Abstract
Alternative polyadenylation (APA) is an evolutionarily conserved mechanism for regulating gene expression. Transcript 3′ end shortening through changes in polyadenylation site usage occurs following T cell activation, but the consequences of APA on gene expression are poorly understood. We previously showed that GU-rich elements (GREs) found in the 3′ untranslated regions of select transcripts mediate rapid mRNA decay by recruiting the protein CELF1/CUGBP1. Using a global RNA sequencing approach, we found that a network of CELF1 target transcripts involved in cell division underwent preferential 3′ end shortening via APA following T cell activation, resulting in decreased inclusion of CELF1 binding sites and increased transcript expression. We present a model whereby CELF1 regulates APA site selection following T cell activation through reversible binding to nearby GRE sequences. These findings provide insight into the role of APA in controlling cellular proliferation during biological processes such as development, oncogenesis and T cell activation
Keywords: Alternative Polyadenylation, CELF1, CUGBP1, Cell Division, mRNA Decay, T Cell Stimulation, GRE
1. Introduction
T cell activation results in a dramatic proliferative response that is coordinated by a complex regulatory network. This cellular response relies on a rapid and accurate change in the T cell transcriptome occurring within hours of T cell receptor-mediated stimulation [1]. Global quantification of mRNA decay rates in resting and stimulated T cells revealed that these gene expression changes are accomplished partly through alterations in the half-life of transcripts [2]. In addition to alterations of the level of transcript expression following T cell stimulation, the sequence content of many transcripts is altered through the mechanisms of alternative splicing [3] and alternative polyadenylation (APA) [4]. APA is an evolutionarily conserved mechanism for regulating the 3′ end lengths of mRNA through utilization of different polyA sites during mRNA maturation [5,6]. Changes in polyA site usage are associated with increased cellular proliferation in development [7], malignancy [4,8] and T cell activation [4]. During T cell activation, APA has been shown to lead to shortened 3′UTRs containing fewer regulatory sequences, including miRNA binding sites [4]. The biochemical mechanisms regulating APA and the relationship between APA and cellular proliferation remain poorly understood
APA is a post-transcriptional regulatory process involving a host of proteins whose regulation and function are incompletely understood. The nuclear maturation of nearly all eukaryotic mRNA involves 3′ endonucleolytic cleavage followed by the untemplated addition of a string of adenines to the 3′ end of the transcript to form a polyA tail [9]. In most transcripts, the polyA tail can be added at one of multiple potential polyadenylation signal sequences in the 3′UTR of the transcript, and the relative usage of different polyA sites is influenced by surrounding sequence elements and the levels of APA related factors [9]. Many polyA sites have a downstream G- and U-rich sequence that is recognized by the protein Cleavage stimulation factor 64kDa (CSTF-64) or it’s tau isoform, CSTF-64τ [10,11]. Up-regulation of CSTF-64 and its binding to the downstream G- and U-rich APA related sequence element may promote the usage of adjacent APA sites through recruitment of additional components of the APA machinery [12–14].
T cell activation results in dramatic changes in transcription and mRNA decay rates of networks of activation gene transcripts within hours of T cell activation [1]. We have previously shown in primary human T cells that the RNA-binding protein CELF1 (also known as CUGBP1) binds to a GU-rich element (GRE) with the sequence UGU[G/U]UGU[G/U]UGU within the 3′UTRs of a network of transcripts involved in cellular proliferation, apoptosis, and post-transcriptional gene regulation [15,16]. Upon binding to the GRE, CELF1 recruits the RNA degradation machinery resulting in the decay of the GRE-harboring transcript. Investigations into the binding targets of CELF1 in resting and stimulated primary human T cells found that the population of CELF1 bound transcripts decreased upon T cell stimulation, and that this was due in part to the activation dependent phosphorylation of CELF1 and a resultant decrease in the affinity of CELF1 for GRE binding sites. [17].
In the present study, we utilized 3′-end RNA sequencing technology to quantify changes in APA after T cell activation. We found that many 3′UTRs shorten via APA within six hours following T cell activation, and that APA preferentially leads to 3′ end shortening of CELF1 target transcripts that encode regulators of cellular division. Among CELF1 target transcripts that undergo 3′ end shortening through APA following T cell activation, we found enrichment of GRE sequences 30–40 nucleotides downstream of 5′ APA sites, suggesting that binding by CELF1 to these sites may regulate transcript shortening. We propose a model whereby binding by CELF1 to GRE sequences immediately downstream of 5′ APA sites prevents utilization of these sites in resting T cells, and therefore downstream APA sites are used. The resulting transcripts contain GRE sequences in their 3′ UTRs and are susceptible to GRE-mediated mRNA decay. Subsequent phosphorylation of CELF1 following T cell activation prevents CELF1 binding to GRE sequences downstream of 5′ APA sites, allowing the 5′ APA sites to be utilized. The shortened transcripts, which encode regulators of cell division, are up-regulated because they no longer contain GRE sequences and are not susceptible to GRE-mediated mRNA decay. This model explains the coordinated regulation of a network of CELF1 target transcripts that regulate cell division following T cell activation. Our findings provide a mechanistic link between APA and cellular proliferation and have implications for immunology, development and cancer biology.
2. Materials and Methods
2.1 Purification of Human T cells
Primary human T cells were purified from peripheral blood mononuclear cells by negative selection using the CD3+ Rosette-Sep antibody cocktail from Stem Cell Technologies as previously described [18]. CD3+ lymphocytes were then isolated through a Ficoll-hypaque cushion (GE Healthcare).
2.2 T cell Stimulation and RNA purification
Purified human T cells from three separate donors were cultured overnight in RPMI 1640 supplemented with 10% fetal bovine serum, 2mM L-glutamine, 100U/mL penicillin G and 100ug/mL streptomycin. Cells were then incubated for the specified length of stimulation in 15cm dishes (5×107 cells/dish) with medium alone or with a combination of immobilized monoclonal antibodies (1ug/mL) directed against the CD3 component of the TCR complex (R&D Systems) and the CD28 co-stimulatory molecule (R&D Systems) as previously described [18]. Following stimulation, T cells were removed from stimulation, washed one time in PBS, and subsequently pelleted by centrifugation. The cells were then lysed using the RNEasy RNA isolation kit from Qiagen, following the manufacturer’s recommandations.
