Significance
Many complex biological phenotypes are multifactorial in nature, yet current strategies for studying biological systems are limited by the inability to generate and track complex combinatorial cellular perturbations in a scalable fashion. Here, we introduce a method, Combinatorial Genetics En Masse (CombiGEM), that allows both facile generation of high-order combinations and high-throughput characterization of pooled populations using next-generation sequencing. We apply CombiGEM to identify genetic combinations that enhance killing of highly antibiotic-resistant bacteria with antibiotics, thus providing potential targets for antimicrobial development. We envision that CombiGEM will be broadly useful for basic science and biotechnology applications, such as complex genetic screening, identification of novel drug targets, interactome mapping, and synthetic circuit characterization.
Keywords: synthetic biology, combination therapy, systems biology, drug resistance, multifactorial genetics
Abstract
New therapeutic strategies are needed to treat infections caused by drug-resistant bacteria, which constitute a major growing threat to human health. Here, we use a high-throughput technology to identify combinatorial genetic perturbations that can enhance the killing of drug-resistant bacteria with antibiotic treatment. This strategy, Combinatorial Genetics En Masse (CombiGEM), enables the rapid generation of high-order barcoded combinations of genetic elements for high-throughput multiplexed characterization based on next-generation sequencing. We created ∼34,000 pairwise combinations of Escherichia coli transcription factor (TF) overexpression constructs. Using Illumina sequencing, we identified diverse perturbations in antibiotic-resistance phenotypes against carbapenem-resistant Enterobacteriaceae. Specifically, we found multiple TF combinations that potentiated antibiotic killing by up to 106-fold and delivered these combinations via phagemids to increase the killing of highly drug-resistant E. coli harboring New Delhi metallo-beta-lactamase-1. Moreover, we constructed libraries of three-wise combinations of transcription factors with >4 million unique members and demonstrated that these could be tracked via next-generation sequencing. We envision that CombiGEM could be extended to other model organisms, disease models, and phenotypes, where it could accelerate massively parallel combinatorial genetics studies for a broad range of biomedical and biotechnology applications, including the treatment of antibiotic-resistant infections.
Antibiotic resistance dramatically reduces the effectiveness of existing treatment options for clinical bacterial infections. Antibiotic-resistant organisms can result in recalcitrant infections that are challenging to treat with existing antibiotics, leading to >20,000 deaths a year (1). Concurrently, the antibacterial discovery pipeline has shrunk dramatically, and ongoing clinical trials for antibacterials often feature derivatives of compounds for which resistance mechanisms already exist (2). Recent attention has centered on drug-resistant Gram-negative bacteria such as carbapenem-resistant Enterobactericeae (CRE) (3, 4). These strains can use degradative enzymes, such as New Delhi metallo-beta-lactamase-1 (NDM-1), which can hydrolyze beta-lactam antibiotics (5). New treatment regimens and adjuvants to combat antibiotic resistance are urgently needed (6, 7).
Previously, we showed that overexpressing single genes from constructs that are delivered into bacteria via engineered bacteriophages could potentiate the lethality of antibiotics in wild-type and quinolone-resistant Escherichia coli carrying a chromosomal gyrA mutation (8). Expression of one of these genes, lexA3, suppressed the SOS network that is normally activated in response to antibiotic treatment (9–12). In addition, we demonstrated that overexpression of the double gene combination csrA + ompF had enhanced antibiotic-potentiating efficacy compared with its individual gene constituents (8). However, this study identified the single genes and the gene pair via a laborious one-by-one testing protocol using a candidate gene approach informed by previous studies (9–14). Based on this strategy, it would take on the order of n2 steps [abbreviated O(n2), using order notation from computer science] to construct all pairwise combinations of n genes and another O(n2) experiments to test their efficacy. The brute-force nature of this approach restricts the scale and combinatorial complexity that can be tested when trying to identify pathways that can modulate antibiotic-resistance phenotypes.
To overcome these limitations, we developed a high-throughput platform for the identification of genetic combinations that potentiate antibiotic activity against clinically relevant drug-resistance determinants. Here, we describe a technology, termed Combinatorial Genetics En Masse (CombiGEM), that enables the rapid generation of high-order barcoded combinations of genetic elements for high-throughput multiplexed characterization based on next-generation sequencing. We applied CombiGEM to identify genetic combinations that enhance the effectiveness of antibiotics against CRE. CombiGEM uses an iterative cloning strategy beginning with a library of barcoded DNA elements (Fig. 1A). Restriction digests of pooled vector and insert libraries, followed by a one-pot ligation reaction, create a library of genetic combinations with the barcodes in close proximity to one another. As a result, the identities of each of the genetic components in a particular construct can be determined by reading the contiguous DNA barcode sequences with high-throughput sequencing. From the newly produced combinatorial library, higher-order combinations can be generated in fixed amounts of time by using the same procedure, rather than in exponentially increasing amounts of time. Therefore, combinatorial libraries can be constructed in a pooled format without needing to barcode each combination within an individual reaction. A similar DNA assembly method was described for optimizing translational rates in metabolic pathways, in which hits were individually sequenced to identify the hit components (15). In contrast, we use high-throughput sequencing to census the distribution of library members within pooled populations in a highly multiplexed manner. By comparing these distributions, we identify changes in the population under different experimental conditions to link combinatorial genotypes to specific phenotypes. We anticipate that CombiGEM will be a broadly useful tool for identifying genetic combinations conferring a desired phenotype and for altering the gene expression profile of a cell.
Fig. 1.
