture S., Szostak J. W. Phylogenetic and genetic evidence for base-triples in the catalytic domain of group I introns. Nature. 1990 Oct 11;347(6293):578–580. doi: 10.1038/347578a0. [DOI] [PubMed] [Google Scholar]
- Michel F., Hanna M., Green R., Bartel D. P., Szostak J. W. The guanosine binding site of the Tetrahymena ribozyme. Nature. 1989 Nov 23;342(6248):391–395. doi: 10.1038/342391a0. [DOI] [PubMed] [Google Scholar]
- Michel F., Jaeger L., Westhof E., Kuras R., Tihy F., Xu M. Q., Shub D. A. Activation of the catalytic core of a group I intron by a remote 3' splice junction. Genes Dev. 1992 Aug;6(8):1373–1385. doi: 10.1101/gad.6.8.1373. [DOI] [PubMed] [Google Scholar]
- Michel F., Netter P., Xu M. Q., Shub D. A. Mechanism of 3' splice site selection by the catalytic core of the sunY intron of bacteriophage T4: the role of a novel base-pairing interaction in group I introns. Genes Dev. 1990 May;4(5):777–788. doi: 10.1101/gad.4.5.777. [DOI] [PubMed] [Google Scholar]
- Michel F., Westhof E. Modelling of the three-dimensional architecture of group I catalytic introns based on comparative sequence analysis. J Mol Biol. 1990 Dec 5;216(3):585–610. doi: 10.1016/0022-2836(90)90386-Z. [DOI] [PubMed] [Google Scholar]
- Peattie D. A. Direct chemical method for sequencing RNA. Proc Natl Acad Sci U S A. 1979 Apr;76(4):1760–1764. doi: 10.1073/pnas.76.4.1760. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Peattie D. A., Herr W. Chemical probing of the tRNA--ribosome complex. Proc Natl Acad Sci U S A. 1981 Apr;78(4):2273–2277. doi: 10.1073/pnas.78.4.2273. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Piccirilli J. A., Vyle J. S., Caruthers M. H., Cech T. R. Metal ion catalysis in the Tetrahymena ribozyme reaction. Nature. 1993 Jan 7;361(6407):85–88. doi: 10.1038/361085a0. [DOI] [PubMed] [Google Scholar]
- Price J. V., Engberg J., Cech T. R. 5' exon requirement for self-splicing of the Tetrahymena thermophila pre-ribosomal RNA and identification of a cryptic 5' splice site in the 3' exon. J Mol Biol. 1987 Jul 5;196(1):49–60. doi: 10.1016/0022-2836(87)90510-9. [DOI] [PubMed] [Google Scholar]
- Pyle A. M., Murphy F. L., Cech T. R. RNA substrate binding site in the catalytic core of the Tetrahymena ribozyme. Nature. 1992 Jul 9;358(6382):123–128. doi: 10.1038/358123a0. [DOI] [PubMed] [Google Scholar]
- Stern S., Moazed D., Noller H. F. Structural analysis of RNA using chemical and enzymatic probing monitored by primer extension. Methods Enzymol. 1988;164:481–489. doi: 10.1016/s0076-6879(88)64064-x. [DOI] [PubMed] [Google Scholar]
- Wang J. F., Downs W. D., Cech T. R. Movement of the guide sequence during RNA catalysis by a group I ribozyme. Science. 1993 Apr 23;260(5107):504–508. doi: 10.1126/science.7682726. [DOI] [PubMed] [Google Scholar]
- Williams K. P., Fujimoto D. N., Inoue T. A region of group I introns that contains universally conserved residues but is not essential for self-splicing. Proc Natl Acad Sci U S A. 1992 Nov 1;89(21):10400–10404. doi: 10.1073/pnas.89.21.10400. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xu M. Q., Shub D. A. The catalytic core of the sunY intron of bacteriophage T4. Gene. 1989 Oct 15;82(1):77–82. doi: 10.1016/0378-1119(89)90032-2. [DOI] [PubMed] [Google Scholar]
- Young B., Herschlag D., Cech T. R. Mutations in a nonconserved sequence of the Tetrahymena ribozyme increase activity and specificity. Cell. 1991 Nov 29;67(5):1007–1019. doi: 10.1016/0092-8674(91)90373-7. [DOI] [PubMed] [Google Scholar]
- Zaug A. J., Kent J. R., Cech T. R. Reactions of the intervening sequence of the Tetrahymena ribosomal ribonucleic acid precursor: pH dependence of cyclization and site-specific hydrolysis. Biochemistry. 1985 Oct 22;24(22):6211–6218. doi: 10.1021/bi00343a027. [DOI] [PubMed] [Google Scholar]
- van der Horst G., Inoue T. Requirements of a group I intron for reactions at the 3' splice site. J Mol Biol. 1993 Feb 5;229(3):685–694. doi: 10.1006/jmbi.1993.1072. [DOI] [PubMed] [Google Scholar]
- von Ahsen U., Davies J., Schroeder R. Non-competitive inhibition of group I intron RNA self-splicing by aminoglycoside antibiotics. J Mol Biol. 1992 Aug 20;226(4):935–941. doi: 10.1016/0022-2836(92)91043-o. [DOI] [PubMed] [Google Scholar]
- von Ahsen U., Noller H. F. Footprinting the sites of interaction of antibiotics with catalytic group I intron RNA. Science. 1993 Jun 4;260(5113):1500–1503. doi: 10.1126/science.8502993. [DOI] [PubMed] [Google Scholar]