2.3 Direct Sequencing of RNA 3′ Ends
Direct RNA sequencing of polyA tails for the determination of polyA sites was performed by Helicos Biosciences as previously described [5]. Briefly, polyA+ RNA was isolated from total cellular RNA using oligo dT coated beads. PolyA+ RNA molecules were then cleaved, and these cleaved fragments were passed through a flow cell containing an oligo-dT coated surface to capture the polyA+ fragments. Following a “fill and lock” step, sequencing is performed using a sequencing-by-synthesis procedure with fluorescently labeled nucleotides.
2.4 Next generation sequencing of T cell RNA
mRNA sequencing libraries were prepared from 1ug of total RNA from using the Truseq RNA library preparation kit from Illumina, following the manufacturers instructions. 100bp single-end reads were acquired using the Illumina HiSeq 2000 sequencing machine. Reads were mapped to the human genome (hg19) using the Bowtie 2.0 algorithm using the default “sensitive” mapping settings, and reporting only uniquely mapping reads [19].
2.5 Determination of Polyadenylation Sites and Sequence Analysis
Sequence reads were aligned to the human genome (hg19) using the SHRiMP2 algorithm with default settings. A list of the coordinates of all human 3′UTR exons was downloaded from the UCSC genome browser, and a pileup view of the reads was generated for each 3′UTR exon using samtools [20–22]. Subsequently, custom C++ algorithms were written to search for sites supported by at least 10 reads. PolyA site locations were then used to determine the genome coordinates of the upstream and downstream 3′UTR regions. Genomic sequences corresponding to these coordinates were extracted using the Galaxy web-based bioinformatics package [23–25].
3. Results
3.1 CELF1 target transcripts are preferentially shortened via APA following T cell activation
We investigated the changes in polyA site usage at six and 24 hours following activation of primary human T cells by performing genome-wide direct sequencing of mRNA 3′ends [5,26]. This technology allowed us to identify the precise location of polyA sites on a genome wide scale and quantitatively assess the relative usage of various polyA sites among alternatively polyadenylated transcripts [5]. We performed this sequencing in resting T cells or T cells that were stimulated with anti-CD3 and anti-CD28 antibodies for six or 24 hours. Sequence reads were mapped to the human genome (GRCh37/HG19) using the SHRiMP 2 algorithm, allowing for up to two mismatches [27]. PolyA sites were included for analysis if they were supported by at least 10 reads and occurred within annotated 3′UTR segments. The number of polyA sites per expressed transcript at each time point was calculated, and we found an average of 1.8 polyA sites per transcript expressed in resting T cells.
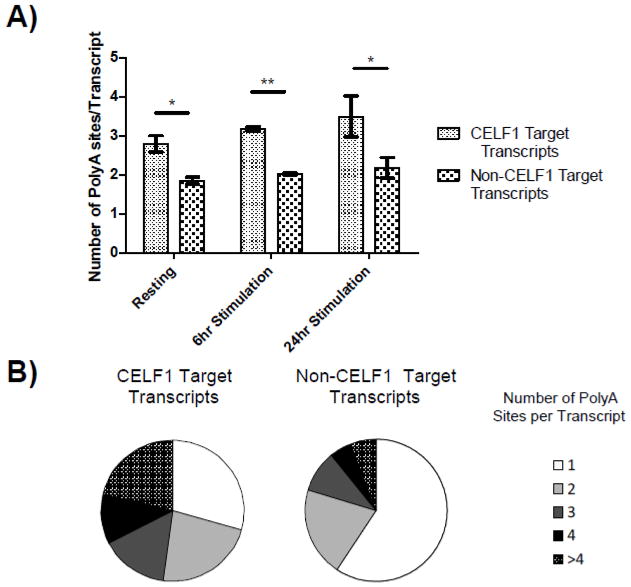
We previously identified CELF1 target transcripts by non-crosslinking RNA-IP followed by microarray analysis in primary human T cells in either the resting state or following six hours of in vitro stimulation with anti-CD3/28 antibodies [17]. In analysis of our polyA site data, we noticed that many GRE-containing CELF1 target transcripts that we previously identified by RNA-IP [17] contained multiple polyA sites. We therefore performed an analysis to determine if polyA sites were enriched in CELF1 target transcripts (Figure 1). CELF1 target transcripts in resting T cells (1280 transcripts) had an average of 2.8 polyA sites per transcript, which was significantly enriched compared to 1.8 polyA sites per transcript in non-CELF1 target transcripts (19,900_transcripts) (p-value 0.02, paired T-test). In T cells that were activated for six or 24 hours, we found that CELF1 target transcripts had an average of 3.2 and 3.5 polyA sites per transcript, compared to 1.9 and 2.1 polyA sites per transcript in non-CELF1 target transcripts, respectively (p-value 0.001 and 0.04 at six and 24 hours, respectively, comparing CELF1 target transcripts to non-CELF1 transcripts, paired T-test). To further characterize the enrichment of polyA sites in CELF1 target transcripts, we graphed the distribution of polyA sites per transcript for CELF1 target transcripts or non-CELF1 target transcripts expressed in resting T cells (Figure 1). CELF1 target transcripts had fewer transcripts with only one polyA site (29% versus 61%) and more transcripts with multiple polyA sites (71% vs 39%). This enrichment of transcripts with two or more polyA sites within CELF1 target transcripts compared to non-CELF1 transcripts was highly statistically significant for each individual donor (p<2.2×10−16, Fisher’s Exact Test, R) as well as the average across donors (p=0.002, Welch Two Sample T-Test, R).
Fig. 1.