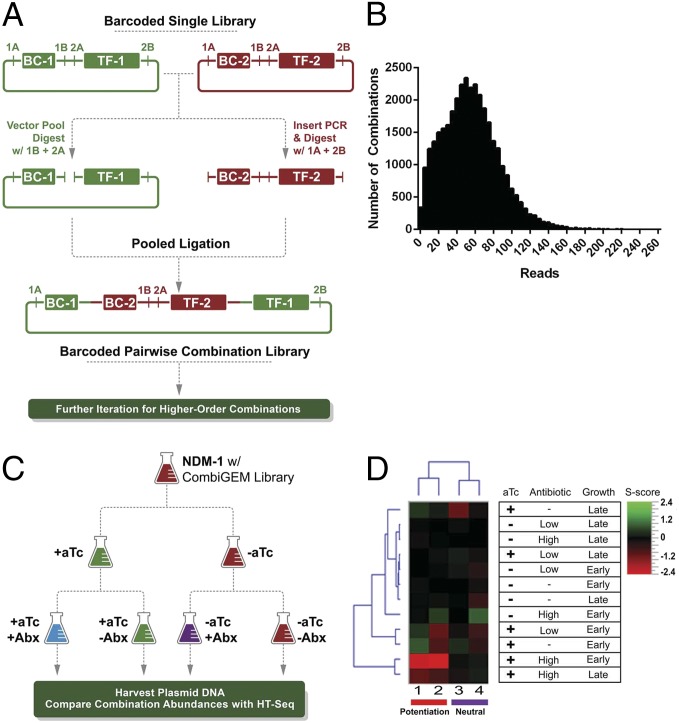
Overview, validation, and application of the CombiGEM technology to identify genetic combinations that combat antibiotic resistance. (A) Outline of the CombiGEM assembly strategy. Transcription factor (TF) expression constructs are barcoded (BC), and four restriction sites (1A, 1B, 2A, 2B) are positioned as shown. The pairs 1A/1B and 2A/2B consist of unique restriction sites that generate compatible overhangs within the pair but are incompatible with the other pair. The barcoded vectors are pooled and digested with enzymes 1B + 2A. Inserts are generated from vectors by PCR and digested with 1A + 2B. A one-pot ligation reaction produces a pairwise combinatorial library, which can be further digested and ligated with the same insert pool to produce higher-order combinations. In this work, barcodes are separated by 6 base pairs. (B) Distribution of high-throughput sequencing reads among combinations in the pairwise library. A total of 1.9 million reads were distributed among 34,554 combinations, recovering 98% of the possible 35,343 pairwise combinations. (C) Experimental outline. E. coli NDM-1 cells containing a CombiGEM library were diluted into cultures with and without anhydrotetracycline (aTc) and grown to middle logarithmic phase. Each culture was then further diluted into cultures with and without antibiotic (Abx), and with and without aTc. DNA from each condition was harvested at early and late logarithmic-growth stages and processed for high-throughput sequencing (HT-seq) with Illumina HiSeq. Comparisons of abundances of barcoded gene combinations among different conditions revealed genotypes that conferred desired phenotypes. (D) Heatmap of S scores for specific hits across experiments. Combinations marked as potentiating combinations represent genetic combinations that drop out upon addition of both ceftriaxone and aTc. Neutral combinations showed neutral S scores in the presence of both ceftriaxone and aTc. Ceftriaxone was applied in two different concentrations: 64 µg/mL (low) and 256 µg/mL (high). Gene pairs correspond to numbers 1, qseB + bolA; 2, lrp + xylR; 3, eCFP + norR; and 4, pdhR + yohL.
Results
Combinatorial Library Generation and Functional Validation.
In this study, 173 E. coli transcription factors (TFs) and 18 fluorescent proteins were placed under the control of the anhydrotetracycline (aTc)-inducible PLtetO promoter (SI Appendix, Fig. S1, and Dataset 1). The TFs chosen represent nearly all E. coli TFs identified in RegulonDB (16). These expression constructs were uniquely barcoded and combined through CombiGEM to yield 35,343 possible pairwise combinations (189 ORFs × 187 ORFs, because some ORFs were rejected due to undesirable cleavage by restriction enzymes). This pairwise library was introduced into an E. coli MG1655 strain by electroporation and sequenced on the Illumina HiSeq platform, which resulted in the recovery of 34,554 combinations—or 98% of all possible combinations—distributed across the population (Fig. 1B). To ensure that inserts remained intact, we isolated and sequenced 50 library members and confirmed that all 50 retained both genetic expression constructs, including the genes, promoters, and ribosome-binding sequences.
To assess expression levels from our combinatorial genetic constructs, including the influence of gene order on expression levels, we measured fluorescence from all four pairwise combinations of GFP and mCherry as well as single GFP and mCherry constructs (SI Appendix, Fig. S2). We observed comparable fluorescence levels for both fluorescent proteins with the GFP–mCherry and mCherry–GFP constructs and approximately double fluorescence signal levels for GFP–GFP and mCherry–mCherry compared with the single gene constructs (SI Appendix, Fig. S3A). We also verified the ability of CombiGEM to scale beyond pairwise libraries through cloning of barcoded GFP and mCherry inserts to reach three-wise and four-wise combinatorial constructs (SI Appendix, Figs. S3 B–D and S4). Flow-cytometry analysis of higher-order combinations showed stepwise increases in fluorescence with each additional fluorescent gene and comparable fluorescence levels regardless of gene order when the number of fluorescence genes was held constant (SI Appendix, Fig. S3D).
CombiGEM Library Screening for Genetic Combinations That Modulate Antibiotic Killing.
We applied CombiGEM to discover TF combinations in our library that could modulate antibiotic activity against drug-resistant NDM-1 E. coli. We first determined the minimal inhibitory concentrations (MICs) for a range of antibiotics against an E. coli NDM-1 strain and found that this MIC profile resembled those of published NDM-1 clinical isolates (SI Appendix, Table S1). This NDM-1 strain was resistant to ceftriaxone, a third-generation cephalosporin with broad-spectrum activity that is used clinically to treat pneumonia, bacterial meningitis, and gonorrhea (17, 18). We then introduced the barcoded pairwise TF library into the E. coli NDM-1 strain by electroporation. We primarily sought to identify TF combinations that could enhance the activity of existing antibiotics to which E. coli NDM-1 was resistant. We subjected pooled NDM-1 strains containing the barcoded pairwise TF combinatorial libraries to a two-by-two matrix of conditions that included media with or without aTc induction and media with or without ceftriaxone treatment (Fig. 1C). We harvested bacterial cultures under each condition at early and late logarithmic-growth stages to observe shifting distributions in the barcoded populations over time. These samples were multiplexed for high-throughput sequencing on the Illumina HiSeq platform (19).
To determine the influence of each combination on NDM-1 viability, we computed statistical S scores to compare each combination’s abundance in a particular experimental condition against its abundance in other experiments (SI Appendix). Combinations that were underrepresented in the presence of aTc + ceftriaxone, compared with the other experimental conditions, suggested that expression of those gene combinations conferred potentiation of ceftriaxone. For validation, we randomly selected potentiating hits from the 200 top-scoring combinations, as well as controls with neutral S scores (Table 1). Hierarchical clustering on the combinations and their scores in each experiment showed clustering by phenotype class (Fig. 1D). To gauge the influence of gene order in our expression construct, we also performed hierarchical clustering on these combinations and their reciprocal configurations. The combinations of the same phenotype clustered together regardless of gene order in the constructs (SI Appendix, Fig. S5).
Table 1.
S and log-ratio scores for the genetic combinations studied
| Genetic combinations | S score | Log-ratio score |
| Potentiating genes (Fig. 2A) | ||
| qseB + bolA | −1.938 | −2.127 |
| lrp + xylR | −1.873 | 0.774 |
| Control genes (Fig. 2B) | ||
| eCFP + norR | −0.164 | −1.120 |
| pdhR + yohL | 0.607 | 1.170 |
| Lethal genes (Fig. 3A) | ||
| torR + metR | −0.523 | −4.336 |
| allR + metJ | −1.053 | −4.188 |
| nhaR + melR | −0.130 | −4.103 |
| malI + yfeT | −0.543 | −3.726 |
| Control genes (Fig. 3B) | ||
| rstA + oxyR | 0.756 | −0.409 |
| cadC + allR | −0.145 | −1.209 |
S scores shown are mean S scores for the aTc + ceftriaxone (256µg/mL) condition taken in early and late logarithmic phases. See SI Appendix for more details.
Validation of Hits via Time-Kill Curves.
To validate the activity of these combinations, we cloned each gene pair and tested their activity individually in time-kill assays against NDM-1 E. coli (Fig. 2). Each strain was grown to early logarithmic phase and treated with and without ceftriaxone and with and without aTc. Aliquots were serially diluted and plated to measure viable cell densities over 10 h after initiation of treatment. Combinations identified as leading to potentiation of killing showed significant amplification of killing by up to 3–5 orders of magnitude in the presence of aTc + ceftriaxone compared with ceftriaxone alone or aTc alone (Fig. 2A). The potentiating combinations also showed significantly greater killing compared with control combinations that were randomly selected for relatively neutral S scores, as well as the NDM-1 E. coli strain with no genetic combinations (Fig. 2B and SI Appendix, Fig. S6). In addition, the gene combinations exhibited significantly higher potentiation of antibiotic activity as pairwise combinations compared with individual genes, potentially indicating unexpected synergy (Fig. 2C). The potentiating combinations conferred a transient lethality, and the cultures did regrow over time. This regrowth was seen in all antibiotic-treated samples and could be due to antibiotic degradation, resistant subpopulations, or other mechanisms that are yet to be determined.
Fig. 2.
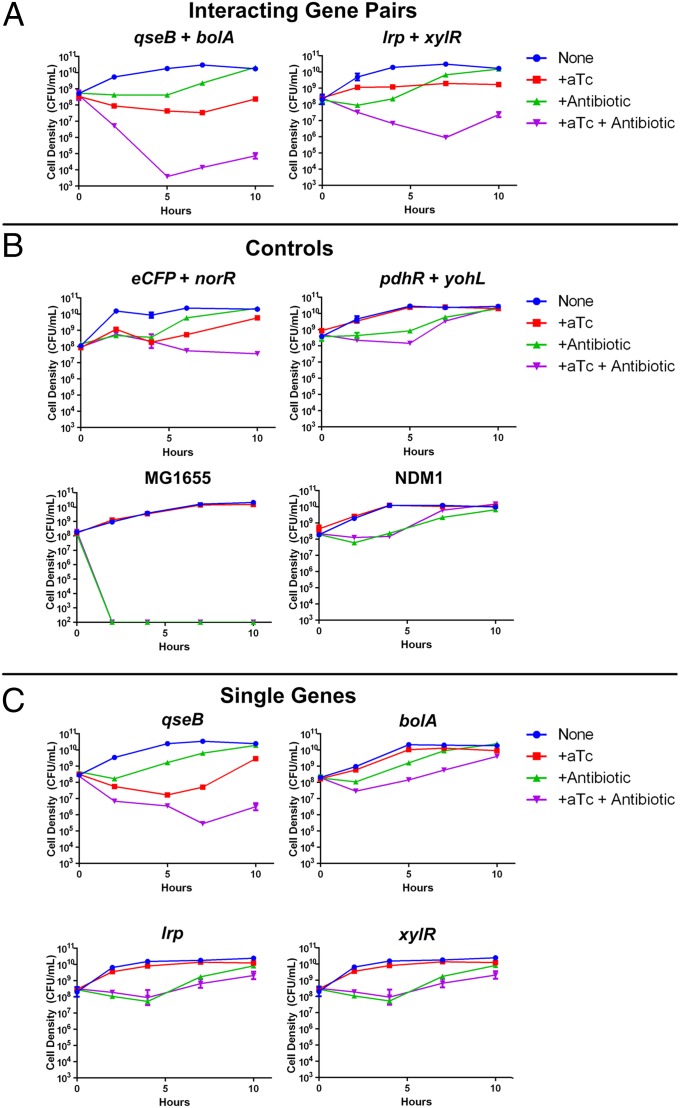
Antibiotic time-kill curves for NDM-1 E. coli with genetic combinations identified through CombiGEM. (A) Interacting gene pairs. Potentiating combinations identified through S-score analysis, qseB + bolA and lrp + xylR, exhibited enhancement of ceftriaxone activity upon induction with aTc, with up to 3–5 orders of magnitude greater killing compared with ceftriaxone alone or aTc alone. (B) Control genetic combinations with neutral S scores showed only minor potentiation of ceftriaxone. E. coli MG1655 cells with no genetic combinations (labeled MG1655) were effectively killed by ceftriaxone down to the limit of detection (100 CFU/mL). NDM-1 E. coli with no genetic combinations (labeled NDM-1) exhibited significantly less killing by ceftriaxone compared with E. coli MG1655. (C) Individual constituent genes of potentiating combinations revealed less effective potentiation of ceftriaxone than the corresponding combinations. In all experiments in this figure, ceftriaxone was administered at a concentration of 192 µg/mL and aTc at 100 ng/mL. Error bars represent SEM (small error bars are obscured by symbols).