CELF1 target transcripts are enriched for polyA sites. A) The average number of experimentally identified polyA sites per transcript was determined for expressed non-CELF1 target transcripts or CELF1 target transcripts in resting T cells and T cells that were stimulated for six or 24 hours. (*= p-value <0.05, ** = p-value <0.01). B) The number of polyA sites per transcript was determined for CELF1 target transcripts and all expressed non-CELF1 target transcripts in resting T cells.
Given the enrichment of multiple polyA sites in CELF1 target transcripts, we hypothesized that CELF1 target transcripts were more likely to shorten through APA following T cell activation compared to non-CELF1 target transcripts. For each T cell donor at each time point, we calculated the average length of each 3′UTR for each transcript, with each length weighted by the number of reads supporting usage of each polyA site (table S1). We then performed a linear regression analysis on the length of each 3′UTR over time and identified 2352 3′UTRs whose length changed over time following T cell activation (regression coefficients different from zero with p<0.05, linear model, R). We found 203 of these transcripts were CELF1 targets and 2149 were non-CELF1 targets. Of the CELF1 targets, 171 (84%) shortened, whereas only 1345 (63%) of the non-CELF1 targets shortened, suggesting that CELF1 targets were highly enriched for 3′ UTRs that shortened after T cell activation (p-value 1×10−10 comparing the enrichment of shortened transcripts among CELF1 targets vs non-CELF1 targets, Fisher’s exact test, R). Thus, CELF1 target transcripts were significantly more likely tobe shortened through APA following T cell activation compared to non-CELF1 targets.
3.2 GREs are enriched downstream of APA sites
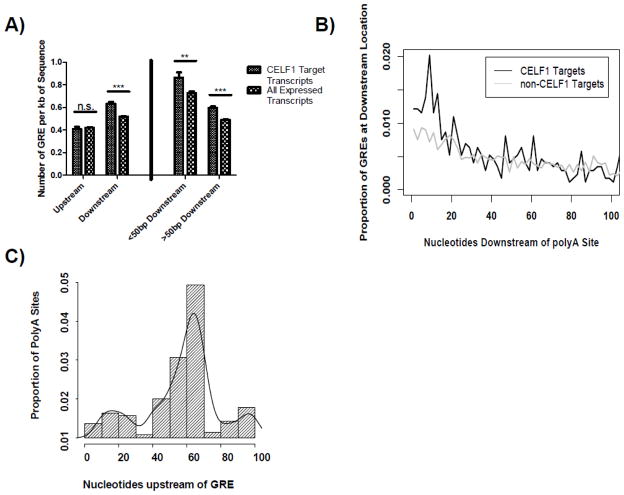
Given that CELF1 target transcripts were enriched for polyA sites and were preferentially shortened by APA following T cell activation, we investigated whether the GRE was enriched downstream of 5′ APA sites in CELF1 target transcripts. We compiled all unique polyA sites from all donors and time points into one master T cell polyA database (table S2). We next used this database to divide each 3′UTR into sequences between the termination codon and the 5′ APA site (upstream region) and from this polyA site to the subsequent polyA site (downstream region). The prevalence of the GRE (defined as the sequence UGU[G/U]UGU[G/U]UGU allowing for one mismatch) was determined for these two regions for CELF1 target transcripts and all transcripts expressed in T cells. As can be seen in Figure 2A, the number of GREs per kb of sequence was similar between CELF1 target transcripts and all expressed non-CELF1 target transcripts in the upstream region (p-value 0.5, Exact Poisson test, R), suggesting that CELF1 targets were not enriched for GRE sequences in this region. In the downstream regions, however, we observed a significant enrichment in the presence of GREs in CELF1 target transcripts compared to non-CELF1 target transcripts (p<0.001, Exact Poisson test, R).
Fig. 2.
GREs are enriched downstream of polyA sites. A) We created a master list of all unique polyA sites from all donors and time points. Using this list we separated 3′UTR sequences into the region from termination codon to the first detected polyA site (upstream) and the region from each polyA site to the subsequent polyA site (downstream). We then determined the number of GREs per kilobase of sequence within these two regions for CELF1 target transcripts and all expressed non-CELF1 target transcripts. We then segregated the downstream regions into sequences less than 50bp or greater than 50bp downstream of detected polyA sites (<50 Downstream and >50 Downstream, respectively). We then calculated the number of GREs per kilobase of sequence within these two regions for CELF1 target transcripts and all expressed non-CELF1 target transcripts (n.s. = not significant, *** = p-value <0.001). B) We determined the distribution of GREs within the first 100nt downstream of polyA sites for CELF1 target transcripts and non-CELF1 targets. C) The 100bp of sequence upstream of each GRE was extracted, and these regions were examined for polyA signals of the sequence A[A/U]UAAA. A histogram of locations of polyA signals is shown. The x-axis represents the number of nucleotides upstream of the GRE where polyA signals were found, and the y-axis represents the proportion of detected polyA signals located at each position.
In the process of polyadenylation, protein CSTF-64 binds to U/GU-rich sequences within the first 50bp downstream of polyA sites and plays a role in polyA site recognition [10]. In order to determine whether the enrichment of GREs in the downstream regions of CELF1 target transcripts occurs within the region immediately downstream of polyA sites, where CSTF-64 binds, we divided the downstream regions into regions less than 50nt downstream and greater than 50nt downstream of the APA site and found enrichment of the GRE within both of these regions (Figure 2A). We further investigated the distribution of the GRE over the 100nt downstream of polyA sites in CELF1 target transcripts and non-CELF1 targets (Figure 2B). We found that the enrichment of the GRE in CELF1 target transcripts occurred largely within the first 20nt downstream of polyA sites. This analysis showed that the CELF1 binding motif occurred preferentially in regions downstream of polyA sites in CELF1 target transcripts, and in particular, we observed a significant enrichment of the GRE in the 20nt immediately downstream of APA sites in CELF1 target transcripts, suggesting a possible functional overlap between the U/GU rich CSTF-64 target sequence and the GRE.