To further study whether CombiGEM could identify genetic combinations across a range of antibiotics, we exposed the pairwise combinatorial library to the beta-lactam imipenem, the combination beta-lactam and -lactamase inhibitor piperacillin-tazobactam, and the aminoglycoside antibiotic gentamicin. Along with ceftriaxone, each antibiotic is used widely in clinical scenarios and is highly resisted by the NDM-1 strain (20–22). We performed hierarchical clustering of the top hits across all conditions based on S scores. Hits for the beta-lactam antibiotics in the presence of aTc clustered together—likely due to their similar mechanisms of action—and separately from gentamicin, likely due to their distinct mechanisms of action (SI Appendix, Fig. S7).
Identifying Lethal Genetic Combinations Through CombiGEM.
We next questioned whether CombiGEM could identify combinations of genes that were significantly lethal upon overexpression but that would also enhance bacterial killing by ceftriaxone. We identified the strongest lethal combinations by calculating the log-ratio of abundances between conditions with and without aTc, averaged across treatments with ceftriaxone and no ceftriaxone (SI Appendix). We selected random combinations from the top 200 hits with the lowest log ratios (Table 1). We discovered combinations that showed up to 6 orders of magnitude greater killing of the bacterial population when used in combination with ceftriaxone compared with ceftriaxone alone (Fig. 3A). Furthermore, in the absence of concomitant antibiotic treatment, hits identified from the log-ratio method showed greater lethality upon induction with aTc (Fig. 3A) compared with hits identified through the S-score method (Fig. 2A).
Fig. 3.
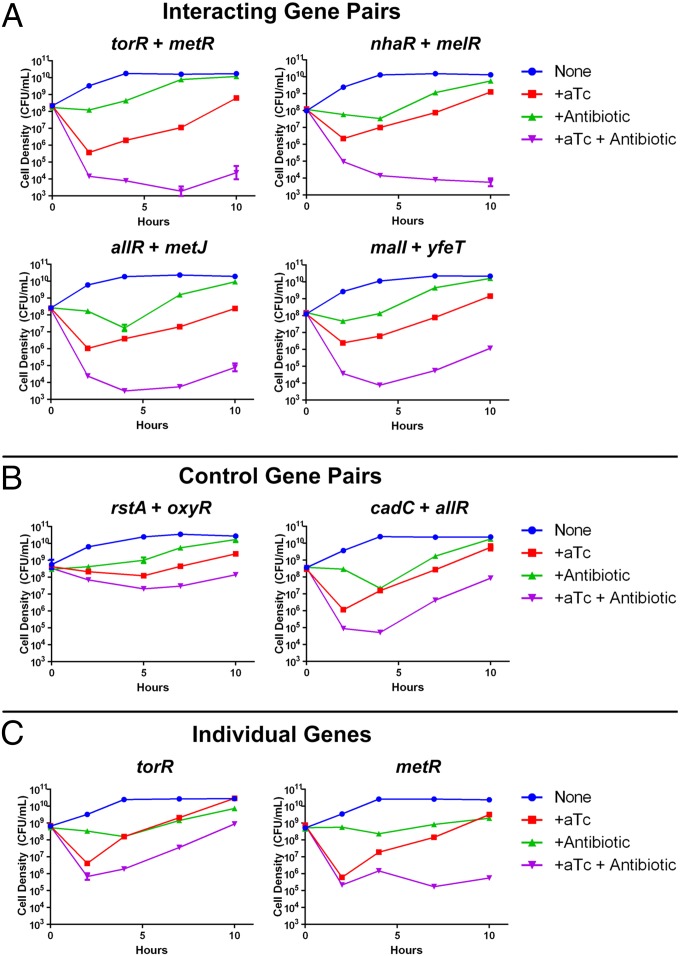
Lethal genetic combinations potentiate antibiotic activity against NDM-1 bacteria. (A) Interacting gene pairs. When used in combination with ceftriaxone, lethal genetic combinations identified via log-ratio analysis achieved reductions of viable cell counts of up to 6 orders of magnitude and suppressed growth for over 10 h in time-kill assays. (B) Control gene pairs. Random combinations with lethal genes, such as rstA + oxyR and cadC + allR, showed significantly less potentiation of antibiotic activity of ceftriaxone and less sustained growth suppression over time compared with lethal combinations identified through CombiGEM. (C) Individual genes. The single gene constituents of the torR + metR combination had weaker effects on bacterial killing compared with the torR + metR combination, when applied together with ceftriaxone. In all experiments in this figure, ceftriaxone was administered at 192 µg/mL and aTc at 100 ng/mL. Error bars represent SEM (small error bars are obscured by symbols).
Because a majority of E. coli ORFs have been reported to show growth inhibition upon induction (23), we constructed random combinations containing previously identified toxic genes to assess whether CombiGEM could identify especially lethal combinations (23). Combinations identified through the log-ratio method showed 1–3 orders of magnitude greater killing at the points of lowest cell density compared with random combinations of toxic genes, as well as longer sustained killing over the period of 10 h tested (Fig. 3 A and B). We also assessed whether the large killing effects of the genetic combinations were enhanced over that of the two constituent genes individually (Fig. 3C and SI Appendix, Fig. S8). Although torR and metR were toxic and increased antibiotic killing when expressed individually, the combination of the two genes resulted in greater lethal effects against NDM-1 bacteria (Fig. 3C). CombiGEM can therefore isolate combinatorial perturbations leading to strong phenotypes that are not ex ante apparent from the combination’s individual constituents.
Gene combinations identified through CombiGEM could be effective adjuvants for antibiotic treatment if delivered, for example, by phage. Previous studies have shown reductions in bacterial resistance (8, 24) and persistence (8) via phage delivery of gene expression constructs, but they relied on candidate gene approaches rather than high-throughput experimental screening as shown here. We packaged combinatorial genetic expression phagemids (identified in Fig. 3A) into M13 phage particles and infected an E. coli EMG2 NDM-1 strain (Fig. 4A). After 30 min of infection, aTc and ceftriaxone were added where appropriate, and viable cell counts were obtained on the bacterial cultures. The induction of gene combinations identified above in conjunction with ceftriaxone led to enhanced bacterial killing of 1–2 orders of magnitude compared with ceftriaxone alone, whereas a control phage showed no improvement (Fig. 4B). These results demonstrate that combinatorial genetic targets identified through this high-throughput approach can increase the killing of drug-resistant bacteria with antibiotic treatment.
Fig. 4.
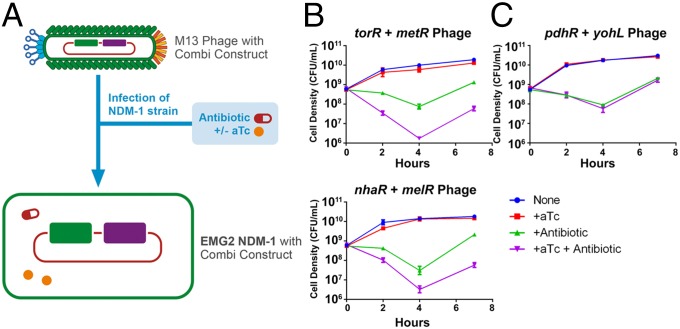
Phagemid-based delivery of genetic combinations enhance killing of NDM-1 E. coli with ceftriaxone. (A) NDM-1 E. coli were infected with M13 phagemids containing combination constructs and treated with antibiotic and aTc where appropriate, which is indicated as time t = 0 on the following graphs. (B) Phagemids containing torR + metR and nhaR + melR, when coadministered with ceftriaxone and induced with aTc, achieved amplified killing of NDM-1 E. coli bacteria. (C) In contrast to the results in B, phagemid-based delivery of a control genetic combination did not result in enhanced killing of NDM-1 E. coli. In all experiments in this figure, ceftriaxone was administered at 192 µg/mL and aTc at 100 ng/mL. Error bars represent SEM (small error bars are obscured by symbols).
Scalability of CombiGEM Assembly to Higher-Order Libraries.
To validate the scalability of the CombiGEM technology to higher-order combinations, we constructed three-wise combination libraries from the pairwise TF library with one-pot reactions. We first produced a lower-complexity library combining 189 × 187 × 18 genes. Out of 636,174 possible combinations, we recovered 582,433 unique combinations (92% of all possible combinations) from ∼30 million Illumina sequencing reads (SI Appendix, Fig. S9A). We also produced a higher-complexity library combining 189 × 187 × 187 genes. Out of 6,609,141 possible gene combinations, we recovered 4,042,316 combinations (61%) from ∼20 million sequencing reads (SI Appendix, Fig. S9B). Greater library coverage could likely be achieved by scaling up bacterial transformations of the libraries and dedicating a greater number of sequencing reads to each sample.
Discussion
In this study, we applied CombiGEM to identify TF combinations conferring strong phenotypes relevant to combating antibiotic resistance, including enhancement of bacterial killing by beta-lactam antibiotics against antibiotic-resistant E. coli NDM-1 strains. Most of the single genes and their combinations identified here have not been reported to directly mediate enhanced killing by antibiotics. For instance, qseB and bolA are implicated in pathways associated with virulence (25) and stress response (26), respectively, and show significantly higher potentiation of antibiotic activity as pairwise combinations compared with the individual genes. Thus, CombiGEM can reveal unexpected interactions between pathways yielding strong phenotypes and empower systems-biology approaches to uncovering network interactions. Future work should explore potential mechanisms of action of these genetic combinations on antibiotic efficacy, which could be due to reduction of antibiotic hydrolysis, influencing drug uptake or efflux, increasing the effectiveness of residual antibiotic, or other effects.
Genetic combinations identified through CombiGEM-based screens could potentially enhance the treatment of bacterial infections by revealing pathways that modulate antibiotic resistance and by providing targets for drug screening. The TFs identified in this study may not be optimal targets themselves for drug screening given the challenges associated with modulating TFs with small molecules (27). However, analysis of the downstream targets modulated by TF combinations using transcriptomic and proteomic approaches could help to identify candidate genes for further study. For example, the transcriptomic profiles of cells with specific genetic combinations could be compared with cells containing the corresponding constituent components as well as wild-type cells to identify genes that are uniquely up- or down-regulated. These genes could then be knocked out, knocked down, or overexpressed individually, and in combination with each other, to map their effects on antibiotic activity.
In future work, CombiGEM could also be used to barcode and screen combinatorial genetic knockdowns based on approaches such as antisense RNA (28) and CRISPR interference (29, 30). The potential advantage of screening for combinatorial knockdowns that enhance antibiotic efficacy is that such candidates could be more easily adapted into drug screening pipelines for small-molecule inhibitor screens. These drugs could then be delivered in combination treatment with antibiotics to improve antimicrobial activity. However, there are associated challenges with delivering combinatorial therapies to target sites at the same time that may need to be overcome (31).
In this work, we primarily studied the effects of beta-lactam antibiotics, but the study can be extended to a greater variety of antibiotics. Furthermore, the antibiotics can be dosed over a range of concentrations, and the samples can be assayed at multiple time points to provide greater insights into the interactions between genetic combinations and potential time-dependent and/or concentration-dependent effects. For example, beta-lactam antibiotics exhibit time-dependent killing, whereas quinolone antibiotics exhibit concentration-dependent killing (32). Moreover, the workflow described in this study identifies combinations generally leading to a drop in representation in the library population and does not explicitly distinguish between bacteriostatic vs. bactericidal activity. However, the protocol could be further adapted to identify combinations with such distinct behaviors, for example by quantifying barcode frequencies relative to standard combinations that have known phenotypes.