Given the relationship between GREs and 5′ APA sites in CELF1 target transcripts, we were interested to determine whether GREs occurring distal to detected polyA sites might be associated with other potential polyA sites. To address this question we extracted GREs found at least 100nt downstream of polyA sites in the CELF1 target transcripts in the downstream regions, and extracted the 100nt proximal to those GREs. We then searched these sequences for polyA signals of the sequence A[A/U]UAAA and plotted the distribution of the locations of detected polyA signals in these sequences (Figure 2C). We found that the distribution of polyA signals upstream of GREs in these regions was non-uniform (p<0.001 of uniform distribution by chi-squared test, R) with a peak occurrence of polyA sites approximately 64nt upstream of the GRE. Combined, these data revealed a distinct spatial distribution between the GRE and upstream polyA signals, further suggesting a functional relationship between the GRE and polyA sites.
3.3 APA regulates a network of CELF1 target transcripts that encode regulators of cell division
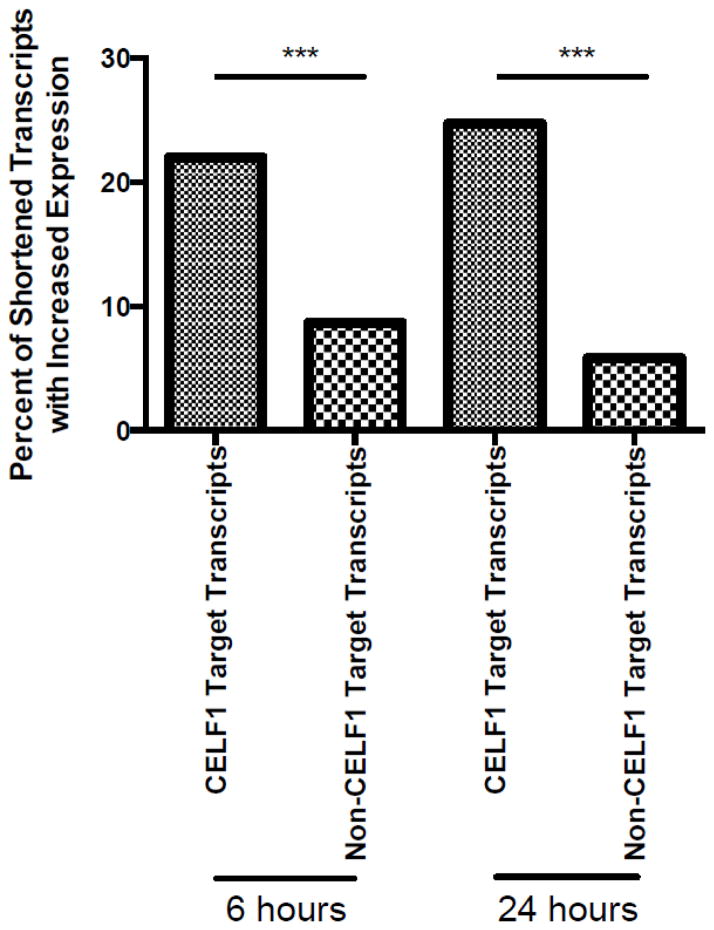
We hypothesized that CELF1 target transcripts whose 3′UTRs were shortened by APA would exhibit an increase in expression following T cell activation due to removal of downstream GREs and resultant decrease in CELF1-mediated mRNA decay. We utilized next generation sequencing to investigate the expression pattern of all T cell transcripts over the first 24 hours of T cell simulation. RNA was isolated from resting T cells or T cells that were stimulated with anti-CD3 and anti-CD28 antibodies for six or 24 hours. Reads were mapped to the human genome (hg19) using Bowtie 2.0 [19], and RPMK expression values were calculated (table S3). We next analyzed the expression patterns of transcripts that shortened over time following T cell activation, which included 171 CELF1 target transcripts (corresponding to 255 unique refseqs) and 1322 non-CELF1 transcripts (corresponding to 1546 refseqs). We found that of these 255 shortened CELF1 target refseqs, 56 (22.0%) were up-regulated six hours following T cell stimulation, and 63 (24.7%) were up-regulated after 24 hours (slope significantly positive with p-value < 0.05, linear model, R). These percentages represented a significant enrichment compared to the non-CELF1 transcripts, of which 135 (8.7%) were up-regulated six hours following T cell stimulation (p-value 8.2×10−9 compared to CELF1 target transcripts, Fisher’sExact Test, R) and 90 (5.8%) were up-regulated after 24 hours (p-value <2.2×10−16 compared to CELF1 target transcripts, Fisher’s Exact Test, R). Thus, CELF1 target transcripts that shortened by alternative polyadenylation during T cell stimulation were significantly more likely to increase their expression compared to non-CELF1 targets (Figure 3). The median fold change in expression compared to resting cells of these up-regulated, shortened CELF1 target transcripts was 1.60 at six hours, and 1.59 at 24 hours. CELF1 target transcripts that exhibited both shortening and up-regulation included numerous transcripts involved in the regulation of cell division and T cell activation. A table of select CELF1 target transcripts that shortened at six or 24 hours following stimulation and were up-regulated compared to resting cells can be found in Table 1. These findings are consistent with a model whereby the 3′ UTRs of a subset of CELF1 target transcripts were shortened by APA following T cell activation, and the resulting removal of GRE sequences stabilized and increased expression of these transcripts.
Fig.3.
CELF1 target transcripts that shorten through APA increase their expression. The percentage of APA shortened CELF1 target transcripts and non-CELF1 target transcripts that statistically significantly increased their expression at six or 24 hours was determined (*** = p-value < 10−8).
Table 1.
CELF1 target transcripts that shortened at six or 24 hours of stimulation and are up-regulated compared to resting cells.
| Gene ID | Gene Name | FC@ 6hrsa | FC @ 24hrsa | p-value 6hrsb | p-value 24hrsb | Function |
|---|---|---|---|---|---|---|
| YME1L1 | YMEl-like 1 ATPase | 1.26 | 1.18 | 0.02 | 0.03 | Cell proliferation |
| EIF4E | Eukaryotic translation initiation factor 4e | 1.77 | 2.31 | 0.02 | 0.01 | Gl/s transition of mitotic cell cycle, Positive regulation of mitotic cell cycle |
| ZC3HE | Zinc finger CCCH-type containing 8 | 1.92 | 2.20 | 0.004 | 0.01 | T cell homeostasis |
| 0GFOD1 | 2-oxoglutarate and iron-dependent oxygenase domain containing 1 | 1.29 | 1.79 | 0.02 | 0.001 | Cell proliferation |
| PPP2R2A | Protein phosphatase 2, regulatory submit B, alpha | 1.22 | 1.21 | 0.05 | 0.01 | G2/M transition of mitotic cell cycle |
| MINA | Myc induced A | 1.36 | 2.35 | 0.03 | 0.004 | Cell proliferation, Ribosome biogenesis |
| NDFIP1 | Nedd4 family interacting protein 1 | 2.61 | 1.31 | 0.002 | 0.09 | Regulation of Tcell proliferation, Regulation of lymphocyte differentiation |
| CD96 | CD96 molecule | 1.73 | 0.93 | 0.03 | 0.73 | Cell adhesion, Regulation of immune response |
| DEK | DEK oncogene | 1.42 | 1.05 | 0.00 | 0.36 | Regulation of double-stranded break repair |
| PDS5A | PDS5 regulator of cohesion maintenance, homolog A (S. cerevisiae) | 1.31 | 1.43 | 0.05 | 0.01 | Mitotic cell cycle |
| ARIH2 | Ariadne RBR E3 ubiquitin protein ligase 2 | 1.11 | 1.43 | 0.07 | 0.01 | Developmental cell growth, hematopoietic stem cell proliferation |
| SEC24A | SEC24 family member A | 1.31 | 1.65 | 0.11 | 0.003 | Antigen processing and presentation of peptide via MHC class 1 |
| ACTR2 | Actin-related protein 2 homolog (yeast) | 0.99 | 1.27 | 0.90 | 0.03 | Cell division, Innate immune response, meiotic cytokinesis |
| RBBP4 | Retinoblastoma binding protein 4 | 0.90 | 1.27 | 0.13 | 0.02 | G2/M transition of mitotic cell cycle |
Fold Change in average expression compared to resting cells
P-value for increased expression compared to resting cells
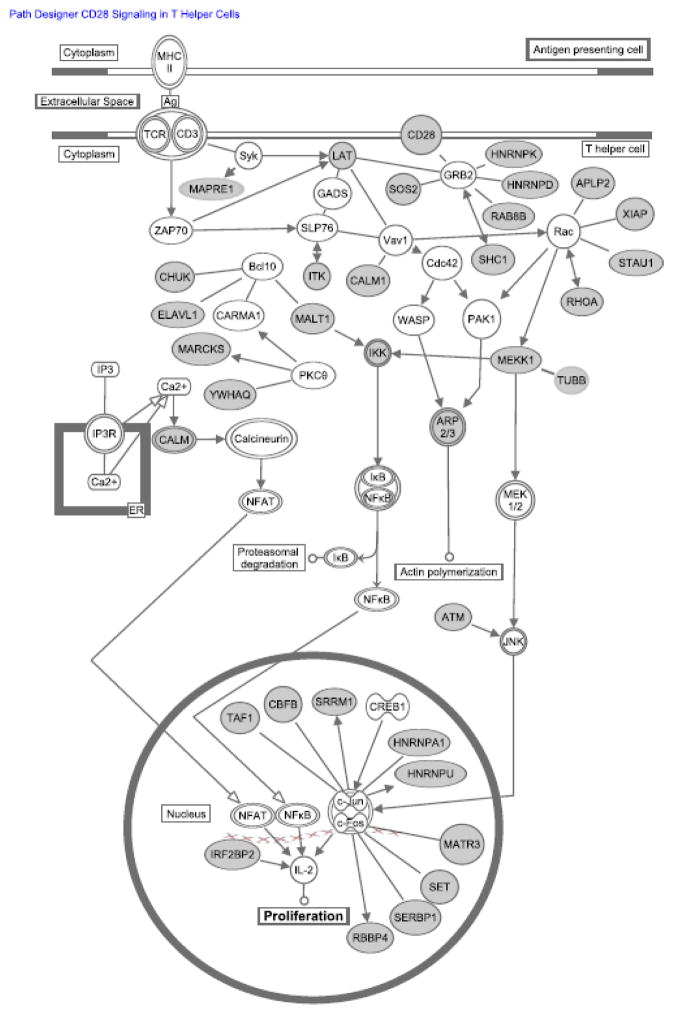
Since CELF1 target transcript expression correlated with change in 3′ UTR length, we investigated the functions of transcripts that were shortened to determine if these transcripts were coordinately regulated following T cell activation as part of a functional gene expression program. We searched for enriched gene ontology terms in the CELF1 target transcripts that shortened at six or 24 hours compared to the list of all CELF1 target transcripts using the online Gene Ontology Enrichment and Visualization Tool (GOrilla, http://cbl-gorilla.cs.technion.ac.il) [28]. We found that the term “Cell Division” was the only enriched term for the list of transcripts shortened via APA at six and 24 hours of stimulation (p<0.001 for both six and 24 hours). Through inspection of this list of shortened CELF1 target transcripts involved in cell division, we found a significant number of transcripts were also involved in T cell activation (figure 4). These findings support a role for APA of CELF1 target transcripts in coordinately regulating the expression of a network of transcripts that control cellular proliferation in response to T cell activation.
Fig. 4.
CELF1 target transcripts that shortened following T cell activation encode regulators of cell division. Transcripts depicted in gray are CELF1 target transcripts that were shortened at six and 24 hours after T cell activation that regulate cellular proliferation through T cell receptor and co-receptor signaling pathways.