More generally, multifactorial genetics are integral to many complex biological phenomena. Discovery of the underlying combinatorial effectors has been limited by current methods but can be enabled by CombiGEM. Differentiation (33, 34) and reprogramming (35) protocols, such as those for inducing pluripotent stem cells (36), require specific combinations of factors that are typically discovered through nonmultiplexed and labor-intensive processes. Similarly, combinatorial drug therapy can achieve greater efficacy than individual drugs in disease management (37), but validating large numbers of drug-target combinations is difficult. Finally, methods such as genome-wide association studies have discovered numerous loci putatively involved in complex diseases, such as heart disease (38) and diabetes (39), but aggregated individual loci usually account for only a minor fraction of the disease heritability (40). Epistatic interactions between genes have been suggested to account for part of this “missing heritability,” (41, 42) but experimentally validating large numbers of such interactions is difficult. In particular, current methods for screening combinatorial genetic perturbations are limited in throughput and complexity. Single-gene overexpression and knockout libraries (23, 43) can be screened for cellular phenotypes but lack combinatorial complexity. Methods such as synthetic genetic arrays (44, 45) or double plasmid transformations enable pairwise gene combination studies but require time-intensive screening of individual clones. Pooled PCR stitching methods, such as Stitch-Seq (46), permit higher-throughput screening of pairwise genetic perturbations for interactome mapping. However, these methods are challenging to scale to three-wise and even higher-order genetic combinations.
In contrast to existing techniques, next-generation sequencing applied to the scalable CombiGEM assembly strategy enables multiplexed tracking of pooled combinatorial populations. By using this strategy, the one-time construction of the initial library containing n members proceeds in linear [or O(n)] time, scaling the library an order higher (e.g., from a pairwise n × n library to a three-wise n × n × n library, and so forth) proceeds in constant [or O(1)] time, and characterization of any of the combinatorial libraries proceeds in O(1) time. The CombiGEM strategy is flexible and can be used to build high-order combinations of other DNA elements, including gene-expression and knockdown (47, 48) constructs, synthetic circuit components, and genomic sequences, all of which are readily available in libraries. We have focused on antibiotic-resistant bacteria in this study, but this approach can also be generalized to study combinatorial genetics in other model organisms, including other bacterial pathogens, simple eukaryotes such as yeast, and human cells. For example, combinatorial genetic studies in Staphylococcus aureus and Pseudomonas aeruginosa could be performed by constructing libraries in E. coli on shuttle vectors, adapting expression constructs with validated promoters and ribosome-binding sequences from the target species, and performing bacterial transformation of these libraries into S. aureus and P. aeruginosa for testing. Factors to consider when extending CombiGEM to other organisms include the availability of tractable genetic tools, the efficiency of DNA delivery, the scale of cell culturing, and thus the possible library complexity that can be screened.
In future work, CombiGEM could also be extended to generate complex interactome datasets (49), screen for combinatorial regulators of desired phenotypes such as differentiation (36), identify potentiators of anticancer drugs (50), enhance pathway engineering (15), and characterize synthetic circuits at greater scales (51–55). The CombiGEM assembly strategy can be adapted to isothermal Gibson assembly (56) or recombinases (57) in situations where the library is incompatible with restriction sites. In summary, by leveraging one-pot combinatorial DNA assembly with highly multiplexed DNA sequencing of combinatorial barcodes, CombiGEM collapses high-dimensional combinatorial genetic studies into straightforward procedures. We envision that high-throughput strategies for probing multifactorial phenotypes and emergent network properties will contribute to our understanding of complex biological systems and their applications.
Methods
Gene expression constructs were created through isothermal Gibson assembly. Further experimental methods are available in SI Appendix. Details of data analysis are also found in SI Appendix.
Supplementary Material
Acknowledgments
This work was supported by the National Institutes of Health New Innovator Award (DP2 OD008435), the Office of Naval Research, the Ellison Foundation New Scholar in Aging Award, and the Henry L. and Grace Doherty Professorship in Ocean Utilization.
Footnotes
Conflict of interest statement: The authors have submitted a patent application for this technology.
This article is a PNAS Direct Submission. M.P.S. is a guest editor invited by the Editorial Board.
This article contains supporting information online at www.pnas.org/lookup/suppl/doi:10.1073/pnas.1400093111/-/DCSupplemental.
References
- 1.Centers for Disease Control and Prevention 2013. Antibiotic resistance threats in the US. Available at www.cdc.gov/drugresistance/threat-report-2013/. Accessed July 21, 2014.