4. Discussion
APA is an evolutionarily conserved process regulating transcript 3′ end length that isutilized in a wide variety of cellular contexts involving cellular proliferation, including cancer and T cell activation. An association between 3′end shortening through APA and cellular proliferation has been described, but mechanisms underlying this association have not previously been elucidated. In the present study, we found that 3′ end shortening through APA was responsible for the coordinate regulation of a network of CELF1 target transcripts that regulate cell division following T cell activation. This result was striking given the previously reported association between cellular proliferation and APA [4,8], and implicates APA as a mechanism for the regulating proliferation through a specific posttranscriptional network involving CELF1 and GREs. Our data suggest that CELF1 target transcripts that encode regulators of cell division are preferentially regulated through 3′ end shorting via APA following T cell activation, and the resulting up-regulation of CELF1 target transcripts promotes cellular proliferation. CELF1 is a ubiquitously expressed protein whose target transcripts regulate cellular proliferation across a number of evolutionarily distinct species [16,29,30]. The evolutionary conservation of CELF1 targets encoding regulators of proliferation, and our finding that CELF1 targets are preferentially shortened through APA, suggests that APA of transcripts in the CELF1 network may regulate proliferation across a number of physiologic processes and cellular contexts.
The spatial relationship between CELF1 binding sites (ie. GREs) and upstream APA sites, prompted us to propose the following hypothetical model through which CELF1 could regulate APA. The polyadenylation factor CSTF-64, and it’s tau isoform, have been shown to bind downstream of polyA sites, and their recently described binding motifs have significant sequence similarity to the GRE [10,11]. Since polyadenylation factor CSTF-64 and CELF1 may both recognize GRE sequences downstream of APA sites, it is possible they compete for binding to the same site. The disruption of CELF1-binding to RNA through activation-dependent phosphorylation of CELF1 following T cell activation may prevent CELF1 binding to 5′ APA sites, whereas increased CSTF-64 expression and function following T cell activation may promote use of 5′ APA sites [12–14](see figure 5). Failure of CELF1 to bind to GRE sequences immediately downstream of APA sites due to CELF1 phosphorylation after T cell activation may leave GRE sites unoccupied by CELF1 and available for binding by CSTF-64, which then directs upstream APA site usage. Future work to validate this model will be crucial to understanding the potential participation of CELF1 in the regulation of APA.
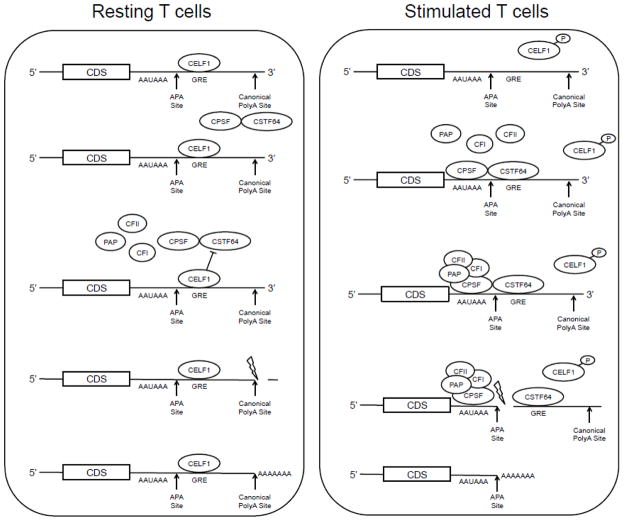
Fig. 5.
Potential role of mechanism of CELF1in regulating APA. In resting T cells, CELF1 binds to the GRE located downstream of 5′ APA sites and may competitively inhibit the binding of CSTF64-CSPF heterodimer to the GRE, preventing utilization of the 5′ APA site. In activated T cells, CELF1 is phosphorylated resulting in decreased RNA-binding activity. CELF1’s reduced binding to the GRE may allow the CSTF64-CSPF heterodimer access to bind the pre-mRNA molecule and recruit the remaining APA machinery to increase utilization of the 5′APA site.
We have previously shown that CELF1-mediated mRNA decay is disrupted by activation-dependent phosphorylation of CELF1 following T cell activation [17]. Here we report 3′end shortening through APA as a second mechanism occurring during T cell activation that interrupts CELF1 function through removal of its RNA-binding site. These mechanisms, which act on different transcript subsets, occur in temporally distinct patterns to finely regulate CELF1 function. CELF1 phosphorylation is a rapid and transient mechanism to interrupt global CELF1 function shortly following T cell activation. In contrast, APA may represent a regulatory mechanism acting later to stabilize a subset of CELF1 target transcripts so they are no longer susceptible to CELF1-mediated degradation. These distinct regulatory phenomena provide a mechanism for the independent regulation of functionally distinct subsets of CELF1 target transcripts over the course of T cell activation.
The results presented here suggest that APA functions to coordinately interrupt CELF1 function through isoform specific exclusion of the GRE following T cell activation. Given the mRNA degradation function of CELF1, the extension of this finding is that loss of the GRE through APA can result in isoform specific up-regulation of transcript expression through mRNA stabilization. These findings are consistent with recently reported results in yeast suggesting that transcript isoforms produced through APA have differing decay rates based on whether or not specific RNA-binding protein target motif are included [31,32]. Our results contribute to the growing body of literature suggesting that post-transcriptional gene regulation occurs in an mRNA isoform specific manner and is dependent on the specific regulatory sequence content included within each transcript isoform.
5. Conclusion
We used next-generation sequencing techniques to evaluate APA 6 hours and 24 hours following T cell activation and found that transcript 3′ end shortening through APA led to coordinated loss of GRE sequences from CELF1 target transcripts involved in cellular proliferation. Among CELF1 target transcripts that underwent APA following T cell activation, we found enrichment of GRE sequences immediately downstream of 5′ APA sites, leading to a proposed model whereby binding by CELF1 to these sites could regulate transcript shortening in the nucleus. In resting T cells, binding by CELF1 to GRE sequences immediately downstream of 5′ APA sites may prevent their utilization, and therefore downstream APA sites are used. The resulting transcripts contain GRE sequences in their 3′ UTRs and are susceptible to GRE-mediated mRNA decay. We propose that subsequent phosphorylation of CELF1 following T cell activation prevents CELF1 binding to GRE sequences downstream of 5′ APA sites, allowing these sites to be utilized for APA. Our results showed that many of these shortened CELF1 target transcripts were up-regulated following T cells activation, perhaps through transcript stabilization because they no longer contain GRE sequences. Overall, our findings provide a mechanistic link between APA and cellular proliferation. Although APA remains incompletely understood, APA is a common phenomenon throughout evolution utilized in a number of processes involving cellular proliferation, including T cell activation, development and oncogenesis. The findings presented here suggest a mechanism underlying the association between proliferation and APA through the coordinated disruption of CELF1-mediated decay of an important network of transcripts that encode regulators of cell division. These findings have implications for understanding coordinate gene regulation at the post-transcriptional level and shed light on the mechanism and biological function of APA.
Supplementary Material
Calculated weighted average lengths of all transcripts from all donors at all time points.
Database of genomic coordinates of experimentally detected polyA sites from all donors at all time points.
RPMK expression values of T cell transcripts throughout the first 24 hours of T cell stimulation.
Highlights.
Global polyadenylation site usage was determined following T cell activation.
CELF1 targets were preferentially shortened via APA following T cell activation.
Shortening of CELF1 targets caused loss of the GRE and increased expression.
CELF1 targets shortened via APA encode regulators of cellular proliferation.
A model for CELF1-mediated regulation of APA was proposed.
Acknowledgments
We thank the Biomedical Genomics Center and the Minnesota Supercomputing Institute at the University of Minnesota for their expertise and services. This work was supported by NIH grants AI057484, AI072068 and AI096925 to P.R.B. D.B. was supported by NIH grant 3T32GM8244-24S2.
Abbreviations
- 3′UTR
3 prime Untranslated Region
- APA
Alternative Polyadenylation
- bp
base pair
- CELF1
CUGBP, Elav-Like Family Member 1
- CUGBP1
CUG-triplet repeat, RNA binding protein 1
- CD3
Cluster of Differentiation 3
- CD28
Cluster of Differentiation 28
- CSTF-64
Cleavage Stimulation Factor 64 kDa subunit
- CSTF-64τ
Cleavage Stimulation Factor 64 kDa subunit tau variant
- G
Guanine
- GRE
GU-Rich Element
- mRNA
Messenger RNA
- nt
nucleotide
- polyA
Polyadenylation
- TCR
T Cell Receptor
- U
Uracil
Footnotes
Author Contributions:
D.B. designed experiments, analyzed data, and wrote the manuscript. C.R. analyzed data and wrote and edited the manuscript. P.R.B. designed experiments and wrote and edited the manuscript.
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could a3ect the content, and all legal disclaimers that apply to the journal pertain.
Contributor Information
Daniel Beisang, Email: beis0051@umn.edu.
Cavan Reilly, Email: cavanr@biostat.umn.edu.
Paul R. Bohjanen, Email: bohja001@umn.edu.
References
- 1.Raghavan A, Dhalla M, Bakheet T, Ogilvie RL, Vlasova IA, Khabar KSA, et al. Patterns of coordinate down-regulation of ARE-containing transcripts following immune cell activation. Genomics. 2004;84:1002–1013. doi: 10.1016/j.ygeno.2004.08.007. [DOI] [PubMed] [Google Scholar]
- 2.Cheadle C, Fan J, Cho-Chung Y, Werner T, Ray J, Do L, et al. Control of gene expression during T cell activation: alternate regulation of mRNA transcription and mRNA stability. BMC Genomics. 2005;6:75. doi: 10.1186/1471-2164-6-75. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Martinez NM, Pan Q, Cole BS, Yarosh CA, Babcock GA, Heyd F, et al. Alternative splicing networks regulated by signaling in human T cells. Rna. 2012;18:1029–1040. doi: 10.1261/rna.032243.112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Sandberg R, Neilson JR, Sarma A, Sharp PA, Burge CB. Proliferating Cells Express mRNAs with Shortened 3′ Untranslated Regions and Fewer MicroRNA Target Sites. Science. 2008;320:1643–1647. doi: 10.1126/science.1155390. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Ozsolak F, Kapranov P, Foissac S, Kim SW, Fishilevich E, Monaghan AP, et al. Comprehensive Polyadenylation Site Maps in Yeast and Human Reveal Pervasive Alternative Polyadenylation. Cell. 2010;143:1018–1029. doi: 10.1016/j.cell.2010.11.020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Li XQ, Du D. RNA polyadenylation sites on the genomes of microorganisms, animals, and plants. PLoS ONE. 2013;8:e79511. doi: 10.1371/journal.pone.0079511. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Ji Z, Lee JY, Pan Z, Jiang B, Tian B. Progressive lengthening of 3′ untranslated regions of mRNAs by alternative polyadenylation during mouse embryonic development. Proceedings of the National Academy of Sciences. 2009;106:7028–7033. doi: 10.1073/pnas.0900028106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Mayr C, Bartel DP. Widespread Shortening of 3′UTRs by Alternative Cleavage and Polyadenylation Activates Oncogenes in Cancer Cells. Cell. 2009;138:673–684. doi: 10.1016/j.cell.2009.06.016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Di Giammartino DC, Nishida K, Manley JL. Mechanisms and Consequences of Alternative Polyadenylation. Molecular Cell. 2011;43:853–866. doi: 10.1016/j.molcel.2011.08.017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Yao C, Biesinger J, Wan J, Weng L, Xing Y, Xie X, et al. Transcriptome-wide analyses of CstF64-RNA interactions in global regulation of mRNA alternative polyadenylation. Proceedings of the National Academy of Sciences. 2012;109:18773–18778. doi: 10.1073/pnas.1211101109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Yao C, Choi EA, Weng L, Xie X, Wan J, Xing Y, et al. Overlapping and distinct functions of CstF64 and CstF64τ in mammalian mRNA 3′ processing. Rna. 2013;19:1781–1790. doi: 10.1261/rna.042317.113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Takagaki Y, Manley JL. Levels of polyadenylation factor CstF-64 control IgM heavy chain mRNA accumulation and other events associated with B cell differentiation. Molecular Cell. 1998;2:761–771. doi: 10.1016/s1097-2765(00)80291-9. [DOI] [PubMed] [Google Scholar]