- 2.Payne DJ, Gwynn MN, Holmes DJ, Pompliano DL. Drugs for bad bugs: Confronting the challenges of antibacterial discovery. Nat Rev Drug Discov. 2007;6(1):29–40. doi: 10.1038/nrd2201. [DOI] [PubMed] [Google Scholar]
- 3.Leverstein-Van Hall MA, et al. Global spread of New Delhi metallo-β-lactamase 1. Lancet Infect Dis. 2010;10(12):830–831. doi: 10.1016/S1473-3099(10)70277-2. [DOI] [PubMed] [Google Scholar]
- 4.Yigit H, et al. Novel carbapenem-hydrolyzing beta-lactamase, KPC-1, from a carbapenem-resistant strain of Klebsiella pneumoniae. Antimicrob Agents Chemother. 2001;45(4):1151–1161. doi: 10.1128/AAC.45.4.1151-1161.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Mochon AB, et al. New Delhi metallo-β-lactamase (NDM-1)-producing Klebsiella pneumoniae: Case report and laboratory detection strategies. J Clin Microbiol. 2011;49(4):1667–1670. doi: 10.1128/JCM.00183-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Pena-Miller R, et al. When the most potent combination of antibiotics selects for the greatest bacterial load: The smile-frown transition. PLoS Biol. 2013;11(4):e1001540. doi: 10.1371/journal.pbio.1001540. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Beardmore RE, Peña-Miller R. Rotating antibiotics selects optimally against antibiotic resistance, in theory. Math Biosci Eng. 2010;7(3):527–552. doi: 10.3934/mbe.2010.7.527. [DOI] [PubMed] [Google Scholar]
- 8.Lu TK, Collins JJ. Engineered bacteriophage targeting gene networks as adjuvants for antibiotic therapy. Proc Natl Acad Sci USA. 2009;106(12):4629–4634. doi: 10.1073/pnas.0800442106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Miller C, et al. SOS response induction by beta-lactams and bacterial defense against antibiotic lethality. Science. 2004;305(5690):1629–1631. doi: 10.1126/science.1101630. [DOI] [PubMed] [Google Scholar]
- 10.Lewin CS, Howard BM, Ratcliffe NT, Smith JT. 4-quinolones and the SOS response. J Med Microbiol. 1989;29(2):139–144. doi: 10.1099/00222615-29-2-139. [DOI] [PubMed] [Google Scholar]
- 11.Little JW, Harper JE. Identification of the lexA gene product of Escherichia coli K-12. Proc Natl Acad Sci USA. 1979;76(12):6147–6151. doi: 10.1073/pnas.76.12.6147. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Cirz RT, et al. Inhibition of mutation and combating the evolution of antibiotic resistance. PLoS Biol. 2005;3(6):e176. doi: 10.1371/journal.pbio.0030176. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Kohanski MA, Dwyer DJ, Hayete B, Lawrence CA, Collins JJ. A common mechanism of cellular death induced by bactericidal antibiotics. Cell. 2007;130(5):797–810. doi: 10.1016/j.cell.2007.06.049. [DOI] [PubMed] [Google Scholar]
- 14.Dwyer DJ, Kohanski MA, Hayete B, Collins JJ. Gyrase inhibitors induce an oxidative damage cellular death pathway in Escherichia coli. Mol Syst Biol. 2007;3:91. doi: 10.1038/msb4100135. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Zelcbuch L, et al. Spanning high-dimensional expression space using ribosome-binding site combinatorics. Nucleic Acids Res. 2013;41(9):e98. doi: 10.1093/nar/gkt151. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Gama-Castro S, et al. RegulonDB version 7.0: Transcriptional regulation of Escherichia coli K-12 integrated within genetic sensory response units (Gensor Units) Nucleic Acids Res. 2011;39(Database issue):D98–D105. doi: 10.1093/nar/gkq1110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Richards DM, Heel RC, Brogden RN, Speight TM, Avery GS. Ceftriaxone. A review of its antibacterial activity, pharmacological properties and therapeutic use. Drugs. 1984;27(6):469–527. doi: 10.2165/00003495-198427060-00001. [DOI] [PubMed] [Google Scholar]
- 18.Norrby SR, Petermann W, Willcox PA, Vetter N, Salewski E. A comparative study of levofloxacin and ceftriaxone in the treatment of hospitalized patients with pneumonia. Scand J Infect Dis. 1998;30(4):397–404. doi: 10.1080/00365549850160710. [DOI] [PubMed] [Google Scholar]
- 19.Meyer M, Kircher M. 2010. Illumina sequencing library preparation for highly multiplexed target capture and sequencing. Cold Spring Harbor protocols 2010(6):pdb prot5448. [DOI] [PubMed]
- 20.Papp-Wallace KM, Endimiani A, Taracila MA, Bonomo RA. Carbapenems: Past, present, and future. Antimicrob Agents Chemother. 2011;55(11):4943–4960. doi: 10.1128/AAC.00296-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Karlowsky JA, Zelenitsky SA, Zhanel GG. Aminoglycoside adaptive resistance. Pharmacotherapy. 1997;17(3):549–555. [PubMed] [Google Scholar]
- 22.Charbonneau P. Review of piperacillin/tazobactam in the treatment of bacteremic infections and summary of clinical efficacy. Intensive Care Med. 1994;20(Suppl 3):S43–S48. doi: 10.1007/BF01745251. [DOI] [PubMed] [Google Scholar]
- 23.Kitagawa M, et al. Complete set of ORF clones of Escherichia coli ASKA library (a complete set of E. coli K-12 ORF archive): Unique resources for biological research. DNA Res. 2005;12(5):291–299. doi: 10.1093/dnares/dsi012. [DOI] [PubMed] [Google Scholar]
- 24.Edgar R, Friedman N, Molshanski-Mor S, Qimron U. Reversing bacterial resistance to antibiotics by phage-mediated delivery of dominant sensitive genes. Appl Environ Microbiol. 2012;78(3):744–751. doi: 10.1128/AEM.05741-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Kostakioti M, Hadjifrangiskou M, Pinkner JS, Hultgren SJ. QseC-mediated dephosphorylation of QseB is required for expression of genes associated with virulence in uropathogenic Escherichia coli. Mol Microbiol. 2009;73(6):1020–1031. doi: 10.1111/j.1365-2958.2009.06826.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Santos JM, Freire P, Vicente M, Arraiano CM. The stationary-phase morphogene bolA from Escherichia coli is induced by stress during early stages of growth. Mol Microbiol. 1999;32(4):789–798. doi: 10.1046/j.1365-2958.1999.01397.x. [DOI] [PubMed] [Google Scholar]
- 27.Berg T. Inhibition of transcription factors with small organic molecules. Curr Opin Chem Biol. 2008;12(4):464–471. doi: 10.1016/j.cbpa.2008.07.023. [DOI] [PubMed] [Google Scholar]