- 13.Martincic K, Campbell R, Edwalds-Gilbert G, Souan L, Lotze MT, Milcarek C. Increase in the 64-kDa subunit of the polyadenylation/cleavage stimulatory factor during the G0 to S phase transition. Proc Natl Acad Sci USA. 1998;95:11095–11100. doi: 10.1073/pnas.95.19.11095. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Shell SA, Hess C, Morris SM, Jr, Milcarek C. Elevated Levels of the 64-kDa Cleavage Stimulatory Factor (CstF-64) in Lipopolysaccharide-stimulated Macrophages Influence Gene Expression and Induce Alternative Poly(A) Site Selection. Journal of Biological Chemistry. 2005;280:39950–39961. doi: 10.1074/jbc.M508848200. [DOI] [PubMed] [Google Scholar]
- 15.Vlasova IA, Tahoe NM, Fan D, Larsson O, Rattenbacher B, SternJohn JR, et al. Conserved GU-Rich Elements Mediate mRNA Decay by Binding to CUG-Binding Protein 1. Molecular Cell. 2008;29:263–270. doi: 10.1016/j.molcel.2007.11.024. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Rattenbacher B, Beisang D, Wiesner DL, Jeschke JC, von Hohenberg M, St Louis-Vlasova IA, et al. Analysis of CUGBP1 Targets Identifies GU-Repeat Sequences That Mediate Rapid mRNA Decay. Mol Cell Biol. 2010;30:3970–3980. doi: 10.1128/MCB.00624-10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Beisang D, Rattenbacher B, Vlasova-St Louis IA, Bohjanen PR. Regulation of CUG-binding Protein 1 (CUGBP1) Binding to Target Transcripts upon T Cell Activation. Journal of Biological Chemistry. 2012;287:950–960. doi: 10.1074/jbc.M111.291658. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Raghavan A, Robison RL, McNabb J, Miller CR, Williams DA, Bohjanen PR. HuA and Tristetraprolin Are Induced following T Cell Activation and Display Distinct but Overlapping RNA Binding Specificities. J Biol Chem. 2001;276:47958–47965. doi: 10.1074/jbc.M109511200. [DOI] [PubMed] [Google Scholar]
- 19.Langmead B, Salzberg SL. Fast gapped-read alignment with Bowtie 2. Nat Meth. 2012;9:357–359. doi: 10.1038/nmeth.1923. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Kent WJ, Sugnet CW, Furey TS, Roskin KM, Pringle TH, Zahler AM, et al. The Human Genome Browser at UCSC. Genome Research. 2002;12:996–1006. doi: 10.1101/gr.229102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Karolchik D, Hinrichs AS, Furey TS, Roskin KM, Sugnet CW, Haussler D, et al. The UCSC Table Browser data retrieval tool. Nucleic Acids Res. 2004;32:493D–496. doi: 10.1093/nar/gkh103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N, et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics. 2009;25:2078–2079. doi: 10.1093/bioinformatics/btp352. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Blankenberg D, Kuster GV, Coraor N, Ananda G, Lazarus R, Mangan M, et al. Galaxy: A Web-Based Genome Analysis Tool for Experimentalists. John Wiley & Sons, Inc; Hoboken, NJ, USA: 2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Giardine B. Galaxy: A platform for interactive large-scale genome analysis. Genome Research. 2005;15:1451–1455. doi: 10.1101/gr.4086505. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Goecks J, Nekrutenko A, Taylor J T Galaxy Team. Galaxy: a comprehensive approach for supporting accessible, reproducible, and transparent computational research in the life sciences. Genome Biol. 2010;11:R86. doi: 10.1186/gb-2010-11-8-r86. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Ozsolak F, Platt AR, Jones DR, Reifenberger JG, Sass LE, McInerney P, et al. Direct RNA sequencing. Nature. 2009;461:814–818. doi: 10.1038/nature08390. [DOI] [PubMed] [Google Scholar]
- 27.David M, Dzamba M, Lister D, Ilie L, Brudno M. SHRiMP2: Sensitive yet Practical Short Read Mapping. Bioinformatics. 2011;27:1011–1012. doi: 10.1093/bioinformatics/btr046. [DOI] [PubMed] [Google Scholar]
- 28.Eden E, Navon R, Steinfeld I, Lipson D, Yakhini Z. GOrilla: a tool for discovery and visualization of enriched GO terms in ranked gene lists. BMC Bioinformatics. 2009;10:48. doi: 10.1186/1471-2105-10-48. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Graindorge A, Le Tonqueze O, Thuret R, Pollet N, Osborne HB, Audic Y. Identification of CUG-BP1/EDEN-BP target mRNAs in Xenopus tropicalis. Nucleic Acids Res. 2008;36:1861–1870. doi: 10.1093/nar/gkn031. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Lee JE, Lee JY, Wilusz J, Tian B, Wilusz CJ. Systematic Analysis of Cis-Elements in Unstable mRNAs Demonstrates that CUGBP1 Is a Key Regulator of mRNA Decay in Muscle Cells. PLoS ONE. 2010;5:e11201. doi: 10.1371/journal.pone.0011201. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Pelechano V, Wei W, Steinmetz LM. Extensive transcriptional heterogeneity revealed by isoform profiling. Nature. 2013;497:127–131. doi: 10.1038/nature12121. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Gupta I, Clauder-Munster S, Klaus B, Jarvelin AI, Aiyar RS, Benes V, et al. Alternative polyadenylation diversifies post-transcriptional regulation by selective RNA-protein interactions. Molecular Systems Biology. 2014;10:719–719. doi: 10.1002/msb.135068. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Calculated weighted average lengths of all transcripts from all donors at all time points.
Database of genomic coordinates of experimentally detected polyA sites from all donors at all time points.
RPMK expression values of T cell transcripts throughout the first 24 hours of T cell stimulation.