- 28.Forsyth RA, et al. A genome-wide strategy for the identification of essential genes in Staphylococcus aureus. Mol Microbiol. 2002;43(6):1387–1400. doi: 10.1046/j.1365-2958.2002.02832.x. [DOI] [PubMed] [Google Scholar]
- 29.Larson MH, et al. CRISPR interference (CRISPRi) for sequence-specific control of gene expression. Nat Protoc. 2013;8(11):2180–2196. doi: 10.1038/nprot.2013.132. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Farzadfard F, Perli SD, Lu TK. Tunable and multi-functional eukaryotic transcription factors based on CRISPR/Cas. ACS Synth Biol. 2013;2(10):604–613. doi: 10.1021/sb400081r. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Mainz D, Borner K, Koeppe P, Kotwas J, Lode H. Pharmacokinetics of lansoprazole, amoxicillin and clarithromycin after simultaneous and single administration. J Antimicrob Chemother. 2002;50(5):699–706. doi: 10.1093/jac/dkf172. [DOI] [PubMed] [Google Scholar]
- 32.Vogelman B, Craig WA. Kinetics of antimicrobial activity. J Pediatr. 1986;108(5 Pt 2):835–840. doi: 10.1016/s0022-3476(86)80754-5. [DOI] [PubMed] [Google Scholar]
- 33.Oldershaw RA, et al. Directed differentiation of human embryonic stem cells toward chondrocytes. Nat Biotechnol. 2010;28(11):1187–1194. doi: 10.1038/nbt.1683. [DOI] [PubMed] [Google Scholar]
- 34.Perrier AL, et al. Derivation of midbrain dopamine neurons from human embryonic stem cells. Proc Natl Acad Sci USA. 2004;101(34):12543–12548. doi: 10.1073/pnas.0404700101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Zhou Q, Brown J, Kanarek A, Rajagopal J, Melton DA. In vivo reprogramming of adult pancreatic exocrine cells to beta-cells. Nature. 2008;455(7213):627–632. doi: 10.1038/nature07314. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Takahashi K, Yamanaka S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell. 2006;126(4):663–676. doi: 10.1016/j.cell.2006.07.024. [DOI] [PubMed] [Google Scholar]
- 37.Al-Lazikani B, Banerji U, Workman P. Combinatorial drug therapy for cancer in the post-genomic era. Nat Biotechnol. 2012;30(7):679–692. doi: 10.1038/nbt.2284. [DOI] [PubMed] [Google Scholar]
- 38.Willer CJ, et al. Newly identified loci that influence lipid concentrations and risk of coronary artery disease. Nat Genet. 2008;40(2):161–169. doi: 10.1038/ng.76. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Frayling TM. Genome-wide association studies provide new insights into type 2 diabetes aetiology. Nat Rev Genet. 2007;8(9):657–662. doi: 10.1038/nrg2178. [DOI] [PubMed] [Google Scholar]
- 40.Zuk O, Hechter E, Sunyaev SR, Lander ES. The mystery of missing heritability: Genetic interactions create phantom heritability. Proc Natl Acad Sci USA. 2012;109(4):1193–1198. doi: 10.1073/pnas.1119675109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Manolio TA, et al. Finding the missing heritability of complex diseases. Nature. 2009;461(7265):747–753. doi: 10.1038/nature08494. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Eichler EE, et al. Missing heritability and strategies for finding the underlying causes of complex disease. Nat Rev Genet. 2010;11(6):446–450. doi: 10.1038/nrg2809. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Soo VW, Hanson-Manful P, Patrick WM. Artificial gene amplification reveals an abundance of promiscuous resistance determinants in Escherichia coli. Proc Natl Acad Sci USA. 2011;108(4):1484–1489. doi: 10.1073/pnas.1012108108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Butland G, et al. eSGA: E. coli synthetic genetic array analysis. Nat Methods. 2008;5(9):789–795. doi: 10.1038/nmeth.1239. [DOI] [PubMed] [Google Scholar]
- 45.Tong AH, et al. Global mapping of the yeast genetic interaction network. Science. 2004;303(5659):808–813. doi: 10.1126/science.1091317. [DOI] [PubMed] [Google Scholar]
- 46.Yu H, et al. Next-generation sequencing to generate interactome datasets. Nat Methods. 2011;8(6):478–480. doi: 10.1038/nmeth.1597. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Root DE, Hacohen N, Hahn WC, Lander ES, Sabatini DM. Genome-scale loss-of-function screening with a lentiviral RNAi library. Nat Methods. 2006;3(9):715–719. doi: 10.1038/nmeth924. [DOI] [PubMed] [Google Scholar]
- 48.Chang K, Elledge SJ, Hannon GJ. Lessons from Nature: MicroRNA-based shRNA libraries. Nat Methods. 2006;3(9):707–714. doi: 10.1038/nmeth923. [DOI] [PubMed] [Google Scholar]
- 49.Yu H, et al. High-quality binary protein interaction map of the yeast interactome network. Science. 2008;322(5898):104–110. doi: 10.1126/science.1158684. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Chan DA, Giaccia AJ. Harnessing synthetic lethal interactions in anticancer drug discovery. Nat Rev Drug Discov. 2011;10(5):351–364. doi: 10.1038/nrd3374. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Sharon E, et al. Inferring gene regulatory logic from high-throughput measurements of thousands of systematically designed promoters. Nat Biotechnol. 2012;30(6):521–530. doi: 10.1038/nbt.2205. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Ye H, et al. Pharmaceutically controlled designer circuit for the treatment of the metabolic syndrome. Proc Natl Acad Sci USA. 2013;110(1):141–146. doi: 10.1073/pnas.1216801110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Davidson EA, Ellington AD. Synthetic RNA circuits. Nat Chem Biol. 2007;3(1):23–28. doi: 10.1038/nchembio846. [DOI] [PubMed] [Google Scholar]
- 54.Wu M, et al. Engineering of regulated stochastic cell fate determination. Proc Natl Acad Sci USA. 2013;110(26):10610–10615. doi: 10.1073/pnas.1305423110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Deans TL, Cantor CR, Collins JJ. A tunable genetic switch based on RNAi and repressor proteins for regulating gene expression in mammalian cells. Cell. 2007;130(2):363–372. doi: 10.1016/j.cell.2007.05.045. [DOI] [PubMed] [Google Scholar]
- 56.Gibson DG, et al. Enzymatic assembly of DNA molecules up to several hundred kilobases. Nat Methods. 2009;6(5):343–345. doi: 10.1038/nmeth.1318. [DOI] [PubMed] [Google Scholar]
- 57.Hartley JL, Temple GF, Brasch MA. DNA cloning using in vitro site-specific recombination. Genome Res. 2000;10(11):1788–1795. doi: 10.1101/gr.143000. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.